Abstract
Zoonotic tuberculosis is a risk for human health, especially when animals are in close contact with humans. Mycobacterium tuberculosis was cultured from several organs, including lung tissue and gastric mucosa, of three captive elephants euthanized in a Swiss zoo. The elephants presented weight loss, weakness and exercise intolerance. Molecular characterization of the M. tuberculosis isolates by spoligotyping revealed an identical profile, suggesting a single source of infection. Multilocus variable-number of tandem-repeat analysis (MLVA) elucidated two divergent populations of bacteria and mixed infection in one elephant, suggesting either different transmission chains or prolonged infection over time. A total of eight M. tuberculosis isolates were subjected to whole-genome sequence (WGS) analysis, confirming a single source of infection and indicating the route of transmission between the three animals. Our findings also show that the methods currently used for epidemiological investigations of M. tuberculosis infections should be carefully applied on isolates from elephants. Moreover the importance of multiple sampling and analysis of within-host mycobacterial clonal populations for investigations of transmission is demonstrated.
Similar content being viewed by others
Introduction
Infections with several members of the Mycobacterium tuberculosis complex (MTBC) can cause tuberculosis (TB) in both captive and free-ranging wildlife species, thus representing a considerable zoonotic risk1,2,3,4. In recent years, because of the broad host range of some MTBC members, numerous reports of MTBC infections in different species, including endangered animals, have been published5,6. Modern phylogenetic analysis defined MTBC into seven major lineages adapted to humans and two lineages adapted to various wild and domestic animal species7. In contrast to the human-adapted lineages, some animal-adapted members of the MTBC can establish independent transmission chains within different animal species8. Some wildlife species are recognized for their role of maintenance host, acting as a reservoir and transmitting the pathogens into domestic livestock, companion animals or human beings5,9. Particular attention has been given to badgers or wild boar as reservoirs of M. bovis, to red deer in the case of M. caprae and wild rodents regarding M. microti. The causative agent of human TB, M. tuberculosis, is an obligate pathogen known to maintain its life cycle of disease and transmission primarily in human populations8. However, infections of M. tuberculosis in a variety of domestic animals such as dogs, cattle, horses and parrots10,11,12,13 as well as in wild animal species such as elephants, rhinoceros, chimpanzee, lesser kudu, tapir and others have been described1,14,15,16. While in some species, such as cattle, M. tuberculosis infections are self-limiting and persistence of the pathogen into the population does not occur without exposure to humans11,17, cases where M. tuberculosis was maintained for a prolonged time within certain wildlife populations have been reported14. However, the dynamics of M. tuberculosis transmission among wild animals remain unclear.
Outbreak
Three captive Asian elephants (Elephas maximus) kept in a Swiss zoo presented unspecific clinical signs such as weight loss, weakness and exercise intolerance. The elephants were wild-born in Indonesia in the early 1960s, and they were moved to Europe at the age of three, four and eight. Living in a well frequented zoo, high contact with visitors exposed the animals to human pathogens over years. Infection with a member of the MTBC was suggested in February 2015 after clinical examination and a positive screen with DPP® VetTB Assay (Chembio Diagnostic Systems, New York, USA)18. This test consists of a lateral flow-based technology that detects serum antibodies specific for antigens MPB83 and ESAT-6/CFP-10 recombinant19. All three elephants resulted positive to the test for both antigens and were euthanized because of their deteriorated general condition between June-July 2015 by the official veterinarian attending the zoo animals. Necropsy and analysis of the specimens was carried out within a diagnostic context, meaning that no animals were killed for the purposes of this research project and ethical approval was not necessary. The privacy rights of the zoo were fully protected and the obtained data were de-identified.
The purpose of the present study was to determine whether the three elephants were infected by a single strain of M. tuberculosis or by multiple ones, identifying the potential index case and assessing the transmission route between the elephants. Additionally, this study aimed to describe the dissemination and microevolution of the bacterium within the elephant’s body.
