Abstract
Long noncoding RNAs (lncRNAs) have been shown to play critical roles in cancer. lncTCF7 (gene symbol: WSPAR) has been reported to maintain stemness in hepatocellular carcinoma (HCC) stem cells. However, little is known about the role of lncTCF7 in glioma. The aim of this study was to identify the role of lncTCF7 in the pathogenesis of glioma. We analysed the relationship of lncTCF7 expression with clinicopathological characteristics in glioma patients. Our results showed that lncTCF7 expression was increased in glioma tissues compared with that in normal brain tissues (P < 0.001). Moreover, lncTCF7 was significantly associated with WHO grade (I–II vs. III–IV; P = 0.006) and tumour size (<3 cm vs. T ≥ 3 cm; P = 0.025). Meanwhile, patients with high lncTCF7 expression levels exhibited markedly worse overall survival prognoses (P < 0.01). Loss of function assays revealed that knockdown of lncTCF7 significantly inhibited glioma cell migration, proliferation and tumorigenicity in vitro and in vivo. Furthermore, we found that hypoxia induced lncTCF7 expression in an autocrine manner through IL-6 in glioma. In conclusion, lncTCF7 may play a vital role in glioma progression and serves as a potential prognostic biomarker in glioma patients, providing new targets for glioma therapy.
Similar content being viewed by others
Introduction
Glioma is the most common primary tumour of the central nervous system and accounts for 81% of malignant brain tumours1. Despite extensive research over the past decades, the prognosis of glioblastoma (grade IV glioma) patients remains poor. However, emerging evidence has suggested that lncRNAs play an important role in cancer pathophysiology, providing new insight into the genetic and molecular mechanisms of cancer2. lncRNAs are defined as transcripts longer than 200 nucleotides that are 5′ capped and 3′ poly-adenylated; however, this class of transcripts has limited coding potential3. lncRNAs exert their functions via diverse mechanisms, including cotranscriptional regulation, modulation of gene expression, scaffolding of nuclear or cytoplasmic complexes, and pairing with other RNAs4. There have been several studies focusing on the role of lncRNAs, including HOTAIR, MALAT1, and H19 5,6,7,8,9, in glioma pathogenesis. However, no article has been published regarding lncTCF7 in glioma.
lncTCF7 can promote liver cancer stem cell (CSC) self-renewal and tumour propagation through activation of Wnt signalling by recruiting the SWI/SNF complex to the TCF7 promoter10. lncTCF7 is highly expressed in HCC tissues and liver CSCs. In addition, lncTCF7 depletion significantly reduced the expression of pluripotent transcription factors Sox2, Nanog, and Oct4, remarkably impairing the generation of the fraction of CD13 + CD133 + cells (CSCs), while lncTCF7 overexpression enhances the tumorigenic capacity of liver CSCs. In this study, we aimed to explore the role of lncTCF7 in glioma progression.
To explore the role of lncTCF7 in glioma, we examined its expression level in 111 glioma tissues and 12 normal brain tissues. We found that the lncTCF7 expression levels are higher in high-grade gliomas than in low-grade gliomas and normal brain tissues. Additionally, lncTCF7 expression is associated with prognosis. Furthermore, the biological function of lncTCF7 in proliferation, cell cycle, and migration was examined in vitro, and its function in tumorigenicity was also investigated in the nude mouse model.
Results
Increased expression of lncTCF7 in Glioma
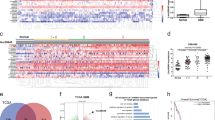
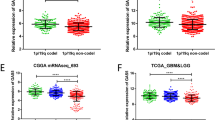
To determine whether lncTCF7 is involved in the development of glioma, we performed qRT-PCR to measure the expression levels of lncTCF7 in 111 glioma tissues. Compared with 12 samples of normal brain tissue, lncTCF7 expression in 111 glioma tissues was increased (P < 0.01, Fig. 1A). The expression of lncTCF7 was higher in high-grade glioma than in low-grade glioma (P < 0.001, Fig. 1A). Additionally, the percentage of high lncTCF7 expression was greater in high-grade glioma than in low-grade glioma (39.5% vs. 57.1%, respectively; Fig. 1B). Next, we examined lncTCF7 expression in glioma cell lines (Fig. 1C) and discovered much higher expression of lncTCF7 in U87 and U251 cell lines than in normal human astrocytes (NHA). Isolation of the nuclear and cytoplasmic fractions in U251 cells and U87 cells indicated that lncTCF7 was mainly located in the nucleus (Fig. 1D). Thus, lncTCF7 is upregulated in glioma and may serve as an oncogene in glioma development and progression.
