Abstract
The structure of pollen grains is related to the reproductive function of the plants. Here, three-dimensional (3D) refractive index maps were obtained for individual conifer pollen grains using optical diffraction tomography (ODT). The 3D morphological features of pollen grains from pine trees were investigated using measured refractive index maps, in which distinct substructures were clearly distinguished and analyzed. Morphological and physiochemical parameters of the pollen grains were quantified from the obtained refractive index (RI) maps and used to quantitatively study the interspecific differences of pollen grains from different strains. Our results demonstrate that ODT can assess the structure of pollen grains. This label-free and rapid 3D imaging approach may provide a new platform for understanding the physiology of pollen grains.
Similar content being viewed by others
Introduction
Pollen grains are the male gametophytes of seed plants and are essential to the life cycle of many plants that play a major role in ecosystems. The study of pollen also has a key role in multiple fields of science. For instance, many crucial perspectives, ranging from evidence of geological history in paleontology1, to plant reproduction2, and to breeding technology in agriculture3, have been advanced by investigations of pollen grains. However, measuring the three-dimensional (3D) structure of individual pollen grains remains challenging, mainly due to the limitations of conventional imaging approaches. Many conventional imaging methods lack the capacity for quantitative 3D measurement, and even advanced 3D imaging methods have limitations or may require highly specialized equipment to image pollen grains, particularly those of conifer plants.
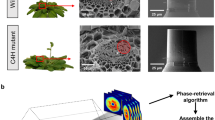
Bright field microscopy and electron microscopy have been conventionally used for imaging pollen grains, yet they have distinct limitations. Bright field microscopy allows convenient real-time observation of pollen grains, yet the resulting images have only provided qualitative 2D information4,5. For high-resolution imaging of conifer pollen grains, electron microscopy has been employed. In previous studies, scanning electron microscopy has provided detailed images of Pinus pollen grains6,7,8,9. However, scanning electron microscopy can only measure the surface attributes of pollen grains that have been prepared in advance using processes such as coating with gold or chromium.
In another study, transmission electron microscopy was employed to study sections of pollen grains of fixed conifer plants6. However, transmission electron microscopy is not capable of measuring entire pollen grains because the specimens are restricted to slices of nanometer-scale thickness, while whole pollen grains are up to tens of micrometers.
3D optical imaging methods have also been exploited to acquire volumetric images of conifer pollen grains. For instance, fluorescence imaging can provide 3D images of fluorescent molecules in conifer pollen grains. In a previous study, 3D imaging of autofluorescence signals from conifer pollen grains was performed10. Studies which imaged conifer pollen grains using an external dye11 and transgenic fluorescent protein12 also suggest that various molecules can be targeted to enhance the 3D imaging of conifer pollen grains. However, fluorescence imaging may chemically alter the sample13, produce photobleaching14, or interfere with the intrinsic pigments of the sample15. It is also noteworthy that fluorescence images can only provide information about the distribution of fluorescent molecules.
Recently, tomographic imaging of Pinus pollen grains was demonstrated using a hard X-ray source16. However, the utilization of hard X-rays creates the risk of radiation damage to the sample, and studies may be hampered by limited access to appropriate X-ray imaging facilities.
Lately, quantitative phase imaging (QPI) methods have emerged, and because they offer quantitative and label-free imaging capability, now play an important role in the study of cellular physiologies17,18. By exploiting the principle of laser interferometry, QPI can quantitatively measure the phase delay of light scattered by a sample. QPI does not rely on an exogenous labeling process, because the phase delay, which is equivalent to the multiplied product of the refractive index (RI) and the thickness of the sample, is determined by the inherent composition of the sample19.
More recently, 3D QPI techniques including optical diffraction tomography (ODT) have been extensively utilized for imaging and analyzing the internal 3D structures of various transparent biological samples20, including phytoplankton21, microalgae22, bacteria23,24,25, yeasts26, mammalian blood cells27,28,29,30,31, and mammalian cells32,33,34,35. ODT is a technique that reconstructs a 3D RI map from light scattered from a sample subjected to illumination at different angles36,37,38. The measured RI is the retardation coefficient of light speed and is related to the density of dipoles in the medium. As a result, using ODT, the structure of a sample can be identified in 3D from the reconstructed RI map.
