Abstract
Serotonergic axons extend diffuse projections throughout various brain areas, and serotonergic system disruption causes neuropsychiatric diseases. Loss of the cytoplasmic region of protocadherin-α (Pcdh-α) family proteins, products of the diverse clustered Pcdh genes, causes unbalanced distributions (densification and sparsification) of serotonergic axons in various target regions. However, which Pcdh-α member(s) are responsible for the phenotype is unknown. Here we demonstrated that Pcdh-αC2 (αC2), a Pcdh-α isoform, was highly expressed in serotonergic neurons, and was required for normal diffusion in single-axon-level analyses of serotonergic axons. The loss of αC2 from serotonergic neurons, but not from their target brain regions, led to unbalanced distributions of serotonergic axons. Our results suggest that αC2 expressed in serotonergic neurons is required for serotonergic axon diffusion in various brain areas. The αC2 extracellular domain displays homophilic binding activity, suggesting that its homophilic interaction between serotonergic axons regulates axonal density via αC2′s cytoplasmic domain.
Similar content being viewed by others
Introduction
Serotonergic somata are located in the midline of the brainstem, and serotonergic neurons project their axons throughout the entire brain and spinal cord1,2,3. The serotonergic system regulates a wide range of brain functions, including circadian rhythm, thermoregulation, response to stress and pain, and feeding, maternal, aggressive and sexual behaviors4.
The ontogeny of the serotonergic projections involves three main steps: the initial orientation of axons along the anterior-posterior axis, the guidance of axons along their main pathways, and the terminal projections to the targets of innervation3,5. Many of the molecular mechanisms underlying these steps have been identified. For example, growth associated protein 43 (GAP-43) and a microtubule-associated protein, stable tubule only polypeptide (STOP) influence the growth of serotonergic axons6,7, and protocadherin-α (Pcdh-α) regulates the distribution of serotonergic axons in many target brain areas8.
The Pcdh-α genes belong to the clustered Pcdh families, which also include the Pcdh-β and Pcdh-γ genes9,10. All of the Pcdh-α genes encode type I transmembrane proteins that have the following distinct domains: six extracellular cadherin domains, a transmembrane domain, and a cytoplasmic domain. The Pcdh-α mRNA consists of a variable exon, encoding extracellular, transmembrane, and juxtamembrane cytoplasmic domains, and three (or four) constant exons, encoding the common cytoplasmic tail9,10. Mutants lacking the common C-terminal cytoplasmic tail exhibit learning defects11 and aberrantly distributed serotonergic axons8. These findings indicate that the common cytoplasmic tail is essential for the normal distribution of serotonergic projections in the target brain regions. In addition, each extracellular domain of Pcdh-α family proteins possesses homophilic binding activity in cell-to-cell interaction12. A diversity of Pcdh-γ proteins is used for the self-repulsion of dendrites, based on isoform-specific homophilic interaction13. However, the role of the divergent Pcdh-α proteins in normal serotonergic projections remains unknown. In the present study, we found that Pcdh-αC2 (αC2), one Pcdh-α isoform, is predominantly expressed by serotonergic neurons among Pcdh-α isoforms, and essential for preventing their axon from being too dense or too sparse and for promoting the diffuse projections of serotonergic axons. Here we propose that serotonergic axons predominantly use αC2 among the Pcdh-α members for appropriate axonal projection.
Results
Deletion of the variable exons α11 to αC2 induces abnormal serotonergic projections, while deletion of α2 to α11 does not
We previously reported that mice in which the Pcdh-α constant region (CR) exon was deleted (Pcdha ∆CR/∆CR) and those lacking the Pcdh-α cytoplasmic tail (Pcdha ∆A/∆A) exhibit abnormal serotonergic projections8. Although these results indicated that Pcdh-α genes are essential for normal serotonergic projections, whether all of the Pcdh-α members or just specific ones are necessary was still unclear. In the current study, we analyzed two different Pcdh-α variable exon-deletion mutant lines: Pcdha del(2-11)/del(2-11) mice, in which the variable exons α2 to α11 were deleted, and Pcdha del(11-C2)/del(11-C2) mice, in which the variable exons α11 to αC2 were deleted14 (Fig. 1a). In the Pcdha del(2-11)/del(2-11) mice, the distribution of serotonergic [serotonin transporter (SERT)-immunopositive] axons in the hippocampus was similar to that of wild-type (WT) mice (Fig. 1b,c). Quantitative analysis of the SERT(+) axon density revealed no significant difference in the density of serotonergic axons in each layer in CA1 and the dentate gyrus (DG) between Pcdha del(2-11)/del(2-11) and WT mice (Fig. 1e). In contrast, the Pcdha del(11-C2)/del(11-C2) mice showed marked densification of serotonergic axons in the stratum lacunosum-moleculare (SLM) of CA1 (Fig. 1b,d). Quantitative analysis of the SERT(+) axon density revealed that in the Pcdha del(11-C2)/del(11-C2) mice, the density was significantly higher in the SLM of CA1 and lower in the stratum oriens (SO) of CA1 and the DG, compared with WT mice (Fig. 1f). These results indicated that the genomic region from the variable exons α12 to αC2 is indispensable for the formation of normal serotonergic projections, whereas the genomic region from the variable exons α2 to α11 is dispensable. Next, we examined the expression level of each Pcdh-α gene by in situ hybridization. In the raphe nuclei of WT mice, the αC2 transcript was strongly expressed, and the αC1 transcript was weakly expressed; in contrast, transcripts for the α10 to α12 genes were not detected (Fig. 1g). These findings suggested that the αC2 transcript is the dominant Pcdh-α transcript in the serotonergic neurons of the raphe nuclei.
Pcdha del(11-C2)/del(11-C2) mice, but not Pcdha del(2-11)/del(2-11) mice, exhibit altered distributions of serotonergic axons. (a) Wild-type (WT) Pcdh-α genes consist of exons (1–12, C1 and C2; green) in a variable region (VR) and exons (CR1–CR3; red) in a constant region (CR). The individual variable exons are transcribed from their own promoters. A Pcdh-α transcript is produced from one variable exon and three or four constant exons by splicing. In the del(2-11) allele, exons α2–α11 were deleted. In the del(11-C2) allele, exons α11, α12, αC1, and αC2 were deleted. (b–d) Serotonergic axons in WT, Pcdha del(2-11)/del(2-11), and Pcdha del(11-C2)/del(11-C2) mice were detected by an anti-serotonin transporter (SERT) antibody. The distribution of serotonergic axons in Pcdha del(2-11)/del(2-11) mice (c) was similar to that in WT mice (b), whereas the serotonergic fibers of Pcdha del(11-C2)/del(11-C2) mice (d) were densified in the stratum lacnosum-moleculare (SLM) of CA1, and sparsified in the stratum oriens (SO) of CA1 and in the dentate gyrus (DG), compared with WT mice. SR, stratum radiatum. Scale bar: (b–d), 500 µm. (e) Quantification of SERT(+) fibers in WT (n = 6) and Pcdha del(2-11)/del(2-11) mice (n = 5). (f) Quantification of SERT(+) fibers in WT (n = 6) and Pcdha del(11-C2)/del(11-C2) mice (n = 5). *p < 0.05, **p < 0.01. Mean ± SEM. (g) Expression analysis by in situ hybridization using probes for α10, α11, α12, αC1, αC2, αCR, and Sert in adjacent coronal sections of the dorsal raphe nucleus of WT and Pcdha del(11-C2)/del(11-C2) mice.
