Abstract
Melilotus is one of the most important legume forages, but the lack of molecular markers has limited the development and utilization of Melilotus germplasm resources. In the present study, 151 M clean reads were generated from various genotypes of Melilotus albus using Illumina sequencing. A total of 19,263 potential EST-SSRs were identified from 104,358 unigene sequences. Moreover, 18,182 primer pairs were successfully designed, and 550 primer pairs were selected using criteria of base repeat type, fragment length and annealing temperature. In addition, 550 primer pairs were screened by using PCR amplification products and used to assess polymorphisms in 15 M. albus accessions. A total of 114 primer pairs were detected as being highly polymorphic, and the average polymorphism information content (PIC) value was 0.79. Furthermore, those 114 polymorphic primer pairs were used to evaluate the transferability to 18 species of the genus Melilotus, and 70 EST-SSR markers were found to be transferable among the 18 Melilotus species. According to the UPGMA dendrogram and STRUCTURE analysis, the 18 Melilotus species were classified into three clusters. This study offers a valuable resource for the genetic diversity and molecular assisted breeding of germplasm resources in the genus Melilotus.
Similar content being viewed by others
Introduction
Melilotus is one of the most important legume forages, and this genus is comprised of 19 annual or biennial species1. All species are native to Eurasia or North Africa2 and diploid with 16 chromosomes (2n = 16) and accessions of Melilotus have two pollination methods, self-pollination and cross-pollination3. Compared with most other pasture crops, members of the genus Melilotus have high seed yields and are adapted to harsh environmental conditions, such as drought, cold and high salinity4,5,6. In addition, it is important for agriculture and animal husbandry, as it is a green manure crop that can be used as a crop fertilizer7. As forage legumes, they have the ability to perform symbiotic nitrogen fixation with species of bacteria species8, and their nitrogen fixation rate is higher than those of many other legumes, making them beneficial for crop rotation9. Previous studies in Melilotus were mainly focused on cultivation techniques10, chemical composition11 and assessment of the agronomic and quality traits12. The genus Melilotus has a single inheritance system, and its relationships with alfalfa and clover were confirmed based on the phylogenetic tree13. However, the molecular markers for the genus Melilotus are limited, hindering its use in genetic and breeding studies. To date, only very limited information has been provided, form 9 SSRs to study the origins of the sweet clover invasion in Alaska14, and 18 SSRs to study the genetic diversity of different species of Melilotus 7. Developing some highly polymorphic EST-SSR markers would allow a better understanding of the genetic diversity in Melilotus, which could facilitate Melilotus breeding programs.
Simple sequence repeats (SSRs) are tandem repeated sequences comprising mono-, di-, tri-, tetra-, penta- or hexa-nucleotide units. Compared with other molecular markers, SSRs have high polymorphism, co-dominance, and locus specificity and are easy to detect15. SSRs are useful tools for studying genetic variation, genetic mapping, and molecular breeding16,17,18,19,20, and they have a high level of transferability between closely related species21. EST-SSRs are a type of molecular marker based on expressed sequence tags; compared with genomic-SSRs, EST-SSRs have a higher level of transferability across related species because EST-SSRs originate from the transcribed regions in genomes and possess conserved sequences among homologous genes22. At present, EST-SSR is widely used in plant genomics research, such as genetic map construction, comparative mapping, genetic diversity evaluation, germplasm identification, and phylogenetic and evolutionary studies23,24,25.
In this study, we aim to (1) develop EST-SSR markers for M. albus using the Illumina HiSeq. 2000 sequencing platform, (2) screen the 550 primer pairs that were selected based on the conditions of base repeat type, fragment length and annealing temperature, by using PCR and PAGE electrophoresis from fifteen accessions of M. albus, (3) detect the transferable and polymorphic EST-SSR markers for the genus Melilotus, and (4) reveal the population structure of Melilotus species.
