Abstract
Acetylcholinesterase (AChE), an enzyme responsible for degradation of acetylcholine, has been identified as a prognostic marker in liver cancer. Although in vivo Ache tumorigenicity assays in mouse are present, no established liver cancer xenograft model in zebrafish using an ache mutant background exists. Herein, we developed an embryonic zebrafish xenograft model using epithelial (Hep3B) and mesenchymal (SKHep1) liver cancer cell lines in wild-type and achesb55 sibling mutant larvae after characterization of cholinesterase expression and activity in cell lines and zebrafish larvae. The comparison of fluorescent signal reflecting tumor size at 3-days post-injection (dpi) revealed an enhanced tumorigenic potential and a reduced migration capacity in cancer cells injected into homozygous achesb55 mutants when compared with the wild-type. Increased tumor load was confirmed using an ALU based tumor DNA quantification method modified for use in genotyped xenotransplanted zebrafish embryos. Confocal microscopy using the Huh7 cells stably expressing GFP helped identify the distribution of tumor cells in larvae. Our results imply that acetylcholine accumulation in the microenvironment directly or indirectly supports tumor growth in liver cancer. Use of this model system for drug screening studies holds potential in discovering new cholinergic targets for treatment of liver cancers.
Similar content being viewed by others
Introduction
The most commonly occurring liver cancer is hepatocellular carcinoma (HCC), with over half a million new cases emerging yearly and a high death rate1,2. Hepatitis B and C virus infections are among the main factors underlying the development of HCC that is also associated with cirrhosis3. Exposure to aflatoxin B, chronic alcohol use, and other factors triggering cirrhosis also are etiologically important4. Moreover, liver cancer patients show differences in prognosis hence, it is important to discover and validate target genes affecting prognosis. Acetylcholinesterase (ACHE) is one such promising target5; changes in AChE amount has been associated with cancer prognosis or stage in different types of cancers including HCC5,6,7. AChE is responsible for enzymatic degradation of acetylcholine (ACh) and its absence or low activity causes high levels of systemic ACh8. ACh or the agonist nicotine binds to nicotinic ACh receptors fine-tuning cellular signal transduction9. As expected, there is an inverse correlation between AChE activity and ACh amount in liver cancer cell lines and in addition a decrease in AChE activity is associated positively with tumor size, multiplicity and TNM (tumor-node-metastasis) stage5. Moreover, in HCC patients with poor prognosis, there is a statistically significant decrease in the AChE amount5. Recombinant AChE treatment decreases cell proliferation in liver cancer cell lines and overexpressing AChE in a nude mice xenograft model decreases tumor size5. In light of these findings decreased AChE amount is associated with tumorigenesis and thus, studying its role in tumor microenvironment has potential implications in cancer diagnosis and therapy. However, currently there is no in vivo xenograft model for evaluating the connection between AChE and liver cancer development and progression in zebrafish.
Xenograft models for liver cancer have been developed in different organisms10,11,12. In mice, ectopic or orthotopic implantation of hepatoma cells from patient tissue or cultured cell lines into athymic (nude) or severe-combined immunodeficient (SCID) mice have been used for drug screening13,14. Use of multiple tumor cell lines has been proposed since there is a possibility of observing different tumor phenotypes13,15,16. Mouse model of xenotransplantation, however, presents challenges such as long tumor development periods and few numbers of cubs in a single born. Recently, the zebrafish has emerged as a model for tumor xenotransplantation studies17,18,19. Having hundreds of transparent offspring enabling ease in imaging and lack of early developmental adaptive immune system are the primary advantages of using the zebrafish embryos for transplantation. In addition, availability of mutant and transgenic strains with different genetic backgrounds might open new avenues for xenotransplantation19,20,21,22,23.
The zebrafish homolog of the human ACHE gene has been well characterized24,25. achesb55 fish have a loss-of-function point mutation changing Ser226 residue to Asn in the catalytic triad of AChE active site resulting in a null phenotype25. Zebrafish ache homozygous mutants are initially motile but paralyzed at around 72 hours post-fertilization (hpf), whereas heterozygote mutants develop normally until adulthood. However, there is no established xenograft model in zebrafish using an ache mutant background in the literature.
Moreover, ALU based transplant DNA quantification methods have been used to quantify tumor load in mammalian xenotransplantation studies26,27,28,29. These methods provide high detection power and accuracy even when considerably lower proportion of human cells are found in host (<1e-4%) as well as with lower DNA amounts (0.1 pg)28. However, there is yet to be an ALU based quantification assay to be developed also in zebrafish.
In the present study, two representative liver cancer cell lines (Hep3B and SKHep1) with differential cholinesterase expression/activity were selected for transplantation. Next, we developed a novel model for comparison of tumorigenesis between phenotypically wild-type and ache mutant sibling embryos. Using microscopic imaging, we observed that ache mutant embryos developed larger tumors than wild-type siblings. Confocal microscopy enabled better visualization of tumor distribution by making use of an antibody against GPF that is stably expressed in Huh7 cells injected to zebrafish embryos. Next, we modified a mammalian ALU xenograft DNA quantification assay for use in single zebrafish embryos injected with cancer cells. Results showed that ache mutants contained higher numbers of tumor cells when compared with ache heterozygotes and homozygous wild-type embryos. Overall our findings indicate that in the absence of ache, excess systemic ACh in zebrafish larvae provides a permissive microenvironment for tumor development directly or indirectly. The effect we observed is relatively independent of the liver cancer cell cholinesterase expression but rather emphasizes differences in the host zebrafish ache expression. This is the first study of an effective xenograft model in zebrafish larvae using a mutant background for liver cancer cell xenotransplantation. This model holds potential for analyzing the tumorigenic capacity of cell lines and cells from primary tumors, and in combination with drug treatment experiments, offers a promising setup for preclinical trials.
Results
A novel xenograft model using liver cancer cells in achesb55 mutant zebrafish
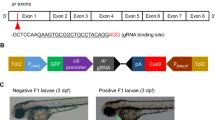
We developed an effective xenograft model using liver cancer cells in ache mutant background (Fig. 1). First, we characterized six different liver cancer cell lines in terms of their acetylcholinesterase/butyrylcholinesterase (ACHE/BCHE) expression and activity (Fig. 1a). Next, we selected for transplantation two different cell lines, namely Hep3B and SKHep1, respectively expressing ACHE/BCHE mRNA in low and high amounts. After staining with live dye DiO (or DiI for ALU quantification), these cells were injected into embryos at 2 days post fertilization (dpf) obtained from heterozygote ache fish in-cross (Fig. 1b,c). Following microinjection, at 6 hours post injection (hpi) embryos positive for tumor seeding were pre-selected, leaving embryos with fluorescent signal outside the injection site (yolk sac) out of further analysis (Fig. 1d). At 3 dpf, we performed touch-evoked tail response test (shortly “tail-test”) to separate ache wild-type and mutant embryos (Fig. 1e). Finally, embryos were fixed and mounted to take images, to measure tumor size and to compare wild-type and mutant siblings in terms of tumorigenesis and tumor cell migration (Fig. 1f). Alternatively, larvae were collected at the 3 dpi to extract DNA for ALU DNA quantification and zebrafish ache genotyping (Fig. 1g).