In this study, we measured and analyzed the 3D structures of pollen grains from Pinus plants using ODT. Characteristic morphological features of the pollen grains were investigated using the RI maps. In addition, we quantified three parameters of individual pollen grains from their RI maps and used them to identify the morphological and chemical properties of the pollen grains.
Result
3D structures of Pinus pollen grains in RI maps
RI maps of individual pollen grains from four types of local Pinus plants were reconstructed using ODT as shown in Fig. 1 (see Methods). Analogous to the sectional images obtained in X-ray computed tomography, 3D RI maps can be visualized at lateral cross-sections (Fig. 2). The bisaccate morphology, which is a commonly shared trait of pollen grains from conifer plants, was clearly visible in the cross-sectional images. Due to the high level of morphological variations between individual pollen grains, an abstract description of the morphology alone is not sufficient to determine the strain of a given pollen grain. 3D rendering images of the entire 3D RI maps visualize the 3D morphologies of the pollen grains (Fig. 2 and Supplementary Video S1–8).
Reconstruction of a three-dimensional (3D) refractive index (RI) map using optical diffraction tomography. (a) A schematic diagram of illumination angle scanning. (b) The optical setup for measurement of the light scattered by the sample. 2 × 2 SMFC: 2-by-2 single-mode fiber optics coupler; C: collimator; TL: tube lens; M: mirror, CL: condenser lens; OL: objective lens, BS: beam splitter; P: polarizer. (c) Image processing for reconstruction of 3D RI maps from angle-scanned holograms. (i) indicates the optical field retrieval process, and (ii) indicates the tomographic reconstruction process. Scale bar = 20 μm.
Three-dimensional (3D) refractive index (RI) maps of individual Pinus pollen grains. Axial section images (upper) and 3D rendering images (lower) of pollen grains from Korean red pine (a), Korean golden pine (b), Japanese red pine (c), and Japanese black pine. For each strain, the left column and the right column presents a whole pollen grain and an empty shell, respectively. Three upper three rows display the sectional RI images at different axial depths. In the lower row, RI between 1.5 and 1.515 is rendered in violet and RI between 1.525 and 1.54 is rendered in yellow-green. Scale bar = 20 μm. Cube length = 60 μm.
In order to demonstrate the 3D imaging capability of ODT, detailed substructures of Pinus pollen grains were further investigated. Structures of exines, or integumentary shells of pollen grains, could be observed in the RI maps of the hollow shells of pollen grains (Fig. 3a). The shell of a pollen grain was distinguished from the medium in the RI map. The shell exhibited RI over 1.53 which was 0.01 higher than the medium RI. The distinction between the medium and the shell in RI provided a sharp outline of each pollen grain.
Characteristic structures of Pinus pollen grains. (a) Axial section images of the hollow shells of pollen grains that exhibit distinguishing exine structures. The sacci (i) are shown to be empty and macroporous exine sacks of refractive index (RI) over 1.53. The cappa (ii) appears as a thick and rigid exine wall of RI near 1.54. The germinal wrinkles (iii) are located at the ventral side of a pollen grain between the two sacci. (b) Axial section images of whole pollen grains. Starch granules (given examples using white arrows) are visualized due to the RI contrast between starch and the rest of the corpus. Scale bar = 20 μm.
The sacci, which are air sacks attached to the pollen grains, were recognizable as paired chambers bulging towards the outside of the pollen grain (Fig. 3a; see the mark (i)). A pair of sacci was separated from the body of the pollen grain with a thin exine screen, while the outer surfaces were composed of the thick yet macroporous walls of the exine. The sacci were depicted as filled with the background medium since the RI level inside the sacks was highly homogeneous near the medium RI.
