Abstract
Saccharomyces cerevisiae and its closely related yeasts undergo mating type switching by replacing DNA sequences at the active mating type locus (MAT) with one of two silent mating type cassettes. Recently, a novel mode of mating type switching was reported in methylotrophic yeast, including Ogataea polymorpha, which utilizes chromosomal recombination between inverted-repeat sequences flanking two MAT loci. The inversion is highly regulated and occurs only when two requirements are met: haploidy and nutritional starvation. However, links between this information and the mechanism associated with mating type switching are not understood. Here we investigated the roles of transcription factors involved in yeast sexual development, such as mating type genes and the conserved zinc finger protein Rme1. We found that co-presence of mating type a1 and α2 genes was sufficient to prevent mating type switching, suggesting that ploidy information resides solely in the mating type locus. Additionally, RME1 deletion resulted in a reduced rate of switching, and ectopic expression of O. polymorpha RME1 overrode the requirement for starvation to induce MAT inversion. These results suggested that mating type switching in O. polymorpha is likely regulated by two distinct transcriptional programs that are linked to the ploidy and transmission of the starvation signal.
Similar content being viewed by others
Introduction
Similar to many other organisms, yeast senses and engages appropriate response programs following changes in their environment, including invasive growth, dimorphic transitions, and sexual differentiation. Spore formation, which is often accompanied by mating/meiosis, is a strategy for survival in harsh environments in many yeast species. Homothallism, which is a characteristics observed in some yeasts, allows mating to occur in populations derived from a single cell and is achieved by switching mating type frequently during vegetative growth in Saccharomyces cerevisiae 1. Mating type in S. cerevisiae is determined by two nonhomologous alleles in a single mating type locus (MAT), MAT a or MATα 1,2. Each of these sequences encodes transcriptional regulators of the two different haploid mating types. Mating type switching is mediated by a site-specific homologous recombination event that replaces one MAT allele with silent DNA sequences, HML a or HMRα which encodes the opposite MAT allele that is located on the same chromosome. A similar mechanism comprising three mating type gene sequences and the site-specific homothallic switching (HO) endonuclease, which generates a double-strand break, are shared among species closely related to S. cerevisiae such as Candida glabrata, Zygosaccharomyces rouxii, and Kluyveromyces lactis 3,4. Interestingly, the HO gene in K. lactis is non-functional, and new mechanisms inducing DNA double-strand breaks have been acquired during evolution5,6. In S. cerevisiae, expression of the HO gene is a trigger to induce mating type switching1. Recently, a novel mode of mating type switching was reported in methylotrophic yeast, including Ogataea polymorpha, which utilizes a chromosomal recombination between inverted-repeat sequences flanking two MAT loci7,8. The inversion is highly regulated and occurs only when cells are haploid and under starvation conditions. However, the links between this information and the mechanism associated with mating type switching remain unknown.
Changes in transcriptional profile underlie the molecular mechanisms that induce mating/meiosis and sporulation in yeast. Genes specifically expressed in a or α cells, namely haploid-specific genes (hsgs), are essential for inducing the mating process, whereas they are repressed in situations where both mating type alleles are present in a single cell (e.g., meiosis-competent diploid cells)1,2. Many hsgs and their expression patterns appear to be common among ascomycetous yeasts, many of which are components of the mating pheromone-signalling pathway9; however, there may exist differences in the regulatory mechanism9. In S. cerevisiae, hsgs are repressed by a heterodimer of homeodomain proteins a1 and α2, whereas K. lactis hsgs are placed under the direct regulation of Rme1, a conserved zinc finger transcription factor. Unlike S. cerevisiae, K. lactis primarily proliferate as haploid, and mating type switching and subsequent mating are tightly connected, both of which occur only under starvation conditions that lead to the spore formation. The starvation signal is incorporated into these programs through induction of the expression of RME1. Additionally, in K. lactis, a1-α2 is involved in the regulation of hsgs expression through repressing RME1. Rme1 was originally identified as a regulator of mating type switching and it directly binds to MAT as well as activates transcription of the KAT1 gene which encodes a domesticated transposase6,10.
Recognition of mating partners is initiated when the pheromone receptor binds to the mating pheromone secreted from cells of the opposite mating type11,12. In the model systems, S. cerevisiae and the fission yeast Schizosaccharomyces pombe, pheromones and pheromone receptor genes are targets of transcriptional regulation by pheromone signalling, which is transmitted through a heterotrimeric G protein and the downstream MAP kinase pathway to ensures the robustness of the signalling and the engagement of cells to the mating process13,14,15. Many of these genes are transcriptionally induced by starvation in yeast species, such as S. pombe, K. lactis, and Candida lusitaniae, where sexual development is restricted under starvation conditions16,17,18,19.
Here, we evaluated the roles of transcription factors involved in sexual development in methylotrophic yeast Ogataea polymorpha, including mating type genes and a conserved transcription factor, Rme1, whose orthologue in K. lactis is involved in mating type switching6,10. Our results indicated that co-expression of a 1 gene in MAT a and α2 gene in MATα were sufficient to prevent mating type switching, suggesting that the combination of mating type genes generates ploidy information for mating type switching. Furthermore, RME1 deletion resulted in reduced mating type switching, whereas its ectopic expression overrode the requirement for starvation for inducing MAT inversion. There results suggested that mating type switching in O. polymorpha is likely regulated by a1-α2 and Rme1.
