Abstract
Belowground interactions between plant roots, mycorrhizal fungi and plant growth-promoting rhizobacteria (PGPR) can improve plant health via enhanced nutrient acquisition and priming of the plant immune system. Two wheat cultivars differing in their ability to form mycorrhiza were (co)inoculated with the mycorrhizal fungus Rhizophagus irregularis and the rhizobacterial strain Pseudomonas putida KT2440. The cultivar with high mycorrhizal compatibility supported higher levels of rhizobacterial colonization than the low compatibility cultivar. Those levels were augmented by mycorrhizal infection. Conversely, rhizobacterial colonization of the low compatibility cultivar was reduced by mycorrhizal arbuscule formation. Single inoculations with R. irregularis or P. putida had differential growth effects on both cultivars. Furthermore, while both cultivars developed systemic priming of chitosan-induced callose after single inoculations with R. irregularis or P. putida, only the cultivar with high mycorrhizal compatibility showed a synergistic increase in callose responsiveness following co-inoculation with both microbes. Our results show that multilateral interactions between roots, mycorrhizal fungi and PGPR can have synergistic effects on growth and systemic priming of wheat.
Similar content being viewed by others
Introduction
The plant immune system can be primed in response to specific signals, released either by biological or chemical agents1. This priming of defences provides plants with an augmented capacity to express basal resistance2, enabling a faster and stronger defensive response against pathogen attack. Symbiotic microorganisms such as arbuscular mycorrhizal fungi (AMF) and plant growth-promoting rhizobacteria (PGPR) can induce systemic resistance to both aerial and soil borne pathogens3,4,5,6. Moreover, the presence of both AMF and PGPR in the rhizosphere is known to be an important determinant of plant health in general7,8,9, that is, of the ability of a plant to carry out its physiological functions to the best of its genetic potential.
The regulation of root microbiome structure, and hence any beneficial effect on the plant, is extremely complex. Apart from environmental conditions (weather, soil nutrient status and physical structure, etc.), interactions between mycorrhizal fungi (from the Glomeromycota phylum), soil bacteria (from several genera such as Pseudomonas, Azotobacter, Bacilus, Azospirillum, etc.) and the plant play a crucial role shaping the microbiome community8,10. For example, host plant genotype strongly influences the extent to which AMF and PGPR colonize the host roots11,12,13. This effect is generally exerted through differences in the profile of plant metabolites exuded by the root that can attract specific organisms to the rhizosphere. For example, strigolactones are known to play a key role in recruiting AMF14 and benzoxazinoids have been shown to induce positive chemotaxis in the case of the PGPR Pseudomonas putida 15.
It can be misleading, however, to consider the chemistry of root exudation, and the signals encoded within, in isolation of the organisms that respond to those signals. This is because the interaction of a specific microorganism with the plant can alter the production of root metabolites16,17, consequently influencing the composition chemoattractants in the resultant root exudates. For example, AMF are responsible for what is known as the mycorrhizosphere effect: the enhanced microbial activity surrounding mycorrhizal roots16,18,19.
In this context, the priming of defences or the induction of resistance could be partially achieved by the combined action of both AMF and PGPR. Support for this hypothesis comes from the recent observation that AMF can enhance the accumulation of the benzoxazinoid, DIMBOA, in plant roots20,21 and thus may amplify positive chemotaxis by the PGPR P. putida, because benzoxazinoids act as semiochemicals for this bacteria15. As a consequence of this discovery, Cameron et al.10 proposed a model to explain how both AMF and PGPR could act together to define the Mycorrhiza-Induced Resistance (MIR), the phenomenon of AM induced protection against biotic stress22. This model postulates that in the earliest phase, the plant root exudes strigolactones that are known to facilitate increased AMF colonisation of the roots14,23. The plant immune system locally responds to the first stages of AMF colonisation, involving a transient activation of salicylic acid (SA)-dependent defences that can prime systemic plant tissues for this type of defence24. In order for the AMF to form a stable symbiosis, the fungus must locally supress these plant defences via hitherto unknown effectors25. Reprogramming of local plant defences, probably including induction of the SA-antagonist abscisic acid (ABA)26, results in changes to the composition of subsequent root exudates and possibly systemic priming of ABA-dependent defences. Finally, during the last phase, the mycorrhizosphere has established and recruited rhizobacteria that can systemically prime jasmonic acid (JA)- and ethylene-dependent defences27. In this way, the immune priming appears as a more complex process that involves a spatio-temporal interplay of different rhizosphere organisms (e.g. mycorrhiza and rhizobacteria) and corresponding host reactions.