Findings
Bacterial isolation from trunk wash
Serial trunk wash and subsequent culturing of the samples obtained was performed monthly for the three months prior to euthanasia for each elephant. Direct MTBC RT-PCR (described below) always resulted negative and growth of MTBC members was not detected. Mycobacterium vaccae and Rhodococcus equi were isolated in two samples at different periods from both Elephant 2 and Elephant 3. Additionally M. nonchromogenicum and M. monacense were isolated from Elephant 2 and Elephant 3, respectively.
Necropsy and M. tuberculosis isolation
All three animals presented multifocal to coalescing yellow-grey necroses of 5 mm to 2 cm in diameter in the lungs (Fig. 1, panel a). The necroses were often mineralized and larger abscesses were encapsulated by fibrosis. All animals showed moderate to severe arthrosis and were mildly anemic. Elephant 2 had severe suppurative discharge from the vagina that was caused by an endometrial cystic hypertrophy with secondary severe pyometra caused by Streptococcus agalactiae. The uterus of Elephant 3 showed histopathological features of an adenocarcinoma and presented metastases in the uterine ligaments (Table 1).
(a) Lung, Asian elephant tuberculosis. Multifocal to coalescing yellow-grey nodules, surrounded by connective tissue and a central caseous necrosis were observed during necropsy. (b) Histopathological examination revealed organized granulomatous inflammation with caseous necrosis, consistent with chronic TB-lesion, hematoxylin and eosin (HE). (c) Inflammation is characterized by numerous epithelioid macrophages (arrow) and several multinucleated Langhans giant cells (arrowhead), HE. (d) Alveolar macrophage with intracellular acid fast rods, Ziehl-Neelsen.
The lesions of the lungs revealed granuloma formations, displaying round lesions with central coagulative necrosis, surrounded by numerous epithelioid macrophages and several multinucleated Langhans giant cells. The giant cells showed a semilunar organization of their nuclei. Further, in the periphery of the granulomas, lymphocytes, plasma cells and proliferation of fibroblasts were conglomerated (Fig. 1, panels b-c). These findings were consistent with chronic TB lesions, indicating that the three elephants were infected over a long period of time. The lymph nodes were atrophied with a regular architecture but no granulomatous alterations were seen. Sinus histiocytosis was observed in the lymph nodes of the lungs. Acid fast bacilli were only found in alveolar macrophages and in histiocytes of the lung lymph nodes and quantified between scanty and 1+ according to the IUATLD and WHO grading scales (Fig. 1, panel d)20.
Spoligotyping and multilocus variable-number of tandem-repeat analysis (MLVA)
A total of eight isolates cultured from different organs (Table 2) were analyzed by spoligotyping and MLVA. Spoligotyping revealed an identical profile for all the isolates, namely SIT276. SIT276 is characterized by the lack of spacers 25–30 and 33–36. MLVA showed two distinct genotypes and a mixed infection in the right lung of Elephant 3. Locus variations were observed in VNTR577 and VNTR4156 by comparing the isolates from Elephant 1 with those generated from Elephant 2 and 3, respectively (Table 2).
Whole-genome sequencing
Genome-wide sequence data for eight isolates cultured from different organs of the three elephants were obtained. The sequencing data has been deposited in the European Nucleotide Archive (EMBL-EBI) under the study ID PRJEB21800 (see Supplementary Table S1). The read depth ranged from 19X to 160X and more than 99% of the MTBC reference genome was covered. The regions not covered primarily included members of the GC-rich and repetitive PE/PPE gene families21. Sequence analysis revealed that all eight isolates were human-adapted strains belonging to MTBC Lineage 422 and sublineage L4.10 also referred to as PGG37. When considering fixed nucleotide differences, the median pairwise distance between any two isolates was 6 SNPs, with a minimum of 0 SNPs and a maximum of 19 SNPs. The widest genetic distance (19 SNPs) was detected between the isolate originating from the gastric mucosa of Elephant 3 and the isolates from Elephant 1’s lungs (Fig. 2). However, the isolate from the pharyngeal swab of Elephant 1 and the isolate from the right lung of Elephant 3 had no fixed SNPs difference.
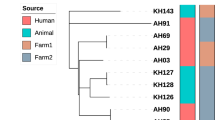
Maximum likelihood phylogeny using the GTR model of evolution obtained with RaxML using a 146 variable positions alignment. M. canettii was used as an outgroup and to root the phylogeny. Nodes supports are indicated by 1,000 bootstrap replicates. The maximum number of SNP differences between isolates from the same and different elephants are indicated in grey.