Results
Mating type genes are not required for the MAT inversion
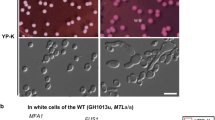
We previously reported α1 is essential for conjugation of α cells with a cells8. Because Hanson et al.7 reported the expression of an a2-like gene in O. polymorpha, we sequenced a2 mRNA and revealed that a predicted intron was indeed removed from mRNA to encode an a2 like protein. In order to examine a possible role of a2 in mating, we crossed a2∆ cells with deletion mutants of α-factor and a-factor receptor genes STE2 and STE3, respectively. Because ste2∆ cells cannot respond to α-factor, homothallic ste2∆ cells are able to mate with cells expressing a identity. Similarly, ste3∆ cells can mate only with α cell. In our mating assay, while a 2∆ cells were able to mate with ste3∆ cells, no mating was observed when crossed with ste2∆ cells (Fig. 1a). This was the reversed pattern of mating capability in α1∆ cells8. These data indicated that O. polymorpha required a2 and α1 for a-, and α-cell identity, respectively (Fig. 1b). Consistently, both a 2 and α1 genes were expressed in mitotically growing a and α cells respectively as well as those cells under starvation condition (Supplementary Fig. S1a).
Mating type genes are not required for the MAT inversion. (a) a 2 gene is required for mating with α cell but not with a cell. Wild-type and a 2∆ strains of ura3–1 (BY4330 and HPH922, respectively) and wild-type, ste2∆ and ste3∆ strains of leu1–1 (HPH22, HPH553, and HPH581, respectively) genotypes were combined on MEMA mating medium and incubated at 30 °C. After 24 h, cells were spread on SD plates to select for Leu+Ura+ diploids. Colony number was counted after 2-day incubation at 37 °C. The average of three independent crosses is shown. Error bars indicate SD. (b) Function of mating type genes in establishing mating type identity. Grey circle represents centromere7. (c) Schematic of the primer designs for PCR reactions specific to I(a)- or A(α)-type chromosomes (MAT PCR analysis). (d) I(a)- or A(α)-type of MAT chromosome was determined by PCR reactions described in (c). PCR reactions to detect I(a)- or A(α)-type chromosomes are designated as a and α and designated a Genomic DNA samples were prepared from the wild-type strain (HPH1047 and HPH1050) and deletion mutants for MAT a1, MAT a2, or MATα (HPK073, HPH1255, and HPK072) before (+N) and after incubation on MEMA medium (−N). CDC28-specific primer set was added to all PCR reactions as controls for the amount of input DNA. Full-length gels are presented in Supplementary Figure S5.
Analysis of RNA expression level revealed that all four mating type genes were expressed mitotically and induced upon starvation (Supplementary Fig. S1). We therefore examined whether MAT gene products contributed to the efficient mating type switching. We observed that mating type switching was not reduced in cells harbouring deletions of each mating type genes (Fig 1c,d), suggesting that MAT genes did not play important roles in the mating type switching.
MAT inversion is inhibited in meiosis-competent diploid cells
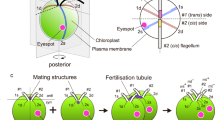
Two sexual differentiation programs, mating and meiosis, are induced by starvation in O. polymorpha 20. Importantly, the mating program should become active only in a- or α- haploid cells and is repressed in a/α diploid cells, whereas initiation of meiosis requires both MAT a and MATα gene products, which makes mating and meiosis mutually exclusive8. Because mating type switching could potentially reduce meiosis efficiency, it might be inhibited in a/α diploid cells. We examined the inversion of the MAT intervening region (MAT inversion) in the A(α)-type chromosome in α haploid and a/α haploid cells. Because it is impossible to distinguish original a/α from α/a diploid cells generated by inverting the MAT-intervening region on both chromosomes, we deleted the centromere proximal inverted repeat (IR2) of the I(a)-type chromosome to prevent the inversion and to obtain a DNA fragment of different size from wild type I(a)-type chromosome in Southern blot analysis (Fig. 2a)8. Inversion efficiency was calculated at ~17.5% in haploid cells, but was below detectable level in a(IR2∆)/α-diploid cells.