Schematic representation of the development of the liver cancer xenograft model in zebrafish ache mutants. (a) Liver cancer cells are grown in culture and characterized for ACHE/BCHE expression and activity. (b) Selected cell lines are harvested and stained with live dye DiO (or DiI for ALU quantification). (c) Embryos from ache heterozygous in-cross are injected with 300 cells into their yolk sac at 2 dpf. (d) At 6 hpi, embryos with positive signal are selected and embryos with signal from outside the injection site are not used. (e) After blind injection, mutant larvae are separated from +/? by a touch-evoked tail response test (tail-test). (f) All larvae are fixed and mounted before tumor size is measured and compared between mutant and wild-type larvae. (g) Alternatively, larvae fixed for ALU based xenograft quantification assay.
Characterization of liver cancer cell lines based on ACHE/BCHE expression and activity
For in vitro characterization of cell line ACHE/BCHE expression, we quantified ACHE and another functionally related gene, BCHE expressions using qPCR (Fig. 2a). In humans, there are two ACHE variants with either exon no.5 retained (ACHEvar4–5, NM_001302621.1) or exon no.6 retained (ACHEvar4-6, NM_001302622.1). We used another set of primers targeting ACHE exons no. 3 and 4 (ACHEex3-4) present in both variants. We found that all three amplicons exhibited correlated expression patterns allowing for estimation of an average ACHE transcript amount (Pearson’s correlation coefficients: rACHE4-6/ACHE4-5 = 0.96, P = 0.0025; rACHE4-5/ACHE3-4 = 0.99, P < 0.0001; rACHE4-6/ACHE3-4 = 0.98, P = 0.0004). ACHE transcripts were present in Hep3B, Huh7, PLC and SKHep1 cell lines. SKHep1 showed the highest relative expression whereas HepG2 had a lower and Snu387 virtually no ACHE expression (Fig. 2a). BCHE, on the other hand, was differentially expressed in PLC, SKHep1, Snu387 and to a lower extent in Huh7 but was almost totally absent in Hep3B and HepG2 (Fig. 2a). Next, we studied AChE/BChE enzymatic activity in our panel of liver cancer cell lines using a modified Ellman method based assay. In this assay Huh7, PLC, SKHep1 and Snu387 showed high enzymatic activities (Fig. 2b). SKHep1 had high levels of both ACHE and BCHE expression hence, expectedly it showed relatively higher enzymatic activity. Hep3B also, exhibited cholinesterase activity although not as highly as SKHep1. Our analyses revealed that total ACHE/BCHE mRNA expression exhibited a positive trend across averaged enzymatic activities (Fig. 2c, R = 0.65, P = 0.16). Additionally, we verified by Western blotting that the studied cancer cell lines exhibited AChE protein expression (Supplementary Figure 1).
ACHE, BCHE expression and enzymatic activity profiling in selected liver cancer cell lines. (a) Human ACHE transcript variants ACHE4-6, ACHE4-5 expression and another set of primers targeting ACHE exons no. 3 and 4 (ACHEex3-4) present in both variants were used for measuring relative ACHE expression by qPCR in the cell lines. Additionally, BCHE expression was measured by qPCR. Relative quantification was performed with respect to the cell lines with the lowest expression and log fold change represented. TPT1 gene was taken as reference and ∆∆Ct method formula was used. (b) AChE enzymatic activity in HCC cell lines was measured using an Ellman method based commercial kit. (c) Mean ACHE/BCHE expression values and enzymatic activities were plotted for observation of correlation. Enzymatic activity numerical values were normalized by division to the highest value. All error bars represent the standard deviation.
Characterization of ache expression and activity in zebrafish
We performed qPCR experiments for determining the embryonic and larval expression levels of ache mRNA expression in zebrafish. We isolated RNA from 1, 2, 3, 6, 12, 24 and 48 hours post fertilization (hpf) embryos and verified ache expression quantitatively (Fig. 3a). We found higher zygotic ache expression at 24 and 48 hpf (also see Supplementary Figure 2a and b for RT-PCR and qPCR, respectively). We then synthesized anti-sense ache in situ probes and performed WISH using larvae at 24 hpf and 48 hpf (Supplementary Figure 2c,d). At 24 hpf ache expression was confined to myotome (Supplementary Figure 2c), whereas at 48 hpf it was restricted to the inner brain where ventrorostral and ventrocaudal clusters were located (Supplementary Figure 2d). These findings were in accord with ache’s established role in early neurogenesis30. We further studied the ache expression at later developmental stages in wild-type embryos up to 5 dpf using qPCR (Fig. 3b; N = 2 experiments with n = 20 embryos/group). This revealed that ache expression was present at all developmental time points studied, with the highest expression observed at 4 dpf.
Temporal gene expression of zebrafish ache during embryonic and larval development. (a) Relative ache expression was quantified by qPCR at selected embryonic stages in AB line (N = 1 experiment, n = 20 embryos per group, standard deviation comes from technical replicates). qPCR reactions were run in duplicates and ∆∆Ct was calculated relative to 48 hpf values and normalized with actb2 expression (b) Relative ache expression was quantified by qPCR at 1–5 dpf (N = 2 experiments, n = 20 embryo/larvae per group). qPCR reactions were run in duplicates and ∆∆Ct was calculated relative to 24 hpf values and normalized with actb2 expression. (c) Normalized O.D. 412 values for groups of embryos. AChE enzymatic activity was almost absent in tail-test selected ache−/− mutant embryos when compared with wild-type ache+/? siblings at 28 °C (P = 0.034) or at 33°C (P = 0.003) (performed in triplicates, n = 8 embryos per group). Error bars show the standard deviation.
We obtained achesb55 founder fish from European Zebrafish Resource Center (EZRC) at 3 dpf and raised them to adulthood. For heterozygous parental fish identification, we performed touch-evoked escape response test (shortly “tail-test”) in embryos obtained from founder fish as described previously25. Zebrafish achesb55 heterozygous (+/−) or wild-type (+/+) (together from here on designated as +/?) siblings were able to swim normally upon touching tails (Supplementary Figure 3, Supplementary Movie 1) while (−/−) homozygous mutant embryos were paralyzed at 72 hpf and were not able to contract their tail muscles due to high levels of ACh in the absence of AChE enzyme25 (Supplementary Figure 3, Supplementary Movie 2). Founder fish producing homozygous mutant embryos were selected and labeled as ache(+/−) heterozygous parents, and ache(−/−) homozygous mutant embryos and ache(+/?) siblings gathered from breeding of these fish were used further in xenotransplantation experiments (Fig. 1). To further verify the reliability of the tail-test and the identity of the heterozygous founder fish, we also measured AChE enzymatic activity in homozygous mutant (ache−/−) embryos identified by tail-test to compare with that of the siblings (ache+/?). As expected, we observed 22-fold higher AChE enzymatic activity in siblings when compared to ache mutant embryos at 28 °C (Fig. 3c). Since, after injection we kept the embryos at 33 °C for tumor development, we also checked AChE activity at this temperature and found a 12-fold higher activity in wild-type embryos (Fig. 3c). Enzymatic activity reflects only that of AChE since zebrafish has no bche gene24.
