Abstract
Nitrate is not only an important nutrient but also a signaling molecule for plants. A few of key molecular components involved in primary nitrate responses have been identified mainly by forward and reverse genetics as well as systems biology, however, many underlining mechanisms of nitrate regulation remain unclear. In this study, we show that the expression of NRT1.1, which encodes a nitrate sensor and transporter (also known as CHL1 and NPF6.3), is modulated by NIN-like protein 7 (NLP7). Genetic and molecular analyses indicate that NLP7 works upstream of NRT1.1 in nitrate regulation when NH4+ is present, while in absence of NH4+, it functions in nitrate signaling independently of NRT1.1. Ectopic expression of NRT1.1 in nlp7 resulted in partial or complete restoration of nitrate signaling (expression from nitrate-regulated promoter NRP), nitrate content and nitrate reductase activity in the transgenic lines. Transcriptome analysis revealed that four nitrogen-related clusters including amino acid synthesis-related genes and members of NRT1/PTR family were modulated by both NLP7 and NRT1.1. In addition, ChIP and EMSA assays results indicated that NLP7 may bind to specific regions of the NRT1.1 promoter. Thus, NLP7 acts as an important factor in nitrate signaling via regulating NRT1.1 under NH4+ conditions.
Similar content being viewed by others
Introduction
Nitrogen is a vital macronutrient for plant growth and development. Plants have evolved a range of mechanisms to adapt to imbalanced nitrogen conditions. In agricultural systems, high-yield of crops relies on application of nitrogen fertilizers. But a large part of nitrogen deposited in the soil can’t be absorbed by plants and is lost to the environment, resulting in severe environmental and ecological pollution1,2. Improving the nitrogen use efficiency (NUE) of crops is the key to solve these problems. Studying the genes and mechanisms involved in regulating nitrogen uptake and assimilation can be a prerequisite for improving NUE of crops, therefore it is of great importance for sustaining agriculture. Nitrate and ammonium are the main nitrogen forms used by plants and most crops, like maize and wheat, take up nitrate as the major nitrogen source3. In addition to its nutrient role, nitrate acts also as a signaling molecule for plants. It regulates the expression levels of many genes, including genes directly involved in nitrate assimilation, namely NIAs, NiR, and some NRTs (short-term processes)4,5,6,7,8,9,10. It is also involved in many adaptive responses of plants11, such as root development and architecture, seed dormancy, flowering time, circadian system, leaf development, stomatal movements, and auxin transportation (long-term processes)12,13,14,15,16,17,18,19,20,21.
Nitrate is taken up into plants by nitrate transporters and high affinity and low affinity nitrate uptake systems have been identified3. Four gene families have been identified that encode nitrate transporters in Arabidopsis: NRT1/PTR (NPF, 53 members), NRT2 (7 members), CLC (7 members), and SLAC1/SLAH (5 members)22. Among these families, NRT1/PTR belongs to the low affinity transport system, and NRT2 belongs to the high affinity transport system22. NRT1.1 (also known as CHL1 and NPF6.3), which belongs to NRT/PTR family, functions in nitrate uptake as both high affinity and low affinity transporter23.
In addition to the nitrate transport systems, genes involved in nitrate signaling have also been identified. Most of these genes were found to function in root architecture or primary nitrate responses14,24,25. The genes functioning in root architecture include the ANR1, the first molecular component isolated by classic molecular genetics approach26, is a MADS box transcription factor and positively regulates lateral root branching under sufficient nitrate condition27,28,29. miR393/AFB3 and NAC4 have been demonstrated to regulate the root system architecture in nitrate signaling using systems approach26,30,31,32. The split-root assays indicated that TCP20 was involved in systemic nitrate signaling for root foraging33. Recently, TCP20 was found to regulate root meristem growth under nitrogen starvation and to interact with NLP6&734. HHO1 and HRS1 are two nitrate-responsive transcription factors isolated by genome-wide analyses35. They function in the repression of primary root growth under both phosphate starvation and nitrate supply conditions.
During last several years, the nitrate regulatory factors involved in the primary nitrate response have been identified. NRT1.1, in addition to its transport function, was identified to work as a nitrate sensor24,36,37,38,39. The study on the crystal structure of NRT1.1 has demonstrated that Thr101 phosphorylation is essential for nitrate transport rate and provides further insights into its transport mechanisms40,41,42. CIPK8 and CIPK23 which belong to CBL-interacting protein kinase family36,43 are important players in responding to primary nitrate. CIPK8 works positively while CIPK23 functions negatively in nitrate regulation36,43. The expression of both CIPK8 and CIPK23 is regulated by NRT1.136,43. Recently, NRG2 which is an essential nitrate regulatory gene was isolated by forward genetics screen44. NRG2 acts as a positive nitrate regulatory factor and modulates NRT1.1 expression and can interact with NLP744. Additionally, several transcription factors were identified to be involved in primary nitrate response, for example, NLP6, NLP7, LBD37/38/39, TGA1, TGA4, and SPL910,45,46,47,48,49. NLP7 is NIN-like protein and acts as an important nitrate positive regulator. NLP7 was isolated by reverse genetics strategy and the nlp7 mutants exhibit a nitrogen-starved phenotype46. The nitrate condition can affect the NLP7′s nuclear retention50. Previous studies have demonstrated that the nitrate response cis-element NRE can be bound by NLPs46,47 and contain a DNA-binding domain RWP-PK and protein-protein interaction domains typeI/II Phox and Ben1p (PB1)34,51. ChIP-chip assays showed that NLP7 could bind 851 genes containing NRT1.1, NRT2.1, LBD37/3850. In addition, overexpression of NLP7 can increase plant biomass, nitrogen uptake, total nitrogen content, and expression levels of genes involved in nitrogen assimilation and signaling52. Moreover, NLP7 can control plant root growth under both N-limited and N-rich conditions46,52. NLP6 also functions positively in nitrate regulation, is retained in the nucleus in nitrate-treated plants34 and can activate the expression of nitrate-responsive genes47. LBD37/38/39 are negative regulators in nitrate signaling. They are involved in primary nitrate response and can affect nitrogen status, growth, and nitrogen-dependent shoot branching26,45. TGA1, TGA4, and SPL9 were isolated by systems approach10,48,49. TGA1 and TGA4 belong to bZIP transcription factor family and TGA1 can bind to the promoters of NRT2.1 and NRT2.248. SPL9 is demonstrated to be a nitrate regulatory hub10.