We previously reported that in Pcdha del(11-C2)/del(11-C2) mice, stochastic and weak expression patterns of α10 are severely altered to αC2-like constitutive and strong expression patterns in several brain regions including the cerebral cortex, hippocampus, and cerebellar Purkinje cells, and that the total expression levels of all of the Pcdh-α isoforms is maintained14. In contrast, in the raphe nuclei of these mice, the expression level of total Pcdh-α transcripts, which was detected by the common Pcdh-α CR exon probe, was markedly lower, and the expression level of α10 was grossly normal (Fig. 1g). These results indicated that the regulation of Pcdh-α gene expression in the raphe nuclei is different from that in the cerebral cortex, hippocampus, and cerebellum.
αC2 is essential for normal serotonergic projections
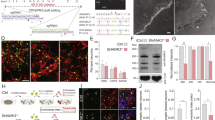
Given that the Pcdha del(11-C2)/del(11-C2) mice exhibited abnormal serotonergic projections and αC2 was strongly expressed in the raphe nuclei, we speculated that αC2 was essential for normal serotonergic projections. To test this possibility, we generated ∆C2 alleles (Fig. 2a). αC2 KO (Pcdha ∆C2/∆C2) mice showed abnormal serotonergic projections in many target brain areas, compared with control mice (Fig. 2b,c). In control mice, serotonergic axons extended diffuse projections throughout various brain areas, including the anterior olfactory nucleus, the olfactory bulb, the cortex, and the superior colliculus (Fig. 2b,e,g,i,k). In contrast, in these regions, αC2 KO mice exhibited both the densification and sparsification of serotonergic axons (Fig. 2c,f,h,j,l). In the spinal cord, the serotonergic axon density in αC2 KO mice was grossly normal, compared with control mice (Fig. 2m,n). In the hippocampus, the density of serotonergic axons in αC2 KO mice was significantly higher in the SLM of CA1, and lower in the SO of CA1 and the DG, compared with WT mice (Fig. 3a–e). We quantified the transcripts of Sert and other Pcdh-α members in whole brains by quantitative reverse transcription (qRT)-PCR, and observed no significant differences in their expression levels between the WT and αC2 KO mice (Fig. 3f). In situ hybridization analysis revealed that the expression levels of α11, α12, αC1, and αCR in αC2 KO mice were similar to those in WT mice in the raphe nuclei (Fig. 3g). In the raphe nuclei of the αC2 KO mice, the expression of αC2 was completely lost, although the expression of αC1 remained (Fig. 3f). These results indicated that αC2 expression is essential for the normal diffusion of serotonergic projections in the target brain regions.
Pcdha ∆C2/∆C2 mice exhibit altered distributions of serotonergic axons. (a) WT and ∆C2 alleles. In the ∆C2 allele, the αC2 exon was deleted. (b–d) Parasagittal sections of WT (b), Pcdha ∆C2/∆C2 (c) and Pcdha ∆CR/∆CR brains (d) were stained with an anti-SERT antibody. Boxed areas (e–l) in (b) and (c) were magnified in the following panels. (e,f) In layer 2 (L2) of the anterior olfactory nucleus, the density of SERT(+) axons in Pcdha ∆C2/∆C2 mice (f) was higher than that in WT mice (e). (g,h) In the granule cell layer (GCL) of the olfactory bulb, SERT(+) axons of Pcdha ∆C2/∆C2 mice (h) showed densification, compared with that of WT (g). EPL, external plexiform layer; GL, glomerular layer; IPL, inner plexiform layer; MCL, mitral cell layer. (i,j) In L1 of the somatomotor cortex, SERT(+) axons of Pcdha ∆C2/∆C2 mice (j) showed densification, compared with that of WT (i). (k, l) In the optic nerve layer (Op) of the superior colliculus, the density of SERT(+) axons of Pcdha ∆C2/∆C2 mice (l) was higher, compared with that of WT mice (k). InG, intermediate gray layer; SuG, superficial gray layer; Zo, zonal layer. (m,n) SERT(+) axons in the spinal cord of Pcdha +/∆C2 (m) and Pcdha ∆C2/∆C2 mice (n). Scale bars: 1 mm (b–d); 100 µm (e–l); 200 µm (m,n).
αC2 protein is required for normal serotonergic projections. (a–d) Serotonergic axons in the hippocampus of WT (a,c) and Pcdha ∆C2/∆C2 (b,d) mice were detected with an anti-SERT antibody. Panels (c) and (d) are higher-magnification images of the boxed areas in (a) and (b), respectively. SP, stratum pyramidale. (e) Quantification of SERT(+) fibers in the hippocampus of WT (n = 4), Pcdha +/∆C2 (n = 3), and Pcdha ∆C2/∆C2 (n = 4) mice. *p < 0.05, **p < 0.01. Mean ± SEM. (f) Quantitative RT-PCR for the Pcdha and Sert genes. There was no significant difference between WT and Pcdha ∆C2/∆C2 KO mice. Mean ± SEM. (g) Expression analysis by in situ hybridization using probes for α11, α12, αC1, αC2, αCR, and Sert in adjacent coronal sections of the dorsal raphe nucleus of WT and Pcdha ∆C2/∆CR mice. Scale bars: (a,b), 200 µm; (c,d) 50 µm; 200 μm (g).
αC2 expression in serotonergic neurons is essential for normal serotonergic projections
Pcdh-α genes are expressed not only in serotonergic neurons, but also in other types of neurons in almost all brain regions, including the hippocampus8. Therefore, it was not clear whether the Pcdh-α expressed by serotonergic neurons or by other neurons in their target brain regions controls the serotonergic innervation. To address this question, we generated brain region-specific Pcdh-α deficient mice using the Cre-loxP system. In the Pcdha flox allele, loxP sequences were inserted upstream and downstream of the Pcdh-α gene cluster (Fig. 4a).