Differential tumorigenic and metastatic capacity of achesb55 mutants and wild-type siblings using Hep3B and SKHep1 cell xenotransplants
Based on qPCR expression analysis, Hep3B, known as a well-differentiated epithelial HCC cell line31, exhibited lower ACHE/BCHE expression/activity while a highly tumorigenic mesenchymal cell-line obtained from liver cancer adenoma32, SKHep1, showed high levels of ACHE/BCHE expression/activity. We developed a novel tumorigenicity model for these two cell lines comparatively, in the ache mutant embryos and healthy wild-type siblings in a microenvironment where ache, a marker for increased tumorigenicity and poor prognosis in human HCC, was depleted (Fig. 1). For this purpose, we injected Hep3B and SKHep1 (300 cells per embryo) to the center of yolk sacs of 2 dpf embryos obtained from achesb55/+ heterozygote in-cross. At 3 dpf, we performed tail-test to separate mutant embryos from wild-type siblings and comparisons for tumorigenicity after cancer cell transplantation was done at 5 dpf using images taken by a fluorescent microscope (Fig. 4). When we compared the tumor size between sb55 mutants and +/? siblings, we found that mutant embryos had significantly larger tumors as evidenced by the amount of fluorescent signal measured in two independent sets of experiments (Fig. 4a–d). In both Hep3B and SKHep1 injected embryos, in ache mutant background there were larger tumors (comparison between +/? and −/− in Fig. 4e,f). This difference in tumor size was independent of the cell lines used. In a Three-Way ANOVA analyzing the effect of phenotype, cell line and experimental sets on tumor size, we found that tumor size showed significant interaction with phenotype (P < 0.001). There was no significant interaction in between the other parameters such as cell line or experimental sets (Pcell-line = 0.757, Pgenotype:cell-line = 0.778, Pgenotype:set = 0.993, Pcell-line:set = 0.210, Pgenotype:cell-line:set = 0.113) yet experimental sets were different in terms of tumor size (Pset = 0.007). Injections to embryos with same genetic background did not differ in tumor size between experimental sets, whereas those with different backgrounds showed a significant difference in between phenotypes (ANOVA followed by Tukey comparisons: P < 0.001 for Hep3B+/? vs. Hep3B−/−, SKHep1+/? vs. SKHep1−/−, SKHep1−/− vs. Hep3B+/? and SKHep1+/? vs. Hep3B−/−). On the other hand, both Hep3B and SKHep1 injected mutants and +/? siblings exhibited comparable number of established tumors whereas migration was enhanced in +/? siblings apparent by a higher number of dispersed cells (Fig. 5a–e, Table 1; +/? = 37.8%, −/− = 6.6%; minimum 5 cells/embryo in the tail posterior to yolk sac taken as positively migrated33). Overall, this suggested that systemic ACh accumulation in the absence of ache function might form a permissive microenvironment for liver cancer cell proliferation and tumor formation.
ache mutant embryos develop larger tumors with both Hep3B and SKHep1 human liver cancer cell xenotransplantation. At 2 dpf, embryos collected from ache+/− in-cross were injected with either Hep3B (a,b) or SKHep1 (c,d) cells. At 3 dpf, tail test was applied for phenotypically separating wild-type (a,c) and mutant embryos (b,d). At 3 dpi, larvae were fixed and imaged for analyzing tumor development (representative of Hep3B and SKHep1 injected groups, three different larvae are shown in a, a’, a”, b, b’, b” and c, c’, c”, d, d’, d”, respectively). Merged images from brightfield and GFP channels were shown. DiO labeled tumor masses can be clearly seen in green. Tumor sizes were measured in ImageJ using the fluorescent signal alone. In total 89 Hep3B injected ache+/?, 26 Hep3B injected ache−/−, 54 SKHep1 injected +/? and 20 SKHep1 injected −/− larvae were analyzed across two independent experiments. Results from each experimental set are separately graphed (e,f). Error bars represent the standard deviation. Bar = 1 mm.
Comparison of metastasis in ache wild-type and mutant larvae. Representative metastatic larva images from Hep3B (a) and SKHep1 (b) injections into ache+/? (n = 154) larvae are shown. Green fluorescent dye DiO labeled migrated cells can be seen both in Hep3B (a’) and SKHep1 (b’) cell injections. Injection of Hep3B (c) and SKHep1 (d) into yolk sac of ache−/− (n = 51) larvae did not show a metastatic phenotype (c’,d’) (when there were more than 5 cells in tail region that embryo was counted as positive for metastasis, see (d’) which was counted as negative for metastasis). (e) Positive metastasis percent in ache wild-type and mutant groups were compared where there were significantly more metastatic larvae in ache wild-type group (P = 0.0002, chi-square test). (f) To see the effect of metastasis on local tumor size, tumor sizes of metastasis positive and negative groups were compared. Metastasis negative larvae (n = 129) showed a trend for larger tumors although non-significant (P = 0.1112, T-test) when compared to metastasis positive larvae (n = 60). n.s = not significant.
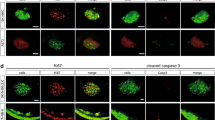
Furthermore, we checked the histology of the xenografted tumors using confocal microscopy by optical sectioning where we used Huh7 HCC cells stably expressing GFP (G12 clone (pre-senescent) transfected with pEGFP-N234). The histology of engrafted Huh7-GFP cells can be clearly seen in the yolk sac of representative ache+/? and ache−/− larvae at 5 dpf (3 dpi) stained with GFP antibodies (in red) (Fig. 6) where cell nuclei were counterstained with DAPI. GFP expressing cancer cells labeled with red could be seen settled in single or multiple foci in the yolk sac of the larva regardless of genotype of the fish (Fig. 6a–f). Furthermore, cancer cells could be clumped or dispersed in the larvae and at times metastasis to the brain and eye could be observed (Fig. 6f). However this method of stable fluorescent marker expression analysis is relatively time consuming and upon inclusion of other markers such as for proliferation or differentiation can be a promising alternative to the live cell staining with DiI/DiO, widely performed in other tumor settings35,36,37.
Optical sectioning and examination of Huh7-GFP xenografted larvae. Larvae (ache+/? and ache−/−) injected with Huh7-GFP cells were stained with GFP primary antibody and Cy3 tagged secondary antibody, and counterstained with DAPI and examined histologically by optical sectioning using a Zeiss LSM 880 confocal microscope. (a–f) At 20× magnification, z-stacks at 10 micron intervals were obtained and consecutive images were Z-projected to obtain these 2D images by maximum intensity projection in ImageJ. Tumor masses can be clearly seen in both images in red. (a’–f’) Same larvae were counterstained with DAPI for marking nuclei. (a”–f”) Merged images were presented. In the fish body schematic, the letters are abbreviations for A: anterior, P: posterior, D: dorsal, V: ventral.
AluYb8 DNA quantification by qPCR confirmed larger tumor volume in ache homozygotes
Prigent et al.28 used an AluYb8 based qPCR method to quantify human progenitor cells transplanted in rodent liver28,38 while the same set of primers were also used in pigs29. We have adopted a similar methodology to quantify human cancer cells in the ache mutant and wild-type zebrafish. In this model AluYb8 repeats were used to detect the human cell contribution in the mixture of human and host cells. We generated the standard curves in the presence of a constant amount of zebrafish DNA with 7-fold serial dilutions of human DNA (10 ng–0.01 pg). Consistent with the Prigent et al.28 the minimum detectable human DNA amount was found to be 0.1 pg. As expected there was a linear trend between the Ct values and the amount of human DNA in the human-zebrafish DNA mixture (Supplementary Figure 4; r = −0.998, P = 2.42e-7). In addition, we did not observe any cross reactivity of the AluYb8 primers with zebrafish DNA (Ct > 35). We did not observe any effect of amount of human DNA on the efficiency of the ache (wild-type and mutant specific) primers (See Supplementary Table 1).