Results
Sequencing and distribution of EST-SSR
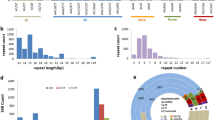
In the present study, we generated 32,939,751, 31,176,000, 32,518,646, 31,470,600 and 35,446,843 raw reads using Illumina sequencing, which included the empty and low-quality reads from the M. albus genotypes N46, N47, N48, N49 and RPh, respectively. After rigorous quality control and data filtering, we generated 30,532,020, 28,785,103, 30,067,146, 29,041,739 and 32,634,517 clean reads, which were deposited into the NCBI SRA database, and obtained 154,458 transcripts and 104,358 unigenes. A total of 19,263 EST-SSR loci were detected from 104,358 unigene sequences, and 18,182 primer pairs were successfully designed. Of these unigenes, 3,063 unigenes contained more than one EST-SSR (Table S1). An average of one EST-SSR was found every 3.99 kb, and the frequency of SSRs was 14.60%. Among the 19,263 potential EST-SSRs, six types of motifs were identified: mononucleotides (12,052, 62.57%), dinucleotides (3,200, 16.61%), trinucleotides (3,654, 18.97%), tetranucleotides (319, 1.66%), pentanucleotides (29, 0.15%), and hexanucleotides (9, 0.05%). EST-SSRs with ten tandem repeats (27.58%) were the most common, and these were followed by eleven, five, six and seven tandem repeats, representing 13.42, 13.29, 11.74, and 5.48%, respectively, while the remaining tandem repeats each accounted for less than 5% of the EST-SSRs (Table 1).
Development, screening, and polymorphism of EST-SSR
In the initial screen of 550 EST-SSR primer pairs were selected based on the conditions of base repeat type, fragment length and annealing temperature by using the genomic DNA of fifteen accessions of M. albus. A total of 351 pairs of primers generated amplification products, while the remaining 199 pairs of primers failed to detect PCR amplification products at multiple annealing temperatures. For the 351 that amplified, 290 pairs of primers were obtained with clear and well-sized amplified products, while the remaining 61 pairs of primers amplified the PCR product bands size is greater or less than the expected fragment size of the primer. Of the 290 pairs of EST-SSR primers capable of amplifying the expected fragment size of the primers, 206 pairs showed polymorphism, while the remaining 84 primers did not (Table S2). Then, 290 EST-SSR primer pairs were selected for further screening based on their polymorphism using sixty individuals of fifteen M. albus accessions. In total, 114 polymorphism primers were selected and used for transferability analysis of the genus Melilotus (Fig. 1, Table S3). Furthermore, to verify the accuracy and authenticity of the PCR amplification bands in this study, PCR products of four pairs of primers (21, 31, 61, 392) were selected and sequenced. As shown in Fig. 2, primer 21 had three-base repeats, (TTC)4, (TTC)5, and (TTC)7; (TCT)4 and (TCT)7 were observed in primer 31; and primer 61 contained tandem repeats of (GATTA)4 and (GATTA)5. (GA)7, (GA)8 and (GA)12 were obtained using primer 392.
Comparative electropherogram analysis of four EST-SSR loci (primers 21, 31, 61 and 392) among different accessions of M. albus. The primer 21 had trinucleotide repeats of (TTC)4, (TTC)5, and (TTC)7. (TCT)4, and (TCT)7 were obtained using primer 31. Primer 61 can generate tandem repeats of (GATTA)4 and (GATTA)5. (GA)7, (GA)8 and (GA)12 were obtained by using primer 392.
Transferability of the newly developed EST-SSR markers
Of the 114 primer pairs, 70 successfully amplified sequence from all accessions of 18 species of the genus Melilotus and showed high polymorphism (Table 2). A total of 411 alleles were detected using 70 EST-SSR loci in 54 Melilotus individuals, ranging from 2 to 11 per locus. Primer 357 yielded the highest number of alleles, and the lowest numbers of alleles were obtained from primers 267 and 383. The observed heterozygosity (H O ) ranged from 0.0 to 1.00 with an average of 0.11. The expected heterozygosity (H E ) ranged from 0.10 to 0.88 with an average of 0.72. The PIC values ranged from 0.10 to 0.87, with an average of 0.69 (Table 2).