Although these nitrate regulatory genes have been identified, our understanding of the nitrate regulatory gene network is still incomplete. For example, both NLP7 and NRT1.1 play essential roles in regulating nitrate signaling and ChIP-chip assay showed that NLP7 might bind NRT1.1, however, their relationship and underlining mechanism remain unclear. In this paper, we investigated the relationship between NRT1.1 and NLP7 in nitrate regulation. Our analyses reveal that NLP7 acts as a positive regulatory factor upstream of NRT1.1 when NH4+ is present and modulates the nitrate signaling function of NRT1.1. NLP7 might function in another pathway to regulate nitrate signaling independent of NRT1.1. In addition, transcriptome data showed that four GO terms related to nitrogen were regulated by NRT1.1 as well as NLP7 in nitrate signaling, providing more evidence to support our above conclusion. Furthermore, the ChIP and EMSA assays indicated that NLP7 could bind to specific regions of the NRT1.1 promoter. Our findings not only further elucidate the relationship between NRT1.1 and NLP7, but also provide insights into the network of the nitrate regulatory genes.
Results
The expression levels of NRT1.1 are modulated by NLP7
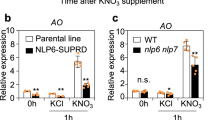
To study the relationship between NLP7 and NRT1.1, the expression levels of NRT1.1 was detected firstly under potassium nitrate and ammonium nitrate conditions. Figure 1a showed that the transcript levels of NRT1.1 in the nlp7 mutants (nlp7-1, nlp7-2, and nlp7-4) were not notably changed under potassium nitrate condition, but was significantly decreased in mutant plants under ammonium nitrate condition (Fig. 1a). This indicates that the expression levels of NRT1.1 can be modulated by NLP7 in the presence of NH4+. In order to test if NLP7 is regulated by NRT1.1, we tested NLP7 expression in chl1-5 and chl1-13 mutants (the nrt1.1 mutants) in potassium nitrate and ammonium nitrate mediums. The expression of NLP7 was not changed in the nrt1.1 mutants (Fig. 1b). This result indicates that NRT1.1 may not regulate the expression of NLP7. We also tested the NRT1.1 expression response to nitrate in WT and the nlp7 mutants. qPCR results showed that the induction of NRT1.1 by nitrate was notably decreased in the nlp7 mutants, indicating that NLP7 affects the response of NRT1.1 to nitrate (Fig. 1c).
The transcript levels of NRT1.1 in the nlp7 plants were reduced when NH4+ is present. (a) Relative expression of NRT1.1 in WT and the nlp7 plants. Seedlings were planted on medium containing 10 mM potassium nitrate or 10 mM ammonium nitrate for 7d for testing NRT1.1′s expression. Error bars mean ± SD of four biological replicates. The significant difference (p < 0.05, t test) is indicated by different letters. (b) Relative expression of NLP7 in WT and the chl1 mutants. The growth conditions of plants are the same as (a). Error bars mean ± SD of four biological replicates. (c) Nitrate response of NRT1.1 in WT and the nlp7 mutants. Seedlings were planted on medium with 2.5 mM ammonium succinate for 7d, then treated with 10 mM KNO3 or KCl for 2 h followed by collecting samples for detecting nitrate induction of NRT1.1. Error bars mean ± SD of four biological replicates. The significant difference (p < 0.05, t test) is indicated by different letters.
NLP7 and NRT1.1 may participate in nitrate signaling in the same pathway
To elucidate the relationship between NLP7 and NRT1.1, the single mutants: nlp7-4 and chl1-13 which contain the nitrate-responsive NRP-YFP transgene, both of which were isolated by our mutant screens described previously39,44 were crossed to obtain the double mutant chl1-13 nlp7-4. The YFP signal levels in the roots of WT, two single mutants, and double mutants were detected. The results showed that the YFP signal of the two single mutants under the ammonium nitrate condition was much weaker than that of WT while the double mutant chl1-13 nlp7-4 exhibited significantly weaker YFP signal than the nlp7-4 mutant and similar to the chl1-13 mutant (Fig. 2a,b). At the same time, we detected the YFP signal of the plants under the potassium nitrate condition without NH4+. The results showed that the YFP levels of the double mutant chl1-13 nlp7-4 were much lower than those of WT and the chl1-13 mutant, and similar to those of the nlp7-4 mutant (Supplementary Fig. S1). Remarkably, the YFP signal of the chl1-13 mutant was mildly weaker than WT. This result is consistent with previous studies demonstrating that NRT1.1 is a key player in the nitrate regulation when NH4+ is present and functions poorly when NH4+ is absent39,44. Because we don’t observe any additive effects in the double mutant, these evidences indicate that NLP7 and NRT1.1 may participate in nitrate signaling in the same pathway.
NLP7 functions in the same nitrate regulation pathway as NRT1.1. (a) Root YFP observation of WT, chl1-13, nlp7-4, and chl1-13 nlp7-4 plants. Fluorescence and light images of seedlings which grown on the medium with 10 mM ammonium nitrate for 5d were captured using a fluorescence microscope. (b) Quantification of root YFP signal levels of WT, chl1-13, nlp7-4, and chl1-13 nlp7-4 plants. The same conditions for plants growth were used as (a). Error bars mean ± SD of sixty biological replicates. The significant difference (p < 0.05, t test) is indicated by different letters. (c) Nitrate content in WT, chl1, nlp7, and chl1-13 nlp7-4 plants. Seedlings grown on the medium with 10 mM ammonium nitrate for 7d were sampled for nitrate content detection. Error bars mean ± SD of four biological replicates. The significant difference (p < 0.05, t test) is indicated by different letters. FW, fresh weight. (d) Nitrate reductase activity in WT, chl1, nlp7, and chl1-13 nlp7-4 plants. The same growth conditions were used as (c). Error bars mean ± SD of four biological replicates. The significant difference (p < 0.05, t test) is indicated by different letters. FW, fresh weight. (e) and (f) Nitrate accumulation in WT, nlp7, chl1-13 and chl1-13 nlp7-4 plants. Seedlings were planted on the medium with 2.5 mM ammonium succinate for 7 d followed by the treatments with different KNO3 concentrations for 2 h (e) or with 5 mM KNO3 for different times (f). The whole plants were used for nitrate content test. Error bars mean ± SD of four biological replicates. The significant difference (p < 0.05, t test) is indicated by different letters.