DNA was extracted from 3 dpi whole larvae and directly used for qPCR. Primers previously defined28,38 were used for AluYb8 detection. We used ache (wild-type specific, S and mutant specific, N) primers for genotyping. DNA A260/A280 ratio on average was 1.89 ± 0.06 and 1.93 ± 0.1 for SKHep1 and Hep3B respectively while we used a value greater than 0.6 for the A260/A230 ratio and 30 ng/μl for concentration as thresholds for DNA quality. We examined the qPCR efficiency by comparing the ache Ct values and the log2 transformed DNA concentration levels of samples (Supplementary Figure 5). We obtained a significant negative relationship suggesting that concentration was a good predictor of ache (host) Ct values (r = −0.57 and P = 5.37e-7; r = −0.43 and P = 9.46e-4 for Hep3B and SKHep1, respectively). On the other hand, DNA concentration did not predict the AluYb8 (human DNA) content significantly (data not shown).
We also optimized the usage of the ache PCR primers for the genotyping of the heterozygote and homozygote zebrafish larvae. For the heterozygotes, we obtained detectable amplification with both primer sets. The correlation coefficient between the Ct values of each allele was highly significant (Supplementary Figure 6). For homozygous wild-type (ache+/+) larvae, the mean Ct difference between ache N - ache S primers was 9.85 (±1.07; SD) and 9.85 (±1.12; SD) while for homozygous mutant larvae (ache−/−) the mean Ct difference between ache N - ache S was −5.65 (±0.64; SD) and −5.7 (±0.8; SD), for SKHep1 and Hep3B, respectively. We considered the samples with an absolute Ct difference 1 and 4 cycles between primer sets as unclassified. With these primers, we successfully identified the genotype of the embryos; and the consistency of results showed the reliability of the genotyping (Supplementary Figure 6).
We next compared the ΔCt values of the AluYb8 primers between different genotypes. Consistently with the imaging results of tumor sizes, the ΔCt values were significantly high in ache+/? larvae compared to ache−/− (one tailed student’s T-test; P value = 3.11e-4, SKHep1; and 8.04e-4, Hep3B). When we compared all genotypes, we have observed that +/+ and +/− was not significantly different (P = 0.99 and P = 0.09 for SKHep1 and Hep3B, respectively) however, ache−/− was different from both ache+/+ and ache+/− (Fig. 7). We did not observe any significant interaction between the cell lines (Two-way ANOVA, P = 0.19).
The comparison of the AluYb8 Ct values between genotypes using a qPCR assay. DNA was extracted from 3 dpi larvae injected with liver cancer cell lines (Hep3B and SKHep1) were amplified by qPCRs with using ache N, ache S, and AluYb8 primers (Supplementary Table 1). The Ct value difference (∆Ct) between ache N and ache S primers were used for the genotyping (Supplementary Figure 8). (a) For Hep3B (n = 18, 28 and 10 for ache+/+, ache+/− and ache−/−) normalized Ct values were significantly higher in ache−/− larvae in comparison to both ache+/− and ache+/+ larvae (P = 0.005 and 0.0001, respectively). (b) For SKHep1 (n = 13, 26 and 16 for ache+/+, ache+/− and ache−/−) normalized Ct values were significantly higher in ache−/− larvae in comparison to both ache+/− and ache+/+ (P = 0.004 and 0.017, respectively).
Discussion
Zebrafish and human liver carcinomas share similar molecular pathways39. Chemical treatment of zebrafish results in liver pathologies like steatohepatitis and hepatocellular carcinoma40,41. Recently, inducible oncogene-dependent transgenic models of HCC have been developed in zebrafish12,42,43. However, obtaining founder fish and growing F1 generation can be tedious and time-consuming. Thus, rapid and amenable procedures for high-throughput drug screening setups in mutant/transgenic zebrafish for treatment of liver cancer is an open avenue. In the present study, we developed a novel zebrafish xenotransplantation model using liver cancer cells with differing molecular and cell behavioral characteristics in ache mutant embryos in comparison with their phenotypically wild-type siblings.
Liver cancers are heterogeneous, hence cultured cell lines can provide opportunities to test functional consequences of differing molecular characteristics. For example liver cancer cell lines show different phenotypes in terms of liver marker expression like AFP and/or their epithelial or mesenchymal characteristics which need to be tested in vivo for their tumor forming and metastatic potentials44,45. In this study, using liver cancer cell lines expressing different amounts of ACHE/BCHE and epithelial or mesenchymal characters, we established a novel xenograft model in wild-type and ache mutant zebrafish larvae at 3 dpi. Previous studies have shown that ache in zebrafish is zygotically transcribed starting between 12–24 hpf 24,46 and enzymatically active. In addition, we have confirmed this using qPCR and in situ experiments and further extended the expression profile investigation to 5 dpf, establishing ache expression also in larval zebrafish.
We also showed that expressions of ACHE splice variants were highly correlated with each other in liver cell lines as a continuum of the findings of Zhao and colleagues47. Our findings suggested that ache mutants exhibited larger tumor volumes in comparison with healthy siblings regardless of the cell line used although they showed differential cholinesterase (ACHE/BCHE) expression.
ACHE expression has been shown to be downregulated in human HCCs5 and lower Ache levels have been implicated also, in cirrhosis induction in a rat model48. Parallel to these findings, we showed that presence of achesb55 mutation in the host enabled enlarged tumor growth after xenotransplantation, suggesting that dysregulation of ACHE expression can be an important factor in liver cancer. ACh and AChE levels have also been associated with several other carcinomas like colon, prostate and breast6,7,49. Therefore, our ache xenograft model can further be tested using other human epithelial cancer cell lines to generalize our findings for liver cancer cells.
The strikingly larger tumor size in ache mutants in comparison to that in ache heterozygotes and/or wild-type embryos indicates that increased ACh levels in the host environment potentially leads to enhanced tumor size. ACh has been reported to stimulate colon and lung cancer cells in an autocrine manner50,51. Here, we provide evidence that systemic ACh accumulation in the absence of ache activity in host zebrafish, may result in enhanced liver cancer cell proliferation and tumor growth in this transplantation model using imaging and Alu-based DNA quantification assays. This observation could indeed be independent of ACHE expression profile in these cells. Another possibility is that the baseline activity of ACHE/BCHE protein in Hep3B might be sufficient so it performs similarly to SKHep1 with a higher ACHE/BCHE activity. The fact that heterozygote zebrafish ache+/− mutants do not exhibit paralysis supports this latter hypothesis. Rescue experiments performed with co-injection of different levels of ache mRNA might further help clarify whether the effects of excessive acetylcholine can be remedied in a dose-dependent manner by increased ache activity in mutant embryos. Future studies also should test the tumor size differences in transplantation of isogenic cell lines in which ACHE mRNA is knocked down or over-expressed within the same background. In addition, RNA or tissue from xenotransplanted tumors can be tested for changes in the expression level of known tumor markers.