Genetic diversity analysis and population structure analysis
As shown in Table 3, the number of polymorphic loci (NPL) for 18 species of Melilotus ranged from 1 (M. sulcatus) to 136 (M. segetalis), and the highest and lowest percentages of polymorphic loci (PPLs) were 33.09% and 0.24%, respectively. The observed number of alleles (na) varied from 1.002 (M. sulcatus) to 1.331 (M. segetalis), and the effective number of alleles (ne), Nei’s gene diversity (h) and Shannon’s Information index (I) values ranged from 1.002 (M. sulcatus) to 1.176 (M. segetalis), 0.001 (M. sulcatus) to 0.112 (M. segetalis) and 0.002 (M. sulcatus) to 0.171 (M. segetalis), respectively.
Using NTSYS-pc.V.2.1 software and the UPGMA method, 58 individual plants of 15 germplasm of M. albus were divided into three clusters (Fig. S1). Cluster I contained five germplasms, PI 478773, PI 508617, PI 342796, PI 662296 and PI 478468. Cluster II contained PI 342765, PI 494706, ZXY06P-1732, Zhongxu-1226 and ZXY07P-3150, and the remaining five accessions were clustered in Cluster III. From the results of the cluster analysis, it can be seen that the individual plants of the germplasm were clustered together, and the genetic similarity coefficients ranged from 0.79 to 0.98. The genetic similarity coefficients were higher, which indicated that the genetic relationship between the 15 germplasms was close.
According to the UPGMA dendrogram (Fig. 3), 18 Melilotus species were classified into three clusters, among which Cluster I contained the 10 species M. albus, M. altissimus, M. dentatus, M. elegans, M. hirsutus, M. officinalis, M. polonicus, M. suaveolens, M. tauricus and M. wolgicus. Cluster II and Cluster III contained four species, respectively, except for the germplasm PI 43597 of M. segetalis and the germplasm PI 317644 of M. spicatus.
The analysis of the genetic structures of 54 individual plants belonging to 18 Melilotus species by using EST-SSR markers was performed with structure software, which was run for K = 2–8. The optimal number of groups was three based on maximum likelihood and delta K (ΔK) values (Fig. 4). Among them, Group I contained 21 individuals belonging to the 7 species M. indicus, M. infestus, M. italicus, M. segetalis, M. siculus, M. speciosus and M. spicatus. Group II contained 18 individuals belonging to the 6 species M. albus, M. dentatus, M. elegans, M. hirsutus, M. officinalis and M. polonicus. Group III contained the remaining 15 individuals, which belonged to the 5 species M. altissimus, M. suaveolens, M. sulcatus, M. tauricus and M. wolgicus.
Genetic structure of 54 individuals for 18 Melilotus species as inferred by STRUCTURE with the EST-SSR marker data set. Histogram of the STRUCTURE analysis for the model with K = 3 (showing the highest ΔK). The smallest vertical barre presents one individual. The assignment proportion of each individual into Cluster I, Cluster II and Cluster III is shown along the y-axis.
Discussion
SSR markers are well known and widely used for genetic diversity analysis, germplasm identification, comparative genetics, phylogenetic relationship, QTL analysis, linkage mapping and marker-assisted selection26,27. Therefore, they are useful for studying genetics and for breeding applications to develop SSR markers from the M. albus transcriptome. In this study, a total of 19,263 potential EST-SSRs were identified from 104,358 Melilotus unigenes, revealing that the abundance of SSRs for Melilotus ESTs was higher than those for alfalfa28, M. truncatula 29, and cupuassu tree30. In total, 18,182 EST–SSR markers were identified from the 19,263 potential EST-SSRs. The study did not analyse the mononucleotide repeats because it was difficult to distinguish single nucleotide repeats from polyadenylation products and single nucleotide stretch errors produced by sequencing28. Here, di- and tri-nucleotide repeats were the most abundant repeats in Melilotus, which is same as in most plants in previous studies. Dinucleotide repeats were found to be the most abundant repeat motif in pistachio21, tea31, rubber tree32 and Neottopterisnidus 33. Tri-nucleotide repeats were the most abundant repeat motif in bread wheat34, Siberian wildrye35, alfalfa28 and castor bean36.