a1-α2 inhibits the MAT inversion. (a) MATα haploid cells (HPH848) and MATα/MAT a(IR2∆) diploid cells (HPH964) were grown in YPDS medium (+N) and transferred to MEMA medium and incubated for 18 hrs (−N). Genomic DNA samples were prepared and digested with EcoRI and subjected to Southern blot analysis using a probe specific for MATα sequence (Left panel). The percentage of I-type chromosomes were calculated by measuring the intensity of A- and I-type bands and presented below the blot. Right panel: schematics of MAT-containing chromosomes with the indicated genotype and predicted fragment sizes for the probe (purple) in Southern blot analysis. (b and c) Genomic DNA was prepared from strains with the indicated genotype (HPH1047, HPH1050, HPH1201, HPK011, HPK084, HPK085, HPK092, and HPK093), and the mating type was determined by MAT PCR analysis as shown in Fig. 1c. +N: cells grown in YPDS medium, −N: cells incubated in MEMA medium for 16 hrs. CDC28-specific primer set was added to all PCR reactions as controls for the amount of input DNA. (d) MAT a IR2∆ haploid strain with ura3-1 ade12-cr3 genotype (HPH1174) and the same strain carrying pHM961 expressing α2 gene (HPH1702) were crossed with wild type strain (HPH22) on NaKG mating medium and incubated at 30 °C. Similarly, MATα IR2∆ haploid strain with ura3-1 leu1-1 genotype (HPH1162) and the same strain carrying pHM964 expressing a 1 gene (HPH1699) were crossed with wild type strain (BY4331). After 24 h, cells were spread on SD + Ura plates to select for Leu+Ade+ diploids. Colony number was counted after 2-days incubation at 37 °C. The average of three independent matings is shown. Error bars indicate SD. Full-length blot and gels are presented in Supplementary Figure 5.
The inhibition of the inversion might have occurred due to the co-presence of the two MAT alleles, rather than ploidy. To investigate this possibility, we introduced a MATα allele at the URA3 locus in a cells. As expected, a cells expressing MATα were unable to undergo the MAT inversion (Fig. 2b). Additionally, the MATα(α2∆) plasmid was unable to inhibit the MAT inversion in a cells, whereas the MATα(α1∆) plasmid inhibited the inversion to the same degree as that observed by the MATα plasmid. We also performed the same experiment in α haploid cells, finding that expression of the a1 gene inhibited the MAT inversion in α haploid cells (Fig. 2c). These results suggested that co-expression of a1 and α2 was sufficient to prevent activation of the mating type switching.
To further investigate the effect of co-expression of a 1 and α2 genes, α2 gene was expressed exogenously in stable a cells (mating type of the strain is unswitchable because of the deletion of IR2 sequences) and crossed with wild type cells. The mating efficiency was severely reduced compared to the parental stable a cells. The equivalent experiment was performed in stable α cells with a similar result, although the extent of the mating inhibition was much weaker (Fig. 2d). These results suggested that co-expression of a1 and α2 had inhibitory effect on mating.
Mating type switching occurs following cessation of mitotic cell division upon nutrient starvation in O. polymorpha
To better understand the physiological conditions that allow the mating type switching to occur in O. polymorpha, the timing of the MAT inversion was investigated more precisely (Fig. 3). Wild type cells grown in yeast extract, peptone, and dextrose medium containing 200 mg/L adenine, leucine, and uracil (YPDS) until the late log phase [optical density at 663 nm (A663) = 10–15] were transferred to minimal medium without nitrogen (NaKG; 0.5% sodium acetate, 1% potassium chloride, and 1% glucose). We observed that the percentage of budded cells remained relatively high (40–45%) over the course of 2.5 h, followed by a gradual decrease to 5% after 10 h following the shift (Fig. 3a). An inverted type of chromosome was first detected at the 6 hr time point when budded cells were dropped and almost no anaphase cells were observed (Fig. 3b). The ratio of inverted chromosomes did not increase to a large degree over the next 24 hrs as judged by the intensity of polymerase chain reaction (PCR) products (Fig. 3b). These results suggested that the chromosomal inversion of the MAT-containing region occurred as cells ceased mitosis. When nocodazole was added into the culture at the time of the shift to NaKG medium, the inverted ratio became lower than that observed in the DMSO control cells (Fig. 3c). This might indicate that the starvation signal by itself was insufficient to induce the MAT inversion and that cell cycle progression was required.
Mating type switching occurs after cessation of cell division under starvation condition. (a) Wild type CBS4329 cells were grown in YPDS medium and transferred to NaKG medium. Samples were collected at different intervals and cell density was measured using a hematocytometer. The remaining of samples were fixed and stained with DAPI to determine anaphase cells (Ana, indicated by magenta) and budding index (Budded cells, indicated by green). (b) Genomic DNA was prepared from samples collected in (a) and subjected to MAT PCR analysis as shown in Fig. 1c. CDC28-specific primers were added to PCR reactions as controls for the amount of input DNA. (c) I(a)-type of CBS4329 cells grown in YPDS medium were incubated in NaKG medium containing either dimethyl sulfoxide or nocodazole for 8 hrs. Genomic DNA samples were prepared and digested with EcoRI and subjected to Southern blot analysis using the probe specific for MATα sequences shown in Fig. 2a. The percentage of A-type chromosomes was calculated by measuring the intensity of A- and I-type bands and presented below the blot. Full-length gels are presented in Supplementary Figure S5.
Pheromone signalling does not induce mating type switching
We then investigated the transcriptional programs activated during nutrient starvation. In O. polymorpha, the pheromone signalling pathway is likely activated under starvation condition because some of pheromone signalling components, such as STE4 and STE18 were transcriptionally induced in malt-extract/maltose mating medium (MEMA) (Supplementary Fig. S2). We found that ste4∆ cells were sterile as predicted, but that the MAT inversion occurred as did in wild-type cells (Fig. 4a). Furthermore, ste2∆ ste3∆ cells, that lacks both a- and α-factor receptors, underwent the MAT inversion (Fig. 4b). Therefore, our results indicated that the mating type switching did not require the activation of the pheromone signalling pathway.