Dorsal telencephalon-specific Pcdh-α KO mice exhibit normal serotonergic projections. (a) In the flox allele, loxP-FRT-neo r -FRT-loxP-Myc-Venus and loxP sequences were inserted upstream of the α1 exon and downstream of the CR3 exon, respectively. Telencephalon-specific Pcdh-α KO mice were homozygous for the Pcdhα flox allele in Emx1 Cre/+. (b,c) In situ hybridization analysis with and αCR probe in the hippocampus. The expression level of the total Pcdh-α mRNA was markedly decreased in the hippocampus of Pcdha flox/flox ; Emx1 Cre/+ mice (c), compared with controls (Pcdha +/+ ; Emx1 Cre/+ mice) (b). (d,e) In the hippocampus, the SERT(+) axonal distributions in the Pcdha flox/flox ; Emx1 Cre/+ mice (e) were normal compared with controls (Pcdha flox/flox, d). (f) Quantification of SERT(+) fibers in the hippocampus of control Pcdha flox/flox (n = 3) and Pcdha flox/flox ; Emx1-Cre mice (n = 3). The density of SERT(+) axons in the hippocampus showed no significant difference. Scale bars, 200 μm (b–e).
In dorsal telencephalon-specific Pcdh-α KO mice (Pcdha flox/flox ; Emx1 +/Cre), Pcdh-α gene expression was lost in almost all of the cells in the hippocampus and cerebral cortex (Fig. 4b,c). In the dorsal telencephalon of Emx1 +/Cre mice, Cre recombinase is expressed in excitatory neurons but not in inhibitory neurons15. Thus, almost all of the cells expressing Pcdh-α in the hippocampus of the dorsal telencephalon-specific Pcdh-α KO mice were thought to be inhibitory neurons. The serotonergic projections in these mice were grossly normal in the hippocampus, compared with control mice (Pcdha flox/flox) (Fig. 4d–f). This result indicated that the expression of Pcdh-α genes in hippocampal excitatory neurons is dispensable for normal serotonergic projection.
Next, we generated serotonergic neuron-specific αC2 KO mice (Pcdha flox/∆C2 ; ePet-Cre) using the ePet-Cre line1 (Fig. 5a). Serotonergic neuron-specific αC2 KO mice (Pcdha flox/∆C2 ; ePet-Cre) showed partial deficiency of αC2 expression in Sert-positive serotonergic neurons [50% of the Sert(+) cells in the median raphe nucleus were αC2(+):23 αC2(+) cells in 47 Sert(+) cells], and displayed abnormal serotonergic projections in the hippocampus, compared with control mice (Pcdha flox/∆C2) (Fig. 5b–g). The density of SERT(+) axons in the serotonergic neuron-specific αC2 KO mice was significantly higher in the SLM of CA1 and lower in the DG, compared with WT mice (Fig. 5h). The serotonergic neuron-specific αC2 KO mice are not simple, because not only biallelic αC2 exons but also hemi-allelic exons from α1 to αC1 are also deleted in the serotonergic neurons. We previously reported that Pcdha +/∆CR mice, which at least lack the expression of α1 to αC1 from the hemi-allele, show normal serotonergic projection in the hippocampus8. These results indicated that the loss of biallelic αC2 expression in serotonergic neurons leads to aberrant serotonergic projection in the hippocampus.
Serotonergic neuron-specific αC2 KO mice exhibit abnormal serotonergic projections. (a) Serotonergic-neuron-specific αC2 KO mice had heterozygous Pcdhα flox and ∆C2 alleles with the ePet-Cre transgene. (b–e) In situ hybridization analysis with Serotonin transporter (Sert) and αC2 probes in the median raphe nuclei using adjacent sections. Almost all the Sert-positive neurons were αC2-positive in the Pcdha flox/∆C2 control (c); in contrast, the αC2-positive neurons were decreased among the Sert-positive neurons of the Pcdha flox/∆C2 ; ePet-Cre mice (e). (f,g) Pcdha flox/∆C2 ; ePet-Cre mice exhibited abnormal SERT(+) axonal distributions in the hippocampus (g) compared with controls (Pcdha flox/∆C2) (f). (h) Quantification of SERT(+) fibers in the hippocampus of control Pcdha flox/∆C2 (n = 3) and Pcdha flox/∆C2 ; ePet-Cre mice (n = 3). *p < 0.05, Mean ± SEM. Scale bars: 100 μm (b–e); 200 μm (f,g).
αC2 protein is required for serotonergic axon diffusion in the hippocampus
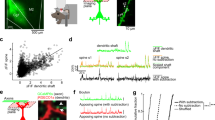
The loss of αC2 causes serotonergic axon densification in the SLM of the hippocampus. To elucidate how the axonal densification occurs morphologically, we sparsely labeled serotonergic axons by injecting an adeno-associated virus vector (AAV-EF1α-DIO-tRFP-WPRE), in which Cre expression induces red fluorescent protein (RFP) expression, into the raphe nuclei of Sert-Cre mice16. Cre(+) cells in the raphe nuclei specifically expressed RFP (Fig. 6a). A portion of serotonergic axons was labeled by RFP in both control (Sert-Cre) and αC2 KO (Pcdha ∆C2/∆C2; Sert-Cre) mice (Fig. 6b–d). We did not find any axons that were in continuous contact with each other, forming thick bundles with multiple axons, or clumps with other single axons in either the control (n = 5 sections, 2 mice) or αC2 KO mice (n = 5 sections, 4 mice). To analyze these axonal routes, we traced the RFP(+) axons passing through a 20-µm inner line (L − 20) in the SLM or a 20-µm outer line (L + 20) in the stratum radiatum (SR) from the boundary between the SLM and SR (L0) (Fig. 6c–f). The ratio of RFP(+) axons crossing L − 20 to those in L + 20 in the αC2 KO mice was higher than that in control mice [control, 82 axons at L − 20, and 71 axons at L + 20 (3 sections from 2 mice); KO, 121 axons at L − 20, and 46 axons at L + 20, Fisher’s exact test, p = 0.0005, Fig. 6g]. This finding was consistent with the result of the SERT-immunopositive intensity experiment (Fig. 3a–e). Next, we analyzed the RFP(+) axons crossing both L − 20 and L + 20. The probability that RFP(+) axons passing through L − 20 also passed through L + 20 in the αC2 KO mice was significantly lower than that in control mice [control, 26 of 82 axons (3 sections from 2 mice); KO, 10 of 121 axons (3 sections from 4 mice), Fisher’s exact test, p < 0.0001, Fig. 6g]. The probability that RFP(+) axons passing through L + 20 also passed through L − 20 in the αC2 KO mice was also significantly lower than that in control mice [control, 26 of 71 axons (3 sections from 2 mice); KO, 10 of 46 axons (3 sections from 4 mice), Fisher’s exact test, p = 0.038, Fig. 6g]. These results indicated that the serotonergic axons in αC2 KO mice were more suppressed from crossing the SLM/SR boundary than those in control mice. These results suggested that αC2 in the serotonergic axons is required for their axonal diffusion.
