Abstract
African swine fever is a devastating viral disease of domestic and wild pigs against which no vaccine or therapy is available. Therefore, we applied the CRISPR (clustered regularly interspaced short palindromic repeats) – Cas9 nuclease system to target the double-stranded DNA genome of African swine fever virus (ASFV). To this end, a permissive wild boar lung (WSL) cell line was modified by stable transfection with a plasmid encoding Cas9 and a guide RNA targeting codons 71 to 78 of the phosphoprotein p30 gene (CP204L) of ASFV. Due to targeted Cas9 cleavage of the virus genome, plaque formation of ASFV was completely abrogated and virus yields were reduced by four orders of magnitude. The specificity of these effects could be demonstrated by using a natural ASFV isolate and escape mutants possessing nucleotide exchanges within the target sequence, which were not inhibited in the Cas9-expressing cell line. Growth of the cell line was not affected by transgene expression which, as well as virus inhibition, proved to be stable over at least 50 passages. Thus, CRISPR-Cas9 mediated targeting of the ASFV p30 gene is a valid strategy to convey resistance against ASF infection, which may also be applied in its natural animal host.
Similar content being viewed by others
Introduction
African swine fever (ASF) is an economically important infectious disease of swine which causes mortality rates of up to 100% in domestic pigs and wild boar. In contrast, infections of African wild pig species (warthogs and bush pigs) are mostly subclinical. Whereas ASF is endemic in Sub-Saharan Africa, previous outbreaks in other parts of the world like South America and Southern Europe could be eliminated, except on the island of Sardinia. However, in 2007 the virus was introduced from Africa to the Caucasian countries Georgia and Armenia. From there it spread via the Russian Federation, Ukraine, and Belarus to the eastern part of the European Union, namely the Baltic states, Poland1,2 and, very recently, the Czech Republic and Romania3.
The causative agent of the disease, African swine fever virus (ASFV), represents the hitherto sole member of the family Asfarviridae4. ASFV is a large (approx. 200 nm) enveloped virus with an icosahedral capsid and two membranes at its inner and outer sides. The 170 to 193 kbp double-stranded DNA genome contains between 150 and 167 open reading frames depending on the isolate, and is predominantly replicated in virus factories located in the cytoplasm of infected cells5. Porcine macrophages are the major natural host cells of ASFV6. However, unlike other known DNA viruses of mammals, it also replicates in several soft tick species7. Ticks play an important role for virus transmission between wild pigs in Africa, but not in Central Europe, where permissive arthropod vectors are absent, and where movement of infected pigs and pig products is the main cause for spread8,9.
Up to now, no efficacious vaccines against ASFV infection are available, although recent attempts to generate e.g. DNA vaccines10, vectored vaccines11, or live attenuated vaccines by defined gene deletions12,13 yielded promising results in laboratory experiments. However, in most cases the considerable variability of the ASFV genome and deduced proteins5 limits the efficacy of potential vaccines, although cross-protection against different ASFV genotypes has been reported for several live vaccine candidates14. Furthermore, several potential antiviral drugs have been evaluated1, and inhibition of ASFV gene expression and replication by specific small interfering RNAs has been shown in vitro15,16.
During recent years, efficient RNA-mediated prokaryotic defence systems like the clustered regularly interspaced short palindromic repeats (CRISPR) - Cas9 nuclease system of Streptococcus pyogenes17,18 have been converted into powerful tools for genome editing in eukaryotes19,20,21. This also opens new possibilities for prevention and control of virus infections22, i.e. by facilitating targeted viral gene deletions or mutations during development of attenuated live vaccines. On the other hand, resistant host organisms can be generated either by knock-out of cellular virus receptor genes, or by constitutive expression of antiviral RNAs by the host. CRISPR/Cas9 mediated receptor targeting has been successful in rendering cells resistant against human immunodeficiency virus (HIV-1) infection in vitro23,24, and swine against the porcine respiratory and reproductive syndrome virus in vivo25,26. However, these pigs were not resistant against ASFV, although the knocked-out macrophage surface protein CD163 has also been considered as a putative ASFV receptor27. Direct targeting of viral genes has been shown to confer virus resistance in plants and insects28,29. Since breeding of clonal transgenic swine modified by using the CRISPR/Cas9 system has been repeatedly shown to be feasible30,31, we decided to attempt the development of ASFV-resistance using this technology.