The cappa, which refers to the thick region of an exine at the dorsal pole of a pollen grain7, was visualized in the RI maps as a robust and smooth exine barrier with an RI near 1.54 (Fig. 3a; see the mark (ii)). The RI map showed that the cappa of a Pinus pollen grain covers a wide area of the dorsal exine. Wrinkles at the ventral poles of the exine shells were depicted in the RI maps as ravines in the exine shells between the air sacks (Fig. 3a; see the mark (iii)). The ends of the wrinkled exine were penetrating towards the centers of the pollen grains; this is an observation which was not revealed in scanning electron microscopy images of conifer pollen grains6,7.
The internal substructures of the pollen grains were investigated using RI maps of the entire pollen grains (Fig. 3b). Unlike the image of a hollow exine shell, the region inside the body of a whole pollen grain, which is occasionally referred to as the corpus, displayed an inhomogeneous distribution of RI. In the region, RI levels both higher and lower than the medium RI were observed. Inside the exine of a whole pollen grain, were multiple granules of 2–5 μm diameter with RI near 1.53. (Fig. 3b; see the white arrows) The granules were found to be embedded in a background of RI lower than the medium RI. The elliptical morphology, the size, and the RI levels of the granules suggested that they were starch granules stored in the pollen grains39,40.
Quantification of parameters using RI maps
Morphological parameters including volume and surface area were quantified by distinguishing the pollen grains from the medium in the RI maps (Fig. 4a and b). The volumes of pollen grains were 33.9 ± 10.6, 43.0 ± 14.4, 21.9 ± 5.5, and 37.4 ± 8.2 pl (mean ± standard deviation) for Korean red pine, golden Korean red pine, Japanese red pine, and Japanese black pine, respectively. The results of the Mann-Whitney U test showed that there were all significant differences (P-value < 0.05) in the volumes of pollen grains between different strains of Pinus plants, except for between golden Korean red pine and Japanese black pine. Surface areas of the pollen grains from the plants in the same order were 7490 ± 1590, 8600 ± 2030, 5230 ± 760, and 7700 ± 1280 μm2. According to the Mann-Whitney U test, the interspecific differences of the surface areas of pollen grains were significant except for between Korean red pine and Japanese black pine. The sphericity index was also quantified for each pollen grain (Fig. 4c). The sphericity index is an indicator for the resemblance of a shape to a sphere. For an individual pollen grain, the sphericity index was calculated from the volume and the surface area of the pollen grain, using the following formula: \(SI={(36\pi {V}^{2})}^{1/3}/S\), where SI, V and S indicate the sphericity index, the volume, and the surface area of the pollen grain respectively. The sphericity indices of the pollen grains from the plants of the identical order were 0.674 ± 0.078, 0.686 ± 0.077, 0.720 ± 0.053, and 0.703 ± 0.058. In the result of the Mann-Whitney U test, sphericity indices of pollen grains were significantly different between Korean red pine and Japanese red pine, and between Korean red pine and Japanese black pine. In addition, information about the distribution of chemical components in pollen grains was quantified from the RI maps of entire pollen grains. Masses of starch, which is a major source of nutrition in pollen grains, were quantified for individual pollen grains (Fig. 4d). Starch, which corresponds to high-RI granules inside the exines of pollen grains, was specifically identified and investigated. Using the density of dry starch, the masses of starch could be quantified. The starch contents were 1.007 ± 1.483, 1.193 ± 1.327, 0.894 ± 0.535, and 0.171 ± 1.671 ng, respectively for pollen grains from Korean red pine, golden Korean red pine, Japanese red pine, and Japanese black pine. Due to the high variation between individual pollen grains, the standard deviation of starch content was comparable to the mean value in every strain we studied. The results of the Mann-Whitney U test indicated there was no significant difference in starch content among the experimentally tested strains of Pinus plants.
Quantification of parameters from Pinus pollen grains. (a) The volumes, (b) surface areas, (c) sphericity indices, and (d) starch contents of individual pollen grains, quantified from the 3D RI maps. Each error bars placed on the left of the scatter plots represent mean ± standard deviation. The symbols *, **, *** and **** refer to significant differences in the Mann-Whitney U test with p-values less than 0.05, 0.01, 0.001 and 0.0001, respectively. The absence of an asterisk refers to a difference of no significance in the Mann-Whitney U test.