The aim of the current study is starting to unravel these complex interactions and test the predictions of the Cameron et al.10 model by investigating: (i) the responses of plant growth to monoxenic colonisation by the plant-beneficial rhizobacterium P. putida KT2440 and the AMF species Rhizophagus irregularis (syn. Glomus intraradices) as well as under co-inoculated conditions; (ii) the effects of R. irregularis on the colonisation of the rhizoplane by P. putida (and vice versa); (iii) the degree of immune priming of plant tissues (assessed by callose deposition as a proxy) under monoxenic colonisation by P. putida KT2440 or R. irregularis as well as under co-inoculated conditions; (iv) the influence of wheat genotype on the root colonisation by P. putida KT2440 and R. irregularis
Results
AMF colonisation
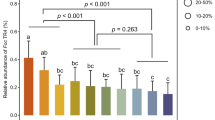
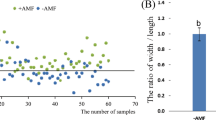
Colonisation of wheat roots by R. irregularis was estimated by calculation of two frequently used indices28: frequency of colonisation of the root system (F) and arbuscule abundance in the root system (A). Regarding frequency of colonisation, the cultivar ‘Mercato’ (which exhibited higher mycorrhizal colonisation), contained more fungal hyphae per unit root length at 14 days than the cultivar ‘Avalon’ (which exhibited lower mycorrhizal colonisation) (Fig. 1A). By 42 days, there was no difference in the fungal hyphae per unit root length between either cultivar (Fig. 1B). Co-inoculation with P. putida KT2440 did not affect the amount of fungal hyphae per unit root length in either cultivar at either time point.
Estimation of colonisation of wheat roots by Rhizophagus irregularis according to Trouvelot et al.28. Two indices were used: frequency of colonisation of the root system (F) at 14 (A) and 42 (B) days post inoculation, and arbuscule abundance in the root system (A) at 14 (C) and 42 (D) days post inoculation. Asterisk indicates significant differences within the same cultivar (Tukey; P < 0.05). Shown are average values (n = 10; ±standard error).
A differential response was also found when arbuscule abundance was measured with virtually no arbuscules being observed in either cultivar at 14 days (Fig. 1C). However by 42 days arbuscules were much more frequently observed with ‘Mercato’ significantly containing a 60% more number of arbuscules per unit root length when compared with ‘Avalon’ (Fig. 1D). At 42 days, P. putida KT2440 significantly reduced the number of arbuscules per unit root length by half in ‘Mercato’. A reduction in number of arbuscules per unit root length was also observed in ‘Avalon’ but this was not significant.
PGPR colonisation
In the cultivar ‘Mercato’, there were significantly more culturable bacteria on the rhizoplane of R. irregularis-infected plants in comparison to non-mycorrhizal plants at 14 days (Fig. 2A, Table 1). By contrast, no other bacteria were detected in the rhizoplane when GFP-tagged P. putida KT2440 was added to the growth substrate. At 14 days, P. putida KT2440 cells were recovered from the rhizoplane in high numbers after being added to the rhizotron at day 0, which was significantly increased by the presence of R. irregularis. By 42 days, the bacterial titre of P. putida KT2440 had declined dramatically, and was significantly lower than numbers of other (non-GFP expressing) culturable bacteria on the rhizoplane of ‘Mercato’. Similarly as for P. putida KT2440, the titre of these other culturable bacteria was significantly enhanced by the presence of R. irregularis when P. putida KT2440 was absent. Although rhizoplane colonization by P. putida KT2440 decreased significantly over time, the proportion of P. putida KT2440 remained significantly high for mycorrhizal roots of ‘Mercato’ relative to the other non-GFP-expressing bacteria at 42 days.
Estimation of bacterial colonisation of wheat roots by Pseudomonas putida KT2440 and other spontaneous bacteria. (A) Number of colony forming units (CFU) per gram of root of wheat cultivar ‘Mercato’ at 14 and 42 days post inoculation. Green bars correspond to GFP-tagged P. putida KT2440 colonies and red bars to unidentified spontaneous growing bacteria. (B) Idem as (A) but for wheat cultivar ‘Avalon’. Shown are average values (n = 10; ±standard error).
In comparison to ‘Mercato’, the rhizoplane of the cultivar ‘Avalon’ supported substantially lower colonization by GFP-expressing P. putida KT2440, but higher colonization by other (non-GFP expressing) culturable bacteria at 14 days (Fig. 2B, Table 1). Furthermore, bacterial colonization of the ‘Avalon’ rhizoplane was un-affected by co-colonization of R. irregularis. By 42 days, no substantial numbers of bacterial colonies (both GFP-expressing P. putida KT2440 and other culturable bacteria) could be isolated from the rhizoplane of ‘Avalon’.
Plant development
Inoculation with R. irregularis and/or P. putida KT2440 had a significant effect on the fresh weight (FW) of the shoots and roots and the root:shoot ratio of wheat plants, which varied both with time and wheat cultivar (Fig. 3, Table 2). Both R. irregularis and P. putida KT2440 increased shoot and root biomass at 14 and 42 days in ‘Mercato’ (Fig. 3A, Table 2). In ‘Avalon’ however, no significant effects of R. irregularis were seen at 14 days for shoots, but P. putida KT2440 caused a significant increase in shoot biomass (Fig. 3B, Table 2). For roots of ‘Avalon’, only R. irregularis caused a significant increase in biomass at 14 days. The combination of R. irregularis and P. putida KT2440 for both roots and shoots was additive at 14 and 42 days for ‘Mercato’. By contrast, ‘Avalon’ shoot biomass was supressed by the combination of R. irregularis and P. putida KT2440 relative to inoculation with P. putida KT2440 alone.
Fresh weight and root:shoot ratio of wheat with different inoculations. (A) Fresh weight of shoot (green) and root (orange) of ‘Mercato’ cultivar at 14 and 42 days post inoculation. Data at 14 days post inoculation are referred to left axis and data at 42 days post inoculation are referred to the right axis. (B) Idem as (A) for ‘Avalon’ cultivar. (C) Root:shoot ratio of ‘Mercato’ cultivar at 14 (white) and 42 (grey) days post inoculation. (D) Idem as (C) for ‘Avalon’ cultivar. Shown are average values (n = 10; ±standard error).