Results
Generation of CRISPR/Cas9 cell lines
To identify and clone suitable CRISPR/Cas9 target sequences of the ASFV genome we used the vector plasmid pX330-U6-Chimeric_BB-CBh-hSpCas920 which has been originally designed for editing of mammalian genomes, and, therefore, encodes nuclear localization signals (NLS) at both ends of the Cas9 open reading frame (ORF). Since ASFV replicates predominantly in the cytoplasm of infected cells5, these signals were removed. In similar approaches, NLS-less Cas9 has been used for successful targeted mutagenesis of vaccinia virus, which also replicates in the cytoplasm32. Furthermore, a neomycin resistance gene was inserted to enable the isolation of stable CRISPR/Cas9 cell lines.
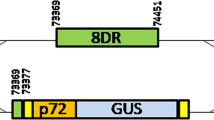
Into the resulting vector pX330-ΔNLS1/2neoR several guide RNA sequences targeting the ORFs of the major capsid protein p72 (B646L), and the viral DNA polymerase (G1211R) of ASFV were inserted at the 5′-end of the trans-activating CRISPR RNA (tracrRNA) gene, but exerted only minor effects on virus replication (not shown). Possibly, the affinity of the Cas9 nuclease to these target sites was suboptimal, or they were highly tolerant against in-frame deletions or insertions resulting from non-homologous end joining. The finally chosen CRISPR/Cas9 guide RNA target sequence (Fig. 1A) encompassed codons 71 to 78 of the ORF encoding the highly immunogenic phosphoprotein p30 (CP204L) of the completely sequenced ASFV strain BA71V (GenBank accession NC_00165933). However, this target sequence is also conserved in the p30 gene of most other ASFV isolates of different genotypes including the viruses currently circulating in Eastern Europe34. Within the viral genome the target sequence was followed by the protospacer adjacent motif (PAM) 5′-NGG-3′, which is required for binding of and cleavage by the Cas9 nuclease of Streptococcus pyogenes (Fig. 1). The corresponding expression plasmid was used for transfection of an ASFV-permissive wild boar lung cell line (WSL), and the obtained neomycin-resistant cell clones were tested by Western blotting for Cas9 expression (Fig. 2A), and for presence of the target specific guide RNA sequences by PCR amplification and sequencing of genomic DNA. Our studies demonstrated that in several cell clones the nuclease and the p30-specific gRNA were stably expressed over many (>50) passages. Nevertheless these cells exhibited similar growth as the parental line, indicating that deleterious off-target reactions of Cas9 nuclease did not occur. This was not surprising, since no sequences matching the chosen gRNA sufficiently21 could be detected within the porcine genome.
Sequence comparison between the ASFV p30 gene-specific guide RNA gene sequence of WSL-gRp30 cells (A), the corresponding viral sequences of ASFV-BA71 and ASFV-Kenya1033 (B), and of two escape mutants of ASFV-Ba71VΔTKdsRed isolated after passage on WSL-gRp30 cells (C). A chromatogram indicating nucleotide peaks and quality is shown above the excerpt of the determined cellular sequence, and the deduced p30 amino acid sequences with position numbers are given below the viral gene fragments. Differences to ASFV-BA71 are coloured, and the targeted 20 nt (vertical lines) as well as the following PAM (red rectangle) are indicated.
(A) Expression of FLAG-tagged Cas9 in WSL-gRp30 cells, and in WSL cells transfected with pX330-ΔNLS1/2neoR was detected by Western blot analyses using an anti-FLAG monoclonal antibody. The expected 161.3 kDa protein is indicated by an arrow. Additional bands detected in transiently expressing cells presumably represent degradation products of Cas9. A parallel blot incubated with an α-tubulin specific monoclonal antibody served as loading control. Molecular masses of marker proteins are indicated. (B) Microscopic fluorescence images showing dsRed-expressing single cells or growing foci and plaques on WSL-gRp30 and WSL cell monolayers of comparable densities at day 5 after infection with the same dilutions of ASFV-BA71VΔTKdsRed or ASFV-Kenya1033ΔCD2vdsRed. Bar indicates 200 µm.
ASFV replication in CRISPR/Cas9 cells
To facilitate infection studies, two ASFV recombinants containing expression cassettes for the red fluorescent protein dsRed were used. The first one, ASFV-BA71VΔTKdsRed, was derived from the cell culture-adapted strain BA71V33 by substitution of the viral thymidine kinase (TK) gene (K196R) by a dsRed expression cassette as described previously35,36. The second mutant, ASFV-Kenya1033ΔCD2vdsRed, was derived from the isolate ASFV-Kenya1033 (publication in preparation) by a similar substitution of the CD2v ORF (EP402R). The CP204L ORF of the Kenyan virus exhibited 87.1% nucleotide sequence identity to that of ASFV-BA71V, resulting in 85.1% identity of the deduced amino acid sequences. However, within the chosen 20 nucleotide (nt) target sequence the Kenyan virus showed 4 base exchanges compared to most other characterised ASFV isolates (Fig. 1B). Remarkably, only one of these alterations led to an amino acid substitution of histidine (H) by glutamine (Q) at position 77 of p30. Because of these mismatches, binding of the guide RNA, and subsequent Cas9 cleavage of the ASFV-Kenya1033 genome appeared unlikely21.