Quantitative determination of lncTCF7 by qRT-PCR in glioma. (A) The relative expression level of lncTCF7 in glioma tissues (n = 111) and normal brain tissues (n = 12) was measured by qRT-PCR. (B) Percentage of glioma patients with high or low lncTCF7 expression level in different WHO grade groups. (C) The lncTCF7 level was evaluated in glioma cells and normal human astrocytes (NHA). NBM, normal brain tissue. (D) Quantification of lncTCF7 in fractionated U251 and U87 cell lysates by qRT-PCR. U6 RNA served as a positive control for nuclear gene expression, and GAPDH served as a control for cytoplasmic gene expression.
lncTCF7 expression is correlated with clinicopathological features in glioma patients
We next analysed the correlation between the expression of lncTCF7 and clinicopathological characteristics of glioma (Table 1). The clinicopathological characteristics of the 111 glioma patients presented in Table 1 showed that high expression of lncTCF7 was significantly correlated with higher WHO grades (III/IV) and larger tumour size (å 3 cm). However, lncTCF7 expression was not associated significantly with other parameters, such as age, gender and tumour location in glioma.
lncTCF7 is an independent unfavourable prognostic factor in glioma
To explore the prognostic value of lncTCF7 expression in glioma patients, we measured the association between lncTCF7 expression and patient survival using Kaplan–Meier analysis. Patients with relatively higher expression of lncTCF7 showed a worse prognosis than those with a lower expression (P < 0.01, Fig. 2). Thus, lncTCF7 is a negative prognostic factor, and high lncTCF7 expression is associated with poor overall survival (OS) in glioma. Furthermore, Cox regression analysis indicated that increased lncTCF7 expression is an independent prognostic factor for glioma patients (Table 2).
Knockdown of lncTCF7 inhibits glioma cell migration
To assess the potential functional role of lncTCF7, two siRNAs were synthesized, and the efficiency was measured in glioma cell lines by qRT-PCR (Fig. 3C). Wound healing assays and transwell assays were employed to detect the effects of si-lncTCF7 on the migratory ability of glioma cells (Fig. 3A,B). As shown in these two assays, knockdown of lncTCF7 reduced cellular migration in glioma cells dramatically, indicating that lncTCF7 is important for glioma cell migration. Similar results of wound healing assays were observed in U251 cells. Considering the tight relationship between the epithelial-mesenchymal transition (EMT) and cellular migration, several EMT-related protein markers were determined by western blotting (Fig. 3D). Knockdown of lncTCF7 significantly decreased the expression of N-cadherin and Vimentin, as well as increased the expression of E-cadherin, which mediates stable intercellular adhesions. These results suggest that knockdown of lncTCF7 suppressed the mesenchymal phenotype and inhibited migration through hindering EMT in glioma cells. Therefore, lncTCF7 may promote glioma progression by enhancing cell migration.
Knockdown of lncTCF7 inhibits the migration ability of glioma cells. (A) Wound healing assay and (B) transwell assay showing U251 and U87 cells with knockdown of lncTCF7, compared with negative controls. (C) The knockdown effects of lncTCF7 were measured by qRT-PCR in glioma cells transfected with siRNA#1, siRNA #2 or its negative controls. (D) Effects of lncTCF7 knockdown by siRNA on the protein expression of EMT markers in U251 and U87 cells.
Knockdown of lncTCF7 inhibits proliferation and induces G1 arrest in glioma cells in vitro
We further examined whether knockdown of lncTCF7 affects the proliferation of glioma cells. U251 and U87 cells were transfected with shRNA#1, and CCK-8 assays (Fig. 4A) were performed to measure the viability at each time point (0 h, 24 h, 48 h, 72 h). Knockdown of lncTCF7 reduced not only the CCK-8 absorbance but also the colony number in colony formation assays (Fig. 4B). Given that cell cycle progression is associated with proliferation, we next used propidium iodide (PI) staining to analyse the effect of lncTCF7 knockdown on glioma cell proliferation by flow cytometry (Fig. 4C). Compared with the respective control groups, lncTCF7 knockdown groups showed a decrease in the percentage of cells in S phase and a marked accumulation in the percentage of cells in the G0/G1 phase in U87 and U251 cells. Meanwhile, the sphere formation capacity of glioma stem cells (GSCs) was also impaired by lncTCF knockdown (Fig. 4E), suggesting that lncTCF7 may have a direct impact on the stemness and tumorigenicity of glioma.