Diffusion of serotonergic axons is suppressed in the hippocampus of αC2 KO mice. (a–f) Viral vector (AAV-EF1α-DIO-tRFP-WPRE) was transferred into serotonergic neurons in the raphe nuclei of Sert-Cre mice. (a) Coronal sections were immuno-stained with an anti-Cre antibody, and RFP (magenta) was expressed in Cre(+) cells (green) in the median raphe nucleus (arrows) of Sert-Cre mice. (b) A portion of SERT(+) axons (green) was RFP-positive (magenta, arrowheads). (c,d) RFP(+) axons in control (Sert-Cre) (c) and αC2 KO (Pcdha ∆C2/∆C2; Sert-Cre) mice (d) were detected in sagittal sections. (e,f) The RFP(+) axons crossing the 20-μm inner line (L − 20) and/or the 20-μm outer line (L + 20) from the SLM/SR boundary (L0) were traced in control (e) and αC2 KO mice (f). Panels (e) and (f) were depicted from panels (c) and (d), respectively. White, magenta, or green lines indicate axons crossing both lines, only L − 20, or only L + 20, respectively (e,f). (g) RFP(+) axons crossing L − 20 and/or L + 20 were counted in control (n = 4 sections, 3 mice) and αC2 KO mice (n = 3 sections, 2 mice). Scale bars: 50 μm (a); 10 μm (b); 40 μm (c–f).
αC2 protein is required for the outward diffusion of serotonergic axons in the olfactory bulb
Layers in the olfactory bulb are separately innervated by median and dorsal raphe nuclei: serotonergic neurons in the median raphe nucleus innervate the glomerular layer (GL), whereas those in the dorsal raphe nucleus innervate the granule cell layer (GCL) to the external plexiform layer (EPL)17. Here we focused on the median raphe-innervating layers. SERT(+) axons in αC2 KO mice were densified in the GCL, and sparsified in the EPL, compared to controls (Fig. 7a–c). To analyze these axonal routes, we traced the SERT(+) axons passing through the 20-µm inner (L − 20) line or 20-µm outer (L + 20) line from the boundary between the GCL and the inner plexiform layer (IPL) (Fig. 7d,e). The probability that SERT(+) axons passing through L − 20 also passed through L + 20 in the αC2 KO mice was significantly lower than in WT mice [WT, 20 of 95 axons (3 sections from 3 mice); KO, 9 of 115 axons (3 sections from 3 mice), Fisher’s exact test, p = 0.0083, Fig. 7f]. In contrast, there was no significant difference in the probability that SERT(+) axons passing through L + 20 also passed through L − 20 between these genotypes (WT, 20 of 94 axons; KO, 9 of 48 axons, Fisher’s exact test, p = 0.83, Fig. 7f). Although we cannot determine the axonal direction (outward or inward) from these results alone, these observations raised two possibilities: either the diffusion of outward axons from the GCL to IPL is suppressed, or the arborization of inward axons from the IPL to GCL is facilitated, in the GCL. Therefore, we analyzed SERT(+) axons in the GCL. In this region, there was no significant difference in the occurrence frequency of end points and branch points of SERT(+) axons between WT and αC2 KO mice (Fig. 7g). These results suggested that the outward diffusion of serotonergic axons from the GCL is suppressed.
Serotonergic axons in the granule cell layer are suppressed from invading the outer layers in αC2 KO mice. (a–c) In the olfactory bulb, serotonergic axons of αC2 KO mice (a) were densified in the granule cell layer (GCL), and sparsified in the internal plexiform layer (IPL), mitral cell layer (MCL), and external plexiform layer (EPL), compared with WT mice (b). Sagittal sections of the dorsal olfactory bulb were stained by an anti-SERT antibody (magenta) and DAPI (blue). In the GCL, the density of SERT(+) axons of αC2 KO mice (n = 5 sections from 3 mice) was significantly higher than that in WT mice (n = 4 sections, 3 mice) (c). In the EPL, the density of SERT(+) axons of αC2 KO mice (n = 6 sections, 3 mice) was significantly lower than that in WT mice (n = 6 sections, 3 mice) (c). (d–f) SERT(+) axons crossing the lines 20-µm within (L − 20) and/or 20-µm outside (L + 20) from the IPL/GCL boundary (L0) were traced (d,e). Panels (d) and (e) were depicted from panels (a) and (b), respectively. White, magenta or green lines indicate axons crossing both lines, only L − 20 or only L + 20, respectively (d,e). Arrowheads indicate end points (d,e). SERT(+) axons crossing L − 20 and/or L + 20 were counted in WT (n = 3 sections, 3 mice) and αC2 KO mice (n = 3 sections, 3 mice) (f). (g) In the GCL, there was no significant difference in the frequency of branch points or of end points between WT and αC2 KO mice. SERT(+) axons in the GCL (50 × 50 × 50 µm) of WT (n = 130 axons from 4 sections, 3 mice) and αC2 KO mice (n = 407 axons from 5 sections, 3 mice) were traced, and their length, branch points, and end points were analyzed. Survival analyses (Log-rank tests) for these axons were performed. (h,i) SERT(+) axons in GCL of αC2 KO mice showed fasciculation (arrows in h and i). Panels (h1)–(h4) and panels (i1)–(i4) indicate the cross sections in panel (h) and (i), respectively. Arrows in h,h1–h4,i, and i1–i4 indicate axon pairs that run parallel to each other. Scale bars, 40 µm (a,b,d,e); 10 µm (h–i).
The somata of granule cells in the GCL were densely clustered in both genotypes (Fig. 7a,c). Serotonergic axons ran through this cell-cluster niche in a manner avoiding the high-density cell clusters. In the GCL soma niche of αC2 KO mice, we frequently found serotonergic axons that ran parallel and partially in contact with each other (Fig. 7h,i). We did not detect this phenotype in WT mice. These results suggest that the loss of αC2 protein reduces the axon-axon repulsion of serotonergic axons in the olfactory bulb. As a result, the reduced axonal repulsion may dampen the outward diffusion of serotonergic axons. Around the GCL/IPL boundary, some SERT(+) axons returned to the GCL without contacting other SERT(+) axons (arrows in Fig. 7b,e). This observation suggests that loss of αC2 protein enhances the attraction of serotonergic axons to the GCL or their repulsion from the IPL. The αC2 protein may mediate not only the trans-homophilic interaction-driven repulsion of serotonergic axons, but also the trans-heterophilic interaction-driven axon guidance in target areas.