Discussion
To summarize, we demonstrated and obtained label-free imaging and analysis of the 3D structures of Pinus pollen grains. From the reconstructed 3D RI maps obtained using ODT, detailed morphological features of individual Pinus pollen grains were identified. Furthermore, quantitative parameters which represent the structures of the pollen grains were quantified using the maps to highlight the variances within and between pollen grains from different strains of Pinus plants.
While the imaging capacity of ODT is remarkable, the technique does not presently allow identification with high chemical specificity, although this limitation can be further improved using other techniques. While RI is an inherent optical property of matter, it cannot be used to determine the chemical composition of a sample alone. Clarifying the relationships between the structures in RI maps and their chemical constituents can be accomplished by inter-calibrating the ODT technique with other techniques that provide high chemical accuracy, including chemically resolving assays and image registration with fluorescence imaging10,41. Advances in high-modality QPI techniques may also provide additional chemical information without labeling. Recently, a method for hyperspectral ODT was developed, demonstrating the reconstruction of multiple RI maps with various wavelengths for illumination42. Since plant tissues have distinct absorption spectra, measuring complex RI depending on frequency can provide additional information that can be used to localize the absorptive components in pollen grains. We note that the present method can also be combined with fluorescence imaging, to access both the morphological and molecular specific information about pollen grains and perform correlative image analysis43,44,45,46.
Birefringence is another property that can be exploited to characterize pollen grains. Crystalline starch, which is one of the nutritional sources in pollen grains, can be detected more precisely using birefringence47, while the measurement of full Jones matrices of microscopic samples has been demonstrated, using QPI and polarization filtering48.
Due to the advantages that ODT possesses, the present method can serve as a new platform for investigating pollen grains, after refinements of the aforementioned limitations. The 3D RI maps are not vulnerable to uncontrolled alterations such as photobleaching and visualize unstained components. Investigating the morphologies of pollen grains using ODT can further provide useful insights into the life cycle of plants because the morphologies influence the mechanical behaviors that play important roles during reproduction. In previous studies, the surface texture and volume of the pollen grains were found to be linked with the aerodynamics during wind pollination of conifer plants49,50. Also, it was also discovered that the structural features of sacci were related to the flotation of pollen grains during fertilization of conifer plant ovules6,51. Therefore, it is likely that label-free identification of pollen structures using ODT will provide useful insights in understanding plant reproduction. For example, ODT may be employed to investigate the structure and the formation of pollen tubes, which have been extensively studied using fluorescence microscopy52,53. ODT also enables rapid examinations of palynological specimens, due to the lack of need for labeling processes. We also expect that use of ODT to identify allergenic pollen grains among airborne particles will allow early caution and prevention against allergic disorders, improving the health of communities54,55,56.
Methods
Sample collection and preparation
The pollen grains were collected during late April and May from blooming pine trees of a local farm. Pollen grains were placed in containers by gently shaking them off the male cones of the trees. Due to the contemporarily humid climate, the pollen grains were dried for approximately 2 hours at room temperature to prevent microbial invasion to the pollen grains. The containers with pollen grains were sealed and preserved in a dark cabinet at room temperature before imaging. In order to prepare pollen grains for imaging, they were suspended in a drop of RI-matching oil (Series A, Cargille Labs, United States) and sandwiched between two cover glasses (C024501, Matsunami Glass Ind. Ltd., Japan). The use of oil enables index matching between the sample and surrounding media, providing the appropriate imaging condition for ODT by significantly reducing light scattering57. The condition for drying and suspending pollen grains in oil does not significantly alter the physiological conditions58,59,60. Imaging was conducted using pollen grains that were not aggregated with other particles.
Image acquisition and 3D RI map reconstruction
Using ODT, the 3D RI map of a sample was reconstructed using holographic images of a sample that had been illuminated from different angles (Fig. 1a). The acquisition of phase and intensity images was performed using an off-axis Mach-Zehnder interferometric setup with a digital micromirror device (DMD) installed (HT-1S, Tomocube Inc., Republic of Korea) (Fig. 1b). The incident angles of illumination were rapidly scanned by projecting a series of binary holograms on the installed DMD61,62. The light scattered by the sample was transmitted to the camera along an imaging system consisting of an objective lens (60×, numerical aperture = 0.8) and a tube lens (f = 175 mm). On the camera plane, the transmitted light interferes with a plane reference beam, which is obliquely incident on the camera. The spatial resolution of the microscope is 166 nm and 1 μm for lateral and axial direction, respectively. The sensitivity of measuring RI values was 5 × 10−4, calculated as the standard deviation of the measured RI values for the area corresponding to the surrounding media. For a single pollen grain, the measurement of scattered fields required less than a second.