The root:shoot ratio of ‘Mercato’, was significantly affected by the presence of P. putida KT2440 (Fig. 3C, Table 2). This was observed at both 14 and 42 days (Fig. 3C, Table 2). The same trend was observed in ‘Avalon’, at 14 days (Fig. 3D, Table 2). However, by 42 days, root:shoot ratio was significantly higher in plants co-infected with R. irregularis and P. putida KT2440 than plants inoculated with P. putida KT2440 alone. All treatments resulted in a lower root:shoot ratio than the control ‘Avalon’ plants at 42 days.
Induction of callose deposition
A calibration curve was developed prior to the experiment, in order to assess the optimal concentration of chitosan for measuring priming of callose deposition. Figure 4 shows that 0.01% (w/v) chitosan solution is the lowest concentration that does not induce callose deposition in un-primed plants, which was selected for further leaf infiltrations to test priming of callose deposition by microbial root inoculations.
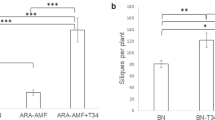
Wheat cultivar and inoculation with R. irregularis and P. putida KT2440 influenced chitosan-induced callose deposition (Fig. 5, Table 3). At 14 days, infiltration of leaves with the buffer control solution did not induce callose deposition in any treatment of either ‘Mercato’ or ‘Avalon’. Similarly, infiltration with the relatively low concentration of 0.01% chitosan did not induce callose deposition in control-treated plants of either cultivar at 14 days. Chitosan treatment of leaves from R. irregularis-colonized and P. putida KT2440 colonised ‘Mercato’ showed relatively low levels of callose deposition, which was of comparable intensity. Interestingly, callose deposition in ‘Mercato’ that had been co-colonised by R. irregularis and P. putida KT2440 showed 10 and 5-fold higher levels of callose deposition compared to leaves from plants colonised by R. irregularis or P. putida KT2440 alone, respectively (Fig. 5A, Table 3). By contrast, co-colonised ‘Avalon’ plants showed similarly low levels of chitosan-induced callose as plants that had been colonised by R. irregularis or P. putida KT2440 alone (Fig. 5B, Table 3). At 42 days, infiltration with 0.01% chitosan failed to induce detectable levels of callose deposition for all cultivar-treatment combinations (data not shown).
Induction of callose deposition in wheat leaves. (A) Percentage of fluorescent pixels of callose relative to the total number of plant material pixels in cultivar ‘Mercato’ at 14 days post inoculation. Callose deposition occurred only in leaves infiltrated with 0.01% chitosan, but not in buffer-infiltrated mock leaves. (B) Idem as (A) for cultivar ‘Avalon’. Shown are average values (n = 10; ±standard error).
Discussion
The rhizosphere and its associated microbiome are key drivers of the health and productivity of crop plants8,29. To understand the underlying mechanisms, we investigated the combined effects of a mycorrhizal fungus (R. irregularis) and plant growth-promoting rhizobacterium (P. putida KT2440) on immune responsiveness and above- and below-ground growth of two cultivars of wheat, ‘Mercato’ and ‘Avalon’ that differed in their ability to form mycorrhizal associations.
Both the AMF, R. irregularis, and the PGPR, P. putida KT2440, exerted a positive effect on wheat growth, however the two microorganisms differed in their relative effects on root and shoot biomass. In isolation, growth promotion by R. irregularis and P. putida is a well-characterised phenomenon, usually attributed to several factors, such as maximizing nutrient and water uptake in the case of AMF7, increasing availability of nutrients and/or production of phytohormones by PGPR30. In the absence of P. putida, and in line with previous reports31, R. irregularis did not change shoot growth relative to shoot root. In contrast to R. irregularis, but in agreement with previous studies of PGPR effects on root:shoot ratios32, root colonisation by P. putida in the absence R. irregularis led to preferential shoot growth, presumably as a reflection of a phytohormonal response related to increased root branching and more efficient nutrient uptake by plant roots. However, when plants were colonised by both AMF and PGPR, the root:shoot ratio was more similar to plants that had been inoculated with AMF only. This effect was similar for both cultivars at 14 and 42 days after inoculation. An intriguing result was the strong effect of PGPR on Avalon biomass despite the low bacterial abundance. A plausible explanation for this could again be related to the composition of root exudates, because such exudates are known to regulate bacterial gene expression33. For example, bacterial biosynthesis of the auxin indole-3-acetic acid (IAA) depends on the supply of exogenous tryptophan, and its presence in exudates from plant roots varies with genotype34. Consequently ‘Avalon’ and ‘Mercato’ may differ in the amount of compounds acting as precursors for the synthesis of plant hormones by the PGPR released in the rhizosphere, and thus affecting the behaviour of P. putida regarding plant growth promotion. This hypothesis warrants further investigation.
Based on our data, and in agreement with earlier studies on other plant species8, colonisation by both PGPR and AMF provides wheat with optimal growth conditions. It then follows that plants are capable of signalling to soil microbes, in order to facilitate a beneficial microbial community structure in the rhizosphere10. Until relatively recently, the chemical messengers deployed by plants into the rhizosphere remained poorly understood. A significant breakthrough came with the realisation that strigolactones induced branching in mycorrhizal hyphae and in so doing, enhancing root colonisation14. Strigolactones, coupled to reciprocal induction of MYC factors, lipochitooligosaccharides produced by the fungus that prime root morphology and chemistry for colonisation by AMF35, are now known to represent early regulators of AM symbiosis in nature.