The pheromone signalling pathway is not required for the MAT inversion. (a) Ste4∆ cells and ste4∆ cells carrying the STE4 plasmid (HPH1268 and HPH1267, respectively) grown in YPDS medium were incubated in NaKG medium for 12 hrs. Genomic DNA was subjected to MAT PCR analysis as shown in Fig. 1c. CDC28-specific primers were added to PCR reactions as controls for the amount of input DNA. (b) STE2 STE3 ku80∆ (HPH1379) and ste2∆ ste3∆ ku80∆ cells (HPH1696) grown in YPDS medium were incubated in NaKG medium for 12 hrs. Genomic DNA was subjected to MAT PCR analysis as shown in Fig. 1c. CDC28-specific primers were added to PCR reactions as controls for the amount of input DNA. Full-length gels are presented in Supplementary Figure S5.
The transcription factor Rme1 is required for efficient inversion and sexual differentiation
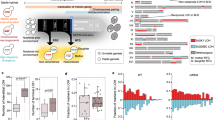
Rme1 is a transcription factor which plays important roles in sexual differentiation in yeast. A homolog of RME1 gene (OpRME1) was found in the O. polymorpha genome. Because the expression of OpRME1 gene was strongly induced upon starvation (Fig. 5a), the roles in sexual cycle were investigated. Deletion of OpRME1 did not completely abolish mating capability, because we were able to backcross the original rme1∆ isolates with a wild-type strain. However, when crossed with wild type cells, rme1∆ cells exhibited >1000-fold less mating efficiency as compared with wild-type cells (Fig. 5b and Supplementary Fig. S3), suggesting that Rme1 was required for mating. We also examined cell cycle arrest upon starvation in rme1∆ cells because G1 arrest is the prerequisite for yeast mating (Fig. 5c). The rme1∆ cells arrested with 1C DNA content as observed in wild type cells, suggesting Rme1 is dispensable for arresting cell cycle in G1. We also examined MAT inversion following starvation and found that it was >50% less frequent compared with that observed in wild-type cells (Figs. 5d, 7.4% of inverted orientation in wild type cells vs. 3.0% in rme1∆ cells). These results suggested that Rme1 played important roles in inducing mating type switching and mating.
Rme1 is required for mating and MAT inversion. (a) RME1 mRNA level increases in response to nutritional starvation. The qPCR analysis was performed with RNAs prepared from wild-type cells grown in YPDS and from the same cells following incubation in NaKG medium. RNA levels were normalized to that of ACT1 RNA. Shown are the averages of three independent PCR reactions. Error bars indicate SD. (b) Mating is severely reduced in rme1∆ cells. Wild-type and rme1∆ strains with ura3-1 genotypes (HPH47 and HPK021, respectively) were combined with wild type cells with leu1-1 genotype (HPH22) on NaKG mating medium and incubated at 30 °C. After 30 h, cells were spread on SD plates to select for Leu+Ura+ diploids. Colony number was counted after 2-days incubation at 37 °C. The average of three independent matings is shown. Error bars indicate SD. (c) Wild type (BY21401) and rme1∆ (HPK187) were grown in YPDS medium and shifted to NaKG medium and incubated for 12 hrs. Cells were fixed with 70% ethanol and DNA content was measured by flowcytometry. (d) MAT inversion was reduced in rme1∆. Genomic DNA was prepared from strains with the indicated genotype (HPK161 and HPK163), digested with EcoRI, and subjected to Southern blot analysis as shown in Fig. 2a. +N: cells grown in YPDS medium, −N: incubated in NaKG medium for 15 hrs. Full-length gels are presented in Supplementary Figure S5.
To further confirm Rme1’s involvement in mating type switching, RME1 was expressed in mitotically growing cells from a strong constitutive TEF1 promoter in O. polymorpha (Fig. 6). RME1-overexpressing cells grown in synthetic defined (SD) medium were a mixture of A(α) and I(a) types, whereas wild-type cells maintained their original MAT chromosome type, indicating that MAT inversion occurred in RME1-overexpressing cells in both A-to-I and I-to-A directions (Fig. 6a). These results suggested that RME1 overexpression was capable of overriding the requirement for nutritional starvation to induce MAT inversion. Moreover, MAT inversion was further induced in RME1-overexpressing cells in starvation medium to a level higher than that observed in wild-type cells under the same condition (Fig. 6b). These results suggested that RME1 expression alone was insufficient for inducing mating type switching, but it might constitute a rate-limiting factor. Moreover, RME1-overexpressing cells exhibited morphological changes that resembled mating projections, as well as signs of meiosis and sporulation (Fig. 6c). Therefore, our findings indicated that RME1 overexpression activated most, if not all, biological programs in response to nutritional starvation, including mating type switching.