Knockdown of lncTCF7 inhibits glioma cell growth and results in the G0/G1 cell cycle G1 arrest. U87 and U251 cells were transfected with lncTCF7 shRNA, and the proliferation were tested by CCK-8 assay (A) and colony formation assay (B,C). U87 and U251 cells were transiently transfected with lncTCF7 shRNAs or NC shRNAs for 48 h, and the cell cycle profiles were examined by flow cytometry. (D) Knockdown of lncTCF7 inhibited TCF7, Sox2 and CyclinD1 expression through hindering Wnt signalling. Full-length images of cropped gels from Figs 3D and 4D are shown in Supplementary Information file. (E) Knockdown of lncTCF7 inhibited sphere formation in glioma stem cells. The right panel presents the statistical results as mean as the means ± SD. Scale bar, 500 μm.
Furthermore, several cell cycle-related proteins were detected by western blotting (Fig. 4D), showing that CyclinD1, one of the direct target genes of Wnt/b-catenin signalling and responsible for G1/S transition11,12,13, was subsequently downregulated after lncTCF7 shRNA transfection. However, c-Myc expression did not decrease. In the meantime, p21, the cyclin-dependent kinase inhibitor which is downregulated by c-Myc, was not changed. These results implied that knockdown of lncTCF7 induced G1 arrest through the downregulation of CyclinD1 rather than c-Myc/p21 by suppressing the Wnt/b-catenin pathway. Moreover, Sox2, another target gene of Wnt/b-catenin signalling, was downregulated by the knockdown of lncTCF7, indicating that the proliferation reduction resulted from cylinD1 downregulation-mediated G1 arrest, as well as Sox2 downregulation-mediated self-renewal and tumorigenicity impairment. Nestin is a protein marker for neural stem cells and glioma stem cells14,15 and is associated with poor clinicopathological features and prognosis in glioma patients16. Knockdown of lncTCF7 decreased the expression of Nestin, indicating lncTCF7 is important for glioma cells to maintain stemness.
Knockdown of lncTCF7 reduces glioma growth in vivo
We further validated the effects of lncTCF7 downregulation in vivo by injecting U87 glioma cells into nude mice. As shown in Fig. 5A, down-regulation of lncTCF7 expression significantly reduced the tumour volume compared with that in the control group.
Hypoxia-induced IL-6 secretion increases lncTCF7 expression
Next, we investigated the mechanisms underlying the upregulation of lncTCF7 in glioma. Hypoxia is an important factor that promotes and maintains the stemness of CSCs and other stem cells17,18. Within GBM, distinct cancer stem cell niches are often located in the perivascular and hypoxic regions19. We detected lncTCF7 expression after hypoxia (1% O2) treatment in both U251 and U87 cell lines. Compared with the normoxia group, lncTCF7 expression was increased under the hypoxia condition in both cell lines (P < 0.01, Fig. 6A). Hypoxia-inducible factors (HIFs) are the main modulators during hypoxia. However, we failed to identify a HRE (HIF-response element) site on the promoter of lncTCF7.
Hypoxia-induced IL-6 secretion increases lncTCF7 expression. (A) Hypoxia increased lncTCF7 expression significantly. Under hypoxia (H) and normoxia (N) conditions, respectively, exogenous IL-6 (20 ng/ml) increased lncTCF7 expression in U87 and U251 cells, while IL6 antibody (Ab, 1 μg/ml) did not inhibit lncTCF7 expression completely. (B) Association between IL-6 and lncTCF7. IL-6 and lncTCF7 expression in 30 glioma tissue specimens was analysed by Fisher’s exact test. (C) Representative images of high-grade and low-grade glioma showed immunofluorescence staining of IL-6. (D) Binding of Stat3 onto the lncTCF7 promoter regions was examined via ChIP assay in glioma cells U251 and U87MG treated with or without IL-6 (20 ng/ml).
While hypoxia can also influence the downstream pathways through modulating inflammatory cytokine secretion20, interleukin 6 (IL6) was reported to induce lncTCF7 expression. IL-6 transcriptionally activated the expression of lncTCF7 in HCC cells by activating STAT321. In both normoxia and hypoxia, IL-6 induced lncTCF7 expression obviously (P < 0.01, Fig. 6A). Chromatin immunoprecipitation (ChIP) assay was performed to examine the interaction between the STAT3 and lncTCF7 promoter. Our results showed that the binding of STAT3 to the lncTCF7 promoter region was similar in both glioma cell lines, especially with the IL-6 (20 ng/ml) stimulus (Fig. 6D).