No phenotypic resistance was detected for the five first-line antimycobacterial antibiotics tested (isoniazid, rifampicin, ethambutol, pyrazinamide and streptomycin). WGS analysis confirmed the susceptibility of the isolates, as none of the sequences contained any drug resistance-conferring mutations, according to Walker and co-workers23.
Discussion
In contrast to the trunk wash results, necropsy and bacteriological culture confirmed shedding of M. tuberculosis based on the positive culture results from the trunk mucosa, tracheobronchial and pharyngeal swab in two out of three elephants. Accurate and robust ante mortem diagnostic tests for elephants are missing and have been limited by challenges associated with sample collection, performance and interpretation. The DPP® VetTB Assay has been accredited for diagnostic purposes by the U.S. Department of Agriculture who assure high confidence of results18. Although the antigens present in this test are reported to be highly specific for the MTBC, they were also reported to be expressed by Nontuberculous Mycobacteria (NTM) species such as M. kansasii and M. marinum 24,25. Similarly, a recent report showed that African elephants infected with M. szulgai cross-reacted with antigen MPB83, leading to false-positive results and to erroneous diagnosis with potentially fatal outcome26. The current gold standard for ante mortem diagnosis of TB infection in elephants is mycobacterial culture from respiratory secretions obtained by trunk wash. However, due to the intermittent shedding of the pathogen, a negative culture does not rule out an infection. Moreover, elephants use their trunks for a variety of activities and culture overgrowth with fungi or NTMs is a common problem27. Another limitation of the current diagnostic gold standard is the relatively slow reporting time, typically 6–8 weeks necessary for culture of MTBC members. While culture results are pending, a potential infected elephant may transmit the infection to other animals or humans that come in close contact. Before the outbreak period, seven additional elephants were present at the zoo with direct contact to the three TB infected animals. Once infection with TB was suspected based on DPP® VetTB Assay results, the remaining herd was separated and kept without direct contact to the three TB infected elephants These elephants were tested at regular intervals by trunk wash sampling without detection of MTBC, and at time of writing, all were alive and in good clinical condition. Although detection of TB was only possible post mortem, the three elephants presented various additional clinical problems, mostly involving the reproductive system and the locomotor activity. Therefore, euthanasia was based on the deteriorated general condition and not uniquely on a suspected TB infection. More controversial is the management of DPP® VetTB Assay positive and trunk-wash negative animals that live in close contact with other animals and do not present any clinical signs.
The higher sensitivity of spoligotyping compared with commercial available methods, e.g. GenoType® MTBC kit (HainLifescience, Nehren, Germany), allowed species determination of the isolates directly from the tissue’s DNA extract. This enables rapid implementation of epidemiological and therapeutic measures and, at the same time, characterization of the MTBC pathogen at the species level. Directly analyzed lung tissue from each elephant revealed three identical spoligotyping profiles, suggesting a single source of infection. However, since the discriminatory power of spoligotyping is too low to infer transmission chains reliably, MLVA offers usually a better alternative28,29. In human TB, double-locus variations observed between different clinical isolates of M. tuberculosis usually reflect infection with distinct clones with no direct epidemiological linkage and therefore belonging to different transmission chains30. However, little is known about the genetic variability of M. tuberculosis within elephants. One possibility is that prolonged infection occurred. This allowed the establishment of different clonal mycobacterial herds showing a marked within-host genetic divergence. The present results indicate therefore the importance of multiple sampling for epidemiological investigation of TB in elephants (Fig. 2). The eight isolates of the present study were classified as M. tuberculosis Lineage 4, belonging to the so-called principal genetic group 3 (PGG3)31, which is geographically one of the most widespread causes of human TB worldwide. Very low genetic distance among the eight isolates was observed overall. The average distance was 6 SNPs, which is below the threshold of 12 SNPs proposed by Walker et al. for epidemiologically linked/unlinked cases32. In detail, according to the proposed thresholds, an epidemiological linkage is expected between isolates differing by five or fewer SNPs, whereas isolates that differ by more than 12 SNPs are supposed to be part of different transmission chains32. The elephant isolates are most likely the result of a transmission chain, potentially