RME1 overexpression induces the MAT inversion in mitotically growing cells. (a) RME1 was expressed in I(a)- and A(α)-type wild-type strains (CBS4329) using a strong constitutive TEF1 promoter. Cells were grown in SD medium supplemented with adenine, leucine, and uracil, and genomic DNA was subjected to MAT PCR analysis as shown in Fig. 1c. (b) A(α)-type wild-type cells carrying PTEF1-RME1 were grown in SD medium (+N), transferred to NaKG medium and incubated for 20 hrs (−N). Genomic DNA was subjected to MAT PCR analysis as shown in Fig. 1c. CDC28-specific primers were included in the reaction as control for input DNA. RME1 -: cells carrying vector (pHM874), OP: cells carrying PTEF1-RME1 (pHM960). (c) Wild-type cells carrying vector (pHM874) or PTEF1-RME1 (pHM960) (OPT1 and OPT2, respectively) were grown on SD plate supplemented with adenine, leucine, and uracil for 3 days at 30 °C. Cells were fixed with ethanol and stained with DAPI. Mating projection-like morphology (orange arrows) and meiotic nuclei (yellow arrow) were evident only in cells carrying PTEF1-RME1. (d) I(a)-type wild type cells (HPH466) carrying either vector (pHM874) or PTEF1-RME1 (pHM960) were grown in SD medium supplemented with adenine, leucine, and uracil or YPDS medium. Genomic DNA was subjected to MAT PCR analysis as shown in Fig. 1c. CDC28-specific primers were included in the reaction as control for input DNA. Full-length gels are presented in Supplementary Figure S5.
Notably, the effect of RME1-overexpression on MAT inversion was not evident in YPDS medium (Fig. 6d). In K. lactis, Rme1 protein levels were 7-fold higher in cells grown in synthetic complete medium as compared with levels observed in cells grown in YPD medium6. Therefore, Rme1 expression might be similarly regulated in O. polymorpha.
MAT inversion is autophagy dependent
Important physiological changes caused by nutritional starvation involve the induction of autophagy. Protein degradation by autophagy provides necessary materials for the cellular response to nutritional starvation, and therefore deletion mutants of core autophagy factors such as ATG1 results in a severe defect in meiosis and sporulation in both S. cerevisiae and S. pombe 21,22,23. Because the mating type switching in O. polymorpha is induced by nutritional starvation, we examined involvement of autophagy in the mating type switching. We found that the MAT-inversion was reduced in atg1∆ and atg13∆ cells, suggesting that the mating type switching is an event downstream of autophagy (Fig. 7a).
MAT inversion is autophagy dependent. (a) Autophagy mutants are MAT inversion deficient. Wild-type, atg1∆, and atg13∆ cells (HPH848, HPH1561, and HPH1556, respectively) were grown in YPDS medium (+N), transferred to MEMA medium and incubated for 15 hrs (−N). Genomic DNA was subjected to MAT PCR analysis as shown in Fig. 1c. (b) Atg1 is dispensable for REM1-induced MAT inversion. Wild-type (BY2140) and atg1∆ cells (HPH1620) carrying either vector (pHM874) or PTEF1-RME1 (pHM960) were grown in YPDS or SD medium supplemented with adenine, leucine, and uracil. Genomic DNA was prepared and subjected to MAT PCR analysis as shown in Fig. 1c. CDC28-specific primers were included in the reaction as control for input DNA. Full-length gels are presented in Supplementary Figure S5.
RME1 transcription was induced in atg1∆ cells as well as in wild type cells upon nutrient starvation, suggesting that Atg1 has a role in mating type switching independent of RME1 expression (Supplementary Fig. S4). If the sole role of autophagy in mating type switching is to supply raw materials for protein production under starvation conditions, mating type switching should occur in the absence of autophagy upon induction in rich medium. As predicted, RME1 overexpression in SD medium induced mating type switching in atg1∆ cells, as well as in wild-type cells (Fig. 7b). Therefore, these results indicated that autophagy likely supports the production of factors reliant upon Rme1-dependent transcription and thereby indirectly involved in activating their functions that are required for MAT inversion.
Rme1 and a1-α2 regulate the MAT inversion through independent pathways
We next examined the relationship between two transcription factors, a1-α2, and Rme1, in mating type switching. We first performed RNA-sequencing analysis in haploid cells and the same cells mimicking a diploid state by expressing other mating type genes from an exogenous locus. We observed that RME1 mRNA expression was induced under starvation conditions in both haploid cells and diploid-mimicking haploid cells (Fig. 8a). This result suggested that a1-α2 unlikely regulated RME1 transcription. Although RME1 mRNA levels increased 3-fold under starvation conditions, Rme1 protein levels increased only 1.5-fold (Fig. 8b), implying that Rme1-dependent functions might be regulated by other means independent of Rme1 levels, such as post-translational modifications or through the activity of other proteins.
a1-α2 functions independently of RME1 expression. (a) a1-α2 does not repress RME1 mRNA levels. A(α)-type haploid cells and I(a)-type haploid cells in the presence or absence of MATα the exogenous URA3 locus (HPH1309, HPH1311, and HPK007, respectively) were grown in YPDS (+N), then shifted to MEMA medium and incubated for 10 hrs (−N). RNA was prepared and subjected to RNA-seq analysis. Relative amount of RME1 RNA is shown. (b) RME1-5flag cells (HPK121) were grown in YPDS (0 hr) following incubation in MEMA medium for the indicated time. Total protein was prepared and Rme1-5flag protein was detected by western blotting using an anti-flag M2 antibody. An anti-actin antibody was used as a loading control. (c) Model of the regulation of the mating type switching and mating in O. polymorpha. Full-length blots are presented in Supplementary Figure S5.