In order to find physiological evidences for the relationship between NLP7 and NRT1.1, we investigated the nitrate content and nitrate reductase activity in plants. Figure 2c showed higher nitrate content in the nlp7 mutants while lower in the chl1 plants than that in WT (Fig. 2c). The double mutant chl1-13 nlp7-4 exhibited significant lower nitrate content than that in WT and the nlp7 plants, and similar to that in the chl1 mutants (Fig. 2c). The nitrate reductase activity of the two single mutants was much lower than that of WT, and the double mutant chl1-13 nlp7-4 showed similar nitrate reductase activity to the chl1-13 mutant (Fig. 2d). In addition, we analyzed the nitrate content in whole seedlings after treatments with different nitrate concentrations (0.25 mM to 20 mM) for 2 h and with 5 mM KNO3 for different times (0 h to 4 h) under the ammonium succinate condition. The results showed that the nlp7 mutants displayed the same nitrate content as WT, but lower in the chl1-13 mutant than in WT under both conditions (Fig. 2e,f). The double mutant chl1-13 nlp7-4 showed the same nitrate content as the chl1-13 mutant (Fig. 2e,f). Because NRT1.1 is epistatic to NLP7 in these experiments, these results indicate that NLP7 and NRT1.1 may be involved in the same pathway and NLP7 may regulate nitrate signaling upstream of NRT1.1.
NRT1.1 can restore the phenotypes of nlp7-4
In order to provide more evidence to support the conclusion that NLP7 regulates nitrate signaling upstream of NRT1.1, we overexpressed NRT1.1 in the nlp7-4 mutant (Supplementary Fig. S2a). The transgenic lines NRT1.1/nlp7-4 exhibited higher expression of NRT1.1 than that in the WT and nlp7 plants (Supplementary Fig. S2b). The YFP signal (from the NRP-YFP transgene) in the NRT1.1/nlp7-4 plants grown under the ammonium nitrate condition was much stronger than that of the nlp7-4 mutant, and weaker than that of WT (Fig. 3a). Quantifying the fluorescence intensity of the NRT1.1/nlp7-4 plants showed that the YFP levels in the NRT1.1/nlp7-4 plants were 70% higher than that in the nlp7-4 mutant and reached 71% of that in WT (Fig. 3b). Then we tested the nitrate content and nitrate reductase activity in the NRT1.1/nlp7-4 plants. As shown in Fig. 3c, the nitrate content in the NRT1.1/nlp7-4 plants was lower than that in the nlp7-4 mutant, and similar to that in WT (Fig. 3c). The nitrate reductase activity in the NRT1.1/nlp7-4 plants was higher than that in the nlp7-4 mutant, and reached the levels of WT (Fig. 3d).
NRT1.1 can restore the phenotypes of nlp7-4. (a) Root YFP observation of WT, nlp7-4, and NRT1.1/nlp7-4 plants. Fluorescence and light images of seedlings which grown on the medium with 10 mM ammonium nitrate for 5d were captured using a fluorescence microscope. (b) Quantification of root YFP signal levels of WT, nlp7-4, and NRT1.1/nlp7-4 plants. The same conditions for plants growth were used as (a). Error bars mean ± SD of sixty biological replicates. The significant difference (p < 0.05, t test) is indicated by different letters. (c) Nitrate content in WT, nlp7-4, and NRT1.1/nlp7-4 plants. The 7-d-old seedlings on the medium with 10 mM ammonium nitrate were used for nitrate content detection. Error bars mean ± SD of four biological replicates. The significant difference (p < 0.05, t test) is indicated by different letters. FW, fresh weight. (d) Nitrate reductase activity in WT, nlp7-4, and NRT1.1/nlp7-4 plants. The growth conditions are the same as (c). Error bars mean ± SD of four biological replicates. The significant difference (p < 0.05, t test) is indicated by different letters. FW, fresh weight. (e) The induction of endogenous genes after nitrate treatment in WT, nlp7-4, and NRT1.1/nlp7-4 plants. Seedlings grown on 2.5 mM ammonium succinate medium for 7d were treated with 10 mM KNO3 or KCl for 2 h. Roots were used for detecting the expression of nitrate inducible genes by qPCR. Error bars mean ± SD of four biological replicates. The significant difference (p < 0.05, t test) is indicated by different letters.
In addition, we examined the transcript levels of NIA1, NiR, NRT2.1, HHO1, and HRS1 which are responsive to nitrate in a short time. We found that the transcript levels of the nitrate inducible genes were significantly higher in the NRT1.1/nlp7-4 plants than that in the nlp7-4 mutant and completely or partially restored to the WT levels (Fig. 3e, Supplementary Fig. S3a), and there was no significant difference between WT and NRT1.1/nlp7-4 plants before nitrate treatment (except for NIA1) (Supplementary Fig. S3b). Taken together, the phenotypes of NRT1.1/nlp7-4 plants were completely or partially recovered to that of WT, further indicating that NLP7 functions upstream of NRT1.1. Since NRT1.1 functions in nitrate signaling pathway dependently on NH4+ while NLP7 is NH4+ independent44, NLP7 may also function in another nitrate signaling pathway independent on NRT1.1 in the absence of NH4+.
Transcriptome analysis of nlp7 and chl1 mutants
To obtain more comprehensive data on the relationship between NLP7 and NRT1.1, we analyzed our transcriptome data of nitrate responses in the WT, nlp7-4 and chl1-13 plants. The experimental process, data quality, and the method of data filtering were described previously44.