Different Cas9 expressing WSL-gRp30 cell clones and parental WSL cells were infected in parallel with serial dilutions of ASFV-BA71VΔTKdsRed and ASFV-Kenya1033ΔCD2vdsRed. Obviously, both viruses were able to enter all cell lines and to initiate viral gene expression, since single red fluorescent cells became visible after 2 to 3 days. Since dsRed expression was under control of the late p72 promoter of ASFV37,38, viral DNA replication also seems to take place in all tested cell lines. However, plaque formation of ASFV-BA71VΔTKdsRed was impaired in several of the transgenic cell clones. One cell line exhibiting an almost complete inhibition of the spread of ASFV-BA71VΔTKdsRed, but permitted efficient plaque formation of ASFV-Kenya1033ΔCD2vdsRed (Fig. 2B, Fig. 3A), was investigated in more detail.
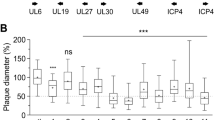
Differential ASFV replication in WSL-gRp30 cells. (A) Plating efficiencies of ASFV-BA71VΔTKdsRed, ASFV-Armenia, ASFV-Kenya1033ΔCD2vdsRed, and ASFV-Kenya1033 were analysed in parallel on WSL (blue bars) and WSL-gRp30 cells (red bars). Mean values of the infectious virus titres (PFU/ml) determined in three experiments and standard deviations are indicated. (B) Growth kinetics of ASFV-BA71VΔTKdsRed and ASFV-Kenya 1033ΔCD2vdsRed on WSL and WSL-gRp30 cells after synchronized infection at an MOI of 0.03. Shown are the mean virus titres at indicated times after infection (h p.i.) and standard deviations of three experiments.
Since BA71V33 is a highly in vitro passaged, avirulent ASFV strain, we also tested a virulent Armenian ASFV isolate from 2007, and the unmodified isolate ASFV-Kenya1033, which were both passaged on WSL cells only few (≤10) times. As expected, plaque assays revealed that replication and spread of ASFV-Armenia, which possesses an identical target sequence to BA71V, were severely inhibited on WSL-gRp30 cells, whereas ASFV-Kenya1033 showed similar replication properties on WSL-gRp30 and WSL cells (Figs 3A, S1).
Furthermore, WSL-gRp30 and normal WSL cells grown in microtiter plates were infected at a multiplicity (MOI) of 0.03 plaque forming units (PFU) per cell of ASFV-BA71VΔTKdsRed or ASFV-Kenya1033ΔCD2vdsRed and incubated for various periods of time at 37 °C. Titration of virus progenies in total cell lysates revealed that both ASFV recombinants exhibited similar replication kinetics on native cells leading to maximum titres of approx. 2 × 106 PFU/ml after 96 h (Fig. 3B). However, on WSL-gRp30 cells titres of ASFV-BA71VΔTKdsRed increased only tenfold compared to the input level, and did not exceed 2 × 102 PFU/ml during the course of the experiment (Fig. 3B). In contrast, replication of ASFV-Kenya1033ΔCD2vdsRed was unaffected on WSL-gRp30 cells (Fig. 3B). One step growth kinetic studies (MOI 3) led to comparable results (not shown).
The almost complete inhibition of spread and productive replication of ASFV-BA71VΔTKdsRed and ASFV-Armenia on WSL-gRp30 cells was considered to be due to specific inhibition by CRISPR/Cas9-mediated cleavage of the virus genome at the p30 gene locus. The secreted and membrane-associated phosphoprotein p30 has been shown to be involved in host cell attachment and entry of ASFV39,40,41, and up to now no corresponding gene deletion mutant could be generated (unpublished results), which strongly indicated that p30 is indispensable for replication of ASFV. The essential function of p30 was confirmed by the present study.
Interestingly, comparative Western blot analyses of lysates of WSL and WSL-gRp30 cells which had been infected with ASFV at a MOI of 5 did not reveal an apparent reduction of the level of p30 caused by CRISPR/Cas9 targeting of CP204L (Fig. S2). This might be due to the very abundant expression of p30 at early times after infection39,40 under control of a strong promoter independent of viral DNA replication42. In contrast, expression of the major capsid protein p72, which is under control of a true late promoter37,38,42 was specifically reduced in WSL-gRp30 cells infected with sensitive ASFV variants (Fig. S2).