Discussion
Conditions under which mating type switching and/or mating occur dictate the dominant and favoured state of ploidy in vegetative cells of homothallic species. Homothallic S. cerevisiae strains that undergo both mating type switching and mating in rich conditions exist mostly as diploid cells, whereas in K. lactis and O. polymorpha cells that are predominantly haploid, both are induced only under nutritionally starved conditions1. Although K. lactis employs essentially the same 3-MAT system as S. cerevisiae for mating type switching, the molecular mechanisms and their regulation differ in order to allow adaption to species-specific life cycles3,6,10. We previously reported that the mating type in O. polymorpha is switched by inverting the intervening region of two MAT loci that reside on the same chromosome, and that homothallism in this yeast relies upon this8. Because an O. polymorpha mating type is stably maintained during vegetative growth and the switching is induced only when the environment becomes nutritionally poor7,8, which engages the sexual differentiation program, this process is likely coordinated with the mating process. Here, we reported that mating type switching in O. polymorpha was regulated by two distinct transcriptional programs: a1-α2 linked to the ploidy and Rme1 transmitting the starvation signal (Fig. 8c).
Determining the link between starvation and MAT inversion is important for understanding the sexual cycle of O. polymorpha. Our observation that RME1-overexpression induced MAT inversion in mitotically growing cells, which occurs only under starvation conditions in wild-type cells, suggested that the starvation signal acted through Rme1 activity. However, Rme1 is unlikely to be the sole factor responsible for inducing MAT inversion, because the inversion efficiency of the rme1∆ strain was reduced by >50% relative to that of the wild-type strain. Notably, autophagy was required for MAT inversion, suggesting that the essential factor(s) inducing MAT inversion need to be synthesized under starvation conditions. However, other transcription factor(s) responsive to starvation signals might also be involved.
Rme1 links starvation, mating, and mating type switching in O. polymorpha in a manner similar to that proposed for K. lactis; however, the composition of MAT loci and the molecular mechanism associated with mating type switching appear to differ between K. lactis and O. polymorpha 6,9,10. While KlRme1 stimulates mating type switching in two ways, one by directly binding to MATα in α cells and the other by activating transcription of the KAT1 gene in a cells, the function of OpRme1 is most likely to activate transcription of downstream target genes. Differences in Rme1 regulation is highlighted by the involvement of a1-α2; whereas KlRME1 transcription was repressed by a1-α2, OpRME1 expression was not9. The molecular target(s) of a1-α2 in O. polymorpha that inhibit MAT inversion remain unknown.
The mechanisms associated with Rme1 regulation in response to starvation remains unknown. Although OpRME1 mRNA levels were strongly induced by starvation, their protein levels were affected only moderately under our experimental conditions. Although regulation of Rme1 levels might account for the difference in Rme1 activity between rich and starvation conditions, there might also be another layer of regulation at the post-translational level, such as nuclear import and protein phosphorylation.
The event that triggers the MAT inversion is currently unclear. One simple scenario might be that a factor such as an endonuclease is induced by starvation to generate DNA lesions. Such genes would be expected to express only in haploid cells within a small window of time after shifting to the starvation conditions, because we did not observe significant increases in the rate of the MAT inversion after prolonged incubation under starvation conditions. However, we have not been successful in identifying such genes in our RNA-seq analysis. Expression profile in rme1∆ cells should be analysed in future. Furthermore, the delay in the timing of the MAT inversion in cell cycle-arrested cells under the starvation condition may suggest that an event associated with cell cycle progression contributes to triggering the MAT inversion in addition to the transcriptional regulations. However, this reduction might be due to the inability to induce the inversion in metaphase when sister chromatids are closely localized and homologous recombination is active. Further analyses are required to clarify these observations.
It might be rational to restrict mating type switching under conditions where mating can occur, because this will increase the population of minor mating type cells in the original population, thereby offering better chances for sexual reproduction, while avoiding energy consumption for conducting the mating type switching event in asexually reproducing conditions. Both of the haploid species, K. lactis and O. polymorpha, employ this strategy; however, the significance of condition-dependent mating type switching should be carefully examined. In O. polymorpha, any culture where one mating type is dominant is capable of mating with a or α cells at similar frequencies, suggesting that the efficiency of mating type switching is not the limiting factor for mating. In the fission yeast S. pombe, where mating is induced by starvation, mating type switching occurs during mitotic growth. Therefore, it is noteworthy that mating efficiency is much higher in S. pombe (~80%) as compared with that observed in K. lactis (~20%) and O. polymorpha (~5%). What determines the overall mating efficiency and how much impact the efficiency of mating type switching has on self-mating frequency would be subjects of future research.
Materials and Methods
Yeast strains and plasmids
Strains and plasmids used in this study are listed in Supplementary Table S1. Unless otherwise indicated, yeast strains were derived from CBS4329 or NCYC495 and were generated by PCR-based methods24. Sequences of primers used in this study are listed in Supplementary Table S2. O. polymorpha cells were transformed by electroporation. The O. polymorpha TEF1 promoter was described previously8,25.