part of a larger cluster with human origin. Based on WGS data and MLVA genotyping, Elephant 3 showed the largest within-host genetic divergence compared with the other two animals, displaying both MLVA profiles in the right lung and a distance of 15 SNPs between the trunk and gastric mucosa isolates compared with the right lung isolate (Fig. 2 and Table 2). Furthermore, the marked divergence of 8 SNPs observed between the isolates from Elephant 1 and Elephant 2 suggests transmission events involving single distinct genotypes originating from Elephant 3, which most likely represent the spreader case. It is intriguing that two of the four isolates from Elephant 3, namely those generated from the gastric– and the trunk mucosa respectively, showed a SNPs distance above the threshold of 12 SNPs if compared with the isolates of Elephant 1 (Fig. 2). This indicates that the analysis of a single sample from Elephant 3, e.g. by trunk wash, would have led to a questionable epidemiological elucidation of the outbreak. Moreover the observed mycobacterial genetic variation within Elephant 3 suggest a particularly prolonged infection, leading to a level of clonal divergence exceeding the proposed maximal limit of single-locus variation seen in human infection and human-to-human transmission. The described results show the high discriminatory power of MLVA between genetically close M. tuberculosis isolates and correlate with the WGS analysis findings. Taking into account the accessibility of MLVA for laboratory with restricted resources and the straightforward performance compared to WGS analysis, MLVA represent a valid and effective technique for genotyping MTBC isolates of animal origin. An exhaustive attempt to delineate the occurred transmission between the three animals however was only possible though WGS analysis, proving a greater resolution compared to MLVA33.
Antituberculosis therapy among captive elephants is not standardized but sporadic successful treatments have been reported34. General condition of diseased animals and management of potentially MTBC shedding patients represent however crucial aspects of antimycobacterial therapy in elephants. All elephant isolates were susceptible to the antimycobacterial agents tested. Recently, novel mutations associated with streptomycin resistance have been reported in humans35. Although all eight genomes showed a non-synonymous mutation in gidB_Ala205Thr, this mutation has also been reported in two susceptible isolates from Belarus35.
Our findings confirm transmission of M. tuberculosis between the three elephants, and from the public health point of view, these animals constituted a potential source for TB infection in humans. On the other hand, since the M. tuberculosis lineage detected is very common in humans, reverse zoonotic transmission from human to elephant most likely occurred, highlighting the importance of protection of this endangered animal species from infection.
Therefore, closer collaboration between public health, veterinary and occupational health officers is recommended.
Material and Methods
Culture and identification of mycobacteria
Trunk wash samples were collected according to the guidelines for the diagnosis and control of TB in captive elephants with slight modifications36. Briefly, 250 ml of sterile saline solution were instilled into one elephant’s nostril, the trunk was lifted in order to distribute the fluid and finally the elephant exhaled the saline in a sterile collection plastic bag. Trunk wash samples were collected for each elephant in a series of three separate days within a one-week period. The collected fluid was centrifuged 15 min at 3000 × g and the sediment decontaminated as previously described37. For the microbiological analysis, two different lung localizations (left and right lung), several lymph nodes (pooled), gastric mucosa, trunk mucosa, pharyngeal- and tracheobronchial swabs were collected from each elephant and processed independently according to Ghielmetti and co-workers37. Decontaminated speciments, including trunk wash fluid, organ samples and swabs were tested using artus® M. tuberculosis PCR Kit with a Rotor-Gene Q device according to the manufacturer’s protocol (QIAGEN, Hilden, Germany) and used as inoculum for culture. Cultures showing growth of suspicious mycobacterial colonies were checked for acid-fast bacilli after Ziehl-Neelsen staining and identified as MTBC by artus® M. tuberculosis PCR. Suspicious mycobacterial colonies yielding negative results by MTBC specific PCR were identified by sequencing the 16S rDNA gene38. Subcultures were established on Middlebrook 7H10 agar plates (Becton, Dickinson & Company, Allschwil, Switzerland) using mycobacteria harvested from 1.5 ml MGIT cultures by centrifugation for 10 min at 13000 x g. To avoid loss of genetic variability among mycobacteria obtained by culture, multiple colonies (5 to 10) from Middlebrook 7H10 agar plates (BD) were picked for DNA extraction. Automated DNA purification was completed on a QIAcube instrument in accordance with the QIAamp cador Pathogen Mini Kit protocol (QIAGEN).