Firstly, we compared the expression of nitrate inducible genes in the WT and the chl1-13 mutant roots after nitrate treatment. The result showed that 152 genes could respond to nitrate in both WT and the chl1-13 mutant (i.e. were unaffected by the chl1-13 mutation, Supplementary Fig. S4a, Supplementary Table S1). In contrast, 438 genes (212 suppressed and 226 induced) lost their nitrate response in the chl1-13 mutant. 47 genes (16 suppressed and 31 induced) showed a gain of nitrate response in the chl1-13 mutant compared with WT. In summary, 485 genes were differentially expressed showed altered nitrate responses in the chl1-13 mutant compared to WT.
We also compared the gene expression between WT and the nlp7-4 mutant in response to nitrate. The results showed that there were 253 genes could respond to nitrate in both WT and the nlp7-4 mutant (Supplementary Fig. S4b, Supplementary Dataset 2). The expression of 337 genes (158 down-regulated and 179 up-regulated) was responsive to nitrate in WT, but not in the nlp7-4 mutant. 191 genes (102 down-regulated and 89 up-regulated) were responsive to nitrate exclusively in the nlp7-4 mutant, but not in WT. In total, 528 genes were differentially expressed in the nlp7-4 mutant.
We compared the 528 differentially expressed genes in the nlp7-4 mutant and 485 differentially expressed genes in the chl1-13 mutant and found 342 genes (64.8% of 528 and 70.5% of 485) were regulated by both NLP7 and NRT1.1 (Fig. 4, Supplementary Dataset 3) while 143 and 186 genes were regulated only by NRT1.1 and NLP7, respectively (Fig. 4, Supplementary Dataset 4). Gene Ontology (GO) analysis of 342 shared genes showed four clusters related to nitrogen, including cellular amino acid biosynthetic process, cellular amino acid metabolic process, response to nitrogen compound, and nitrogen compound transport (Table 1). In these four clusters, genes involved in amino acid synthesis and nitrate transport were enriched (Supplementary Dataset 5). These data indicate extensive overlap in the target genes modulated by NLP7 and NRT1.1 and provide additional support that both NLP7 and NRT1.1 work in the same pathway in nitrate signaling.
The 186 and 143 genes exclusively regulated by NLP7 and NRT1.1, respectively (Supplementary Dataset 4) were further investigated by GO analysis. Among the 186 genes regulated by NLP7, there are two nitrogen-related clusters (response to nitrogen compound and nitrogen compound transport) (Table 2), including genes involved in amino acid transport and nitrate transport (Table 3). Among the 143 genes modulated by NRT1.1, three nitrogen-related clusters were found (response to nitrogen compound, cellular response to nitrogen compound, and response to nitrate) (Table 4). In the three clusters, gene involved in nitrate transport was enriched (Table 5). These data imply that NLP7 may function in nitrate signaling pathway independent of NRT1.1.
NLP7 can bind to the promoter of NRT1.1
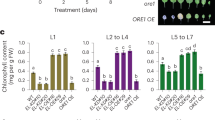
Previous ChIP-chip test indicated that NRT1.1 was one of the NLP7 targets50. To investigate further the binding of NLP7 to NRT1.1, we carried out a ChIP assay using the transgenic plants expressing pNLP7:NLP7-GFP fusion protein in nlp7-1 mutant. The ChIP-qPCR assay was performed using a series of primers designed to cover the NRT1.1 promoter region (Fig. 5a) and a sample without antibody was used as a negative control. The results showed that NLP7-specific enrichments were much higher in two regions of NRT1.1 promoter in the sample than in negative control, and the two regions were −1537 to −1334bp, and −686 to −465bp, respectively (Fig. 5b). ChIP-PCR results confirmed the binding of NLP7 to the two regions (Fig. 5c, Supplementary Fig. 5). These results indicate that NLP7 can bind to the promoter of NRT1.1 in vivo. We also did the ChIP-qPCR and ChIP-PCR assays with the seedlings expressing p35S::NLP7-GFP fusion protein in nlp7-1 plant. The results also showed that NLP7 could bind to NRT1.1 promoter in vivo (Supplementary Fig. S6b,c). To provide more evidences to the binding of NLP7 to NRT1.1 promoter, electrophoretic mobility shift assay (EMSA) was performed. The result showed that NLP7 could bind to the Q4-3 and Q4-4 regions of NRT1.1 promoter in vitro (Fig. 5d), implying that NLP7 may bind to the specific regions of NRT1.1 promoter. However, no activity was found for the binding of NLP7 to the Q8 region of NRT1.1 promoter (Supplementary Fig. S7).
NLP7 binds to the promoter of NRT1.1. (a) Segmentation schematic diagram of the promoter of NRT1.1. Q1-Q10 represent different fragments of NRT1.1 promoter. (b) ChIP-qPCR assay shows that NLP7 binds to NRT1.1 promoter in vivo. ChIP assays were performed with nlp7-1 seedlings expressing pNLP7:NLP7-GFP fusion protein grown on 10 mM ammonium nitrate medium for 7 d. A sample without antibody was used as the negative control for the ChIP-qPCR (n = 3). (c) ChIP-PCR assay shows that NLP7 binds to NRT1.1 promoter in vivo. ChIP assays were performed with the same seedlings as in (b). Input was used as positive control and a sample without antibody was used as the negative control. Q4 and Q8 are the promoter sites of NRT1.1 showing the binding activity with NLP7 and images come from the checking gel shown in the Supplementary Fig. 5. (d) EMSA assay shows that NLP7 binds to NRT1.1 promoter in vitro. Red arrow indicated the positions of protein-DNA complexes caused by binding of NLP7 to DNA probes. The NRE probe was used as a positive control. The DNA probes were listed in Supplementary Dataset 6.
Discussion
NRT1.1 have been characterized to be a nitrate transceptor in previous reports36 and can regulate the expression of nitrate regulatory genes CIPK8 and CIPK2336,43. Moreover, NRT1.1 can be phosphorylated by CIPK23 at amino acid Thr-101 under low nitrate concentration36. NLP7 is another key nitrate regulator46,47,50. As a transcription factor, it can bind to the nitrate response cis-elements (NREs)47. ChIP-chip and ChIP-qPCR assays have illustrated that NLP7 binds to many genes functioning in the nitrate regulation and assimilation, for instance NRT2.1, NRT2.2, and LBD3747,50. It has been reported that NLP7 could bind NRT1.1, but it is not known if NLP7 binds to the promoter of and regulates NRT1.1.