Likewise, plants can produce chemical signals that elicit behavioural response of PGPRs10,15. For example, benzoxazinoid metabolites in root exudates of maize have been shown to induce positive chemotaxis in P. putida 15. Recent evidence suggests that benzoxazinoid production in the roots is enhanced by mycorrhizal infection20,21. Provided such changes in root benzoxazinoid composition result in enhanced exudation of these chemicals, they could promote a new equilibrium in microbiome composition16,19, commonly referred to as the “mycorrhizosphere effect”10,18. Based on this concept, we predicted that wheat plants colonised by AMF should attract greater numbers of resistance-inducing P. putida KT2440 than non-mycorrhizal plants.
Using GFP-tagged P. putida KT2440 as a marker for rhizoplane colonization by beneficial rhizobacteria, we recovered circa double the number of green-fluorescent CFUs from the rhizoplane of the cultivar ‘Mercato’ (which exhibited higher mycorrhizal colonisation) at 14 days, while this effect was not present in the cultivar ‘Avalon’ (which exhibited lower mycorrhizal colonisation). Other culturable rhizobacteria (e.g. non-GFP expressing) were also enhanced by the presence of mycorrhiza in ‘Mercato’ at 14 days. By 42 days, GFP-tagged P. putida numbers had declined in the rhizoplane of ‘Mercato’. This is unsurprising considering that benzoxazinoids are known to decline in root exudates of cereals with time post-germination15,36. Intriguingly, lower numbers of fluorescent CFUs were recovered from the rhizoplane of the less mycorrhizal cultivar ‘Avalon’ at 14 days. Furthermore, rhizoplane bacterial numbers from ‘Avalon’ dropped to almost undetectable levels by 42 days post-inoculation. Whether this is a function of a generic reduction in root exudates or, more specifically, a reduction in exudation of strigolactones and benzoxazinoids in ‘Avalon’ warrants further investigation. This question becomes especially relevant when considering that such specific changes in root exudation chemistry could account for the low mycorrhizal and low bacteria colonization phenotypes that we have observed in the ‘Avalon’ cultivar. It is also possible that ‘Avalon’ produces a water-soluble metabolite capable of inhibiting P. putida KT2440 growth. For example, catechin and compounds mimicking N-acyl homoserine lactone have been shown to interfere and inhibit bacterial quorum-sensing signals and hence, affecting biofilm formation and population density37,38.
Our rhizotron studies also revealed that P. putida significantly reduced arbuscule density in roots of ‘Mercato’. The mechanistic basis for this remains unclear. However, there are two distinct but mutually non-exclusive explanations: 1) P. putida is known to solubilise complex inorganic P (for example see Das et al.39), thereby increasing P supply to the host plant. It is also well established that high cellular P concentrations down-regulate strigolactone production40. This, in turn, would reduce AMF root colonisation, since hyphal-branching directly influences the degree of host colonisation by AMF14,23. 2) P. putida KT2440 has been shown to prime plant defences and induce systemic resistance41,42,43, which could dampen AMF infection. Successful establishment of the AM symbiosis is linked to down-regulation of local defences, most likely as a consequence of AMF effectors that are released during the early stages of the interaction. Since P. putida KT2440 colonization occurred before AMF colonization (Figs 1 and 2), it is plausible that the corresponding immune priming by P. putida antagonises the action of susceptibility-inducing effectors by AMF.
In addition to the effects of AMF and PGPR on rhizoplane colonization by other microbes and plant growth, AMF and PGPR can also systemically prime their host plants for augmented plant defence. This induced resistance cannot simply be attributed to improved plant nutrition as a result of AMF and PGPR colonisation of the roots44, but is rather a function of modulation of the host immune system by the symbionts10,45. This mycorrhiza-induced resistance (MIR) shares characteristics with pathogen-induced systemic-acquired resistance (SAR), such as priming of salicylic SA-dependent genes, and rhizobacterial induced systemic resistance (ISR), often characterized as priming of JA-dependent defences and cell wall defences46. This led Cameron et al.10 to propose that MIR might in fact be the additive product of AMF-induced and PGPR-induced priming mechanisms.
To test this hypothesis, we used callose deposition as a well-characterized marker for plant immune responsiveness to pathogen-associated molecular patterns, such as chitosan47. Callose, a β-glucan polysaccharide, facilitates reinforcement of the cell wall against attack and is thought to act as a matrix for the immobilisation of plant defence components, such as phytoalexins, reactive oxygen species, and cell-wall reinforcing enzymes (e.g. peroxidases)47,48. Leaf infiltration with 0.01% chitosan, which does not induce callose in un-primed plants (Fig. 4), resulted in detectable levels of callose deposition (quantified by epifluorescence) in both ‘Mercato’ and ‘Avalon’ at 14 days after inoculation with either R. irregularis, or P. putida KT2440. However, upon colonisation by both the AMF and PGPR, callose deposition in ‘Mercato’ was 10-fold greater than the AMF treatment alone, and 5-fold greater than the PGPR treatment alone. Hence, co-colonization by PGPR and AMF leads to synergistic levels of immune priming in AMF-responsive wheat. Conversely, callose deposition of co-inoculated ‘Avalon’ plants (which exhibited lower mycorrhizal colonisation) had no synergistic effect on callose deposition, suggesting fundamental differences in the signalling pathway leading to systemic immune priming. A threshold in PGPR population density has been shown as necessary for inducing effective resistance against phytopathogens49, so a minimum bacterial population could also be needed in order to synergistically prime the plant’s defences. Nevertheless, future work should be performed in order to completely validate this hypothesis using complementary techniques that allow identification of primed responses.