The phase and amplitude of the scattered light can be retrieved from the recorded interference pattern, and then processed for reconstruction of the 3D RI map (Fig. 1c). Reconstruction of the RI maps was conducted using the principle previously proposed36. Detailed descriptions of the reconstruction algorithm can be found elsewhere27,63.
Using ODT, 3D RI maps of a total of 92 entire pollen grains and 175 hollow shells of pollen grains were reconstructed. From Korean red pine trees (Pinus densiflora), a total of 6 pollen grains and 113 shells were imaged. From golden Korean red pine trees (Pinus densiflora Aurea), 17 pollen grains and 37 shells were imaged. From Japanese red pine trees (Pinus densiflora for. multicaulis), 38 pollen grains and 12 shells were imaged. From Japanese black pine trees (Pinus thunbergii), 31 pollen grains and 13 shells were imaged.
Parameter extraction
The structures of pollen grains from species of Pinus plants were statistically investigated by retrieving several properties from the measured 3D RI maps. In each 3D RI map, the volume occupied by the pollen grain was determined using RI contrast. The volume of the pollen grain was obtained by multiplying the unit volume of a voxel and the total number of voxels in the occupied region. The surface area was calculated as the area of the outermost surface of the occupied region. The sphericity index of a pollen grain was calculated from the volume and the surface area. A higher sphericity index indicates a stronger resemblance to a sphere. The region occupied by starch granules was also segmented using RI contrast, in a manner similar to how the pollen grain was separated from the background.
Statistical analyses
To quantify the interspecific differences between distributions of pollen properties, the Mann-Whitney U test was used. For each parameter, the p-values from the Mann-Whitney U test were used as indicators of interspecific similarities of the parameters. P-values were chosen to efficiently connote the statistical differences because a p-value from the Mann-Whitney U test is the probability that a pair of statistics arises from an identical population.
References
Lewin, R. Fragile Forests Implied by Pleistocene Pollen. Science 226, 36–37, https://doi.org/10.1126/science.226.4670.36 (1984).
Knight, T. M. et al. Pollen limitation of plant reproduction: pattern and process. Annu. Rev. Ecol. Evol. Syst. 36, 467–497 (2005).
Shivanna, K. R. & Sawhney, V. K. Pollen biotechnology for crop production and improvement. (Cambridge University Press, 1997).
deWin, A. H. N. et al. Development and cellular organization of Pinus sylvestris pollen tubes. Sex Plant Reprod 9, 93–101, https://doi.org/10.1007/s004970050015 (1996).
Derksen, J., Li, Y. Q., Knuiman, B. & Geurts, H. The wall of Pinus sylvestris L-pollen tubes. Protoplasma 208, 26–36, https://doi.org/10.1007/Bf01279072 (1999).
Runions, C. J., Rensing, K. H., Takaso, T. & Owens, J. N. Pollination of Picea orientalis (Pinaceae): saccus morphology governs pollen. Am J Bot 86, 190–197 (1999).
Bagnell, C. R. Species Distinction among Pollen Grains of Abies, Picea, and Pinus in Rocky-Mountain Area (a Scanning Electron-Microscope Study). Rev Palaeobot Palyno 19, 203–220 (1975).
Bykowska, J. & Klimko, M. Pollen Morphology of Pinus Mugo Turra X Pinus Sylvestris L. Hybrids and Parental Species in an Experimental Culture. Acta Biol Cracov Bot 57, 149–160, https://doi.org/10.1515/abcsb-2015-0009 (2015).
Shi, J., Cui, M., Yang, L., Kim, Y.-J. & Zhang, D. Genetic and biochemical mechanisms of pollen wall development. Trends in plant science 20, 741–753 (2015).