Discussion
Here we found that, among the Pcdh-α genes, αC2 was dominantly expressed in the serotonergic neurons of the raphe nuclei, and that the loss of only αC2, and not of α2 to α11, caused unbalanced distributions (densification and sparsification) of serotonergic axon density in various brain regions, including the hippocampus. The abnormalities of the serotonergic projections in Pcdha del(11-C2)/del(11-C2) and Pcdha ∆C2/∆C2 mice were very similar to those in Pcdha ∆CR/∆CR mice8. Thus, the diversity of the Pcdh-α genes is not required for the normal distribution of serotonergic axons. We also analyzed dorsal telencephalon-specific Pcdh-α KO mice and serotonergic neuron-specific αC2 KO mice. The former showed a normal distribution of serotonergic projections, whereas the latter showed unbalanced distributions similar to those of the αC2 KO mice. Analyses of serotonergic projections at the single-axon level revealed that the serotonergic-axon diffusion in αC2 KO mice is suppressed. These results indicated that the αC2 expression in serotonergic neurons is essential for the normal diffuse pattern of serotonergic axons in various target regions.
Recently, Chen et al. also analyzed Pcdh-α function in serotonergic system18. (1) They analyzed the expression of Pcdh-α, -β, and -γ by single-cell RNA-seq in serotonergic neurons, and showed that αC2 is intensely expressed in serotonergic neurons. (2) They showed that loss of αC1 and αC2 causes clumping of serotonergic axons in the hippocampus. (3) They showed that loss of Pcdhα in serotonergic neurons causes clumping of serotonergic axons in the hippocampus although loss of Pcdhα in the hippocampal neurons do not. (4) They individually labeled serotonergic axons in serotonergic neuron-specific Pcdh-α KO mice, and found clumped axons in the hippocampus and substantia innominata. From these results, they concluded that αC2 is required for axonal tiling and assembly of serotonergic axons.
Similar abnormal axonal clumps appear in Ia afferent axons in the spinal cord of Pcdh-γ KO mice19, and in triple KO mutants in γC3 to γC5 20. Interestingly, γC3 to γC5 are constitutively expressed genes similar to αC2. However, analyses of conditional Pcdh-γ KO mice revealed that Pcdh-γ expressed in both Ia afferents and their targets (interneurons in the spinal cord) is required to prevent the clumping of projections during the formation of Ia afferent terminals19. In the serotonergic projection, αC2 was required not in the target neurons, but in the serotonergic neurons themselves. Therefore, the mechanism underlying the inhibition of axonal densification in serotonergic axons by αC2 is distinct from that in Ia afferents by Pcdh-γC3–C5.
We previously reported that the cytoplasmic tail of Pcdh-α is essential for normal serotonergic projection8. The cytoplasmic domain of αC2 is known to bind cytoplasmic signaling proteins of the focal adhesion kinase (FAK) family, FAK and PYK221. The FAK family contributes to signaling cascades that regulate the growth cone motility, and is activated by several axon-guidance cues, including brain-derived neurotrophic factor (BDNF)22,23. Interestingly, BDNF can induce serotonergic axonal growth in the brain24,25. Stathmin family proteins, microtubule-destabilizing proteins, also bind to the cytoplasmic domain of Pcdh-α26. These proteins are involved in axonal elongation and branching27, and their phosphorylation is induced by BDNF28,29. Therefore, BDNF and stathmin family proteins are candidate molecules involved in the αC2 signaling cascade that controls serotonergic projections.
Glia cell line-derived neurotrophic factor (GDNF) enhances the neurite elongation of serotonergic neurons in vitro 30. GDNF family receptor, which is expressed in serotonergic neurons31, forms a receptor complex with Ret tyrosine kinase, an essential molecule for GDNF signaling. Ret directly binds to α4, and enhances the phosphorylation of α4′s cytoplasmic domain, in a GDNF dosage-dependent manner26. This binding stabilizes both Ret and α4. In serotonergic neurons, the α4 expression level is considerably lower than that of αC2. Nevertheless, GDNF and Ret are also candidates for regulating αC2’s function in the serotonergic projections.
A similar situation is seen for the Down syndrome cell adhesion molecule (DSCAM), which belongs to the immunoglobulin superfamily. Two closely related proteins (Dscam and Dscaml1) are expressed in vertebrates, and 19,008 proteins are generated by distinct combinations of alternative splicings of their homolog in Drosophila 32. Each DSCAM homophilically mediates negative cell-cell interactions33, and these homophilic interactions are responsible for dendrite and axonal self-avoidance. In mice, Dscam is expressed in most retinal ganglion cells and prevents the fasciculation of their dendrites by self-avoidance. Similarly, Dscaml1 is expressed in rod bipolar cell dendrites and prevents the fasciculation and clumping of their dendrites and cell bodies by self-avoidance34. Thus, homophilic negative interactions between molecules of a single isoform among many diverse Dscams are sufficient to induce repulsion and prevent fasciculation in homotypic dendrites and axons. The diversity of the Pcdh-α genes is not required for the normal diffuse innervation of serotonergic axons. Therefore, our study could not reveal the role of the diversity of clustered Pcdh-α genes, but can support the possibility that the diversity of Pcdh-α isoforms mediates the axon and/or dendrite repulsion of cortical and hippocampal neurons, and of Purkinje cells of the cerebellum, which show diverse expression patterns14,35, like the Drosophila DSCAMs.
Azmitia and Segal analyzed serotonergic axonal routes in detail36. From the observation, they speculated that serotonergic projection is regulated by epiphytic guidance, that is, they rely on another group of fibers for their structural support. We found many serotonergic axons crossing GCL/IPL boundary in the olfactory bulb. Because granule cells in GCL project their dendrites to EPL, their may guide serotonergic axons. We found serotonergic axons returning to GCL from GCL/IPL boundary without entering IPL in αC2 KO mice (arrows in Fig. 7b,e). These results raise a possibility that αC2 in serotonergic axons is required to attach the other fibers and to find their appropriate routes and final target areas.
In this study, we reported that αC2 in serotonergic neurons enables serotonergic axons to extend diffuse projections in various brain areas. Understanding the mechanism by which αC2 promotes diffuse projections in the appropriate brain target regions should shed light on the cellular and molecular mechanisms important for homotypic axonal repulsion and serotonin-related developmental disorders.
Methods
Animal experiments
All of the experimental procedures were in accordance with the Guide for the Care and Use of Laboratory Animals of the Science Council of Japan and were approved by the Animal Experiment Committee of Osaka University or the National Institute of Genetics. Male or female mice in adulthood (>3 months old) were used in the axon-tracing analyses. In the other experiments, male or female mice at postnatal day 21 were used.
Generation of Pcdh-α mutant mice
Mice possessing the Pcdha del(2-11), Pcdha del(2-11), or Pcdha flox alleles were described previously14,37,38. To generate dorsal telencephalon-specific Pcdh-α KO mice, we crossed Pcdha flox mice with Emx1-Cre KI-∆Neo mice15. To generate serotonergic neuron-specific Pcdh-α KO mice, we crossed Pcdha flox mice with ePet-Cre mice1.