In this paper, our results revealed that the transcript levels of NRT1.1 were down-regulated in the nlp7 plants when NH4+ is present. The nitrate induction of NRT1.1 was reduced as well in the mutants (Fig. 1). These data indicate that the expression and induction of NRT1.1 can be modulated by NLP7 in the presence of NH4+. Our genetic analysis showed that the nitrate-responsive YFP signal, nitrate content, nitrate reductase activity, and nitrate uptake in the double mutant chl1-13 nlp7-4 were strongly reduced than those in WT and close to those in chl1-13 mutant (Fig. 2). These results indicate that NLP7 participates in nitrate signaling in the same pathway as NRT1.1 and NLP7 may work upstream of NRT1.1. We also overexpressed the NRT1.1 in the nlp7-4 mutant and found that the YFP signal, nitrate content, nitrate reductase activity, and the induction levels of the endogenous genes after nitrate treatment in the NRT1.1/nlp7-4 plants were completely or partially recovered to WT levels (Fig. 3). Even though the levels of NRT1.1 mRNA were higher in the transgenic plants, one was similar to WT levels and both lines gave similar results. These results demonstrate that NLP7 regulates nitrate signaling upstream of NRT1.1.
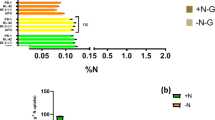
We found that the nitrate content was significant higher in the nlp7 mutants than in WT (Fig. 2c), consistent with previous studies46,52. The reason for the higher nitrate content in the mutants may come from higher nitrate uptake and/or lower nitrate reduction. We tested the nitrate accumulation and found no significant difference between the nlp7 mutants and WT (Fig. 2e,f), similar to the report that nitrate uptake activity of the nlp7-1 mutant at 5 mM 15NO3− external concentration was the same as that of WT52. We also tested the total nitrogen content and similar levels were found in the nlp7 mutants and WT, indicating that the nitrate import was not changed in the nlp7 plants (Supplementary Fig. S8). However, the nitrate reductase activity in the nlp7 plants was much lower than that in WT (Fig. 2d). These findings indicate that the higher nitrate content in the nlp7 mutants is caused by the reduced nitrate assimilation, rather than altered nitrate uptake. In addition, it has been reported that the nitrate content is lower and the nitrate uptake ability is reduced in chl1 mutants. Interestingly, our data revealed that the nitrate reductase activity was notably depressed in the chl1 mutants, indicating that the reduced nitrate content in the chl1 mutants may result from both decreased nitrate uptake and reduction.
NRT1.1 is an essential component of nitrate uptake and nitrate signaling, respectively36. But its function in nitrate signaling has been found to work mainly when NH4+ is present39,44. In this paper, we found that the expression of NRT1.1 could be modulated by NLP7 when NH4+ is present (Fig. 1a) and the nitrate uptake of nlp7 mutants was not affected (Fig. 2e,f, Supplementary Fig. S8). Combining with our results that NLP7 works upstream of NRT1.1, these findings indicate that NLP7 regulates the nitrate signaling function but does not affect the transport function of NRT1.1. NRG2, another important nitrate regulatory gene, has been reported to modulate the expression of NRT1.1 in both presence and absence of NH4+, indicating that NRG2 can affect both nitrate transport and signaling functions of NRT1.144. These findings demonstrate that both NRG2 and NLP7 can regulate NRT1.1 but in different ways. NRT1.1′s functioning as a regulatory gene is NH4+-dependent, while NLP7 is NH4+-independent and NRT1.1 can restore the phenotype of the nlp7-4 mutant (Fig. 3). Accordingly, these findings suggest that NLP7 functions in nitrate signaling independent of NRT1.1 in the absence of NH4+.
Our RNA-sequencing analysis showed that many genes were regulated by both NLP7 and NRT1.1 in response to nitrate (Fig. 4). GO analysis of these genes revealed that there were four nitrogen-related clusters, including cellular amino acid biosynthetic process, cellular amino acid metabolic process, response to nitrogen compound, and nitrogen compound transport, modulated by both NLP7 and NRT1.1 (Table 1). These data provide further evidences that NLP7 modulates nitrate regulation in the same pathway as NRT1.1 under these conditions.
Since NLP7 is a transcription factor and ChIP-chip test has shown that it could bind NRT1.150, we performed ChIP-qPCR, ChIP-PCR, and EMSA to investigate the binding activity of NLP7 to the promoter of NRT1.1. These results indicate that NLP7 can bind to the promoter of NRT1.1 (Fig. 5, Supplementary Figs S5, S6). ChIP assays showed that NLP7 could bind the Q4 and Q8 region of NRT1.1 promoter (Fig. 5b,c, Supplementary Figs S5, S6), while EMSA assay revealed that NLP7 could bind to Q4 region, but not Q8 region of NRT1.1 promoter (Fig. 5d, Supplementary Fig. S7), implicating that NLP7 may bind Q8 region indirectly.
These results were obtained from the conditions with both ammonium and nitrate. Since ammonium is crucial for the regulation of NRT1.1 by NLP7, the presence of NH4+ might be also important for the binding of NLP7 to the promoter of NRT1.1. Without NH4+, the NLP7 protein may not bind to the promoter of NRT1.1, resulting in the lost regulation function of NLP7 to NRT1.1.
Taken together, in the primary nitrate response, NLP7 regulates nitrate signaling upstream of NRT1.1 in the presence of NH4+ and may modulate an NRT1.1-independent pathway in the absence of NH4+ in nitrate signaling. Moreover, NLP7 may regulate NRT1.1 via binding to the promoter of NRT1.1. Therefore we propose the working model as shown in Fig. 6. NLP7 plays a key role through regulating NRT1.1 in the presence of NH4+ while through an NRT1.1-independent pathway in nitrate signaling when NH4+ is absent. NLP7 can interact with NRG2 and NRG2 works upstream of NRT1.1 when NH4+ is present and/or absent. NRT1.1 regulates the transcript levels of CIPK8 and CIPK23 and can be phosphorylated by CIPK23 to switch its nitrate affinity. The genes regulated by NLP7 in the nitrate signaling pathway without NH4+ need to be investigated. The characterization of the regulation relationship of NLP7 and NRT1.1 has provided insights into the regulatory mechanisms of the genes functioning in nitrate regulation.