Yeast growth conditions and general methods
Yeast strains were grown in YPDS or SD medium supplemented with appropriate amino acids and nucleotides26. All experiments were performed at 30 °C unless otherwise indicated. Mating and meiosis were induced on MEMA or NaKG plate at 30 °C7,8.
Flowcytometry and microscopy
Yeast cells were prepared for flow cytometry as described27. DNA was stained with propidium iodide and DNA content was measured by FACSCalibur (BD Biosciences, NJ, USA). To visualize DNA, yeast cells were fixed with 70% ethanol, washed with phosphate-buffered saline (PBS), and incubated in PBS containing 1 µg/ml 4′6,-diamidino-2-phenylindole (DAPI). Images were acquired using Eclipse80i (Nicon Co., Tokyo, Japan) equipped with Cool SNAP EZ (Photometrics, AZ, USA) and driven by Meta Vue ver. 7.7.1.0 (Molecular Devices, Inc., CA, USA). Adobe Photoshop (Adobe Systems, Inc., San Jose, CA, USA) was used to mount images.
Determination of A(α)- or I(a)-type
Orientation of the region between IR1 and IR2 [A(α) or I(a)-type] were determined by PCR as previously described with the primers listed in Supplementary Table S2 8. For Southern blot analysis, O. polymorpha genomic DNA was prepared using a standard protocol. Briefly, DNA was digested with EcoRI restriction enzyme before electrophoresis. A standard protocol was used for blotting and hybridization. DNA probes were prepared, and detection was performed using the AlkPhos direct labeling and detection system with CDP-Star (GE Healthcare, Pittsburgh, PA, USA). Signal intensities were quantified with ImageJ version 1.47 (National Institute of Health, Bethesda, MD, USA) and Photoshop (Adobe Systems, San Jose, CA, USA) was used to mount the images.
Semi-quantitative mating
Yeast strains of leu1–1, ura3–1, or ade12-cr3 genotypes were grown at 30 °C in YPDS until the A663 was between 0.5 and 8 . Cells were washed with PBS, diluted to A663 = 1.0, and a 10-µL cell suspension of the two strains was mixed on a nitrocellulose-membrane filter that was placed on a MEMA or NaKG plate and incubated for 20–30 hrs at 30 °C. Cells were resuspended in PBS, and dilutions were plated on SD plates supplemented with leucine or uracil and adenine or on unsupplemented SD plates that were incubated for 2 days at 37 °C. The mating percentage was calculated as the number of colonies on unsupplemented plates divided by the number on adenine and leucine- or uracil-supplemented plates (i.e., whichever had fewer colonies).
RNA analysis
Total RNA was isolated from O. polymorpha as previously described, treated with DNase I, and then further purified using the RNeasy Plus kit or RNeasy MinElute Cleanup Kit (Qiagen, Valencia, CA, USA). A total of 800 ng-1 µg RNA was used to synthesize cDNA with SuperScriptIII (Invitrogen, Carlsbad, CA, USA) or Reverse Tra Ace qPCR RT Master Mix (Toyobo Co., Ltd., Osaka, Japan) according to the manufacturer’s protocol, and a 0.1–1 µl cDNA reaction mixture was used in a qPCR reaction with the primers listed in Supplementary Table S2.
RNAs prepared from wild type a, α, and a cells carrying MATα were sequenced using HiSeq system (Illumina Inc., San Diego, CA, USA). The cDNA libraries were prepared with a TruSeq DNA Sample Preparation v2 Kit (Illumina Inc.) according to the manufacturer’s protocol. The reference sequence contains only ORF sequences of O. polymorpha based on The JGI Genome Portal28,29,30. Supplementary information Dataset 1 shows the result of gene expression analysis.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Haber, J. E. Mating-type genes and MAT switching in Saccharomyces cerevisiae. Genetics 191, 33–64 (2012).
Herskowitz, I. A regulatory hierarchy for cell specialization in yeast. Nature 342, 749–757 (1989).
Butler, G. et al. Evolution of the MAT locus and its Ho endonuclease in yeast species. Proceedings of the National Academy of Sciences 101, 1632–1637 (2004).
Herman, A. & Roman, H. Allele specific determinants of homothallism in Saccharomyces lactis. Genetics 53, 727–740 (1966).
Rajaei, N., Chiruvella, K. K., Lin, F. & Aström, S. U. Domesticated transposase Kat1 and its fossil imprints induce sexual differentiation in yeast. Proc. Natl. Acad. Sci. USA 111, 15491–15496 (2014).
Barsoum, E., Martinez, P. & Astrom, S. U. Alpha3, a transposable element that promotes host sexual reproduction. Genes & Development 24, 33–44 (2010).
Hanson, S. J., Byrne, K. P. & Wolfe, K. H. Mating-type switching by chromosomal inversion in methylotrophic yeasts suggests an origin for the three-locus Saccharomyces cerevisiae system. Proc. Natl. Acad. Sci. USA 111, E4851–8 (2014).
Maekawa, H. & Kaneko, Y. Inversion of the chromosomal region between two mating type loci switches the mating type in Hansenula polymorpha. PLoS Genet 10, e1004796 (2014).