To remove the initiation codon of αC2, we generated the SBC2 targeting vector by inserting a PCR-amplified 5′ homologous arm of a 317-base region downstream of the αC2 initiation codon (5.5 kb, from 5′-catactacctgattgtccatc-3′ to 5′-gctgcctctgccagggcc-3′), a sleeping beauty cassette (2.5 kb, SalI-loxP–IR/DR-L–loxP–neomycin-resistance (neor)–loxP–IR/DR-R-SalI)39, and a PCR-amplified 5′ homologous arm of a 7-base region upstream of the αC2 initiation codon (5.5 kb, from 5′-cgaggcgctgtgcgagcag-3′ to 5′-ggtgtctttaaaactttaaatgat-3′), into pMC1DT-A 40 (Supplemental Fig. 1a,b). The linearized vector was introduced into TT2 ES cells by electroporation41. Using standard methods, we obtained targeted recombinants and their chimeric offspring. We confirmed them by Southern blot analysis with a 5′ probe (synthesized by the primer set, 5′-tacacactgaaagttaaggc-3′ and 5′-tgtcactccatctactcc-3′), and 3′ probe (synthesized by the primer set, 5′-gaattagtggttagaacctcaccc-3′ and 5′-taggaccaacctaaccacaagaccc-3′) after NheI and ApaLI digestion, respectively (Supplemental Fig. 1c,d). To remove the neor sequence from the SBC2 allele, we crossed the SBC2 mice with Sycp-Cre mice14, and obtained mice carrying the SBC2∆Neo allele (Supplemental Fig. 1a). To completely remove the coding region of the αC2 exon from the SBC2∆Neo allele by Cre-mediated trans-allelic recombination, we crossed female mice (C57BL/6 J) with male mice carrying the Sycp-Cre transgene, SBC2∆Neo allele, and G1 allele, in which the loxP sequence was inserted between the αC2 exon and CR1 exon42, and obtained mice carrying the ∆C2 allele (Supplemental Fig. 1a). Pcdha +/∆C2 and Pcdha ∆C2/∆C2 mice were fertile. Their genotypes were confirmed by PCR using tail chip DNA. The bands of the WT (967 bp) and ∆C2 alleles (424 bp) were obtained using primers (PcdhaC2(−120):F, 5′-gtagtgcgtgagaggtgaag-3′; PcdhaC2 (+847):R 5′-ggtccgaggcattcaacttc-3′; and G1-GTP-R1, 5′-gcccaggatggctcaaattc-3′) (Supplemental Fig. 1e).
In situ hybridization, histochemistry, and quantitative RT-PCR
In situ hybridization, histochemistry, and quantitative RT-PCR were performed by previously described methods8,14. Quantitative RT-PCR for the Sert transcript used the primer pair: 5′-catcagccctctgtttctcc-3′ and 5′-cttaagtgtccctggagtgc-3′. Control plasmid containing the Sert gene was constructed by inserting a PCR amplicon (5′-ctgtcattggctatgccgtg-3′ and 5′-cttaagtgtccctggagtgc-3′) into a pCR4-Blunt-TOPO vector (Thermo Fisher Scientific).
Analysis of serotonergic fiber density
The distribution of serotonergic axons using an anti-serotonin transporter (SERT) antibody (HTT-N77, a generous gift from Dr. Masahiko Watanabe, Hokkaido University, Japan), and the SERT-immunopositive fiber density in the hippocampus were analyzed as described previously with small modifications8. All photographs were taken with a DP-72 CCD camera (Olympus) mounted on a BX51 microscope (Olympus). The resolution of the photographs used to analyze serotonergic axon density was 1101 pixel/mm (Fig. 1e,f), or 1643 pixel/mm (Figs 3e, 4f and 5h). Prism software (GraphPad Software, Inc.) was used for the statistical analyses. Statistical comparisons among three mouse lines were done with one-factor ANOVA and Bonferroni’s post hoc test. Statistical comparisons between two mouse lines were done with the Mann-Whitney U test. The criterion for significance was p < 0.05.
Analyses of serotonergic axons at the single-axon level
To label serotonergic neurons that project to the hippocampus, we injected the adeno-associated virus vector pK207_AAV-EF1α-DIO-tRFP-WPRE into the median raphe nucleus (angle: 25°, AP: −4.7 mm, ML: 0.2~0.6 mm, DC 5.0 mm from bregma) of mice that were anesthetized by a Univentor 410 Anesthetisia Unit, using a stereotaxic apparatus (Narishige). 5HTT-Cre (Sert-Cre) mice16 and Sert-Cre;Pcdha ∆C2/∆C2 mice were used as control and experimental groups, respectively. pK207 was constructed by inserting tRFP (pTurboRFP-N, evrogen) into the AscI/NheI site of pK168_AAV-EF1α-DIO-tTA-tRFP-WPRE (Adgene 85039)43. Virus was prepared as described previously43. More than 4 weeks after the virus injection, mice were transcardially perfused with 4% paraformaldehyde/0.1 M phosphate buffer (PB). The brains were removed and immersed in 30% sucrose/0.1 M PB, and 100-μm-thick sagittal sections in the hippocampus and 100-μm-thick coronal sections in the raphe nuclei were prepared with a microtome (Yamato Kohki). The sections were incubated in rabbit anti-SERT (1:1000, Frontier Institute) [and mouse anti-Cre antibodies (1:1000, Millipore) in some cases] in phosphate buffed saline (PBS) containing 1.5% normal goat serum and 0.25% Triton X-100 for >2 days. After washing with PBS, the sections were incubated in Alexa Fluor 488 anti-rabbit IgG (1:1000) [and Alexa Fluor 647 anti-mouse IgG antibody in some cases] and 4′,6-diamidino-2-phenylindole (DAPI, 1 μg/mg) for >2 days, and washed with PBS. From the sections, z-stack images were captured with a confocal microscope (Leica). In the images, the SLM/SR boundary (L0) was determined by the density difference in DAPI(+) nuclei (high in SLM, low in SR), and lines 20 μm away from the SLM/SR boundary were drawn on both the inner (L − 20) and outer (L + 20) sides. Z-stack images were smoothed by the 3D Gaussian lowpass filter function of ImageJ. All RFP(+) axons crossing both lines in one set of z-stack images (depth: 100 μm) were traced with the ImageJ plugin software, Simple Neurite Tracer44. For the statistical analysis, Fisher’s exact test was used.