Fifteen M. albus accessions were used to screen the 550 pairs of primer for PCR amplification, and clear bands were detected with 351 pairs. This was fewer than for Siberian wildrye35 but higher than alfalfa28 and rubber tree32. Among these 351 primer pairs that produced amplification, 290 pairs of primers were obtained with clear and well-sized amplified products, while the remaining 61 primer pairs amplified PCR product bands that were larger or smaller than the expected. These deviations may be due to the large insertions or to variation in the repeat number, lack of specificity or assembly error and the existence of introns37,38. However, 114 of those 290 primer pairs were polymorphic among 60 individuals of 15 M. albus accessions. In the tested accessions, the percentage of polymorphic loci was higher than the results for Siberian wildrye35,39. The high levels of polymorphism observed may be due to the M. albus materials selected for screening the primers.
In this study, 114 newly developed EST-SSR markers were used to evaluate the transferability of 18 species of the genus Melilotus. In total, 70 of the 114 primer pairs successfully amplified in all species and obtained stable transferability (61.4%), which is higher than the rates obtained in Siberian wildrye (49.11%)35 and Cucumis (12.7%)40. The high rate of transferability of EST-SSRs was may be because all genotypes belonged to the genus Melilotus, which means that the EST-SSR markers are derived from transcribed regions that are conserved across species. Additionally, the EST-SSRs have higher transferability than SSRs were from untranscribed regions41. Molecular markers are useful for evaluating the genetic diversity in crop species. The genetic diversity estimated by EST-SSR loci was supported by the high values of N A , H O , H E and PIC. In this study, the avenge values of N A , H O , H E and PIC were 5.87, 0.11, 0.72 and 0.69, respectively. These values were lower than those seen in our previous studies in Melilotus 7. This may be because the expressed sequences, from which EST-SSR are derived, are highly conserved. EST-SSR markers detected a lower rate of polymorphism than the genomic SSRs42,43. However, the average values of H E and PIC were higher than in previous studies in Elymus 35, alfalfa (M. sativa L.)28 and millet (Setariaitalica L.)44. However, the value of H O was low, with an average of 0.11. Therefore, to analyse the genetic diversity and promote genetic breeding programs in future studies, it is necessary to improve the heterozygosity of Melilotus accessions7. As shown in the UPGMA results, M. albus, M. altissimus, M. dentatus, M. elegans, M. hirsutus, M. officinalis, M. polonicus, M. tauricus, M. wolgicus and M. suaveolens were clustered into single group, which is consistent with the results of previous studies13. However, in a previous study, M. dentatus and M. tauricus were not clustered into the same group as the other species7. Among the 40 Melilotus germplasms, the association between the clustering pattern and the geographical distribution was less significant. The result may be due to the small number of accessions from each geographical location used in this study. The similar results have been reported in alfalfa45 and drumstick46. These results revealed that genetic distance cannot be the only criterion for genetic differentiation of populations. Moreover, the genetic structure revealed some species showing admixture between group I and group II, while species showed less admixture in group III. Therefore, it is important to use more EST-SSR loci and more individual plants to reveal the relationships among Melilotus species in future studies.
In the present study, we developed a large number of EST-SSR markers for Melilotus from transcriptome data. A total of 104,358 unigenes were generated, and 19,263 EST-SSRs were identified. For these EST-SSRs, 18,182 primer pairs were successfully designed, providing an important foundation for molecular marker development in Melilotus. Of these EST-SSRs, 550 were selected on the basis of base repeat type, fragment length and annealing temperature for further validation. A total of 114 primer pairs detected high polymorphism among 15 accessions of M. albus. In addition, 70 showed transferability of 114 polymorphic primer pairs among 18 Melilotus species. The results suggest that these 70 primer pairs will be useful in future studies of Melilotus population structure, genetic diversity, molecular assisted selection, QTL analysis, and evaluation of germplasm accessions. This study offers a valuable resource for the genetic diversity and molecular-assisted breeding of germplasm resources in the genus Melilotus.