Characterization of resistant virus mutants
CRISPR/Cas9 targeting of essential genes in other, e.g. herpesviruses or HIV-1 genomes led to the occurrence of escape mutants resulting from small in-frame deletions after repair of the cleaved DNA, or from proliferation of spontaneous virus variants, which are no longer recognised by the specific guide RNAs43,44. In contrast, consecutive (>10fold) passages of ASFV-BA71VΔTKdsRed at low dilutions on WSL-gRp30 cells or repeated splitting of the abortively infected cells did not lead to enrichment of virus variants capable of more efficient replication in 8 of 10 experimental assays (see below). Thus, neither repair of the Cas9-cleaved DNA, nor spontaneous mutations resulted in viruses possessing a modified functional p30 gene. This indicates that the targeted region of p30 (amino acids 71 to 78) is critical for function.
However, in two out of ten passage experiments of ASFV-BA71VΔTKdsRed infected WSL-gRp30 cells, plaque-forming virus could be reisolated, and PCR amplification and sequencing of the targeted p30 gene region revealed one or two nucleotide exchanges leading to one or two amino acid substitutions (Fig. 1C). Furthermore, several natural ASFV isolates from Kenya, Uganda, Tanzania and Burundi exhibit four base substitutions within the target region of the p30 gene which are identical to those found in the Kenyan virus used in this study (Fig. 1B). Three of them affect wobble bases at third codon positions and are translationally silent. The last one however, although also located at the third position of the codon, leads to a non-conservative amino acid substitution of histidine (H) by glutamine (Q) (Fig. 1B). At present it is not clear whether these substitutions are functionally tolerated, or depend on compensatory alterations elsewhere in p30 or within interacting proteins of the corresponding ASFV strains or mutants.
Discussion
In summary, our experiments provide the first proof-of-principle that ASFV replication can be suppressed in cells expressing a CRISPR/Cas9 system targeting an essential virus gene. This novel approach is important because of the absence of other prophylactic or therapeutic measures against this lethal porcine infection1, and can now be applied to create Cas9 and guide RNA-expressing pigs by transgenesis. Previous studies have already shown the feasibility of stable expression of antiviral CRISPR/Cas9 systems in plants29, as well as in the silkworm Bombyx mori28. Different transgenes encoding e.g. human proteins could be also permanently expressed in pigs during attempts to make them suitable as organ donors for xenotransplantation30,45. To this end, fetal swine fibroblasts were stably transfected with the desired constructs, tested for stable transgene expression in vitro and injected into enucleated oocytes. After fusion and activation the embryos were transferred into the uterus of synchronized sows, and the desired transgenic animals were born. In a similar manner, we will attempt to generate pigs expressing ASFV-specific CRISPR/Cas9 systems. Since swine fibroblasts are in general hardly infectable by ASFV (results not shown), CRISPR/Cas9-expressing transgenic animals have to be generated first, before their macrophages as the natural ASFV host cells6 can be tested for susceptibility. If they exhibit significantly reduced levels of ASFV replication upon in vitro infection, the respective animals will be tested for resistance by challenge infections with virulent ASFV.
Our data show that escape mutations within the targeted part of the ASFV p30 gene are infrequent, and that expression of the p30 gene-specific gRNA leads to an approx.10,000fold reduction of virus replication in WSL cells. Thus it appears likely that a similar effect in transgenic pigs would prevent fatal infection with a matching virus. Innate and adaptive immune responses should then be able to eliminate the virus before escape mutants accumulate to significant levels. Furthermore, only few virulent ASF viruses possessing an altered p30 gene have been detected in the past (see above). Future studies have to demonstrate, whether a modified guide RNA matching this altered p30 gene sequence specifically and efficiently inhibits replication of the respective viruses, and whether cell lines expressing Cas9 nuclease together with both gRNAs are resistant against all natural ASFV variants. Moreover, like in herpesvirus-infected cells43 simultaneous targeting of additional virus genes by the CRISPR/Cas9 system could further increase its efficacy, in particular against multiple ASFV strains and isolates. To achieve this, we have started to adapt a multiplex CRISPR/Cas9 vector system46.