Sagittal sections of the olfactory bulb were also stained with the anti-SERT antibody and DAPI with the same method. From these sections, z-stack images were captured with the confocal microscope. In the images, the GCL/IPL boundary was determined by the density difference of DAPI(+) nuclei (high in GCL, low in IPL), and lines 20 μm apart from the GCL/IPL boundary were drawn on both the inner (L − 20) and outer (L + 20) sides. All SERT(+) axons crossing both lines in one set of z-stack images (depth: 50 μm) were traced with Simple Neurite Tracer. The length of SERT(+) axons in boxes (50 × 50 × 50 μm) in the GCL and EPL was traced and measured with Simple Neurite Tracer. For the statistical analysis, Welch’s t-test was used. In the analyses of branch and end points in boxes (50 × 50 × 50 μm) in the GCL, an axon connecting with branch points was regarded as one axon, and its length was defined as its sum length. In the survival analyses of branch (or end) points, axons with branch (or end) points were dealt with as an event (death), and the other axons were regarded as being censored. For the statistical analyses, the log-rank test was used.
References
Scott, M. M. et al. A genetic approach to access serotonin neurons for in vivo and in vitro studies. Proc Natl Acad Sci USA 102, 16472–16477, https://doi.org/10.1073/pnas.0504510102 (2005).
Deneris, E. S. Molecular genetics of mouse serotonin neurons across the lifespan. Neuroscience 197, 17–27, https://doi.org/10.1016/j.neuroscience.2011.08.061 (2011).
Kiyasova, V. & Gaspar, P. Development of raphe serotonin neurons from specification to guidance. Eur J Neurosci 34, 1553–1562, https://doi.org/10.1111/j.1460-9568.2011.07910.x (2011).
Gaspar, P., Cases, O. & Maroteaux, L. The developmental role of serotonin: news from mouse molecular genetics. Nat Rev Neurosci 4, 1002–1012, https://doi.org/10.1038/nrn1256 (2003).
Lidov, H. G. & Molliver, M. E. An immunohistochemical study of serotonin neuron development in the rat: ascending pathways and terminal fields. Brain research bulletin 8, 389–430 (1982).
Donovan, S. L., Mamounas, L. A., Andrews, A. M., Blue, M. E. & McCasland, J. S. GAP-43 is critical for normal development of the serotonergic innervation in forebrain. J Neurosci 22, 3543–3552, doi:20026295 (2002).
Fournet, V. et al. The deletion of the microtubule-associated STOP protein affects the serotonergic mouse brain network. J Neurochem 115, 1579–1594, https://doi.org/10.1111/j.1471-4159.2010.07064.x (2010).
Katori, S. et al. Protocadherin-alpha family is required for serotonergic projections to appropriately innervate target brain areas. J Neurosci 29, 9137–9147, https://doi.org/10.1523/JNEUROSCI.5478-08.2009 (2009).
Kohmura, N. et al. Diversity revealed by a novel family of cadherins expressed in neurons at a synaptic complex. Neuron 20, 1137–1151 (1998).
Wu, Q. & Maniatis, T. A striking organization of a large family of human neural cadherin-like cell adhesion genes. Cell 97, 779–790 (1999).
Fukuda, E. et al. Down-regulation of protocadherin-alpha A isoforms in mice changes contextual fear conditioning and spatial working memory. Eur J Neurosci 28, 1362–1376, https://doi.org/10.1111/j.1460-9568.2008.06428.x (2008).
Thu, C. A. et al. Single-cell identity generated by combinatorial homophilic interactions between alpha, beta, and gamma protocadherins. Cell 158, 1045–1059, https://doi.org/10.1016/j.cell.2014.07.012 (2014).
Lefebvre, J. L., Kostadinov, D., Chen, W. V., Maniatis, T. & Sanes, J. R. Protocadherins mediate dendritic self-avoidance in the mammalian nervous system. Nature 488, 517–521, https://doi.org/10.1038/nature11305 (2012).
Noguchi, Y. et al. Total expression and dual gene-regulatory mechanisms maintained in deletions and duplications of the Pcdha cluster. J Biol Chem 284, 32002–32014, https://doi.org/10.1074/jbc.M109.046938 (2009).
Iwasato, T. et al. Cortex-restricted disruption of NMDAR1 impairs neuronal patterns in the barrel cortex. Nature 406, 726–731, https://doi.org/10.1038/35021059 (2000).
Arakawa, H. et al. Thalamic NMDA receptor function is necessary for patterning of the thalamocortical somatosensory map and for sensorimotor behaviors. J Neurosci 34, 12001–12014, https://doi.org/10.1523/JNEUROSCI.1663-14.2014 (2014).
Steinfeld, R., Herb, J. T., Sprengel, R., Schaefer, A. T. & Fukunaga, I. Divergent innervation of the olfactory bulb by distinct raphe nuclei. J Comp Neurol 523, 805–813, https://doi.org/10.1002/cne.23713 (2015).
Chen, W. V. et al. Pcdhalphac2 is required for axonal tiling and assembly of serotonergic circuitries in mice. Science 356, 406–411, https://doi.org/10.1126/science.aal3231 (2017).
Prasad, T. & Weiner, J. A. Direct and Indirect Regulation of Spinal Cord Ia Afferent Terminal Formation by the gamma-Protocadherins. Front Mol Neurosci 4, 54, https://doi.org/10.3389/fnmol.2011.00054 (2011).
Chen, W. V. et al. Functional significance of isoform diversification in the protocadherin gamma gene cluster. Neuron 75, 402–409, https://doi.org/10.1016/j.neuron.2012.06.039 (2012).
Chen, J. et al. alpha- and gamma-Protocadherins negatively regulate PYK2. J Biol Chem 284, 2880–2890, https://doi.org/10.1074/jbc.M807417200 (2009).
Woo, S., Rowan, D. J. & Gomez, T. M. Retinotopic mapping requires focal adhesion kinase-mediated regulation of growth cone adhesion. J Neurosci 29, 13981–13991, https://doi.org/10.1523/JNEUROSCI.4028-09.2009 (2009).
Myers, J. P., Robles, E., Ducharme-Smith, A. & Gomez, T. M. Focal adhesion kinase modulates Cdc42 activity downstream of positive and negative axon guidance cues. J Cell Sci. https://doi.org/10.1242/jcs.100107 (2012).
Mamounas, L. A., Blue, M. E., Siuciak, J. A. & Altar, C. A. Brain-derived neurotrophic factor promotes the survival and sprouting of serotonergic axons in rat brain. J Neurosci 15, 7929–7939 (1995).
Mamounas, L. A. et al. BDNF promotes the regenerative sprouting, but not survival, of injured serotonergic axons in the adult rat brain. J Neurosci 20, 771–782 (2000).
Schalm, S. S., Ballif, B. A., Buchanan, S. M., Phillips, G. R. & Maniatis, T. Phosphorylation of protocadherin proteins by the receptor tyrosine kinase Ret. Proc Natl Acad Sci USA 107, 13894–13899, https://doi.org/10.1073/pnas.1007182107 (2010).