In conclusion, both AMF and PGPR can act additively on plant growth promotion, presumably due to complementary impacts on soil nutrient solubilisation and uptake. Moreover, co-colonization by AMF and PGPR appeared to have strongly synergistic effects on priming of host immunity, suggesting involvement of multiple defence pathways. This then supports a mechanistic explanation for the observations that MIR can be effective against both biotrophic and necrotrophic pathogens, which are resisted by different types of plant basal defences50,51,52.
Materials and Methods
Plant material and cultivation
We used two cultivars of wheat (Triticum aestivum L.) ‘Mercato’ and ‘Avalon’ that we had previously determined to differ in the extent of mycorrhizal colonisation seen in their root; Mercato exhibiting high colonisation levels and Avalon exhibiting low colonisation levels (and see results). Seeds were germinated in the dark at 20 °C and high humidity in between two rockwool blocks following the procedure described by Gurney et al.53. Seedlings were transferred after 7 days to rhizotrons built with 250 × 250 mm square petri dishes filled with sterile vermiculite, onto which was placed a 35 μm mesh (Plastok Associated, Birkenhead, UK)53. This kept the roots growing down between the mesh and the dish cover, allowing root observation during the experiment. Rhizotrons were covered with a black plastic sleeve for preventing light to reach the roots. Plants were grown in a controlled environment greenhouse room, with 12 h photoperiod and a day:night temperature of 20:18 °C. Rhizotrons were irrigated daily with 30 ml of 40% Long Ashton solution54 but lacking phosphorus. Fourteen days after sowing, plants were inoculated either with in vitro cultured spores of Rhizophagus irregularis isolate 0955, Pseudomonas putida KT2440 or both, leaving non-inoculated plants as control. Controls were treated with matching volumes of water and buffer. MgSO4 buffer does not contain P and MSR medium contains 0.41 g/l KH2PO4, so considering dilution with water and the rhizotrons volume, there were a final amount of 0.728 ppm of PO4 per rhizotron (available P in soil is usually around 20–50 ppm). At sampling dates (14 and 42 days post inoculation) fresh weight was determined for each plant. For further analysis and measurements, at least six plants per treatment and sampling date were used (totalling 96 plants).
AMF species and cultivation
Rhizophagus irregularis isolate 09 was used as inoculum for AMF colonisation studies. The fungus was maintained and replicated using a monoxenic culture with transformed carrot (Daucus carota L.) roots according to the method established by Declerck et al.56. Infected roots were kept growing in 150 mm diameter petri dishes containing modified Strullu-Romand (MRS) agar medium. For inoculation of wheat plants in rhizotrons, the content of a petri dish was blended and mixed with 50 ml sterile distilled water. The concentration of spores was measured and adjusted to approximately 500–750 spores per ml, and 5 ml were added on the root of each plant in the rhizotrons.
Bacterial strain and cultivation
A green fluorescent protein (GFP)-tagged Pseudomonas putida KT2440 strain previously described by Neal et al.15 was used for inoculation experiments. Stocks were stored at −80 °C and fresh cultures were used for inoculation. Bacteria were grown overnight in agitation (150 rpm) at 28 °C in M9 minimal salt medium supplemented with 0.1% glucose. Plants were inoculated with 3 ml of 108–109 bacterial suspensions in 10 mM MgSO4 buffer on the root surface.
AMF and PGPR colonisation assays
Fourteen days after sowing, wheat plants were inoculated as described previously, giving four different treatments: non-inoculated control, AMF (R. irregularis) inoculated, PGPR (P. putida KT2440) inoculated, AMF and PGPR inoculated. Fourteen and forty-two days after inoculation (i.e. twenty-eight and fifty-six days after sowing), root samples were collected for assessing mycorrhizal and rhizobacterial colonisation.
For AMF quantification, root samples were fixed in 50% ethanol for at least 24 h at 4 °C. Then, roots were immersed in 10% KOH and subjected to an autoclave cycle (15 min, 121 °C, 15 psi), following stain with trypan blue during 20 min (0.4 g trypan blue + 50 g phenol + 50 ml lactic acid + 100 ml glycerol + 50 ml distilled water). After destaining 30 min in 50% glycerol, roots were washed, squashed and mounted on slides with 50% glycerol. At least ten fragments of 1 cm length were mounted on each slide for estimation of mycorrhizal colonisation according to Trouvelot et al.28 (see also Dodd et al.57). This method allows for the calculation of the frequency of colonisation of the root system (F) and arbuscule abundance in the root system (A).
Rhizobacteria colonisation was estimated by shaking root fragments during 20 min at 200 rpm in 10 mM MgSO4 buffer (10 ml per root g) and plating several dilutions onto petri dishes containing M9 minimal salt medium supplemented with 0.1% glucose. Plates were incubated in darkness at 28 °C during 48 h. Total number of GFP-expressing P. putida KT2440 colonies were determined using a transilluminator, whereas the other non-fluorescent bacteria colonies were counted under natural light. Data were expressed as number of colony forming units (CFU) per g of root fresh weight.