In addition, the IL-6-lncTCF7 axis was also validated in patient samples. As shown in Fig. 6B,C, a higher expression level of lncTCF7 was accompanied by higher IL-6 expression, and the association analysis revealed that lncTCF7 expression was associated with IL-6 expression (P < 0.01, Fig. 6B). However, the IL-6-neutralizing antibody was not sufficient to block lncTCF7 expression (Fig. 6A), indicating that other compensate pathways may be involved in lncTCF7 regulation and that anti-IL-6 therapy alone may promote resistance.
All together, these data showed that hypoxia-induced IL-6 secretion increases lncTCF7 expression, which is important for the migration and proliferation of glioma cells (Fig. 7).
Discussion
Advances in human transcriptome analysis has revealed that less than 2% of the transcriptional output encodes proteins, and the remaining 98% serve as different classes of non-coding RNAs22. lncRNA, as a new class of non-coding RNAs, has attracted the increased attention of researchers.
Recent studies have investigated the regulatory roles of lncTCF7 in liver and lung cancer progression. Yanying Wang et al. first found lncTCF7 as a key regulator of liver cancer stem cell self-renewal and tumour propagation through interfering with Wnt signalling10. Jun Wu et al. found that STAT3, which is activated by exogenous IL-6, can directly bind to the promoter regions of lncTCF7 to activate its transcription. They provided evidence that this aberrant IL-6/STAT3/ lncTCF7 signalling axis promotes HCC aggressiveness through EMT induction21. The role of lncTCF7 was also investigated in lung cancer. lncTCF7 was proven to promote the invasiveness and self-renewal of NSCLC cells23.
The Wnt signalling pathway controls a myriad of biological processes throughout development and the adult life of all animals24. Cell proliferation and migration are two aspects in which Wnt signalling plays a vital role.
Increased β-catenin can initiate the transcriptional activation of proteins such as CyclinD1 and c-Myc, which control the G1-to-S phase transition in the cell cycle25. In addition, Wnt signalling is an inducer of EMT and can promote cell migration by interfering with cell adhesion26. Therefore, it is not difficult to understand that Wnt signalling underlies a wide range of diseases in humans, especially cancer. Increased expression of Wnt ligand proteins were observed in the development of glioblastoma, oesophageal cancer and ovarian cancer27. In glioma, the Wnt signalling pathway is also one of the key signalling pathways for development and progression28,29. Meanwhile, recent studies have revealed that many lncRNAs could regulate the Wnt/b-catenin signalling pathway in several types of cancer. Yun Cui’s study showed that lncRNA SNHG1 promotes lung cancer tumorigenesis and progression via the miR-101-3p/SOX9/Wnt/b-catenin axis30. Zheying Zhang et al. revealed that lncRNA CASC11 promotes colorectal cancer cell proliferation and metastasis by associating with hnRNP-K and activating WNT/β-catenin signalling31. Hongying Liu et al. demonstrated that the knockdown of lncRNA UCA1 inhibits Wnt/b-catenin signalling and increases tamoxifen sensitivity in breast cancer32.
In the present study, we first found that lncTCF7 was overexpressed in glioma tissues and U251 and U87 cell lines, and its expression was positively correlated with advanced tumour stage and tumour size. Meanwhile, increased lncTCF7 expression predicted an unfavourable prognosis in glioma patients. Furthermore, the loss of function assay revealed that knockdown of lncTCF7 significantly inhibited glioma cell migration, proliferation and tumorigenicity through upregulating E-cadherin or suppressing TCF7/Sox2 and CyclinD1 expression.
Hypoxia is one of the important characteristics of solid tumours, including glioma, and is associated with worse patient survival and therapeutic resistance. Recently, dozens of lncRNAs have been reported to be dysregulated in hypoxia and exert diverse functions in tumour biology33. IL6 is a potent cytokine that regulates cell growth and differentiation and plays an important role in the immune response34,35. Increased or deregulated expression of IL6 significantly contributes to the pathogenesis of various human diseases, including tumours. Consistent with Jun Wu’s study21, we found that IL-6 induced lncTCF7 expression in glioma cells (Fig. 6). Based on our previous study20, we investigated whether the IL-6 blocking antibody can serve as a potential therapy targeting the IL-6/lncTCF7/Wnt pathway. However, the IL-6 antibody did not reduce lncTCF7 expression to basal levels (Fig. 6A), indicating the presence of another compensatory pathway involving other pro-inflammatory cytokines, such as macrophage migration inhibitory factor (MIF) and transforming growth factor beta (TGF-β). Further research into the regulation of lncTCF7 may provide a solution for anti-IL-6 therapy resistance.