Punyasena, S. W., Tcheng, D. K., Wesseln, C. & Mueller, P. G. Classifying black and white spruce pollen using layered machine learning. New Phytol 196, 937–944, https://doi.org/10.1111/j.1469-8137.2012.04291.x (2012).
Anderhag, P. & Hepler, P. K. & Lazzaro, M. D. Microtubules and microfilaments are both responsible for pollen tube elongation in the conifer Picea abies (Norway spruce). Protoplasma 214, 141–157, https://doi.org/10.1007/Bf01279059 (2000).
Tian, L. N., Seguin, A. & Charest, P. J. Expression of the green fluorescent protein gene in conifer tissues. Plant Cell Rep 16, 267–271 (1997).
Dixit, R. & Cyr, R. Cell damage and reactive oxygen species production induced by fluorescence microscopy: effect on mitosis and guidelines for non-invasive fluorescence microscopy. Plant J 36, 280–290 (2003).
Song, L. L., vanGijlswijk, R. P. M., Young, I. T. & Tanke, H. J. Influence of fluorochrome labeling density on the photobleaching kinetics of fluorescein in microscopy. Cytometry 27, 213–223 (1997).
Zhou, X., Carranco, R., Vitha, S. & Hall, T. C. The dark side of green fluorescent protein. New Phytol 168, 313–322, https://doi.org/10.1111/j.1469-8137.2005.01489.x (2005).
Li, Q. et al. Pollen structure visualization using high-resolution laboratory-based hard X-ray tomography. Biochem Biophys Res Commun 479, 272–276, https://doi.org/10.1016/j.bbrc.2016.09.058 (2016).
Lee, K. et al. Quantitative phase imaging techniques for the study of cell pathophysiology: from principles to applications. Sensors (Basel) 13, 4170–4191, https://doi.org/10.3390/s130404170 (2013).
Popescu, G. Quantitative phase imaging of cells and tissues. (McGraw Hill Professional, 2011).
Kemper, B. & von Bally, G. Digital holographic microscopy for live cell applications and technical inspection. Applied optics 47, A52–A61 (2008).
Kim, K. et al. Optical diffraction tomography techniques for the study of cell pathophysiology. Journal of Biomedical Photonics & Engineering 2 (2016).
Lee, S. et al. High-resolution 3-D refractive index tomography and 2-D synthetic aperture imaging of live phytoplankton. Journal of the Optical Society of Korea 18, 691–697 (2014).
Jung, J. et al. Label-free non-invasive quantitative measurement of lipid contents in individual microalgal cells using refractive index tomography. bioRxiv, 226480 (2017).
Cotte, Y. et al. Marker-free phase nanoscopy. Nature Photonics 7, 113–117 (2013).
Kim, T. I. et al. Antibacterial Activities of Graphene Oxide–Molybdenum Disulfide Nanocomposite Films. ACS Applied Materials & Interfaces 9, 7908–7917 (2017).
Bennet, M., Gur, D., Yoon, J., Park, Y. & Faivre, D. A Bacteria‐Based Remotely Tunable Photonic Device. Advanced Optical Materials 5 (2017).
Habaza, M., Gilboa, B., Roichman, Y. & Shaked, N. T. Tomographic phase microscopy with 180 rotation of live cells in suspension by holographic optical tweezers. Optics letters 40, 1881–1884 (2015).
Kim, K. et al. High-resolution three-dimensional imaging of red blood cells parasitized by Plasmodium falciparum and in situ hemozoin crystals using optical diffraction tomography. J Biomed Opt 19, 011005, https://doi.org/10.1117/1.JBO.19.1.011005 (2014).
Lee, S. et al. Refractive index tomograms and dynamic membrane fluctuations of red blood cells from patients with diabetes mellitus. Scientific Reports 7 (2017).
Yoon, J. et al. Identification of non-activated lymphocytes using three-dimensional refractive index tomography and machine learning. Sci Rep 7, 6654, https://doi.org/10.1038/s41598-017-06311-y (2017).