Metastasis might partially contribute to observed tumor size difference between +/? and −/− larvae due to potentially larger tumor bulk in the yolk of metastasis-negative larvae (Fig. 5f, not significant). Alternatively, but not mutually exclusively, secondary factors, e.g., increased inflammation and edema might lead to tumors growing faster. Potentially the yolk area in which the tumors can grow might exhibit differences between the ache mutants and wild- type/heterozygote population resulting in easier growth due to larger size and/or differential levels of nutrients. Our comparative analysis of yolk sac size indicated that when not injected with tumor cells, mutant embryos have 11 to 25% larger yolk sac sizes compared to siblings at 3 and 5 dpf, respectively (Supplementary Figure 7). Given that when injected with cancer cells the resulting tumors are up to 3 folds larger in mutant embryos, the observed yolk sac size difference could not solely explain the enlarged tumor size. In addition, the body length of mutant larvae was decreased (about 0.5 mm at 5 dpf) in comparison with ache wild-type larvae, indicating a general developmental delay (Supplementary Figure 8). Furthermore, we observed an increase in dispersion of injected cells to tail in wild-type embryos in comparison with mutants. One explanation for the presence of lower number of dispersed cancer cells in ache mutants could be faulty circulation since inhibition of ache activity by organophosphate inhibitors has been shown to result in cardiac edema and decreased blood flow although the vasculature developed normally52. The positive green signal coming from DiO labeled dispersed cells mostly coincided with a signal obtained from the red channel potentially due to autofluorescence of iridophores53. Hence, it is possible that the dispersed cells use a similar route where iridophores migrate to reach their final physiological target locations and/or the red signal has bled through from the green. Xenotransplantation of GFP-labeled Huh7 cells in the +/? and −/− ache siblings was a first step taken in the present study for better visualization of the xenotransplants of liver cancer cells through confocal microscopy. Future studies should include optimization of multi-color staining as well as use of ache mutant hosts expressing fli-GFP for better tracking of the cells.
Human tumor cells originating from breast, adenocarcinoma, melanoma, pancreas, colorectum, glioma, ovary, prostate, and leukemia have been successfully xenotransplanted into zebrafish host previously (for a review see Konantz et al.54). Although there are different sites for injection (e.g. duct of cuvier, perivitelline or peritoneal cavities), the yolk is the most widely used location for transplantation. The number of injected cells ranges from as low as 40 up to 2000 cells. Recently, cancer stem cells from the HepG2 liver cancer cell line has been shown to form tumors and tail metastasis in an adult zebrafish xenograft model55. In a related study, HCC xenografts in zebrafish embryos were produced using Bel-7402 hepatoma cell line; and the effect of ADAM15 recombinant human disintegrin domain (rhddADAM15) was analyzed on tumor growth and metastasis56. In our study, we managed to implant tumors from human liver cancer cell lines in zebrafish yolk sac within a time frame as short as 5 dpf. Tumor sizes were estimated using 2D measurements commonly used in the literature54,56,57. Since the model uses zebrafish embryos, it is possible to screen drugs in a 96 well or even 384-well format because of the smaller body size and transparency of the embryos. We have the advantage of using human cells that more specifically reveal the molecular characteristics of clinical cases; and it would also be possible to use primary HCC cells, e.g. patient-derived xenografts58, as well as murine tumor cell lines or healthy hepatocytes in this in vivo setup. These above-mentioned arguments pinpoint the advantages of drug screening for liver cancer in the zebrafish ache model. Indeed in a similar recent study, the dynamics of a novel HCC cell line xenografted in zebrafish larvae was described demonstrating neo-angiogenesis and potential of a proteasome inhibitory drug to decrease tumor size and cell proliferation57.
In the present study, we also adopted a primate specific AluYb8-based quantification method, previously used in rodents and pigs, for detecting Hep3B and SKHep1 cell line DNA in the zebrafish xenograft model. To the best of our knowledge this is the first time to apply such methodology in zebrafish. Imaging and probe based assays provide semi-quantitative and/or highly variable data. However, a qPCR based model enables to detect and compare the results from single genotyped embryos. The variability in DNA quality and quantity affects the qPCR results but this can be tested and normalized using appropriate controls. For this study, we used quality control thresholds to eliminate low DNA quality samples and performed genotyping for ache in the same assay; thus, this allowed comparison of homozygote mutants against both the heterozygotes and homozygous wild-type embryos. Our findings suggested that the observed increase in tumor size was unique to homozygous mutants and no significant dose effect was apparent for the cell lines studied. We used the ΔCt measurements (human cell DNA normalized with zebrafish ache DNA) in our model to reduce the variability in the DNA concentration for estimating human DNA concentration. In future studies, different DNA extraction protocols might be tested to increase consistency of DNA concentration. In our model, we used DNA mixes from human cancer cells and zebrafish embryos for the standard curve generation. In the literature, it was reported that DNA mix based standard curves tended to underestimate the cell number in the rodent host28,59. Cell mixes may provide a more refined model for determination of the number of cells in the future.
In conclusion, we have developed a novel xenograft model in zebrafish embryos using a mutant background, i.e., the achesb55 mutation, for human liver cancer cells. In ache mutants, the absence of host ache enzymatic activity directly or indirectly resulted in enlarged tumor size, increased tumor load, and in part decreased metastasis to tail region. The contribution of the yolk environment, vasculature, edema, and possible inflammation should be pursued and better characterized. Moreover, our mutant xenograft model and the AluYb8-based quantification method hold great potential to follow effects of cholinergic as well as known (e.g. Sorafenib) and novel drugs in an in vivo setup as a promising screening platform.
Methods
Ethics statement
The experimental protocols in this study were approved by Bilkent University Laboratory Animals Local Ethics Committee (Protocol no: 176, Decree no: 2013/48). All the applied methods were performed in accordance with the relevant guidelines and regulations.
Animal care and handling
Zebrafish achesb55 mutant and AB lines (obtained from European Zebrafish Research Center (EZRC)) were raised and staged according to the standard protocols60. The adult fish were maintained at 28 °C in aquaria with 14 hours of day and 10 hours of night cycles (ZebTEC, Italy). Embryos were kept in E3 medium at 28 °C until they reached the desired embryonic stage.
DNA extraction
We used single 5 dpf embryos for the DNA extraction. Embryos were incubated in DNA extraction buffer (100 mM Tris-Cl, 10 mM EDTA, 200 mM NaCl and 0.5% SDS) and proteinase K (200 ug/ml). Isopropanol and 70% EtOH were used for the DNA precipitation and wash61.
cDNA synthesis and quantitative real-time PCR
Total RNA from liver cancer cell lines and zebrafish embryos/larvae were isolated using Trizol reagent according to manufacturer’s instructions (Invitrogen, USA). cDNA was produced using oligodT primers from 1μg of total RNA with RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher Scientific, USA).
PCR primers were designed in Primer3 software for gene specific RT-PCR experiments (Supplementary Table 1)62. Quantitative real-time PCR was performed as described previously using 2 × SYBR green mix (Roche, Switzerland) for human ACHE variants, BCHE, AluYb8 and zebrafish ache44. Delta-delta Ct (∆∆Ct) method was applied for relative quantification using human TPT1 and zebrafish actb2 or elfa as reference genes63.
Delta Ct method was used for the relative quantification of the AluYb8 with using zebrafish ache gene as a reference. Due to the copy number difference between homozygous and heterozygous DNA, a value of 1, representing the 2-fold difference, was added to the Ct values of the primary primers (ache S for wild-type and ache N for mutant) for homozygous while for heterozygotes the mean values of both primers were used.
For the AluYb8 primers as Walker et al.38 stated there could be two peaks in the melting curve analysis. We have confirmed that there was only one amplicon with the expected length (226 bp) upon running randomly selected qPCR products on the agarose gel although double Tm values were present in the qPCR results. Samples containing only the dimer were determined based on Tm analysis and excluded.