Materials and Methods
Plant materials and DNA extraction
Seeds from Melilotus species were obtained from the National Gene Bank of Forage Germplasm (NGBFG, China) and the National Plant Germplasm System (NPGS, USA) as summarized in Tables 4 and 5. Fifteen accessions of M. albus were used to screen EST-SSR markers for polymorphism and to assess genetic diversity (Table 4), and each accession contained four individual plants. Forty accessions of eighteen species in Melilotus genus were collected to evaluate the transferability of these newly developed EST-SSR markers to other related species and to analyse genetic diversity among Melilotus species (Table 5), and each species contained three individual plants from different accessions. Genomic DNA was extracted from the young leaves using the sodium dodecyl sulfate (SDS) method47. The extracted DNA was detected by agarose gel electrophoresis. The samples were diluted with ddH2O to 50 ng/μL and stored at −20 °C.
Detection of the EST-SSR markers and primer design
SSRs were detected in the assembled unigenes using the Simple Sequence Repeat Identification Tool program (MicroSatellite), and the SSRs were considered to contain mono-, di-, tri-, tetra-, penta-, and hexa-nucleotides with minimum repeat numbers of ten, six, five, five and five, respectively. The EST-SSR primers were designed using BatchPrimer3, and the designed EST-SSR primers were synthesized by Shanghai Sangon Biological Engineering Technology (Shanghai, China).
Primer selection and PCR conditions
A total of 550 primer pairs were selected according to the conditions of base repeat type (except for single bases), fragment length (150–200 bp) and annealing temperature (55–60 °C). PCR and electrophoresis were performed to screen EST-SSR primer pairs for polymorphism using Melilotus species. PCR amplifications were performed in a final volume of 10 μL containing 1 μL genomic DNA (50 ng/μL), 4.9 μL 2 × Reaction Mix (dNTPs at 500 μM each, 20 mM Tris–HCl, 100 mM KCl, 3 mM MgCl2), 0.1 μL 2.5 U/μL Golden DNA Polymerase, 1.0 μL of each primer (4 μM each) and 2.0 μL double distilled water. PCR cycling conditions were 3 min at 94 °C, 35 cycles of 30 s at 94 °C, 30 s at the annealing temperature, 30 s at 72 °C and a final extension step of 7 min at 72 °C. The PCR products were subjected to electrophoresis on 6.0% non-denaturing polyacrylamide gels and stained using silver dye. In addition, the DL500 DNA marker was used to determine the sizes of the PCR products.
Sequencing of PCR amplification products
To verify the accuracy and authenticity of the PCR amplification products, we selected some PCR amplification products for sequencing by Shanghai Sangon Biological Engineering Technology (Shanghai, China). PCR products should be selected according to the presence of single bands and high amplification efficiency.
Data analysis
The number of alleles of each EST-SSR locus was calculated based on presence (1) or absence (0). The observed heterozygosity (H O ), expected heterozygosity (H E ), and polymorphic information content (PIC) were calculated as previously reported48. The program POPGENE 3249 was used to calculate the number of polymorphic loci (NPL), the percentage of polymorphic loci (PPL), the observed number of alleles (na), the effective number of alleles (ne), Nei’s gene diversity (h) and Shannon’s Information index (I) values. With the help of SAHN-clustering in NTSYSpc-v.2.1 software, Nei’s unbiased genetic distance and UPGMA were used for cluster analysis to generate a dendrogram50. The model-based approach implemented was used to subdivide the individuals into different subgroups in the program STRUCTURE 2.351.
Data availability
The RNA-seq data supporting the results of this article are available at NCBI under BioProject with accession PRJNA331091.
References
Outreach, S. Managing cover crops profitably. Sustainable Agriculture Network Handbook (1992).
Aboel-Atta, A.-M. I. I. RAPD and ISSR Variation in Melilotus indica (L.) All. and M. siculus (Turra) BG Jacks. (Leguminosae). Acad. J. Plant Sci. 2, 113–118 (2009).
Brenner, D. M. Methods for Melilotus germplasm regeneration. Plant Genetic Resources Newsletter 141, 51–55 (2005).