Spoligotyping and MLVA
Spoligotyping was performed by using a commercial microarray system with integrated data analysis (Alere Technologies, Jena, Germany)37. For each elephant, one lung sample was tested directly by spoligotyping prior to culturing. Additionally, all isolates obtained by culture were analyzed individually. The spoligotype patterns were assigned according to the international nomenclature (SITVIT database). MLVA was performed using the standard 24 markers panel used for M. tuberculosis typing according to Supply et al.30. In addition, two hypervariable loci (VNTR2163a and VNTR3232) were included. The single markers were amplified by end-point PCR and subsequently analyzed using a capillary electrophoresis device37.
Whole-genome sequencing and sequence analysis
Whole-genome sequencing (WGS) was performed using the Nextera XT library preparation kit and the HiSeq 2500 Illumina sequencer. The FastQ files containing the raw paired-end reads were processed using a python pipeline developed in house as follows. The reads were first adapter- and quality- trimmed with Trimmomatic v0.3339. Reads lower than 20 bp were not kept for the downstream analysis. Overlapping paired-end reads were then merged with SeqPrep (https://github.com/jstjohn/SeqPrep). The resulting filtered reads were mapped to a hypothetical reconstructed MTBC ancestor22 with BWA v0.7.1240. Duplicated reads were marked by the MarkDuplicates module of Picard v 2.1.1 (https://github.com/broadinstitute/picard). The RealignerTargetCreator and IndelRealigner modules of GATK v.3.4.041 were used to perform local realignment of reads around indels. SNPs were called with Samtools v1.242 and VarScan v2.4.143 using the following thresholds: minimum mapping quality of 20, minimum base quality at a position of 20 and minimum read depth at a position of 7X. SNPs were considered fixed at a frequency of ≥90% and alleles were considered ancestral when the SNP frequency was ≤10%. Furthermore, SNPs were called only if the alternative basecall was supported by at least five reads and without strand bias. All variants were annotated using snpEff v4.1144, in accordance with the M. tuberculosis H37Rv reference annotation (AL123456.3). SNPs falling in regions with at least 50 bp identity to other regions in the genome were excluded from the analysis45.
The variable SNPs alignment was obtained by concatenating the SNP calls present in the variant calling file of each genome, using the IUPAC nucleotide ambiguity codes for heterozygous calls. A position was considered variable if at least one genome had a SNP at that position. Called deletions and positions not called according to the minimum threshold of 7 were encoded as gaps. Positions for which the proportion of gaps exceeded 50% were excluded from the alignment. The variable SNPs alignment was used to infer a maximum likelihood phylogeny using RaxML 8.2.846, applying the general time-reversible (GTR) model of evolution. The confidence of nodes was assessed by bootstrapping 1000 pseudoreplicates, and M. canettii was used as an outgroup and to root the phylogeny. Number of SNPs difference between every pairwise comparison was obtained with Mega647, reporting only fixed SNPs within a genome.
Drug susceptibility testing (DST)
The World Health Organization and the U.S. Centre for Disease Control and Prevention recommend the use of liquid culture systems for DST and for improving time to detection48,49,50. To provide an effective prophylactic drug regimen for contact animals that live close to the positive patients, phenotypic antimycobacterial susceptibility testing was performed. For each elephant, one cultured isolate was tested for drug resistance using the Bactec MGIT 960 SIRE (streptomycin STR, isoniazid INH, rifampicin RMP and ethambutol EMB) Kits and the 960 PZA (pyrazinamide PZA) Kit according to manufacturer’s instructions (BD).
Data Availability
The datasets generated and analysed during the current study are available in the European Nucleotide Archive (EMBL-EBI) under the study ID PRJEB21800. http://www.ebi.ac.uk/ena/data/view/PRJEB21800.