Proposed model for the nitrate regulatory genes in the nitrate signaling. NLP7 regulates nitrate signaling upstream of NRT1.1 when NH4+ is present and may regulate an NRT1.1-independent pathway in the absence of NH4+ in nitrate signaling. NLP7 interacts with NRG2 and NRG2 works upstream of NRT1.1 when NH4+ is present and/or absent. NRT1.1 regulates CIPK8 and CIPK23 at the transcript level. CIPK23 can phosphorylate NRT1.1. These genes are all involved in regulating the expression of nitrate responsive genes.
Methods
Plant materials
Arabidopsis thaliana Col-0 was used as the ecotype of wild-type in this research Homozygous transgenic seeds with the NRP-YFP construct39 and carrying p35S::NLP7-GFP or pNLP7:NLP7-GFP50, the mutant lines chl1-536, chl1-13 containing the NRP-YFP construct39, nlp7-1, nlp7-246, nlp7-4 containing the NRP-YFP construct44 were described previously. Transgenic plants carrying p35S::NRT1.1 cDNA-FLAG were obtained by floral dipping53 of nlp7-4 mutants.
Growth and treatment conditions
To test gene expression in plants, seedlings grown on 10 mM ammonium nitrate or potassium nitrate medium (aseptic hydroponics) for 7 days were collected for qPCR analysis. To test YFP signal of the plants containing the NRP-YFP construct, seedlings were cultured on mediums containing 10 mM ammonium nitrate or potassium nitrate for 5 days. The fluorescence was observed under a fluorescence microscope (Nikon Eclipse Ti-S) and ImageJ was used to quantify the YFP signal levels of plants. For the nitrate uptake assays, seedlings were grown on medium containing 2.5 mM ammonium succinate (aseptic hydroponics) for 7 days followed by treatments with different nitrate concentrations (0.25 mM to 20 mM) for 2 h or with 5 mM KNO3 for different times (0 h to 4 h). For qPCR analysis of nitrate responsive genes, seedlings were cultured as described44. For nitrate content test and nitrate reductase activity assays, seedlings were cultured on medium containing 10 mM ammonium nitrate (aseptic hydroponics) for 7 days.
qPCR analysis
Total RNA was extracted from plants using an ultrapure RNA kit (CWBIO). cDNA samples were carried out using a EasyScript One-Step gDNA Removal and cDNA Synthesis SuperMix Kit (TransGen Biotech). The UltraSYBR Green Mixture qPCR kit (CWBIO) was used in the qPCR reaction. The real-time PCR was carried out using VIIA7 (Applied Biosystems). We used TUB2 (At5g62690) as the internal control.
Nitrate content, nitrate reductase (NR) activity and total nitrogen content assays
Nitrate in the plants was extracted with the method as previously described44 and the samples were measured with the hydrazine-sulfate method7,39 using the machine AutoAnalyzer 3 (SEAL). Nitrate reductase activity was measured mainly based on the method described previously54,55. Briefly, about 0.1 g fresh samples were frozen by liquid nitrogen and broken into powder by a RETCH MM4000. 1 mL of extraction buffer (25 mM phosphate buffer, pH7.5, 5 mM cysteine, 5 mM EDTA) was added into the samples and then centrifuged at 4 °C and 4000 g for 5 min. 100 µL supernatant was transferred into a new 1.5-mL tube, then 375 µL 0.1 M KNO3-phosphate buffer and 125 µL 2 mg/mL NADH2 were added. The reactions were incubated at 25 °C for 30 min. After 250 µL of 1% sulfanilic acid and 250 µL of 0.2% α-naphthylamine were added and incubated at room temperature for 20 min, the OD540 value was measured. 100 µL of extraction buffer was used as a control. The following equation was used to calculate nitrate reductase activity: A = C × V1/V2/W/t (A, nitrate reductase activity; C, nitrite amount deduced from the regression equation; V1, the volume of extraction reagent; V2, volume of supernatant added into the reaction; W, sample weight; t, reaction time). The concentrations of NO2− used in standard curve were at the amount from 0.00435 to 0.0435µmol and regression equation was deduced from the basis of standard curve. Total nitrogen content was tested with the machine AutoAnalyzer 3 (SEAL). Weighed samples (~0.1 g DW) were digested using H2SO4-H2O2 and the samples were measured using the machine AutoAnalyzer 3 (SEAL).
Transcriptome analysis
The transcriptome analysis was performed using RNA sequencing technology. Methods for RNA sequencing assay and data filtering were described previously44. GO enrichment analysis of the data was performed using Omicshare (http://www.omicshare.com/tools/Home/Soft/gogsea). We have deposited the RNA sequencing data into National Center for Biotechnology Information database (accession number SRP067979, www.ncbi.nlm.nih.gov/sra)44.
ChIP assay
The transgenic plants carrying NLP7-GFP were planted under 10 mM ammonium nitrate condition for 7 days, the whole seedlings were used for ChIP assay. The ChIP experiment was carried out following the procedure described previously56,57. All of the primers used in ChIP-qPCR or ChIP-PCR are listed in Supplementary Dataset 6.
EMSA
The DNA probes were labelled by using Biotin 5′ and the DNA probes were listed in the Supplementary Dataset 6. The NLP7 protein was obtain by the transient transformation of Nicotiana benthamiana with the 35Spro:: NLP7-FLAG construct. The EMSA assays were performed using the Lightshift Chemiluminescent EMSA kit (Beyotime Biotechnology).
Statistical analysis
Statistically significant differences were computed based on the Student’s t-tests.
References
Hirel, B., Tétu, T., Lea, P. J. & Dubois, F. Improving nitrogen use efficiency in crops for sustainable agriculture. Sustainability 3, 1452–1485 (2011).