Luo, K. et al. Genotypic Variation in a Breeding Population of Yellow Sweet Clover (Melilotus officinalis). Fron. Plant Sci.7 (2016).
Rogers, M. E. et al. Diversity in the genus Melilotus for tolerance to salinity and waterlogging. Plant Soil 304, 89–101 (2008).
Emada, A. S. Melilotus indicus (L.) All., a salt-tolerant wild leguminous herb with high potential for use as a forage crop in salt-affected soils. Flora - Morphology, Distribution, Functional Ecology of Plants 204, 737–746 (2009).
Wu, F. et al. Analysis of genetic diversity and population structure in accessions of the genus. Melilotus. Industrial Crops & Products 85, 84–92 (2016).
Bromfield, E. S. P. et al. Ensifer, Phyllobacterium and Rhizobium species occupy nodules of Medicago sativa (alfalfa) and Melilotus alba (sweet clover) grown at a Canadian site without a history of cultivation. Microbiology 156, 505–520 (2010).
Stickler, F. C. & Johnson, I. J. Dry Matter and Nitrogen Production of Legumes and Legume Associations in the Fall of the Seeding Year1. Agron. J. 51, 135–137 (1959).
Conn, J. S. & Seefeldt, S. S. Invasive white sweetclover (Melilotus officinalis) control with herbicides, cutting, and flaming. Invasive Plant Science & Management 2, 270–277 (2014).
Quijano-Celis, C. E., Pino, J. A. & Morales, G. Chemical Composition of the Leaves Essential Oil of Melilotus officinalis (L.) Pallas from Colombia. J. Essent. Oil Bear. Pl. 13, 313–315 (2010).
Luo, K., Di, H. Y., Zhang, J. Y., Wang, Y. R. & Li, Z. Q. Preliminary evaluation of agronomy and quality traits of nineteen Melilotus accessions. Prataculturalence 31, 2125–2134 (2014).
Di, H. et al. Interspecific Phylogenic Relationships within Genus Melilotus Based on Nuclear and ChloroplastDNA. Plos One 10, e0132596 (2014).
Winton, L. M., Krohn, A. L. & Conn, J. S. Microsatellite markers for the invasive plant species white sweetclover (Melilotus alba) and yellow sweetclover (Melilotus officinalis). Mol. Ecol. Notes 7, 1296–1298 (2007).
Song, Y. P. et al. Differences of EST-SSR and genomic-SSR markers in assessing genetic diversity in poplar. Forest Science and Practice 14, 1–7 (2012).
Mohammadreza, N. et al. Comparison of genetic variation among accessions of Aegilops tauschii using AFLP and SSRmarkers. Genet. Resour. Crop Ev. 54, 237–240 (2007).
Gupta, P. K., Langridge, P. & Mir, R. R. Marker-assisted wheat breeding: present status and future possibilities. Mol. Breeding 26, 145–161 (2010).
Prasanna, B. M., Kevin, P., Marilynl, W. & Xie, C. X. Molecular marker-assisted breeding options for maize improvement in Asia. Mol. Breeding 26, 339–356 (2010).
Salem, K. F. M., Varshney, R. K., Röder, M. S. & Börner, A. EST-SSR based estimates on functional genetic variation in a barley (Hordeum vulgare L.) collection from Egypt. Genet. Resour. Crop Ev. 57, 515–521 (2010).
Li, H. et al. Development and genetic mapping of microsatellite markers from whole genome shotgun sequences in Brassica oleracea. Mol. Breeding 28, 585–596 (2011).
Ziya, M. E., Kafkas, S., Khodaeiaminjan, M. N. Ç. & Gözel, H. Genome survey of pistachio (Pistacia vera L.) by next generation sequencing: Development of novel SSR markers and genetic diversity in Pistacia species. Bmc Genomics 17, 998 (2016).
Wu, J., Cai, C., Cheng, F., Cui, H. & Zhou, H. Characterisation and development of EST-SSR markers in tree peony using transcriptome sequences. Mol. Breeding 34, 1853–1866 (2014).