Callose quantification
Fourteen and forty-two days after inoculation, samples from the youngest expanded leaf were collected from every plant for quantification of callose deposition. Leaf segments (2 cm) were infiltrated with 0.01% chitosan solution in 0.2% acetic acid (pH 5.7), and mock treatments with 0.2% acetic acid solution only. Vacuum infiltration was performed at −60 kPa for 5 min, and then the leaf segments were incubated on moistened filter paper in sealed petri dishes and collected for callose staining 24 h after infiltration. Leaf segments were fixed and destained in 100% ethanol at 4 °C for at least 48 h until tissue turned translucent. Then, they were washed and incubated in 0.07 M phosphate buffer (pH = 9) for 30 minutes, and finally stained overnight in darkness with 0.05% aniline blue in phosphate buffer. Samples were observed under epifluorescence (UV excitation: 330–385 nm; emission: 420 nm) using a BX51 Olympus microscope (Olympus Optical Ltd, London, UK) and pictures were taken with a connected DP71 digital camera. Callose was quantified by the number of deposition pixels relative to the total number of plant material pixels, using GIMP 2.8 software.
Statistical analysis
Differences between treatments with microorganisms were analysed with two-ways ANOVA considering two factors (AMF inoculation and PGPR inoculation) using Statistix software (version 8.1). Variables were log-transformed when necessary to meet assumptions of normality and homogeneity. At least six replicates per treatment were used.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Walters, D. R., Ratsep, J. & Havis, N. D. Controlling crop diseases using induced resistance: challenges for the future. J. Exp. Bot. 64, 1263–1280 (2013).
Ahmad, S., Gordon-Weeks, R., Pickett, J. & Ton, J. Natural variation in priming of basal resistance: from evolutionary origin to agricultural exploitation. Mol. Plant Pathol. 11, 817–827 (2010).
Cordier, C., Pozo, M. J., Barea, J. M., Gianinazzi, S. & Gianinazzi-Pearson, V. Cell defense responses associated with localized and systemic resistance to Phytophthora parasitica induced by an arbuscular mycorrhizal fungus. Mol. Plant Microbe In. 11, 1017–1028 (1998).
Jetiyanon, K. & Kloepper, J. W. Mixtures of plant growth promoting rhizobacteria for induction of systemic resistance against multiple plant diseases. Biol. Control 24, 285–291 (2002).
Lioussanne, L. The role of the arbuscular mycorrhiza-associated rhizobacteria in the biocontrol of soilborne phytopathogens. Span. J. Agric. Res. 8(S1), 51–61 (2010).
D’Alessandro, M. et al. Volatiles produced by soil-borne endophytic bacteria increase plant pathogen resistance and affect tritrophic interactions. Plant Cell Environ. 37, 813–826 (2013).
Cameron, D. Arbuscular mycorrhizal fungi as (agro)ecosystem engineers. Plant Soil 333, 1–5 (2010).
Berendsen, R. L., Pieterse, C. M. J. & Bakker, P. A. H. M. The rhizosphere microbiome and plant health. Trends Plant Sci. 17, 478–486 (2012).
Berta, G. et al. Maize development and grain quality are differentially affected by mycorrhizal fungi and growth-promoting psedomonad in the field. Mycorrhiza 24, 161–170 (2014).
Cameron, D. D., Neal, A. L., van Wees, S. C. M. & Ton, J. Mycorrhiza-induced resistance: more than the sum of its parts? Trends Plant Sci. 18, 539–545 (2013).
Azcon, R. & Ocampo, J. A. Factors affecting the vesicular-arbuscular infection and mycorrhizal dependency of thirteen wheat cultivars. New Phytol. 87, 677–685 (1981).
Aira, M., Gómez-Brandón, M., Lazcano, C., Baath, E. & Domínguez, J. Plant genotype strongly modifies the structure and growth of maize rhizosphere microbial communities. Soil Biol. Biochem. 42, 2276–2281 (2010).
An, G. H. et al. How does arbuscular mycorrhizal colonization vary with host plant genotype? An example based on maize (Zea mays) germplasms. Plant Soil 327, 441–453 (2010).
Akiyama, K., Matsuzaki, K. & Hayashi, H. Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi. Nature 435, 824–827 (2005).
Neal, A. L., Ahmad, S., Gordon-Weeks, R. & Ton, J. Benzoxazinoids in root exudates of maize attract Pseudomonas putida to the rhizosphere. PLOS One 7, e35498, https://doi.org/10.1371/journal.pone.0035498 (2012).
Marschner, P., Crowley, D. E. & Higashi, R. M. Root exudation and physiological status of a root-colonizing fluorescent pseudomonad in mycorrhizal and non-mycorrhizal pepper (Capsicum annuum L.). Plant Soil 189, 11–20 (1997).
Laparre, J. et al. Combining metabolomics and gene expression analysis reveals that propionyl- and butyryl- carnitines are involved in late stages of arbuscular mycorrhizal symbiosis. Mol. Plant 7, 554–566 (2014).
Linderman, R. G. Mycorrhizal interactions with the rhizosphere microflora: the mycorrhizosphere effect. Phytopathol. 78, 366–371 (1988).