In modified approaches, the CRISPR/Cas9 system should also facilitate targeted deletion of CP204L and other essential or nonessential ASFV genes from the virus genome. This can be supported by generation of trans-complementing cell lines which express the affected proteins from codon-modified ORFs. Such ASFV recombinants could assist in elucidation of gene functions, and might be also suitable as attenuated or single-cycle live vaccines.
Methods
Cells and viruses
The ASFV-permissive wild boar lung cell line WSL was cultivated in Iscove′s Modified Dulbecco’s Medium with Ham’s F-12 Nutrient Mix and 10% fetal bovine serum as described35. For stable introduction of CRISPR/Cas9 plasmids the K2 Transfection System (Biontex) was used. Two days after transfection the cells were trypsinised, seeded into 96 well plates and maintained in medium containing 500 µg/ml G418. Single resistant cell clones detected after 2 to 3 weeks were further propagated.
ASFV-BA71VΔTKdsRed was derived from the Vero cell-adapted strain BA71V33 by substitution of codons 7 to 137 of the viral thymidine kinase (TK) gene (K196R) by the dsRed ORF (recloned from plasmid pDsRed-Express, Clontech) under control of the ASFV major capsid protein (p72) promoter. The virus was isolated after transfection of WSL cells with a corresponding transfer plasmid and subsequent wild-type ASFV infection, followed by selection of TK-negative recombinants on TK-negative WSL cells in the presence of 25 µg/ml bromodeoxyuridine as described35,36. ASFV-Kenya1033ΔCD2vdsRed, was derived from ASFV-Kenya1033 (GenBank submission in preparation) by substitution of the CD2v ORF (EP402R) from codon 77 to the translational stop codon (386) by the same dsRed expression cassette, and repeated plaque purification of fluorescent cell foci after transfer plasmid transfection and virus infection. Furthermore, an Armenian ASFV isolate from 2007 was used in this study, which, like ASFV-Kenya1033, had been adapted by ≤10 serial passages to more efficient replication in WSL cells. Reporter gene expression by the respective virus recombinants was detected by fluorescence microscopy (Axio Vert. A1 with software ZEN 2, Zeiss).
Construction of the CRISPR/Cas9 expression plasmid
The two NLS within the Cas9 ORF of vector pX330-U6-Chimeric_BB-CBh-hSpCas920 were removed by site directed mutagenesis (QuikChange II XL kit, Agilent), using the complementary primer pairs C9ΔNLS1-MF/MR and C9ΔNLS2-MF/MR (Table 1). Furthermore, a neomycin resistance gene under control of the simian virus 40 (SV40) early promoter was inserted into the obtained plasmid. To this end, it was linearized by digestion with PciI, and ligated with a 1661 bp AseI fragment of pcDNA3.1(+) (Thermo Fisher Scientific) after Klenow fill-in of the non-compatible single-stranded overhangs, resulting in plasmid pX330-ΔNLS1/2neoR. Finally, the overlapping complementary oligonucleotide primers gRp30B-F and gRp30B-R (Table 1) were hybridised and inserted into the BbsI-digested vector, resulting in pX330-ΔNLS1/2neoR-gRp30.
DNA preparation, PCR, and sequence analyses
Total DNA of infected and uninfected WSL cells was prepared as described47. To confirm presence of the ASFV-specific guide RNA gene, a 246 bp fragment of genomic cell DNA was amplified by PCR using primers X330gRR-F and X330-gRR-R (Table 1) and Pfx DNA polymerase (Thermo Fisher Scientific), and sequenced with the same primers using the BigDye Terminator v1.1 cycle sequencing kit, and a 3130 Genetic Analyzer (Applied Biosystems). The targeted p30 gene region of the ASFV genome was analysed by PCR amplification and sequencing of a 639 bp fragment with primers CP204L-PSF and CP204L-PSR (Table 1). Results were evaluated with the Geneious® software package in version 10.2.3 (Biomatters).
Western blot analyses
Proteins of WSL and WSL-gRp30 cell lysates were separated by discontinuous sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), transferred to nitrocellulose membranes, and blots were subsequently incubated as described48. The anti-FLAG M2 monoclonal antibody (Sigma-Aldrich), and the α-tubulin specific monoclonal antibody B-5-1-2 (Sigma-Aldrich) were used at dilutions of 1: 5,000 or 1: 20,000, respectively. Binding of peroxidase-conjugated anti-mouse IgG antibodies (Jackson ImmunoResearch) was detected by chemiluminescence (Clarity Western ECL Substrate, Bio-Rad), and recorded (VersaDoc 4000 MP, Bio-Rad).