Poulain, F. E. & Sobel, A. The microtubule network and neuronal morphogenesis: Dynamic and coordinated orchestration through multiple players. Mol Cell Neurosci 43, 15–32, https://doi.org/10.1016/j.mcn.2009.07.012 (2010).
Cardinaux, J. R., Magistretti, P. J. & Martin, J. L. Brain-derived neurotrophic factor stimulates phosphorylation of stathmin in cortical neurons. Brain Res Mol Brain Res 51, 220–228 (1997).
Jeanneteau, F., Deinhardt, K., Miyoshi, G., Bennett, A. M. & Chao, M. V. The MAP kinase phosphatase MKP-1 regulates BDNF-induced axon branching. Nat Neurosci 13, 1373–1379, https://doi.org/10.1038/nn.2655 (2010).
Ducray, A. et al. GDNF family ligands display distinct action profiles on cultured GABAergic and serotonergic neurons of rat ventral mesencephalon. Brain Res 1069, 104–112, https://doi.org/10.1016/j.brainres.2005.11.056 (2006).
Sarabi, A., Hoffer, B. J., Olson, L. & Morales, M. Glial cell line neurotrophic factor-family receptor alpha-1 is present in central neurons with distinct phenotypes. Neuroscience 116, 261–273, doi:S0306452202005596 [pii] (2003).
Zipursky, S. L. & Sanes, J. R. Chemoaffinity revisited: dscams, protocadherins, and neural circuit assembly. Cell 143, 343–353, https://doi.org/10.1016/j.cell.2010.10.009 (2010).
Wojtowicz, W. M. et al. A vast repertoire of Dscam binding specificities arises from modular interactions of variable Ig domains. Cell 130, 1134–1145, https://doi.org/10.1016/j.cell.2007.08.026 (2007).
Fuerst, P. G. et al. DSCAM and DSCAML1 function in self-avoidance in multiple cell types in the developing mouse retina. Neuron 64, 484–497, https://doi.org/10.1016/j.neuron.2009.09.027 (2009).
Esumi, S. et al. Monoallelic yet combinatorial expression of variable exons of the protocadherin-alpha gene cluster in single neurons. Nat Genet 37, 171–176, https://doi.org/10.1038/ng1500 (2005).
Azmitia, E. C. & Segal, M. An autoradiographic analysis of the differential ascending projections of the dorsal and median raphe nuclei in the rat. J Comp Neurol 179, 641–667, https://doi.org/10.1002/cne.901790311 (1978).
Hasegawa, S. et al. Distinct and Cooperative Functions for the Protocadherin-alpha, -beta and -gamma Clusters in Neuronal Survival and Axon Targeting. Front Mol Neurosci 9, 155, https://doi.org/10.3389/fnmol.2016.00155 (2016).
Tarusawa, E. et al. Establishment of high reciprocal connectivity between clonal cortical neurons is regulated by the Dnmt3b DNA methyltransferase and clustered protocadherins. BMC Biol 14, 103, https://doi.org/10.1186/s12915-016-0326-6 (2016).
Takeda, J., Keng, V. W. & Horie, K. Germline mutagenesis mediated by Sleeping Beauty transposon system in mice. Genome Biol 8(Suppl 1), S14, https://doi.org/10.1186/gb-2007-8-s1-s14 (2007).
Yagi, T. et al. A novel negative selection for homologous recombinants using diphtheria toxin A fragment gene. Anal Biochem 214, 77–86, https://doi.org/10.1006/abio.1993.1459 (1993).
Yagi, T. et al. A role for Fyn tyrosine kinase in the suckling behaviour of neonatal mice. Nature 366, 742–745, https://doi.org/10.1038/366742a0 (1993).
Hasegawa, S. et al. The protocadherin-alpha family is involved in axonal coalescence of olfactory sensory neurons into glomeruli of the olfactory bulb in mouse. Mol Cell Neurosci 38, 66–79, https://doi.org/10.1016/j.mcn.2008.01.016 (2008).
Luo, W. et al. Supernova: A Versatile Vector System for Single-Cell Labeling and Gene Function Studies in vivo. Sci Rep 6, 35747, https://doi.org/10.1038/srep35747 (2016).
Longair, M. H., Baker, D. A. & Armstrong, J. D. Simple Neurite Tracer: open source software for reconstruction, visualization and analysis of neuronal processes. Bioinformatics 27, 2453–2454, https://doi.org/10.1093/bioinformatics/btr390 (2011).
Acknowledgements
We thank the members of the KOKORO Biology Laboratory and Division of Neurogenetics for assistance and discussions. We are grateful to E. S. Deneris (Case Western Reserve University) for the ePet-Cre mouse, to M. Watanabe (Hokkaido University) for the anti-SERT antibody, and to K. Senzaki (University of Tsukuba) for the generation of targeting vectors. This work was supported by JSPS KAKENHI Grant Numbers 15K14889, 13J04498, and 22700369 (to S.K.); JSPS KAKENHI Grant Numbers 16H06459, 15H04263, and 15H01454 (to T.I.); JSPS KAKENHI Grant Numbers 19100006 (Grant-in-Aid for Scientific Research (S)), 23115513 (Innovative Areas “Mesoscopic Neurocircuitry”), Collaborative Research (A) of National Institute of Genetics, a Grant-in-Aid for Scientific Research on Innovative Areas (Comprehensive Brain Science Network) from the Ministry of Education, Science, Sports, and Culture of Japan (MEXT), and JST-CREST (to T.Y.).
Author information
Authors and Affiliations
Contributions
S.K. and T.Y. planned the experiments and S.K., T.I., and T.Y. wrote the manuscript. S.K. performed histological and data analyses. Y. N.-K., A.O., Y.K., K.S., and T.H. produced Pcdha mutant mice. T.I. produced Emx1-Cre and 5HTT-Cre mice. W.L. contributed to the virus experiment. T.Y. supervised the experiments.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Katori, S., Noguchi-Katori, Y., Okayama, A. et al. Protocadherin-αC2 is required for diffuse projections of serotonergic axons. Sci Rep 7, 15908 (2017). https://doi.org/10.1038/s41598-017-16120-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-16120-y
This article is cited by
-
Development of FRET-based indicators for visualizing homophilic trans interaction of a clustered protocadherin
Scientific Reports (2021)
-
Wiring the Brain by Clustered Protocadherin Neural Codes
Neuroscience Bulletin (2021)
-
Pcdhβ deficiency affects hippocampal CA1 ensemble activity and contextual fear discrimination
Molecular Brain (2020)
-
An epigenome-wide association study of early-onset major depression in monozygotic twins
Translational Psychiatry (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.