References
Michalak, K. et al. Mycobacterium tuberculosis infection as a zoonotic disease: transmission between humans and elephants. Emerg Infect Dis. 4, 283–287, https://doi.org/10.3201/eid0402.980217 (1998).
Montali, R. J., Mikota, S. K. & Cheng, L. I. Mycobacterium tuberculosis in zoo and wildlife species. Rev Sci Tech. 20, 291–303 (2001).
Murphree, R., Warkentin, J. V., Dunn, J. R., Schaffner, W. & Jones, T. F. Elephant-to-human transmission of tuberculosis, 2009. Emerg Infect Dis. 17, 366–371, https://doi.org/10.3201/eid1703.101668 (2011).
Zlot, A. et al. Diagnosis of Tuberculosis in Three Zoo Elephants and a Human Contact - Oregon, 2013. MMWR Morb Mortal Wkly Rep. 64, 1398–1402, https://doi.org/10.15585/mmwr.mm6452a2 (2016).
Chandranaik, B. M. et al. Mycobacterium tuberculosis Infection in Free-Roaming Wild Asian Elephant. Emerg Infect Dis. 23, 555–557, https://doi.org/10.3201/eid2303.161439 (2017).
Miller, M. A., Buss, P. E., van Helden, P. D. & Parsons, S. D. Mycobacterium bovis in a Free-Ranging Black Rhinoceros, Kruger National Park, South Africa, 2016. Emerg Infect Dis. 23, 557–558, https://doi.org/10.3201/eid2303.161622 (2017).
Stucki, D. et al. Mycobacterium tuberculosis lineage 4 comprises globally distributed and geographically restricted sublineages. Nat Genet. 48, 1535–1543, https://doi.org/10.1038/ng.3704 (2016).
Brites, D. & Gagneux, S. Co-evolution of Mycobacterium tuberculosis and Homo sapiens. Immunol Rev. 264, 6–24, https://doi.org/10.1111/imr.12264 (2015).
Prodinger, W. M., Eigentler, A., Allerberger, F., Schonbauer, M. & Glawischnig, W. Infection of red deer, cattle, and humans with Mycobacterium bovis subsp. caprae in western Austria. J Clin Microbiol. 40, 2270–2272 (2002).
Lyashchenko, K. P. et al. Pulmonary Disease due to Mycobacterium tuberculosis in a Horse: Zoonotic Concerns and Limitations of Antemortem Testing. Vet Med Int. 2012, 642145, https://doi.org/10.1155/2012/642145 (2012).
Ocepek, M., Pate, M., Zolnir-Dovc, M. & Poljak, M. Transmission of Mycobacterium tuberculosis from human to cattle. J Clin Microbiol. 43, 3555–3557, https://doi.org/10.1128/JCM.43.7.3555-3557.2005 (2005).
Ribeiro, M. G. et al. Pre-Multidrug-Resistant Mycobacterium tuberculosis Infection Causing Fatal Enteric Disease in a Dog from a Family with History of Human Tuberculosis. Transbound Emerg Dis. https://doi.org/10.1111/tbed.12513 (2016).
Schmidt, V., Schneider, S., Schlomer, J., Krautwald-Junghanns, M. E. & Richter, E. Transmission of tuberculosis between men and pet birds: a case report. Avian Pathol. 37, 589–592, https://doi.org/10.1080/03079450802428901 (2008).
Michel, A. L. et al. Mycobacterium tuberculosis at the human/wildlife interface in a high TB burden country. Transbound Emerg Dis. 60(Suppl 1), 46–52, https://doi.org/10.1111/tbed.12099 (2013).
Miller, M., Michel, A., van Helden, P. & Buss, P. Tuberculosis in Rhinoceros: An Underrecognized Threat? Transbound Emerg Dis., https://doi.org/10.1111/tbed.12489 (2016).
Zachariah, A. et al. Mycobacterium tuberculosis in Wild Asian Elephants, Southern India. Emerg Infect Dis. 23, 504–506, https://doi.org/10.3201/eid2303.161741 (2017).