Booth, L. N., Tuch, B. B. & Johnson, A. D. Intercalation of a new tier of transcription regulation into an ancient circuit. Nature 468, 959–963 (2010).
Barsoum, E., Rajaei, N. & Astrom, S. U. RAS/cyclic AMP and transcription factor Msn2 regulate mating and mating-type switching in the yeast Kluyveromyces lactis. Eukaryotic Cell 10, 1545–1552 (2011).
Merlini, L., Dudin, O. & Martin, S. G. Mate and fuse: how yeast cells do it. Open Biology 3, 130008 (2013).
Raudaskoski, M. & Kothe, E. Basidiomycete mating type genes and pheromone signaling. Eukaryotic Cell 9, 847–859 (2010).
Jones, S. K. Jr. & Bennett, R. J. Fungal mating pheromones: Choreographing the dating game. Fungal Genetics and Biology 48, 668–676 (2011).
Nakayama, N., Miyajima, A. & Arai, K. Common signal transduction system shared by STE2 and STE3 in haploid cells of Saccharomyces cerevisiae: autocrine cell-cycle arrest results from forced expression of STE2. EMBO J. 6, 249–254 (1987).
Herskowitz, I. MAP kinase pathways in yeast: review for mating and more. Cell 80, 187–197 (1995).
Kitamura, K. & Shimoda, C. The Schizosaccharomyces pombemam2 gene encodes a putative pheromone receptor which has a significant homology with the Saccharomyces cerevisiae Ste2 protein. EMBO J. 10, 3743–3751 (1991).
Tanaka, K., Davey, J., Imai, Y. & Yamamoto, M. Schizosaccharomyces pombe map3 + encodes the putative M-factor receptor. Molecular and Cellular Biology 13, 80–88 (1993).
Sherwood, R. K., Scaduto, C. M., Torres, S. E. & Bennett, R. J. Convergent evolution of a fused sexual cycle promotes the haploid lifestyle. Nature 506, 387–390 (2014).
Reedy, J. L., Floyd, A. M. & Heitman, J. Mechanistic plasticity of sexual reproduction and meiosis in the Candida pathogenic species complex. Curr. Biol. 19, 891–899 (2009).
Hansen, H. & Hollenberg, C. P. in Nonconventional Yeasts in Biotechnology (ed. Wolf, K.) 293–311 (Springer Berlin Heidelberg, 1996). https://doi.org/10.1007/978-3-642-79856-6_9.
Kamada, Y. et al. Tor-mediated induction of autophagy via an Apg1 protein kinase complex. The Journal of Cell Biology 150, 1507–1513 (2000).
Kohda, T. A. et al. Fission yeast autophagy induced by nitrogen starvation generates a nitrogen source that drives adaptation processes. Genes Cells 12, 155–170 (2007).
Mukaiyama, H., Nakase, M., Nakamura, T., Kakinuma, Y. & Takegawa, K. Autophagy in the fission yeast Schizosaccharomyces pombe. FEBS Lett. 584, 1327–1334 (2010).
Lu, S. F., Tolstorukov, I. I., Anamnart, S., Kaneko, Y. & Harashima, S. Cloning, sequencing, and functional analysis of H-OLE1 gene encoding delta9-fatty acid desaturase in Hansenula polymorpha. Appl. Microbiol. Biotechnol. 54, 499–509 (2000).
Kiel, J. A., Titorenko, V. I., van der Klei, I. J. & Veenhuis, M. Overproduction of translation elongation factor 1-alpha (eEF1A) suppresses the peroxisome biogenesis defect in a Hansenula polymorphapex3 mutant via translational read-through. FEMS Yeast Research 7, 1114–1125 (2007).
Sherman, F. Getting started with yeast. Meth. Enzymol. 194, 3–21 (1991).
Hutter, K. J. & Eipel, H. E. Microbial determination by flowcytometry. J. Gen. Microbiol. 113, 369–375 (1979).
Grigoriev, I. V. et al. The genome portal of the Department of Energy Joint Genome Institute. Nucleic Acids Res. 40, D26–D32 (2011).
Nordberg, H. et al. The genome portal of the Department of Energy Joint Genome Institute: 2014 updates. Nucleic Acids Res. 42, D26–31 (2014).
Acknowledgements
We thank Dr. K Wolfe for communicating unpublished observations. We appreciate the technical assistance from The Research Support Center, Research Center for Human Disease Modeling, Kyushu University Graduate School of Medical Sciences. The work was supported by the Endowed Chair Program of the Institute for Fermentation, Osaka (IFO), Japan (YK) and JSPS KAKENHI Grant Number JP24570214 (HM).
Author information
Authors and Affiliations
Contributions
H.M. and K.Y. designed the study. K.Y., H.M. and T.N.M.T. performed the experiments. H.M. and K.Y. analysed the data. Y.K. and K.T. provided materials and funding. H.M. wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yamamoto, K., Tran, T.N.M., Takegawa, K. et al. Regulation of mating type switching by the mating type genes and RME1 in Ogataea polymorpha . Sci Rep 7, 16318 (2017). https://doi.org/10.1038/s41598-017-16284-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-16284-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.