Virus replication studies
For determination of plating efficiencies WSL-gRp30 and parental WSL cells were grown overnight in microtiter plates, and inoculated in parallel with serial dilutions of ASFV-BA71VΔTKdsRed, ASFV-Armenia, ASFV-Kenya1033ΔCD2v dsRed, or ASFV-Kenya1033 by centrifugation for 1 h at 600 × g and 25 °C. Then the inoculum was replaced by semisolid medium containing 6 g/ml methyl cellulose, and incubation was continued at 37 °C. After 5 days plaques and foci of infected cells were counted to calculate infectious virus titres (PFU/ml). For determination of growth kinetics, WSL-gRp30 and normal WSL cells grown in microtiter plates were inoculated as above at a MOI of 0.03 with ASFV-BA71VΔTKdsRed or ASFV-Kenya1033ΔCD2vdsRed. After removal of the inoculum, the cells were washed and incubated under normal medium at 37 °C. Immediately thereafter, and after 24, 36, 48, 72, 96, 144, and 192 h single plates containing cells plus medium were frozen at −80 °C. Finally, the cells were thawed at 37 °C and virus progenies in total cell lysates were analysed by titration on normal WSL cells.
Biosafety
The Friedrich-Loeffler-Institut is licensed by the competent German authority to work with African swine fever virus, and all experiments with infectious ASFV were performed in improved biosafety level (BSL)−3 laboratories, which fulfill BSL-4 standards with respect to air filtration and effluent treatment.
Data availability
All relevant data generated or analysed during this study are included in this published article.
References
Galindo, I. & Alonso, C. African swine fever virus: A review. Viruses 9, https://doi.org/10.3390/v9050103 (2017).
Guinat, C. et al. Transmission routes of African swine fever virus to domestic pigs: current knowledge and future research directions. Vet Rec 178, 262–267, https://doi.org/10.1136/vr.103593 (2016).
ADNS. ADNS disease overview, https://ec.europa.eu/food/sites/food/files/animals/docs/ad_adns_outbreaks-per-disease.pdf (2017).
Virus Taxonomy: Ninth Report of the International Committee on Taxonomy of Viruses. (ed. King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ) (Academic Press, 2012).
Dixon, L. K., Chapman, D. A., Netherton, C. L. & Upton, C. African swine fever virus replication and genomics. Virus research 173, 3–14, https://doi.org/10.1016/j.virusres.2012.10.020 (2013).
Alcami, A., Carrascosa, A. & Vinuela, E. Interaction of African swine fever virus with macrophages. Virus research 17, 93–104 (1990).
Kleiboeker, S. B., Scoles, G. A., Burrage, T. G. & Sur, J. H. African swine fever virus replication in the midgut epithelium is required for infection of ornithodoros ticks. Journal of virology 73, 8587–8598 (1999).
Costard, S., Mur, L., Lubroth, J., Sanchez-Vizcaino, J. M. & Pfeiffer, D. U. Epidemiology of African swine fever virus. Virus research 173, 191–197, https://doi.org/10.1016/j.virusres.2012.10.030 (2013).
Jori, F. et al. Review of the sylvatic cycle of African swine fever in sub-Saharan Africa and the Indian ocean. Virus research 173, 212–227, https://doi.org/10.1016/j.virusres.2012.10.005 (2013).
Lacasta, A. et al. Expression library immunization can confer protection against lethal challenge with African swine fever virus. Journal of virology 88, 13322–13332, https://doi.org/10.1128/JVI.01893-14 (2014).
Lokhandwala, S. et al. Adenovirus-vectored novel African swine fever virus antigens elicit robust immune responses in swine. PLoS One 12, e0177007, https://doi.org/10.1371/journal.pone.0177007 (2017).
O’Donnell, V. et al. Simultaneous deletion of the 9GL and UK genes from the African swine fever virus Georgia 2007 isolate offers increased safety and protection against homologous challenge. Journal of virology 91, e01760–01716, https://doi.org/10.1128/JVI.01760-16 (2017).
Reis, A. L. et al. Deletion of African swine fever virus interferon inhibitors from the genome of a virulent isolate reduces virulence in domestic pigs and induces a protective response. Vaccine 34, 4698–4705, https://doi.org/10.1016/j.vaccine.2016.08.011 (2016).
Monteagudo, P. L. et al. BA71DeltaCD2: A new recombinant live attenuated African swine fever virus with cross-protective capabilities. Journal of virology, https://doi.org/10.1128/JVI.01058-17 (2017).
Freitas, F. B., Frouco, G., Martins, C., Leitao, A. & Ferreira, F. In vitro inhibition of African swine fever virus-topoisomerase II disrupts viral replication. Antiviral research 134, 34–41, https://doi.org/10.1016/j.antiviral.2016.08.021 (2016).