Eisenberg, T. et al. Mycobacterium tuberculosis exposure of livestock in a German dairy farm: implications for intra vitam diagnosis of bovine tuberculosis in an officially tuberculosis-free country. Epidemiol Infect. 144, 724–731, https://doi.org/10.1017/S0950268815001995 (2016).
Lyashchenko, K. P. et al. Tuberculosis in elephants: antibody responses to defined antigens of Mycobacterium tuberculosis, potential for early diagnosis, and monitoring of treatment. Clin Vaccine Immunol. 13, 722–732, https://doi.org/10.1128/CVI.00133-06 (2006).
Roos, E. O. et al. Test performance of three serological assays for the detection of Mycobacterium bovis infection in common warthogs (Phacochoerus africanus). Vet Immunol Immunopathol. 182, 79–84, https://doi.org/10.1016/j.vetimm.2016.10.006 (2016).
Reider, H., Van Deun, A. & Kam, K. Grading scales for bright field (Ziehl-Neelsen) and fluorescence microscopy, Priorities for Tuberculosis Bacteriology Services in Low-income countries. International Union Against Tuberculosis and Lung Disease. (2007).
Brennan, M. J. & Delogu, G. The PE multigene family: a ‘molecular mantra’ for mycobacteria. Trends Microbiol. 10, 246–249 (2002).
Comas, I. et al. Human T cell epitopes of Mycobacterium tuberculosis are evolutionarily hyperconserved. Nat Genet. 42, 498–503, https://doi.org/10.1038/ng.590 (2010).
Walker, T. M. et al. Whole-genome sequencing for prediction of Mycobacterium tuberculosis drug susceptibility and resistance: a retrospective cohort study. Lancet Infect Dis. 15, 1193–1202, https://doi.org/10.1016/S1473-3099(15)00062-6 (2015).
Vordermeier, H. M. et al. Assessment of cross-reactivity between Mycobacterium bovis and M. kansasii ESAT-6 and CFP-10 at the T-cell epitope level. Clin Vaccine Immunol. 14, 1203–1209, https://doi.org/10.1128/CVI.00116-07 (2007).
Waters, W. R. et al. Immune responses to defined antigens of Mycobacterium bovis in cattle experimentally infected with Mycobacterium kansasii. Clin Vaccine Immunol. 13, 611–619, https://doi.org/10.1128/CVI.00054-06 (2006).
Lacasse, C. et al. Two cases of atypical mycobacteriosis caused by Mycobacterium szulgai associated with mortality in captive African elephants (Loxodonta africana). J Zoo Wildl Med. 38, 101–107, https://doi.org/10.1638/06-051.1 (2007).
Mikota, S. K. & Maslow, J. N. Tuberculosis at the human-animal interface: an emerging disease of elephants. Tuberculosis (Edinb). 91, 208–211, https://doi.org/10.1016/j.tube.2011.02.007 (2011).
Comas, I., Homolka, S., Niemann, S. & Gagneux, S. Genotyping of genetically monomorphic bacteria: DNA sequencing in Mycobacterium tuberculosis highlights the limitations of current methodologies. PLoS One. 4, e7815, https://doi.org/10.1371/journal.pone.0007815 (2009).
Vadwai, V., Shetty, A., Supply, P. & Rodrigues, C. Evaluation of 24-locus MIRU-VNTR in extrapulmonary specimens: study from a tertiary centre in Mumbai. Tuberculosis (Edinb). 92, 264–272, https://doi.org/10.1016/j.tube.2012.01.002 (2012).
Supply, P. et al. Proposal for standardization of optimized mycobacterial interspersed repetitive unit-variable-number tandem repeat typing of Mycobacterium tuberculosis. J Clin Microbiol. 44, 4498–4510, https://doi.org/10.1128/JCM.01392-06 (2006).
Sreevatsan, S. et al. Restricted structural gene polymorphism in the Mycobacterium tuberculosis complex indicates evolutionarily recent global dissemination. Proc Natl Acad Sci USA 94, 9869–9874 (1997).
Walker, T. M. et al. Whole-genome sequencing to delineate Mycobacterium tuberculosis outbreaks: a retrospective observational study. Lancet Infect Dis. 13, 137–146, https://doi.org/10.1016/S1473-3099(12)70277-3 (2013).