In summary, we demonstrated that lncTCF7 is a negative prognostic factor in glioma and that down-regulation of lncTC7 expression inhibits glioma cell migration, proliferation and tumourigenicity potential by suppressing the Wnt signalling pathway. Furthermore, hypoxia-induced IL-6 secretion increases lncTCF7 expression. Therefore, the hypoxia/IL-6/lncTCF7 axis may be a potential target for glioma therapy.
Materials and Methods
Patients samples
This study was performed in accordance with relevant guidelines and was approved by the Ethics Committee of Qilu Hospital. Informed consent for both study participation and publication was obtained from all patients. One hundred eleven glioma tissues were obtained from patients with glioma who underwent initial surgery at Qilu Hospital. The specimens were snap frozen in liquid nitrogen for real-time PCR. Overall survival (OS) was defined as the interval between the dates of surgery and death, and 74 patients were followed up successfully.
RNA extraction and quantitative real-time PCR
Total RNA was extracted using TRIzol (Invitrogen) according to the manufacturer’s protocol. Total RNA was reverse transcribed into cDNA using the ReverTraAce qRT-PCR kit (FSQ-101; Toyobo). Real-time PCR was performed using the Lightcycler 480 SYBR GREEN I MASTER (Roche). The primers used for lncTCF7 and GAPDH were as follows: GAPDH-F: 5′-GCACCGTCAAGGCTGAGAAC-3′, R: 5′-TGGTGAAGACGCCAGTGGA-3′; lncTCF7-F: 5′-AGGAGTCCTTGGACCTGAGC-3′, R: 5′-AGTGGCTGGCATATAACCAACA-3′. All of the data for each sample were collected in triplicate. Standard curves were generated, and the relative amount of lncTCF7 was normalized to the amount of GAPDH (2−ΔCt).
lncTCF7 knockdown and transfection
Both siRNA and shRNA were used for lncTCF7 knockdown. We employed siRNA in migration assay and western blotting, while used shRNA to get a sustained knockdown effect in proliferation-related assay. The effective shRNAs were as follows10: lncTCF7 1#:5′-AGCCAACATTGTTGGTTAT-3′, 2# 5′-CACCTAGGTGCTCACTGAA-3′; the corresponding siRNAs were designed by Gene Pharma (Shanghai, China): lncTCF7 1#:5′-AGCCAACAUUGUUGGUUAUTT-3′, 2#:5′-CACCUAGGUGCUCACUGAATT-3′; the control siRNA (negative control, NC) was as follows: 5′-UUCUCCGAACGUGUCACGUTT-3′.
For siRNAs transfection, cells were seeded into 6-well plates at 106 cells per well. 24 h later, cells were transfected with siRNAs or the negative control sequences using Lipofectamine 2000 Transfection Reagent (11668–019, Life Technologies, CA, USA) according to the manufacturer’s instructions. The medium was replaced 6 h after the transfection. Transfected cells were harvested for other experiments after another 36 h.
For knockdown with shRNAs, shRNAs were constructed into the pcDNA3.1 plasmid. Cells were seeded in a 6-well plate at a concentration of 1.5 × 106 cells per well and allowed overnight growth to reach 70–80% confluency. Cells were then transfected with the mixture of 1 μg of plasmid DNA and 8 μl of Lipofectamine 2000 in 1.5 ml of serum-free medium. Fetal bovine serum (200 μl/well) was added at 6 h post-transfection. At 24 h post-transfection, the medium was replaced by complete medium and cultured up to 48 h after transfection for proliferation-related assay.
For xenograft assay, U87 cells transfected with pcDNA3.1- lncTCF7 shRNA were selected with G418. At 48 h post-transfection, medium containing G418 was added and the medium was replaced every 2 days. After 3 passages, U87 cells with stable knockdown of lncTCF7 were obtained for xenograft assay.