Keita, D., Heath, L. & Albina, E. Control of African swine fever virus replication by small interfering RNA targeting the A151R and VP72 genes. Antiviral therapy 15, 727–736, https://doi.org/10.3851/IMP1593 (2010).
Horvath, P. & Barrangou, R. CRISPR/Cas, the immune system of bacteria and archaea. Science 327, 167–170, https://doi.org/10.1126/science.1179555 (2010).
Wiedenheft, B., Sternberg, S. H. & Doudna, J. A. RNA-guided genetic silencing systems in bacteria and archaea. Nature 482, 331–338, https://doi.org/10.1038/nature10886 (2012).
Cho, S. W., Kim, S., Kim, J. M. & Kim, J. S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nature biotechnology 31, 230–232, https://doi.org/10.1038/nbt.2507 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823, https://doi.org/10.1126/science.1231143 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Soppe, J. A. & Lebbink, R. J. Antiviral goes viral: Harnessing CRISPR/Cas9 to combat viruses in humans. Trends Microbiol. https://doi.org/10.1016/j.tim.2017.04.005 (2017).
Hou, P. et al. Genome editing of CXCR4 by CRISPR/cas9 confers cells resistant to HIV-1 infection. Sci Rep 5, 15577, https://doi.org/10.1038/srep15577 (2015).
Kang, H. et al. CCR5 disruption in induced pluripotent stem cells using CRISPR/Cas9 provides selective resistance of immune cells to CCR5-tropic HIV-1virus. Mol Ther Nucleic Acids 4, e268, https://doi.org/10.1038/mtna.2015.42 (2015).
Whitworth, K. M. et al. Gene-edited pigs are protected from porcine reproductive and respiratory syndrome virus. Nature biotechnology 34, 20–22, https://doi.org/10.1038/nbt.3434 (2016).
Burkard, C. et al. Precision engineering for PRRSV resistance in pigs: Macrophages from genome edited pigs lacking CD163 SRCR5 domain are fully resistant to both PRRSV genotypes while maintaining biological function. PLoS pathogens 13, e1006206, https://doi.org/10.1371/journal.ppat.1006206 (2017).
Popescu, L. et al. Genetically edited pigs lacking CD163 show no resistance following infection with the African swine fever virus isolate, Georgia 2007/1. Virology 501, 102–106, https://doi.org/10.1016/j.virol.2016.11.012 (2016).
Chen, S. et al. Transgenic clustered regularly interspaced short palindromic repeat/Cas9-mediated viral gene targeting for antiviral therapy of Bombyx mori nucleopolyhedrovirus. Journal of virology 91, e02465–02416, https://doi.org/10.1128/JVI.02465-16 (2017).
Zaidi, S. S., Tashkandi, M., Mansoor, S. & Mahfouz, M. M. Engineering plant immunity: Using CRISPR/Cas9 to generate virus resistance. Front Plant Sci 7, 1673, https://doi.org/10.3389/fpls.2016.01673 (2016).
Petersen, B. & Niemann, H. Molecular scissors and their application in genetically modified farm animals. Transgenic Res 24, 381–396, https://doi.org/10.1007/s11248-015-9862-z (2015).
Ryu, J. & Lee, K. In Methods in Molecular Biology Vol. 1605 (ed Lee K.) Ch. Zygotic Genome Activation, 231–244 (Humana Press, 2017).
Yuan, M. et al. Efficiently editing the vaccinia virus genome by using the CRISPR-Cas9 system. Journal of virology 89, 5176–5179, https://doi.org/10.1128/JVI.00339-15 (2015).
Yanez, R. J. et al. Analysis of the complete nucleotide sequence of African swine fever virus. Virology 208, 249–278 (1995).
Chapman, D. A. et al. Genomic analysis of highly virulent Georgia 2007/1 isolate of African swine fever virus. Emerging infectious diseases 17, 599–605, https://doi.org/10.3201/eid1704.101283 (2011).
Portugal, R., Martins, C. & Keil, G. M. Novel approach for the generation of recombinant African swine fever virus from a field isolate using GFP expression and 5-bromo-2′-deoxyuridine selection. J Virol Methods 183, 86–89, https://doi.org/10.1016/j.jviromet.2012.03.030 (2012).
Keil, G. M. & Giesow, K. & Portugal, R. A novel bromodeoxyuridine-resistant wild boar lung cell line facilitates generation of African swine fever virus recombinants. Archives of virology 159, 2421–2428, https://doi.org/10.1007/s00705-014-2095-2 (2014).