Gardy, J. L. et al. Whole-genome sequencing and social-network analysis of a tuberculosis outbreak. N Engl J Med. 364, 730–739, https://doi.org/10.1056/NEJMoa1003176 (2011).
Simpson, G. et al. Mycobacterium tuberculosis Infection among Asian Elephants in Captivity. Emerg Infect Dis. 23, 513–516, https://doi.org/10.3201/eid2303.160726 (2017).
Wollenberg, K. R. et al. Whole-Genome Sequencing of Mycobacterium tuberculosis Provides Insight into the Evolution and Genetic Composition of Drug-Resistant Tuberculosis in Belarus. J Clin Microbiol. 55, 457–469, https://doi.org/10.1128/JCM.02116-16 (2017).
Animal and Plant Health Inspection Service. Guidelines for the Control of Tuberculosis in Elephants, https://www.aphis.usda.gov/aphis/ourfocus/animalwelfare/SA_Animal_Welfare_News/CT_USDA_to_Consider_Guidelines_for_the_Control_of_Tuberculosis_in_Elephants (2010).
Ghielmetti, G. et al. Epidemiological tracing of bovine tuberculosis in Switzerland, multilocus variable number of tandem repeat analysis of Mycobacterium bovis and Mycobacterium caprae. PLoS ONE. 12, e0172474, https://doi.org/10.1371/journal.pone.0172474 (2017).
Lane, D. J. In Nucleic acid techniques in bacterial systematics (ed E. Stackebrandt) Ch. 16S/23S rRNA sequencing, 115–175 (John Wiley & Sons., 1991).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 30, 2114–2120, https://doi.org/10.1093/bioinformatics/btu170 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 25, 1754–1760, https://doi.org/10.1093/bioinformatics/btp324 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303, https://doi.org/10.1101/gr.107524.110 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 25, 2078–2079, https://doi.org/10.1093/bioinformatics/btp352 (2009).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576, https://doi.org/10.1101/gr.129684.111 (2012).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin). 6, 80–92, https://doi.org/10.4161/fly.19695 (2012).
Stucki, D. et al. Standard Genotyping Overestimates Transmission of Mycobacterium tuberculosis among Immigrants in a Low-Incidence Country. J Clin Microbiol. 54, 1862–1870, https://doi.org/10.1128/JCM.00126-16 (2016).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 30, 1312–1313, https://doi.org/10.1093/bioinformatics/btu033 (2014).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol Biol Evol. 30, 2725–2729, https://doi.org/10.1093/molbev/mst197 (2013).
World Health Organization. Use of liquid TB culture and drug susceptibility testing (DST) in low and medium income settings. (Geneva, 2007).
Kent, P. T. & Kubica, G. T. Public health mycobacteriology: a guide for the level III laboratory. (Atlanta, 1985).
Tenover, F. C. et al. The resurgence of tuberculosis: is your laboratory ready? J Clin Microbiol. 31, 767–770 (1993).
Acknowledgements
Calculations were performed at sciCORE (http://scicore.unibas.ch/) scientific computing core facility at the University of Basel. Work in our group is supported by the Swiss National Science Foundation (grants 310030_166687, IZRJZ3_164171 and IZLSZ3_170834), the European Research Council (309540-EVODRTB), and SystemsX.ch.
Author information
Authors and Affiliations
Contributions
G.G. and S.G. supervised and conceived the project; M.R. directed and completed necropsy and histology, U.F. isolated the mycobacteria and performed drug susceptibility testing, G.G. identified the isolates and performed spoligotyping and MLVA, J.F. provided library preparation for WGS, M.C., D.S. and C.L. developed the pipeline and performed WGS analysis, M.C. designed the variable SNPs alignment, G.G. and M.C. wrote the manuscript with the contribution of all co-authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ghielmetti, G., Coscolla, M., Ruetten, M. et al. Tuberculosis in Swiss captive Asian elephants: microevolution of Mycobacterium tuberculosis characterized by multilocus variable-number tandem-repeat analysis and whole-genome sequencing. Sci Rep 7, 14647 (2017). https://doi.org/10.1038/s41598-017-15278-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15278-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.