Shrivastava, D., Verma, P. & Bhatia, S. Expanding the repertoire of microsatellite markers for polymorphism studies in Indian accessions of mung bean (Vigna radiata L. Wilczek). Mol. Biol. Rep. 41, 5669–5680 (2014).
Raveendar, S. et al. Cross-amplification of Vicia sativa subsp. sativa microsatellites across 22 other Vicia species. Molecules 20, 1543–1550 (2015).
Bhawna, Abdin, M. Z., Arya, L. & Verma, M. Transferability of cucumber microsatellite markers used for phylogenetic analysis and population structure study in bottle gourd (Lagenaria siceraria (Mol.) Standl.). Appl. Biochem. Biotec. 175, 1–18 (2015).
Saha, M. C., Cooper, J. D., Mian, M. A., Chekhovskiy, K. & May, G. D. Tall fescue genomic SSR markers: development and transferability across multiple grass species. Theor. Appl. Genet. 113, 1449–1458 (2006).
Cavagnaro, P. F. et al. Genome-wide characterization of simple sequence repeats in cucumber (Cucumis sativus L.). BMC Genomics 11, 569 (2010).
Wang, Z., Yan, H., Fu, X., Li, X. & Gao, H. Development of simple sequence repeat markers and diversity analysis in alfalfa (Medicago sativa L.). Mol. Biol. Rep. 40, 3291 (2013).
Eujayl, I. et al. Medicago truncatula EST-SSRs reveal cross-species genetic markers for Medicago spp. Theor. Appl. Genet. 108, 414–422 (2004).
Lucas, F. D. S. et al. First Microsatellite Markers Developed from Cupuassu ESTs: Application in Diversity Analysis and Cross-Species Transferability to Cacao. Plos One 11, e0151074 (2016).
Sharma, R. K., Bhardwaj, P., Negi, R., Mohapatra, T. & Ahuja, P. S. Identification, characterization and utilization of unigene derived microsatellite markers in tea (Camellia sinensis L.). BMC Plant Biol. 9, 53 (2009).
Feng, S. P., Li, W. G., Huang, H. S., Wang, J. Y. & Wu, Y. T. Development, characterization and cross-species/genera transferability of EST-SSR markers for rubber tree (Hevea brasiliensis). Mol. Breeding 23, 85–97 (2009).
Jia, X., Deng, Y., Sun, X., Liang, L. & Su, J. De novo assembly of the transcriptome of Neottopteris nidus using Illumina paired-end sequencing and development of EST-SSR markers. Mol. Breeding 36, 1–12 (2016).
Gupta, P. K. et al. Transferable EST-SSR markers for the study of polymorphism and genetic diversity in bread wheat. Molecular Genetics & Genomics 270, 315–323 (2003).
Zhou, Q. et al. Development and cross-species transferability of EST-SSR markers in Siberian wildrye (Elymus sibiricus L.) using Illumina sequencing. Sci. Rep. 6, 20549 (2016).
Qiu, L., Yang, C., Bo, T., Yang, J. B. & Liu, A. Exploiting EST databases for the development and characterization of EST-SSR markers in castor bean (Ricinus communis L.). BMC Plant Biol. 10, 1–10 (2010).
Wang, Z. et al. De novo assembly and characterization of root transcriptome using Illumina paired-end sequencing and development of cSSR markers in sweet potato (Ipomoea batatas). BMC Genomics 11, 726 (2010).
Wei, W. et al. Characterization of the sesame (Sesamum indicum L.) global transcriptome using Illumina paired-end sequencing and development of EST-SSR markers. BMC Genomics 12, 451 (2011).
Lei, Y., Zhao, Y., Yu, F., Li, Y. & Dou, Q. Development and characterization of 53 polymorphic genomic-SSR markers in Siberian wildrye (Elymus sibiricus L.). Conserv. Genet. Resour. 6, 861–864 (2014).
Fernandezsilva, I. et al. Bin mapping of genomic and EST-derived SSRs in melon (Cucumis melo L.). Theoretical and Applied Genetics 118, 139–150 (2008).