Hur, J., Kim, K., Lee, S., Park, H. & Park, Y. Melittin-induced alterations in morphology and deformability of human red blood cells using quantitative phase imaging techniques. Sci Rep 7, 9306, https://doi.org/10.1038/s41598-017-08675-7 (2017).
Merola, F. et al. Tomographic flow cytometry by digital holography. Light: Science & Applications 6, e16241 (2017).
Kim, K. et al. Three-dimensional label-free imaging and quantification of lipid droplets in live hepatocytes. Scientific reports 6, 36815 (2016).
Villone, M. M. et al. Full-angle tomographic phase microscopy of flowing quasi-spherical cells. Lab Chip 18, 126–131, https://doi.org/10.1039/c7lc00943g (2017).
Yang, S. A., Yoon, J., Kim, K. & Park, Y. Measurements of morphological and biophysical alterations in individual neuron cells associated with early neurotoxic effects in Parkinson’s disease. Cytometry Part A 91, 510–518 (2017).
Kim, T. et al. White-light diffraction tomography of unlabelled live cells. Nature Photonics 8, 256–263 (2014).
Wolf, E. Three-dimensional structure determination of semi-transparent objects from holographic data. Optics Communications 1, 153–156 (1969).
Debailleul, M., Georges, V., Simon, B., Morin, R. & Haeberlé, O. High-resolution three-dimensional tomographic diffractive microscopy of transparent inorganic and biological samples. Optics letters 34, 79–81 (2009).
Lauer, V. New approach to optical diffraction tomography yielding a vector equation of diffraction tomography and a novel tomographic microscope. Journal of Microscopy 205, 165–176 (2002).
Wolf, M. J., Ruggles, V. J. & Macmasters, M. M. Refractive indices of wheat starch granules at various moisture levels determined with an interference microscope. Biochim Biophys Acta 57, 135–142 (1962).
Fernando, D. D., Lazzaro, M. D. & Owens, J. N. Growth and development of conifer pollen tubes. Sex Plant Reprod 18, 149–162, https://doi.org/10.1007/s00497-005-0008-y (2005).
Schmidt, M. A. & Herman, E. M. Suppression of Soybean Oleosin Produces Micro-Oil Bodies that Aggregate into Oil Body/ER Complexes. Mol Plant 1, 910–924, https://doi.org/10.1093/mp/ssn049 (2008).
Jung, J., Kim, K., Yoon, J. & Park, Y. Hyperspectral optical diffraction tomography. Opt. Express 24, 2006–2012, https://doi.org/10.1364/OE.24.002006 (2016).
Kim, K. et al. Correlative three-dimensional fluorescence and refractive index tomography: bridging the gap between molecular specificity and quantitative bioimaging. Biomedical Optics Express 8, 5688–5697 (2017).
Shin, S., Kim, D., Kim, K. & Park, Y. Super-resolution three-dimensional fluorescence and optical diffraction tomography of live cells using structured illumination generated by a digital micromirror device. arXiv preprint arXiv:1801.00854 (2018).
Chowdhury, S., Eldridge, W. J., Wax, A. & Izatt, J. A. Structured illumination multimodal 3D-resolved quantitative phase and fluorescence sub-diffraction microscopy. Biomedical Optics Express 8, 2496–2518 (2017).
Schürmann, M. et al. Three‐dimensional correlative single‐cell imaging utilizing fluorescence and refractive index tomography. Journal of biophotonics (2017).
Franchi, G. G., Bellani, L., Nepi, M. & Pacini, E. Types of carbohydrate reserves in pollen: Localization, systematic distribution and ecophysiological significance. Flora 191, 143–159 (1996).
Park, J., Yu, H., Park, J. H. & Park, Y. LCD panel characterization by measuring full Jones matrix of individual pixels using polarization-sensitive digital holographic microscopy. Opt Express 22, 24304–24311, https://doi.org/10.1364/OE.22.024304 (2014).
Schwendemann, A. B. et al. Aerodynamics of saccate pollen and its implications for wind pollination. Am J Bot 94, 1371–1381, https://doi.org/10.3732/ajb.94.8.1371 (2007).