Cell proliferation and colony formation assay
Cell proliferation was detected by the Cell Counting Kit-8 (CCK-8). 48 h after transfection with shRNA, U251 or U87 (2.0 × 103) cells were plated into 96-well plates and were cultured for 1, 2, 3 or 4 days. Next, 10 μl of CCK-8 (Dojindo, Japan) in 100 μl of culture medium was added to each well and was incubated for a further 1 h at 37 °C. The absorbance was measured at a wavelength of 450 nm. For the colony formation assay, 2000 cells per well were seeded into a six-well plate after transfection. Approximately 10 days later, the colonies were fixed in 4% methanol for 30 min and were stained with 1% crystal violet overnight, and then the number of colonies was counted.
Cell cycle analysis
For cell cycle analysis, after the transfected cells described above were seeded on six-well plates for 48 h, the cells were collected by centrifugation and then were fixed with 70% ethanol at 4 °C overnight. Finally, the cellular DNA content was analysed by flow cytometry after propidium iodide (PI) staining (BD C6, BD Biosciences) using ModFit LT v2.0 software.
Chromatin immunoprecipitation (ChIP) assay
ChIP assays were performed using the EZ-Magna ChIP kit (Millipore, Temecula, CA, USA) according to the manufacturer’s instructions. In brief, U251 or U87MG cells were fixed in 1% formaldehyde for 10 min at room temperature and were sonicated to prepare chromatin fragments. The genomic DNA fragments were immunoprecipitated with antibodies against phosphor-Stat3(TyR705) (Cell Signaling Technology) or normal rabbit IgG at 4 °C overnight. After crosslinking reversal and DNA clean-up, purified precipitated DNA was analysed by quantitative real-time PCR using primers targeting the lncTCF7 promoter region: 5′-AGCCAGACAGAAGAGTGGA-3′and 5′-TGGGATGGGGATGTCAGAAC-3′.
Sphere formation assay
Glioma stem cells were cultured from the primary cells of one glioma patient sample in serum-free Neurobasal-A medium (Invitrogen) containing 1 × penicillin G and streptomycin supplemented with 20 ng/ml epithelial growth factor (Peprotech), 10 ng/ml fibroblast growth factor-2 (Peprotech), and B27 (Invitrogen) at 37 °C in 5% CO2. Additionally, these cells were identified as glioma stem cells by immunofluorescence with antibodies against Nestin and Sox2. After digestion into single cells, glioma stem cells with the scrambled sequence or shRNA were seeded onto six-well plates. After one week, the numbers of spheres were counted under a stereomicroscope.
Xenograft growth in nude mice
Animal studies were performed in accordance with the national guidelines of the Institutional Animal Care and Use Committee. For subcutaneous injection models, 106 U87 cells were implanted into mice (male BALB/c nude mice), aged 4 weeks, at the posterior dorsal flank region (n = 3 per group). The tumours were measured every other day. All experimental procedures were approved by the ethical committee of Qilu Hospital. The mice were maintained under standard conditions according to the institutional guidelines for animal care.
Statistical analysis
Data analyses were conducted using SPSS 20.0 and GraphPad Prism 6. Data are presented as the means ± SD. Chi-square test was used to evaluate the relationship between the lncTCF7 expression profiles and clinicopathological features. Comparisons between groups were analysed using Student’s test or ANOVA. Kaplan–Meier analysis was used to evaluate the probability of patient survival, and distributions were evaluated using the log-rank test. Cox regression models were used to calculate hazard ratios and identify factors affecting survival. For all statistical analyses, P < 0.05 was considered statistically significant.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Ostrom, Q. T. et al. The epidemiology of glioma in adults: a “state of the science” review. Neuro-oncology 16, 896–913, https://doi.org/10.1093/neuonc/nou087 (2014).
Schmitt, A. M. & Chang, H. Y. Long Noncoding RNAs in Cancer Pathways. Cancer cell 29, 452–463, https://doi.org/10.1016/j.ccell.2016.03.010 (2016).
Quinn, J. J. & Chang, H. Y. Unique features of long non-coding RNA biogenesis and function. Nature reviews. Genetics 17, 47–62, https://doi.org/10.1038/nrg.2015.10 (2016).
Ulitsky, I. & Bartel, D. P. lincRNAs: genomics, evolution, and mechanisms. Cell 154, 26–46, https://doi.org/10.1016/j.cell.2013.06.020 (2013).
Zhang, K. HOTAIR promotes glioblastoma cell cycle progression in an EZH2 dependent manner. oncotarget (2014).