Cloning, probe preparation and whole-mount in situ hybridization
Zebrafish ache coding sequence specific PCR fragment was amplified and T-A cloned into pGEMT-easy vector (Promega, Madison, USA). Plasmids were linearized by restriction enzyme digestion, purified and digoxygenin-labeled anti-sense RNA was produced using MAXIscript SP6/T7 in vitro transcription kit (Invitrogen, USA). Eluted probe was purified using LiCl method. ache whole-mount in situ hybridization (WISH) was performed using standard protocols64.
AChE enzymatic activity assay
For measuring AChE/BChE enzymatic activity, an Ellman method based protocol was used as described by the manufacturer (Abnova, Taiwan). This assay was not AChE specific but rather measured the activity of both AChE and BChE (Abnova, personal communication). Briefly, homogenates from 5 × 106 liver cancer cells (grown on 96 mm dishes) or zebrafish embryos (8 embryos per sample in triplicates at 3 dpf, grown at 28 or 33 °C) were prepared by passing samples 20 times through a syringe needle and subsequent centrifugation and supernatant collection. Homogenates were pipetted in triplicates and O.D. 412 was measured on an ELISA reader (BioTek, USA). Data from three measurements per group were averaged and activity was calculated as stated in the manufacturer’s protocol (Abnova, Taiwan).
Liver cancer cell line maintenance
Liver cancer cell lines Hep3B, HepG2, Huh7, PLC, SKHep1 and Snu387 were maintained as previously described44. Briefly, Hep3B, HepG2, Huh7, PLC and SKHep1 lines were cultured in low glucose DMEM while Snu387 line was cultured in RPMI, and in both cases supplemented with 10% fetal bovine serum, 100 U/mL penicillin-streptomycin and 0.1 mM non-essential amino acids. In addition, Huh7 cells stably overexpressing GFP were supplemented with 200 μg/mL geneticin G-418 sulfate (Thermo Fisher Scientific, USA) and no additional antibiotics34. Cells were grown at 37 °C in an incubator supplied with 5% CO2. Cells reaching confluency (every 3 to 4 days) were split into new plates containing fresh media.
Xenografting into achesb55 embryos and tumorigenesis comparison with siblings
Xenotransplantation was performed using Hep3B and SKHep1 cells as well as Huh7-GFP cells as previously described for other cell types65,66. Briefly, cells were grown on T-75 cell culture flasks to confluency, trypsinized, and then counted using a hemocytometer before collecting 3 × 106 cells for each sample. Hep3B or SKHep1 cells were washed once with PBS and then incubated with fluorescent dyes, for DiO (final 200 mM) at 37 °C for 20 minutes or for DiI (final 25 μg/ml; for AluYb8 experiment) at 37 °C for 5 minutes and following at 4°C for 15 minutes. Subsequently, cells were washed once with FBS and twice with PBS. Finally, cells were dissolved in 50 µl PBS containing a final 0.025% phenol red. At 2 dpf embryos were dechorionated using Pronase (Sigma, USA) and anesthetized in 0.4 mg/ml Tricaine (Fluka, USA). Cell suspension in approximately 10 nl volume containing approximately 300 cells was injected per embryo using an automated injector (Eppendorf FemtoJet, Germany). After injection of cells, embryos were screened using a Leica MZ10F stereo microscope at 6 hpi. Fish with fluorescent signal at the injection site were kept for further analysis while fish having signal outside of injection area or showing no signal were removed from the analysis. At 3 dpf, embryos were screened for the presence of paralysis, separated according to phenotype (Supplementary Figure 3). Embryos were incubated at 33 °C for 3 days after injection (dpi) at which they were fixed overnight at 4 °C in 4% paraformaldehyde; washed three times with PBS and then mounted in 50% glycerol. Fluorescent and brightfield images were taken using a Leica MZ10F stereo microscope (Leica, Switzerland) and analyzed using ImageJ software (NIH)67.
Tumor size from each embryo measured in ImageJ using freehand polygonal selection tool and the area encircled obtained as pixels for each image at the same magnification (x3.2). Additionally, segmentation was performed in ImageJ using threshold function for automated measurement of tumor area. Averaged manual-automatic tumor area measurements were used to construct graphs. For yolk sac measurements, a new set of embryos at 3 dpf and 5 dpf were bred. For tumor measurements fluorescent images and for yolk sac measurements brightfield images were used. Yolk sac measurements were performed manually using freehand polygonal selection tool in ImageJ (Supplementary Information for details). For quantification of metastasis, larvae with local tumor and migrated cells out of yolk were analyzed. Metastatic cell numbers were counted and larvae with more than 5 cells beyond yolk were counted as positive for metastasis33.
Immunofluorescent staining and confocal imaging
At 3 dpi (5 dpf) the larvae injected with Huh7-GFP cells, as described for Hep3B and SKHep1 cells, were fixed using 4% paraformaldehyde (PFA) for 1 hour at room temperature and further permeabilized with cold methanol for 1 hour at 4 °C. Then, larvae were washed three times with PBS-Tween (0.3%) and blocked in PBDT (PBS, DMSO, Tween) supplemented with 2% normal sheep serum for 30 mins at room temperature. Rabbit phospho-histone H3 (ser10) (PH3) (Cell Signaling #9701) and mouse anti-GFP antibody (Cell Signaling, #2955) diluted 1:400 in blocking buffer and incubated with larvae overnight at 4 °C. After six washes with PBDT for a total of 2 hours, larvae were incubated with both anti-rabbit FITC and anti-mouse Cy3 tagged secondary antibody, respectively for PH3 and GFP proteins (Jackson’s Immuno Research, 711-096-152 and 715-167-003, respectively) diluted 1:400 in blocking buffer. Following six PBDT washes for a total of 2 hours, cells were stained with DAPI and washed three times in PBS-T. Finally, larvae were mounted in glycerol and imaged using a Zeiss LSM 880 confocal microscope (Zeiss, Germany). For tumor-focused images 20× objective was used and z-stacks at 10 micron intervals were taken using ZEN software (Zeiss, Germany) for each channel. 4 to 5 consecutive z-stacks were projected using ImageJ software’s Z-project function. Since the green GFP signal/background was not completely washed away it coincided with the green PH3 signal/puncta. Therefore, only images from the blue (DAPI) and red (GFP antibody) channels were used for merging to report the distribution of tumor cells in larvae.
Statistical analyses
For two sample comparisons, student’s T-test was used while for multi-sample comparisons. One-Way ANOVA followed by Tukey’s test was performed in R68. Effects of genotype, cell line and experimental sets on tumor size were analyzed by a Three-Way ANOVA in R. To determine the interaction of genotypes with ALU Ct values (for human tumor DNA quantification) student’s T-test, One-Way ANOVA followed by Tukey’s test or Two-Way ANOVA in Matlab was performed (where indicated).
Data availability
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Tazi, E. M., Essadi, I., M’rabti, H., Touyar, A. & Errihani, H. Systemic treatment and targeted therapy in patients with advanced hepatocellular carcinoma. N. Am. J. Med. Sci. 3, 167–175 (2011).
Lu, J.-W. et al. Zebrafish as a disease model for studying human hepatocellular carcinoma. World J. Gastroenterol. 21, 12042–58 (2015).