Grega, L. et al. Aerodynamic Characteristics of Saccate Pollen Grains. Int J Plant Sci 174, 499–510, https://doi.org/10.1086/668694 (2013).
Leslie, A. B. Flotation preferentially selects saccate pollen during conifer pollination. New Phytol 188, 273–279, https://doi.org/10.1111/j.1469-8137.2010.03356.x (2010).
Edlund, A. F., Swanson, R. & Preuss, D. Pollen and stigma structure and function: the role of diversity in pollination. The Plant Cell 16, S84–S97 (2004).
Chae, K. & Lord, E. M. Pollen tube growth and guidance: roles of small, secreted proteins. Annals of botany 108, 627–636 (2011).
Subiza, J. et al. Allergenic Pollen and Pollinosis in Madrid. J Allergy Clin Immun 96, 15–23 (1995).
Mandal, J., Chakraborty, P., Roy, I., Chatterjee, S. & Gupta-Bhattacharya, S. Prevalence of allergenic pollen grains in the aerosol of the city of Calcutta, India: a two year study. Aerobiologia 24, 151–164 (2008).
Ghufran, M. A., Hamid, N., Ali, A. & Ali, S. M. Prevalence of Allergenic Pollen Grains in the City of Islamabad, Pakistan and Its Impact on Human Health. Pak J Bot 45, 1387–1390 (2013).
bin Mat Yunus, W. M. & bin Abdul Rahman, A. Refractive index of solutions at high concentrations. Applied optics 27, 3341–3343 (1988).
Lush, W. M., Grieser, F. & Wolters-Arts, M. Directional Guidance of Nicotiana alataPollen Tubes in Vitro and on the Stigma. Plant Physiology 118, 733–741 (1998).
Runions, C. J., Rensing, K. H., Takaso, T. & Owens, J. N. Pollination of Picea orientalis (Pinaceae): saccus morphology governs pollen buoyancy. American Journal of Botany 86, 190–197 (1999).
Bykowska, J. & Klimko, M. Pollen morphology of Pinus mugo Turra X Pinus sylvestris L. hybrids and parental species in an experimental culture. Acta Biologica Cracoviensia s. Botanica 57, 149–160 (2015).
Shin, S., Kim, K., Yoon, J. & Park, Y. Active illumination using a digital micromirror device for quantitative phase imaging. Optics Letters 40, 5407–5410 (2015).
Lee, K., Kim, K., Kim, G., Shin, S. & Park, Y. Time-multiplexed structured illumination using a DMD for optical diffraction tomography. Optics Letters 42, 999–1002, https://doi.org/10.1364/OL.42.000999 (2017).
Lim, J. et al. Comparative study of iterative reconstruction algorithms for missing cone problems in optical diffraction tomography. Opt. Express 23, 16933–16948, https://doi.org/10.1364/Oe.23.016933 (2015).
Acknowledgements
This work was supported by KAIST, BK21+ program, Tomocube Inc., and National Research Foundation of Korea (2015R1A3A2066550, 2017M3C1A3013923, 2014K1A3A1A09063027).
Author information
Authors and Affiliations
Contributions
G.K. and Y.K.P. designed the study. G.K. performed the experiments and analyzed data. S.Y.L. and S.S. provide analysis methods and analyzed data. All authors wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
Mr. Shin and Prof. Park have financial interests in Tomocube Inc., a company that commercializes ODT and QPI instruments and is one of the sponsors of the work.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, G., Lee, S., Shin, S. et al. Three-dimensional label-free imaging and analysis of Pinus pollen grains using optical diffraction tomography. Sci Rep 8, 1782 (2018). https://doi.org/10.1038/s41598-018-20113-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-20113-w
This article is cited by
-
Towards standardisation of automatic pollen and fungal spore monitoring: best practises and guidelines
Aerobiologia (2022)
-
Bipolar pollen germination in blue spruce (Picea pungens)
Protoplasma (2019)
-
Measurements of three-dimensional refractive index tomography and membrane deformability of live erythrocytes from Pelophylax nigromaculatus
Scientific Reports (2018)
-
Super-resolution three-dimensional fluorescence and optical diffraction tomography of live cells using structured illumination generated by a digital micromirror device
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.