Scott, K. D. et al. Analysis of SSRs derived from grape ESTs. Theor. Appl. Genet. 100, 723–726 (2000).
Chabane, K., Ablett, G. A., Cordeiro, G. M., Valkoun, J. & Henry, R. J. EST versus genomic derived microsatellites for genotyping wild and cultivated barley. Genet. Resour. Crop Ev. (2005).
Cho, Y. G. et al. Diversity of microsatellites derived from genomic libraries and GenBank sequences in rice (Oryza sativa L.). Theor. Appl. Genet. 100, 713–722 (2000).
Zhao, W. et al. Development and use of novel SSR markers for molecular genetic diversity in Italian millet (Setaria italica L.). Genes & Genom. 34, 51–57 (2012).
Zhang, Z., Min, X. & Wang, Z. et al. Genome-wide development and utilization of novel intron-length polymorphic (ILP) markers in Medicago sativa. Mol. Breeding 37(7), 87 (2017).
Kumar Ganesan, S., Singh, R., Choudhury, D. R., Bharadwaj, J., Gupta, V. & Singode, A. Genetic diversity and population structure study of drumstick (Moringa oleifera lam.) using morphological and SSRmarkers. Ind Crop Prod 60, 316–325 (2014).
Shan, Z., Hong-Liang, W. U., Cheng-Lei, L. I., Chen, H. & Qi, W. U. Improved SDS method for general plant genomic DNA extraction. Guangdong Agricultural Sciences 38, 113–115 (2011).
Chung, J. W., Kim, T. S., Suresh, S., Lee, S. Y. & Cho, G. T. Development of 65 novel polymorphic cDNA-SSR markers in common vetch (Vicia sativa subsp. sativa) using next generation sequencing. Molecules 18, 8376 (2013).
Yeh, F. C. Population genetic analysis of codominant and dominant markers and quantitative traits. Belg.j.bot 129 (1997).
Rohlf, F. J. NTSYS-pc - Numerical Taxonomy and Multivariate Analysis System. Version 2.1 (2005).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol. Ecol. Resour. 155, 945–959 (2000).
Acknowledgements
This work was financially supported by grants from National Basic Research Program (973) of China (2014CB138704), the National Natural Science Foundation of China (31572453), Special Fund for Agro-scientific Research in Public Interest (20120304205), Program for Changjiang Scholars and Innovative Research Team in University (IRT_17R50), Fundamental Research Funds for the Central Universities (lzujbky-2016-10), and the 111 Project (B12002).
Author information
Authors and Affiliations
Contributions
Zhuanzhuan Yan, Fan Wu, Yanrong Wang and Jiyu Zhang conceived the topic. Zhuanzhuan Yan, Fan Wu, Kai Luo, Yufeng Zhao, Qi Yan and Yufei Zhang performed the experiments. Zhuanzhuan Yan, Fan Wu, Kai Luo and Jiyu Zhang analyzed all statistical data. Zhuanzhuan Yan wrote the manuscript. All authors read and approved the final version.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yan, Z., Wu, F., Luo, K. et al. Cross-species transferability of EST-SSR markers developed from the transcriptome of Melilotus and their application to population genetics research. Sci Rep 7, 17959 (2017). https://doi.org/10.1038/s41598-017-18049-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-18049-8
This article is cited by
-
Genome survey sequencing-based SSR marker development and their validation in Dendrocalamus longispathus
Functional & Integrative Genomics (2023)
-
Development of Novel Genic SSR Markers and Their Transferability Across the Genus Pistacia Species and Comparison of In Silico Genomic SSRs and Genic SSRs in Pistachio
Plant Molecular Biology Reporter (2023)
-
EST-SSR marker-based genetic diversity and population structure analysis of Indian Curcuma species: significance for conservation
Brazilian Journal of Botany (2021)
-
Genome-wide development and application of miRNA-SSR markers in Melilotus genus
Physiology and Molecular Biology of Plants (2021)
-
Development of EST-SSR markers and association mapping with floral traits in Syringa oblata
BMC Plant Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.