O’Brien, J. A. et al. Nitrate Transport, Sensing, and Responses in Plants. Mol. Plant 9, 837–856 (2016).
Crawford, N. M. & Glass, A. D. Molecular and physiological aspects of nitrate uptake in plants. Trends Plant Sci. 3, 389–395 (1998).
Wang, R., Guegler, K., LaBrie, S. T. & Crawford, N. M. Genomic analysis of a nutrient response in Arabidopsis reveals diverse expression patterns and novel metabolic and potential regulatory genes induced by nitrate. Plant Cell 12, 1491–1509 (2000).
Wang, R., Okamoto, M., Xing, X. & Crawford, N. M. Microarray analysis of the nitrate response in Arabidopsis roots and shoots reveals over 1,000 rapidly responding genes and new linkages to glucose, trehalose-6-phosphate, iron, and sulfate metabolism. Plant Physiol. 132, 556–567 (2003).
Scheible, W.-R. et al. Genome-wide reprogramming of primary and secondary metabolism, protein synthesis, cellular growth processes, and the regulatory infrastructure of Arabidopsis in response to nitrogen. Plant Physiol. 136, 2483–2499 (2004).
Wang, R. et al. Genomic analysis of the nitrate response using a nitrate reductase-null mutant of Arabidopsis. Plant Physiol. 136, 2512–2522 (2004).
Gutiérrez, R. A. et al. Qualitative network models and genome-wide expression data define carbon/nitrogen-responsive molecular machines in Arabidopsis. Genome Biol. 8, R7 (2007).
Wang, R., Xing, X. & Crawford, N. Nitrite acts as a transcriptome signal at micromolar concentrations in Arabidopsis roots. Plant Physiol. 145, 1735–1745 (2007).
Krouk, G., Mirowski, P., LeCun, Y., Shasha, D. E. & Coruzzi, G. M. Predictive network modeling of the high-resolution dynamic plant transcriptome in response to nitrate. Genome Biol. 11, 1 (2010).
Stitt, M. Nitrate regulation of metabolism and growth. Curr. Opin. Plant Biol. 2, 178–186 (1999).
Vidal, E. A. & Gutierrez, R. A. A systems view of nitrogen nutrient and metabolite responses in Arabidopsis. Curr. Opin. Plant Biol. 11, 521–529 (2008).
Krouk, G., Crawford, N. M., Coruzzi, G. M. & Tsay, Y.-F. Nitrate signaling: adaptation to fluctuating environments. Curr. Opin. Plant Biol. 13, 265–272 (2010).
Alvarez, J. M., Vidal, E. A. & Gutiérrez, R. A. Integration of local and systemic signaling pathways for plant N responses. Curr. Opin. Plant Biol. 15, 185–191 (2012).
Walch-Liu, P. et al. Nitrogen regulation of root branching. Ann. Bot. 97, 875–881 (2006).
Alboresi, A. et al. Nitrate, a signal relieving seed dormancy in Arabidopsis. Plant, Cell Environ. 28, 500–512 (2005).
Marín, I. C. et al. Nitrate regulates floral induction in Arabidopsis, acting independently of light, gibberellin and autonomous pathways. Planta 233, 539–552 (2011).
Roenneberg, T. & Rehman, J. Nitrate, a nonphotic signal for the circadian system. FASEB J. 10, 1443–1447 (1996).
Rahayu, Y. S. et al. Root-derived cytokinins as long-distance signals for NO3−-induced stimulation of leaf growth. J. Exp. Bot. 56, 1143–1152 (2005).
Wilkinson, S., Bacon, M. A. & Davies, W. J. Nitrate signalling to stomata and growing leaves: interactions with soil drying, ABA, and xylem sap pH in maize. J. Exp. Bot. 58, 1705–1716 (2007).
Krouk, G. et al. Nitrate-regulated auxin transport by NRT1. 1 defines a mechanism for nutrient sensing in plants. Dev. cell 18, 927–937 (2010).
Krapp, A. et al. Nitrate transport and signalling in Arabidopsis. J. Exp. Bot. 65, 789–798 (2014).
Liu, K. H. & Tsay, Y. F. Switching between the two action modes of the dual‐affinity nitrate transporter CHL1 by phosphorylation. EMBO J. 22, 1005–1013 (2003).
Medici, A. & Krouk, G. The primary nitrate response: a multifaceted signalling pathway. J. Exp. Bot. 65, 5567–5576 (2014).
Forde, B. G. Nitrogen signalling pathways shaping root system architecture: an update. Curr. Opin. Plant Biol. 21, 30–36 (2014).
Vidal, E. A., Álvarez, J. M., Moyano, T. C. & Gutiérrez, R. A. Transcriptional networks in the nitrate response of Arabidopsis thaliana. Curr. Opin. Plant Biol. 27, 125–132 (2015).
Zhang, H. & Forde, B. G. An Arabidopsis MADS box gene that controls nutrient-induced changes in root architecture. Science 279, 407–409 (1998).
Gan, Y., Bernreiter, A., Filleur, S., Abram, B. & Forde, B. G. Overexpressing the ANR1 MADS-box gene in transgenic plants provides new insights into its role in the nitrate regulation of root development. Plant Cell Physiol. 53, 1003–1016 (2012).
Remans, T. et al. The Arabidopsis NRT1. 1 transporter participates in the signaling pathway triggering root colonization of nitrate-rich patches. Proc. Natl. Acad. Sci. USA 103, 19206–19211 (2006).
Vidal, E. A. et al. Nitrate-responsive miR393/AFB3 regulatory module controls root system architecture in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 107, 4477–4482 (2010).
Vidal, E. A., Moyano, T. C., Riveras, E., Contreras-López, O. & Gutiérrez, R. A. Systems approaches map regulatory networks downstream of the auxin receptor AFB3 in the nitrate response of Arabidopsis thaliana roots. Proc. Natl. Acad. Sci. USA 110, 12840–12845 (2013).
Vidal, E. A., Álvarez, J. M. & Gutiérrez, R. A. Nitrate regulation of AFB3 and NAC4 gene expression in Arabidopsis roots depends on NRT1. 1 nitrate transport function. Plant Signal. Behav. 9, e28501 (2014).