Zhang, J. X. et al. HOTAIR, a cell cycle-associated long noncoding RNA and a strong predictor of survival, is preferentially expressed in classical and mesenchymal glioma. Neuro-oncology 15, 1595–1603, https://doi.org/10.1093/neuonc/not131 (2013).
Han, Y. et al. Tumor-suppressive function of long noncoding RNA MALAT1 in glioma cells by downregulation of MMP2 and inactivation of ERK/MAPK signaling. Cell death & disease 7, e2123, https://doi.org/10.1038/cddis.2015.407 (2016).
Ma, K. X. et al. Long noncoding RNA MALAT1 associates with the malignant status and poor prognosis in glioma. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine 36, 3355–3359, https://doi.org/10.1007/s13277-014-2969-7 (2015).
Jia, P. et al. Long non-coding RNA H19 regulates glioma angiogenesis and the biological behavior of glioma-associated endothelial cells by inhibiting microRNA-29a. Cancer letters 381, 359–369, https://doi.org/10.1016/j.canlet.2016.08.009 (2016).
Wang, Y. et al. The long noncoding RNA lncTCF7 promotes self-renewal of human liver cancer stem cells through activation of Wnt signaling. Cell stem cell 16, 413–425, https://doi.org/10.1016/j.stem.2015.03.003 (2015).
Tetsu, O. & McCormick, F. [beta]-Catenin regulates expression of cyclin D1 in colon carcinoma cells. Nature 398, 422–426 (1999).
Shtutman, M. et al. The cyclin D1 gene is a target of the beta-catenin/LEF-1 pathway. Proceedings of the National Academy of Sciences of the United States of America 96, 5522–5527 (1999).
Lin, S. Y. et al. Beta-catenin, a novel prognostic marker for breast cancer: its roles in cyclin D1 expression and cancer progression. Proceedings of the National Academy of Sciences of the United States of America 97, 4262–4266, https://doi.org/10.1073/pnas.060025397 (2000).
Dahlstrand, J., Collins, V. P. & Lendahl, U. Expression of the Class VI Intermediate Filament Nestin in Human Central Nervous System Tumors. Cancer research 52, 5334–5341 (1992).
Jin, X., Jin, X., Jung, J. E., Beck, S. & Kim, H. Cell surface Nestin is a biomarker for glioma stem cells. Biochemical and biophysical research communications 433, 496–501, https://doi.org/10.1016/j.bbrc.2013.03.021 (2013).
Lv, D. et al. Nestin Expression Is Associated with Poor Clinicopathological Features and Prognosis in Glioma Patients: an Association Study and Meta-analysis. Molecular neurobiology 54, 727–735, https://doi.org/10.1007/s12035-016-9689-5 (2017).
Keith, B. & Simon, M. C. Hypoxia-inducible factors, stem cells, and cancer. Cell 129, 465–472, https://doi.org/10.1016/j.cell.2007.04.019 (2007).
Mohyeldin, A., Garzón-Muvdi, T. & Quiñones-Hinojosa, A. Oxygen in Stem Cell Biology: A Critical Component of the Stem Cell Niche. Cell stem cell 7, 150–161, https://doi.org/10.1016/j.stem.2010.07.007 (2010).
Schonberg, D. L., Lubelski, D., Miller, T. E. & Rich, J. N. Brain tumor stem cells: Molecular characteristics and their impact on therapy. Molecular aspects of medicine 39, 82–101, https://doi.org/10.1016/j.mam.2013.06.004 (2014).
Xue, H. et al. A novel tumor-promoting mechanism of IL6 and the therapeutic efficacy of tocilizumab: Hypoxia-induced IL6 is a potent autophagy initiator in glioblastoma via the p-STAT3-MIR155-3p-CREBRF pathway. Autophagy 12, 1129–1152, https://doi.org/10.1080/15548627.2016.1178446 (2016).
Wu, J. et al. Long noncoding RNA lncTCF7, induced by IL-6/STAT3 transactivation, promotes hepatocellular carcinoma aggressiveness through epithelial-mesenchymal transition. Journal of experimental & clinical cancer research: CR 34, 116, https://doi.org/10.1186/s13046-015-0229-3 (2015).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108, https://doi.org/10.1038/nature11233 (2012).
Wu, J. & Wang, D. Long noncoding RNA TCF7 promotes invasiveness and self-renewal of human non-small cell lung cancer cells. Human cell, https://doi.org/10.1007/s13577-016-0147-5 (2016).