El-Serag, H. B., Marrero, J. A., Rudolph, L. & Reddy, K. R. Diagnosis and Treatment of Hepatocellular Carcinoma. Gastroenterology 134, 1752–1763 (2008).
Schlaeger, C. et al. Etiology dependent molecular mechanisms in human hepatocarcinogenesis. Hepatology 47, 511–520 (2008).
Zhao, Y. et al. Acetylcholinesterase, a key prognostic predictor for hepatocellular carcinoma, suppresses cell growth and induces chemosensitization. Hepatology 53, 493–503 (2011).
Novotny, A. et al. Is acetylcholine a signaling molecule for human colon cancer progression? Scand. J. Gastroenterol. 46, 446–455 (2011).
Battisti, V. et al. Cholinesterase activities and biochemical determinations in patients with prostate cancer: Influence of Gleason score, treatment and bone metastasis. Biomed. Pharmacother. 66, 249–255 (2012).
Dvir, H., Silman, I., Harel, M., Rosenberry, T. L. & Sussman, J. L. Acetylcholinesterase: From 3D structure to function. Chem. Biol. Interact. 187, 10–22 (2010).
Paleari, L., Grozio, A., Cesario, A. & Russo, P. The cholinergic system and cancer. Semin. Cancer Biol. 18, 211–217 (2008).
Heindryckx, F., Colle, I. & Van Vlierberghe, H. Experimental mouse models for hepatocellular carcinoma research. International Journal of Experimental Pathology 90, 367–386 (2009).
Mizgirev, I. & Revskoy, S. Generation of clonal zebrafish lines and transplantable hepatic tumors. Nat. Protoc. 5, 383–94 (2010).
Li, Z. et al. Inducible and repressable oncogene-addicted hepatocellular carcinoma in Tet-on xmrk transgenic zebrafish. J. Hepatol. 56, 419–425 (2012).
Huynh, H., Soo, K. C., Chow, P. K. H., Panasci, L. & Tran, E. Xenografts of human hepatocellular carcinoma: a useful model for testing drugs. Clin. Cancer Res. 12, 4306–4314 (2006).
Senturk, S. et al. Transforming growth factor-beta induces senescence in hepatocellular carcinoma cells and inhibits tumor growth. Hepatology 52, 966–974 (2010).
Matsuo, M., Sakurai, H. & Saiki, I. ZD1839, a selective epidermal growth factor receptor tyrosine kinase inhibitor, shows antimetastatic activity using a hepatocellular carcinoma model. Mol Cancer Ther 2, 557–561 (2003).
Kelland, L. R. Of mice and men: values and liabilities of the athymic nude mouse model in anticancer drug development. Eur. J. Cancer 40, 827–36 (2004).
Haldi, M., Ton, C., Seng, W. L. & McGrath, P. Human melanoma cells transplanted into zebrafish proliferate, migrate, produce melanin, form masses and stimulate angiogenesis in zebrafish. Angiogenesis 9, 139–51 (2006).
Taylor, A. M. & Zon, L. I. Zebrafish Tumor Assays: The State of Transplantation. 6, 339–346 (2010).
Lee, L. M. J., Seftor, E. A., Bonde, G., Cornell, R. A. & Hendrix, M. J. C. The fate of human malignant melanoma cells transplanted into zebrafish embryos: assessment of migration and cell division in the absence of tumor formation. Dev. Dyn 233, 1560–70 (2005).
Berghmans, S. et al. Tp53 Mutant Zebrafish Develop Malignant Peripheral Nerve Sheath Tumors. Proc. Natl. Acad. Sci. USA 102, 407–12 (2005).
Nicoli, S., Ribatti, D., Cotelli, F. & Presta, M. Mammalian tumor xenografts induce neovascularization in zebrafish embryos. Cancer Res. 67, 2927–31 (2007).
Liu, S. & Leach, S. D. Zebrafish models for cancer. Annu. Rev. Pathol. 6, 71–93 (2011).
Marques, I. J. et al. Metastatic behaviour of primary human tumours in a zebrafish xenotransplantation model. BMC Cancer 9, 128 (2009).
Bertrand, C. et al. Zebrafish acetylcholinesterase is encoded by a single gene localized on linkage group 7. Gene structure and polymorphism; molecular forms and expression pattern during development. J. Biol. Chem. 276, 464–74 (2001).
Behra, M. et al. Acetylcholinesterase is required for neuronal and muscular development in the zebrafish embryo. Nat. Neurosci. 5, 111–8 (2002).
Becker, M. et al. Sensitive PCR method for the detection and real-time quantification of human cells in xenotransplantation systems. Br. J. Cancer 87, 1328–1335 (2002).
Malek, A., Catapano, C. V., Czubayko, F. & Aigner, A. A sensitive polymerase chain reaction-based method for detection and quantification of metastasis in human xenograft mouse models. Clin. Exp. Metastasis 27, 261–271 (2010).
Prigent, J. et al. Human progenitor cell quantification after xenotransplantation in rat and mouse models by a sensitive qPCR assay. Cell Transplant. 24, 1639–1652 (2015).
Abellaneda, J. M. et al. Validation of a quantitative polymerase chain reaction method for human alu gene detection in microchimeric pigs used as donors for xenotransplantation. Transplant. Proc. 47, 132–135 (2015).
Hanneman, E. & Westerfield, M. Early expression of acetylcholinesterase activity in functionally distinct neurons of the zebrafish. J. Comp. Neurol. 284, 350–361 (1989).
Zhao, H. et al. Epithelial-Mesenchymal Transition Predicts Sensitivity to the Dual IGF-1R/IR Inhibitor OSI-906 in Hepatocellular Carcinoma Cell Lines. Molecular Cancer Therapeutics 11, 503–513 (2012).
Eun, J. R. et al. Hepatoma SK Hep-1 cells exhibit characteristics of oncogenic mesenchymal stem cells with highly metastatic capacity. PLoS One 9 (2014).
Teng, Y. et al. Evaluating human cancer cell metastasis in zebrafish. BMC Cancer 13, 453 (2013).
Ozturk, N. et al. Reprogramming of replicative senescence in hepatocellular carcinoma-derived cells. Proc. Natl. Acad. Sci. USA 103, 2178–2183 (2006).
Harfouche, R. et al. Nanoparticle-mediated targeting of phosphatidylinositol-3-kinase signaling inhibits angiogenesis. Angiogenesis 12, 325–38 (2009).
Thomas, C. et al. ERβ1 represses basal-like breast cancer epithelial to mesenchymal transition by destabilizing EGFR. Breast Cancer Res. 14, R148 (2012).
Yang, X.-J. et al. A novel zebrafish xenotransplantation model for study of glioma stem cell invasion. PLoS One 8, e61801 (2013).
Walker, J. A. et al. Human DNA quantitation using Alu element-based polymerase chain reaction. Anal. Biochem. 315, 122–128 (2003).
Siew, H. L. & Gong, Z. Modeling liver cancer using zebrafish: A comparative oncogenomics approach. Cell Cycle 5, 573–577 (2006).
Amali, A. A. et al. Thioacetamide induced liver damage in zebrafish embryo as a disease model for steatohepatitis. J. Biomed. Sci. 13, 225–232 (2006).
Mizgireuv, I. V. & Revskoy, S. Y. Transplantable tumor lines generated in clonal zebrafish. Cancer Res. 66, 3120–5 (2006).