Guan, P. et al. Nitrate foraging by Arabidopsis roots is mediated by the transcription factor TCP20 through the systemic signaling pathway. Proc. Natl. Acad. Sci. USA 111, 15267–15272 (2014).
Guan, P. et al. Interacting TCP and NLP transcription factors control plant responses to nitrate availability. Proc. Natl. Acad. Sci. USA 114, 2419–2424 (2017).
Medici, A. et al. AtNIGT1/HRS1 integrates nitrate and phosphate signals at the Arabidopsis root tip. Nat. Commun. 6 (2015).
Ho, C.-H., Lin, S.-H., Hu, H.-C. & Tsay, Y.-F. CHL1 functions as a nitrate sensor in plants. Cell 138, 1184–1194 (2009).
Remans, T. et al. The Arabidopsis NRT1.1 transporter participates in the signaling pathway triggering root colonization of nitrate-rich patches. Proc. Natl. Acad. Sci. USA 103, 19206–19211 (2006).
Forde, B. G. & Walch-Liu, P. Nitrate and glutamate as environmental cues for behavioural responses in plant roots. Plant, Cell Environ. 32, 682–693 (2009).
Wang, R., Xing, X., Wang, Y., Tran, A. & Crawford, N. M. A genetic screen for nitrate regulatory mutants captures the nitrate transporter gene NRT1. 1. Plant Physiol. 151, 472–478 (2009).
Parker, J. L. & Newstead, S. Molecular basis of nitrate uptake by the plant nitrate transporter NRT1. 1. Nature 507, 68–72 (2014).
Sun, J. et al. Crystal structure of the plant dual-affinity nitrate transporter NRT1. 1. Nature 507, 73–77 (2014).
Tsay, Y.-F. Plant science: how to switch affinity. Nature 507, 44–45 (2014).
Hu, H. C., Wang, Y. Y. & Tsay, Y. F. AtCIPK8, a CBL‐interacting protein kinase, regulates the low‐affinity phase of the primary nitrate response. Plant J. 57, 264–278 (2009).
Xu, N. et al. The Arabidopsis NRG2 protein mediates nitrate signaling and interacts with and regulates key nitrate regulators. Plant Cell 28, 485–504 (2016).
Rubin, G., Tohge, T., Matsuda, F., Saito, K. & Scheible, W.-R. Members of the LBD family of transcription factors repress anthocyanin synthesis and affect additional nitrogen responses in Arabidopsis. Plant Cell 21, 3567–3584 (2009).
Castaings, L. et al. The nodule inception‐like protein 7 modulates nitrate sensing and metabolism in Arabidopsis. Plant J. 57, 426–435 (2009).
Konishi, M. & Yanagisawa, S. Arabidopsis NIN-like transcription factors have a central role in nitrate signalling. Nat. Commun. 4, 1617 (2013).
Alvarez, J. M. et al. Systems approach identifies TGA1 and TGA4 transcription factors as important regulatory components of the nitrate response of Arabidopsis thaliana roots. Plant J. 80, 1–13 (2014).
Wang, J.-W., Czech, B. & Weigel, D. miR156-regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell 138, 738–749 (2009).
Marchive, C. et al. Nuclear retention of the transcription factor NLP7 orchestrates the early response to nitrate in plants. Nat. Commun. 4, 1713 (2013).
Schauser, L., Wieloch, W. & Stougaard, J. Evolution of NIN-like proteins in Arabidopsis, rice, and Lotus japonicus. J. Mol. Evol. 60, 229–237 (2005).
Yu, L.-H. et al. Overexpression of Arabidopsis NLP7 improves plant growth under both nitrogen-limiting and-sufficient conditions by enhancing nitrogen and carbon assimilation. Sci. Rep. 6 (2016).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method forAgrobacterium‐mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Hageman, R. & Hucklesby, D. [45] Nitrate reductase from higher plants. Methods Enzymol. 23, 491–503 (1971).
Radin, J. In vivo assay of nitrate reductase in cotton leaf discs. Plant Physiol. 51, 332–336 (1973).
Lee, J. et al. Analysis of transcription factor HY5 genomic binding sites revealed its hierarchical role in light regulation of development. Plant Cell 19, 731–749 (2007).
Lin, R. et al. Transposase-derived transcription factors regulate light signaling in Arabidopsis. Science 318, 1302–1305 (2007).
Acknowledgements
We would like to thank Dr. Anne Krapp for the pNLP7:NLP7-GFP/nlp7-1 and p35S:NLP7-GFP/nlp7-1 seeds. This work was supported by the National Natural Science Foundation of China (grant number 31670247), Taishan Scholar Foundation, and Funds of Shandong “Double Tops” Program (grant number SYL2017YSTD01) to Y.W., Natural Science Foundation of Shandong (grant number BS2014NY003 to S. Q.).
Author information
Authors and Affiliations
Contributions
Y.W., N.C., and L.Z. designed the project. L.Z., W.Z., Y.Y. and N.L. conducted the experiments. The data were analyzed by L.Z., Z. Li, S.Q., and Y.W. The manuscript was written by Y.W., N.C., and L.Z. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, L., Zhang, W., Yang, Y. et al. The Arabidopsis NLP7 gene regulates nitrate signaling via NRT1.1–dependent pathway in the presence of ammonium. Sci Rep 8, 1487 (2018). https://doi.org/10.1038/s41598-018-20038-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-20038-4
This article is cited by
-
Multiple mechanisms are involved in the alleviation of ammonium toxicity by nitrate in cucumber (Cucumis sativus L.)
Plant and Soil (2024)
-
Genomic characterization of a nematode tolerance locus in sugar beet
BMC Genomics (2023)
-
Two nitrate sensors, how many more?
Nature Plants (2022)
-
The source, level, and balance of nitrogen during the somatic embryogenesis process drive cellular differentiation
Planta (2022)
-
The MdABI5 transcription factor interacts with the MdNRT1.5/MdNPF7.3 promoter to fine-tune nitrate transport from roots to shoots in apple
Horticulture Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.