Garcia-Escudero, R. & Vinuela, E. Structure of African swine fever virus late promoters: Requirement of a TATA sequence at the initiation region. Journal of virology 74, 8176–8182 (2000).
Rodriguez, J. M. & Salas, M. L. African swine fever virus transcription. Virus research 173, 15–28, https://doi.org/10.1016/j.virusres.2012.09.014 (2013).
Afonso, C. L. et al. Characterization ofp30, a highly antigenic membrane and secreted protein of African swine fever virus. Virology 189, 368–373 (1992).
Prados, F. J., Vinuela, E. & Alcami, A. Sequence and characterization of the major early phosphoprotein p32 of African swine fever virus. Journal of virology 6, 2475–2485 (1993).
Gomez-Puertas, P. et al. The African swine fever virus proteins p54 and p30 are involved in two distinct steps of virus attachment and both contribute to the antibody-mediated protective immune response. Virology 243, 461–471 (1998).
Portugal, R. S., Bauer, A. & Keil, G. M. Selection of differently temporally regulated African swine fever virus promoters with variable expression activities and their application for transient and recombinant virus mediated gene expression. Virology 508, 70–80, https://doi.org/10.1016/j.virol.2017.05.007 (2017).
van Diemen, F. R. et al. CRISPR/Cas9-mediated genome editing of herpesviruses limits productive and latent infections. PLoS pathogens 12, e1005701, https://doi.org/10.1371/journal.ppat.1005701 (2016).
Lebbink, R. J. et al. A combinational CRISPR/Cas9 gene-editing approach can halt HIV replication and prevent viral escape. Sci Rep 7, 41968, https://doi.org/10.1038/srep41968 (2017).
Niemann, H. & Petersen, B. The production of multi-transgenic pigs: update and perspectives for xenotransplantation. Transgenic Res 25, 361–374, https://doi.org/10.1007/s11248-016-9934-8 (2016).
Sakuma, T., Nishikawa, A., Kume, S., Chayama, K. & Yamamoto, T. Multiplex genome engineering in human cells using all-in-one CRISPR/Cas9 vector system. Sci Rep 4, 5400, https://doi.org/10.1038/srep05400 (2014).
Fuchs, W. & Mettenleiter, T. C. DNA sequence and transcriptional analysis of the UL1 to UL5 gene cluster of infectious laryngotracheitis virus. The Journal of general virology 77, 2221–2229 (1996).
Pavlova, S. P., Veits, J., Mettenleiter, T. C. & Fuchs, W. Live vaccination with an H5-hemagglutinin-expressing infectious laryngotracheitis virus recombinant protects chickens against different highly pathogenic avian influenza viruses of the H5 subtype. Vaccine 27, 5085–5090, https://doi.org/10.1016/j.vaccine.2009.06.048 (2009).
Acknowledgements
The technical help of Anja Landmesser, Katrin Giesow and Antje Frenzel is greatly appreciated. The Armenian and Kenyan ASFV isolates were kindly provided by Sandra Blome and Richard Bishop, respectively. This work has been supported by an intramural grant of the FLI.
Author information
Authors and Affiliations
Contributions
B.P., G.M.K., H.N., and T.C.M. formed the concept to target the ASFV genome by using CRISPR/Cas9 technology. G.M.K. further provided the parental WSL cell line, and the reporter gene-labelled mutant of the Kenyan ASF virus. A.H. conducted most of the other experiments, and analysed the results. W.F. conceived the experiments, and wrote the draft manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hübner, A., Petersen, B., Keil, G.M. et al. Efficient inhibition of African swine fever virus replication by CRISPR/Cas9 targeting of the viral p30 gene (CP204L). Sci Rep 8, 1449 (2018). https://doi.org/10.1038/s41598-018-19626-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-19626-1
This article is cited by
-
The non-classical major histocompatibility complex II protein SLA-DM is crucial for African swine fever virus replication
Scientific Reports (2023)
-
Transcriptome profile of spleen tissues from locally-adapted Kenyan pigs (Sus scrofa) experimentally infected with three varying doses of a highly virulent African swine fever virus genotype IX isolate: Ken12/busia.1 (ken-1033)
BMC Genomics (2022)
-
Recent advances of the biological and biomedical applications of CRISPR/Cas systems
Molecular Biology Reports (2022)
-
Development and application of a colloidal-gold dual immunochromatography strip for detecting African swine fever virus antibodies
Applied Microbiology and Biotechnology (2022)
-
The first genotype II African swine fever virus isolated in Africa provides insight into the current Eurasian pandemic
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.