Gupta Sood, S. Chemotactic response of plant-growth-promoting bacteria towards roots of vesicular-arbuscular mycorrhizal tomato plants. FEMS Microbiol. Ecol. 45, 219–227 (2003).
Song, Y. Y. et al. Induction of DIMBOA accumulation and systemic defense responses as a mechanism of enhanced resistance of mycorrhizal corn (Zea mays L.) to sheath blight. Mycorrhiza 21, 721–731 (2011).
Walker, V. et al. Variation of secondary metabolite levels in maize seedling roots induced by inoculation with Azospirillum, Pseudomonas and Glomus consortium under field conditions. Plant Soil 356, 151–163 (2012).
Pozo, M. J. & Azcón-Aguilar, C. Unraveling mycorrhiza-induced resistance. Curr. Opin. Plant Biol. 10, 393–398 (2007).
Besserer, A. et al. Strigolactones stimulate arbuscular mycorrhizal fungi by activating mitochondria. PLoS Biol. 4, e226, https://doi.org/10.1371/journal.pbio.0040226 (2006).
Gallou, A., Mosquera, H. P. L., Cranenbrouck, S., Suárez, J. P. & Declerck, S. Mycorrhiza induced resistance in potato plantlets challenged by Phytophthora infestans. Physiol. Mol. Plant Pathol. 76, 20–26 (2011).
Kloppholz, S., Kuhn, H. & Requena, N. A secreted fungal effector of Glomus intraradices promotes symbiotic biotrophy. Curr. Biol. 21, 1204–1209 (2011).
Ton, J., Flors, V. & Mauch-Mani, B. The multifaceted role of ABA in disease resistance. Trends Plant Sci. 14, 310–317 (2009).
Van der Ent, S., Van Wees, S. C. M. & Pieterse, C. M. J. Jasmonate signaling in plant interactions with resistance-inducing beneficial microbes. Phytochemistry 70, 1581–1588 (2009).
Trouvelot, A., Kough, J. L. & Gianinazzi-Pearson, V. Mesure du taux de mycorhization VA d’un système radiculaire. Recherche de méthodes d’estimation ayant une signification fonctionnelle in Physiological and genetical aspects of mycorrhizae (eds Gianinazzi-Pearson, V. & Gianinazzi, S.) 217–221 (INRA Press, 1986).
Chaparro, J. M., Sheflin, A. M., Manter, D. K. & Vivanco, J. M. Manipulating the soil microbiome to increase soil health and plant fertility. Biol. Fert. Soils 48, 489–499 (2012).
Vessey, J. K. Plant growth promoting rhizobacteria as biofertilizers. Plant Soil 255, 571–586 (2003).
Maherali, H. Is there an association between root architecture and mycorrhizal growth response? New Phytol. 204, 192–200 (2014).
Bashan, Y. & Dubrovsky, J. G. Azospirillum spp. participation in dry matter partitioning in grasses at the whole plant level. Biol. Fert. Soils 23, 435–440 (1996).
Vacheron, J. et al. Plant growth-promoting rhizobacteria and root system functioning. Front. Plant Sci. 4, 356, https://doi.org/10.3389/fpls.2013.00356 (2013).
Kamilova, F. et al. Organic acids, sugars, and L-tryptophane in exudates of vegetables growing on stonewool and their effects on activities of rhizosphere bacteria. Mol. Plant Microbe In. 193, 250–256 (2006).
Maillet, F. et al. Fungal lipochitooligosaccharide symbiotic signals in arbuscular mycorrhiza. Nature 469, 58–63 (2011).
Huang, Z., Haig, T., Wu, H., An, M. & Pratley, J. Correlation between phytotoxicity on annual ryegrass (Lolium rigidum) and production dynamics of allelochemicals within root exudates of an allelopathic wheat. J. Chem. Ecol. 29, 2263–2279 (2003).
Teplitski, M., Robinson, J. B. & Bauer, W. D. Plants secrete substances that mimic bacterial N-acyl homoserine lactone signal activities and affect population density-dependent behaviors in associated bacteria. Mol. Plant Microbe In. 13, 637–648 (2000).
Vandeputte, O. M. et al. Identification of catechin as one of the flavonoids from Combretum albiflorum bark extract that reduces the production of quorum-sensing-controlled virulence factors in Pseudomonas aeruginosa PAO1. Appl. Environ. Microbiol. 6, 243–53 (2010).
Das, K., Katiyar, V. & Goel, R. ‘P’ solubilization potential of plant growth promoting Pseudomonas mutants at low temperature. Microbiol. Res. 158, 359–362 (2003).
Yoneyama, K. et al. Strigolactones, host recognition signals for root parasitic plants and arbuscular mycorrhizal fungi, from Fabaceae plants. New Phytol. 179, 484–494 (2008).
Matilla, M. A. et al. Pseudomonas putida KT2440 causes induced systemic resistance and changes in Arabidopsis root exudation. Env. Microbiol. Rep. 2, 381–388 (2010).
Neal, A. L. & Ton, J. Systemic defense priming by Pseudomonas putida KT2440 in maize depends on benzoxazinoid exudation from the roots. Plant Signal. Behav. 8(1), e22655, https://doi.org/10.4161/psb.22655 (2013).