Clevers, H. & Nusse, R. Wnt/beta-catenin signaling and disease. Cell 149, 1192–1205, https://doi.org/10.1016/j.cell.2012.05.012 (2012).
Davidson, G. & Niehrs, C. Emerging links between CDK cell cycle regulators and Wnt signaling. Trends in cell biology 20, 453–460, https://doi.org/10.1016/j.tcb.2010.05.002 (2010).
Heuberger, J. & Birchmeier, W. Interplay of cadherin-mediated cell adhesion and canonical Wnt signaling. Cold Spring Harbor perspectives in biology 2, a002915, https://doi.org/10.1101/cshperspect.a002915 (2010).
Anastas, J. N. & Moon, R. T. WNT signalling pathways as therapeutic targets in cancer. Nature reviews. Cancer 13, 11–26, https://doi.org/10.1038/nrc3419 (2013).
Zhang, K., Zhang, J., Han, L., Pu, P. & Kang, C. Wnt/beta-catenin signaling in glioma. Journal of neuroimmune pharmacology: the official journal of the Society on NeuroImmune Pharmacology 7, 740–749, https://doi.org/10.1007/s11481-012-9359-y (2012).
Hu, B. et al. Epigenetic Activation of WNT5A Drives Glioblastoma Stem Cell Differentiation and Invasive Growth. Cell 167, 1281–1295 e1218, https://doi.org/10.1016/j.cell.2016.10.039 (2016).
Cui, Y. et al. Upregulated lncRNA SNHG1 contributes to progression of non-small cell lung cancer through inhibition of miR-101-3p and activation of Wnt/beta-catenin signaling pathway. Oncotarget, https://doi.org/10.18632/oncotarget.14854 (2017).
Zhang, Z. et al. Long non-coding RNA CASC11 interacts with hnRNP-K and activates the WNT/beta-catenin pathway to promote growth and metastasis in colorectal cancer. Cancer letters 376, 62–73, https://doi.org/10.1016/j.canlet.2016.03.022 (2016).
Tan, M. et al. Knockdown of Long Non-Coding RNA UCA1 Increases the Tamoxifen Sensitivity of Breast Cancer Cells through Inhibition of Wnt/β-Catenin Pathway. PloS one 11, e0168406, https://doi.org/10.1371/journal.pone.0168406 (2016).
Choudhry, H., Harris, A. L. & McIntyre, A. The tumour hypoxia induced non-coding transcriptome. Molecular aspects of medicine 47-48, 35–53, https://doi.org/10.1016/j.mam.2016.01.003 (2016).
Sansone, P. & Bromberg, J. Targeting the interleukin-6/Jak/stat pathway in human malignancies. Journal of clinical oncology: official journal of the American Society of Clinical Oncology 30, 1005–1014, https://doi.org/10.1200/JCO.2010.31.8907 (2012).
Kumari, N., Dwarakanath, B. S., Das, A. & Bhatt, A. N. Role of interleukin-6 in cancer progression and therapeutic resistance. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine 37, 11553–11572, https://doi.org/10.1007/s13277-016-5098-7 (2016).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (Nos. 81571284, 91542115, 81372719, and 81172403), Taishan Scholars of Shandong Province of China (No. ts201511093), and Promotive Research Fund for Excellent Young and Middle-aged Scientists of Shandong Province(BS2015SW008).
Author information
Authors and Affiliations
Contributions
Xiao G. and Xing G. performed the experiments and analysed the data. H.X., W.Q., Xiaofan G., J.Z., M.Q. and T.L. provided assistance in some experiments and helped with the data collection and analyses. G.L., J.S., L.D. and Q.L. conceived and designed the experiments and supervised the project. Xiao G., Xing G. and H.X. wrote the paper. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gao, X., Guo, X., Xue, H. et al. lncTCF7 is a negative prognostic factor, and knockdown of lncTCF7 inhibits migration, proliferation and tumorigenicity in glioma. Sci Rep 7, 17456 (2017). https://doi.org/10.1038/s41598-017-17340-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-17340-y
This article is cited by
-
Transcriptional expression of ZICs as an independent indicator of survival in gliomas
Scientific Reports (2021)
-
Signaling in and out: long-noncoding RNAs in tumor hypoxia
Journal of Biomedical Science (2020)
-
Hypoxia-induced lncRNA PDIA3P1 promotes mesenchymal transition via sponging of miR-124-3p in glioma
Cell Death & Disease (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.