Sun, L., Nguyen, A. T., Spitsbergen, J. M. & Gong, Z. Myc-Induced Liver Tumors in Transgenic Zebrafish Can Regress in tp53 Null Mutation. PLoS One 10, e0117249 (2015).
Nguyen, A. T., Koh, V., Spitsbergen, J. M. & Gong, Z. Development of a conditional liver tumor model by mifepristone-inducible Cre recombination to control oncogenic krasV12 expression in transgenic zebrafish. Sci. Rep. 6, 19559 (2016).
Avci, M. E., Konu, O. & Yagci, T. Quantification of SLIT-ROBO transcripts in hepatocellular carcinoma reveals two groups of genes with coordinate expression. BMC Cancer 8, 392 (2008).
Yuzugullu, H. et al. Canonical Wnt signaling is antagonized by noncanonical Wnt5a in hepatocellular carcinoma cells. Mol. Cancer 8, 90 (2009).
Allebrandt, K. V., Rajesh, V., Andermann, P. & Layer, P. G. (01) Expression of acetylcholinesterase (AChE) and aryl acylamidase (AAA) during early zebrafish embryogenesis. in Chemico-Biological Interactions 157-158, 353–355 (2005).
Zhao, Y. et al. Acetylcholinesterase, a key prognostic predictor for hepatocellular carcinoma, suppresses cell growth and induces chemosensitization. Hepatology 53, 493–503 (2011).
García-Ayllón, M. S. et al. Changes in liver and plasma acetylcholinesterase in rats with cirrhosis induced by bile duct ligation. Hepatology 43, 444–453 (2006).
Bernardi, C. C. et al. Amplification and deletion of the ACHE and BCHE cholinesterase genes in sporadic breast cancer. Cancer Genet. Cytogenet. 197, 158–165 (2010).
Cheng, K. et al. Acetylcholine release by human colon cancer cells mediates autocrine stimulation of cell proliferation. Am J Physiol Gastrointest Liver Physiol 295, G591–7 (2008).
Song, P. et al. Synthesis of acetylcholine by lung cancer. Life Sci. 72, 2159–2168 (2003).
Lin, C. C., Hui, M. N. Y. & Cheng, S. H. Toxicity and cardiac effects of carbaryl in early developing zebrafish (Danio rerio) embryos. Toxicol. Appl. Pharmacol. 222, 159–168 (2007).
Higdon, C. W., Mitra, R. D. & Johnson, S. L. Gene Expression Analysis of Zebrafish Melanocytes, Iridophores, and Retinal Pigmented Epithelium Reveals Indicators of Biological Function and Developmental Origin. PLoS One 8 (2013).
Konantz, M. et al. Zebrafish xenografts as a tool for in vivo studies on human cancer. Ann. N. Y. Acad. Sci. 1266, 124–37 (2012).
Zhang, B. et al. Zebrafish xenotransplantation model for cancer stem-like cell study and high-throughput screening of inhibitors. Tumour Biol. 35, 11861–9 (2014).
Hou, Y. et al. Recombinant disintegrin domain of ADAM15 inhibits the proliferation and migration of Bel-7402 cells. Biochem. Biophys. Res. Commun. 435, 640–645 (2013).
Tonon, F. et al. Rapid and cost-effective xenograft hepatocellular carcinoma model in Zebrafish for drug testing. Int. J. Pharm. 515, 583–591 (2016).
Huynh, H. et al. Brivanib alaninate, a dual inhibitor of vascular endothelial growth factor receptor and fibroblast growth factor receptor tyrosine kinases, induces growth inhibition in mouse models of human hepatocellular carcinoma. Clin Cancer Res 14, 6146–6153 (2008).
McBride, C., Gaupp, D. & Phinney, D. G. Quantifying levels of transplanted murine and human mesenchymal stem cells in vivo by real-time PCR. Cytotherapy 5, 7–18 (2003).
Kimmel, C. B., Ballard, W. W. & Kimmel, S. R. Stages of embryonic development of the zebrafish. Am. J (1995).
Westerfield, M. The Zebrafish Book. A Guide for the Laboratory Use of Zebrafish (Danio rerio), 5th Edition. Univ. Oregon Press. Eugene (2007).
Rozen, S. & Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–386 (2000).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Thisse, C. & Thisse, B. High-resolution in situ hybridization to whole-mount zebrafish embryos. Nat. Protoc. 3, 59–69 (2008).
Marques, I. J. et al. Metastatic behaviour of primary human tumours in a zebrafish xenotransplantation model. BMC Cancer 9, 128 (2009).
Belo, A. I., van der Sar, A. M., Tefsen, B. & van Die, I. Galectin-4 Reduces Migration and Metastasis Formation of Pancreatic Cancer Cells. PLoS One 8, e65957 (2013).
Rasband, W. ImageJ [Software]. U. S. Natl. Institutes Heal. Bethesda, Maryland, USA//imagej.nih.gov/ij/ (2015).
R Development Core Team, R. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing 1 (2011).
Acknowledgements
We thank Tulay Arayici for zebrafish breeding and care and Dr. Gulcin Cakan-Akdogan and Dr. Gökhan Dalgın for helpful discussions. We thank DPT KANILTEK (granted to M.O.) for the establishment of the Bilkent University Zebrafish Facility. We thank Dr. Güneş Özhan and Dr. Özgür Şahin for sharing immunofluorescence antibodies and chemicals. M.E.A. was supported by a TUBITAK-2232 (The Scientific and Technological Research Council of Turkey) Post-doctoral Fellowship under the advisorship of O.K.; M.M.A. helped fund parts of the research through monetary support from her EMBO Installation Grant.
Author information
Authors and Affiliations
Contributions
M.E.A., O.K., and M.O. identified the research problem and M.E.A. designed and performed all experiments dealing with model development, injections and analyses for imaging as well as some of the in situ and RT-qPCR studies. Injections and analyses for AluYb8 qPCR studies were performed by AGK and ST under the advisorship of O.K., and M.E.I., A.G.K. and S.T. contributed in AChE activity, RT-qPCR, and/or whole-mount in situ hybridization experiments. O.K., M.M.A., and M.O. gave guidance in the overall progression of the project at various points and R.C.A. contributed in the design of Huh7-GFP xenograft and immunofluorescence experiments. M.E.A., O.K., A.G.K., and S.T. performed relevant statistical analyses and wrote the manuscript at large. All authors read, revised, and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Avci, M.E., Keskus, A.G., Targen, S. et al. Development of a novel zebrafish xenograft model in ache mutants using liver cancer cell lines. Sci Rep 8, 1570 (2018). https://doi.org/10.1038/s41598-018-19817-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-19817-w
This article is cited by
-
Protein liposomes-mediated targeted acetylcholinesterase gene delivery for effective liver cancer therapy
Journal of Nanobiotechnology (2021)
-
Modeling oncolytic virus dynamics in the tumor microenvironment using zebrafish
Cancer Gene Therapy (2021)
-
Zebrafish Xenotransplantation Models for Studying Gene Function and Drug Treatment in Hepatocellular Carcinoma
Journal of Gastrointestinal Cancer (2021)
-
UBE2S activates NF-κB signaling by binding with IκBα and promotes metastasis of lung adenocarcinoma cells
Cellular Oncology (2021)
-
lncRNA HOTAIR overexpression induced downregulation of c-Met signaling promotes hybrid epithelial/mesenchymal phenotype in hepatocellular carcinoma cells
Cell Communication and Signaling (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.