Planchamp, C., Glauser, G. & Mauch-Mani, B. Root inoculation with Pseudomonas putida KT2440 induces transcriptional and metabolic changes and systemic resistance in maize plants. Front. Plant Sci. 5, 719, https://doi.org/10.3389/fpls.2014.00719 (2015).
Fritz, M., Jakobsen, I., Lyngkjær, M. F., Thordal-Christensen, H. & Pons-Kühnemann, J. Arbuscular mycorrhiza reduces susceptibility of tomato to Alternaria solani. Mycorrhiza 16, 413–419 (2006).
Jung, S. C., Martinez-Medina, A., Lopez-Raez, J. A. & Pozo, M. J. Mycorrhiza-induced resistance and priming of plant defenses. J. Chem. Ecol. 38, 651–664 (2012).
Van Wees, S. C. M., Van der Ent, S. & Pieterse, C. M. J. Plant immune responses triggered by beneficial microbes. Curr. Opin. Plant Biol. 11, 443–448 (2008).
Luna, E. et al. Callose deposition: a multifaceted plant defense response. Mol. Plant Microbe In. 24, 183–93 (2011).
Ellinger, D. & Voigt, C. A. Callose biosynthesis in Arabidopsis with a focus on pathogen response: what we have learned within the last decade. Ann. Bot. 114, 1349–1358 (2014).
Raaijmakers, J. M., Leeman, M., van Oorschot, M. M. P., van der Sluis, I. & Schipper, B. Dose-response relationships in biological control of fusarium wilt of radish by Pseudomonas spp. Phytopathol. 85, 1075–1081 (1995).
Thomma, B. P. et al. Separate jasmonate-dependent and salicylate-dependent defense-response pathways in Arabidopsis are essential for resistance to distinct microbial pathogens. Proc. Nat. Acad. Sci. USA 95, 15107–15111 (1998).
Ton, J., Van Pelt, J. A., VanLoon, L. C. & Pieterse, C. M. J. Differential effectiveness of SA-dependent and jasmonate- and ethylene-dependent induced resistance in Arabidopsis. Mol. Plant MicrobeIn. 15, 27–34 (2002).
Rahman, T. A. E., Oirdi, M. E., Gonzalez-Lamothe, R. & Bouarab, K. Necrotrophic pathogens use the salicylic acid signalling pathway to promote disease development in tomato. Mol. Plant Microbe In. 25, 1584–1593 (2012).
Gurney, A. L., Slate, J., Press, M. C. & Scholes, J. D. A novel form of resistance in rice to the angiosperm parasite Striga hermonthica. New Phytol. 169, 199–208 (2006).
Hewitt, E. J. Sand and water culture methods used in the study of plant nutrition. (Commonwealth Agricultural Bureaux, 1966).
Kiers, E. T. et al. Reciprocal rewards stabilize cooperation in the mycorrhizal symbiosis. Science 333, 880–882 (2011).
Declerck, S., Strullu, D. G. & Plenchette, C. Monoxenic culture of the intraradical forms of Glomus sp. isolated from a tropical ecosystem: a proposed methodology for germplasm collection. Mycologia 90, 579–585 (1998).
Dodd, J. C., Clapp, J. P. & Zhao, B. Mycorrhiza Manual prepared for the Workshop Arbuscular mycorrhizal fungi in plant production systems: detectikon, taxonomy, conservation and ecophysiology. https://www2.dijon.inra.fr/mychintec/Protocole/protoframe.html (2001).
Acknowledgements
This work has been supported by the EU FP7 project 326788 “MYCOCROP” to A.P.d.L., D.D.C. and J.T. and a Technology Strategy Board (Innovate UK) grant to D.D.C. (award number: TS/I001751/1). J.T.’s research activities are supported by a grant from the European Research Council (ERC; award number: 309944; ‘PRIME-A-PLANT’) and a Research Leadership Award from the Leverhulme Trust (award number: RL-2012-042). D.D.C. is supported by a Royal Society University Research Fellowship (award numbers: UF090328 and UF140597) and the Grantham Foundation for the Protection of the Environment. We are very grateful to Prof. Toby Kiers (VU Amsterdam) for kindly providing the mycorrhizal isolate.
Author information
Authors and Affiliations
Contributions
A.P.d.L. performed the research, data collection, analysis and interpretation, and wrote the manuscript; S.T., I.J. and D.P.P. helped with data collection and performing the research; J.T. and D.D.C. designed the research, helped with data analysis, performed interpretation of results, and wrote and corrected the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pérez-de-Luque, A., Tille, S., Johnson, I. et al. The interactive effects of arbuscular mycorrhiza and plant growth-promoting rhizobacteria synergistically enhance host plant defences against pathogens. Sci Rep 7, 16409 (2017). https://doi.org/10.1038/s41598-017-16697-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-16697-4
This article is cited by
-
Root Exudates and Their Significance in Abiotic Stress Amelioration in Plants: A Review
Journal of Plant Growth Regulation (2024)
-
Fostering plant growth performance under drought stress using rhizospheric microbes, their gene editing, and biochar
Environmental Geochemistry and Health (2024)
-
Assessing the functional diversity of rhizobacteria from cacao by partitioning root and shoot biomasses
Applied Microbiology and Biotechnology (2023)
-
Mycorrhiza, Charcoal, and Rocket Salad Powder as Eco-friendly Methods for Controlling Broomrape Weed in Inter-planted Faba Bean with Flax
Journal of Soil Science and Plant Nutrition (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.