Abstract
The relationship between evolutionary rates and gene expression in model plant orthologs is well documented. However, little is known about the relationships between gene expression and evolutionary trends in Arachis orthologs. We identified 7,435 one-to-one orthologs, including 925 single-copy and 6,510 multiple-copy sequences in Arachis duranensis and Arachis ipaënsis. Codon usage was stronger for shorter polypeptides, which were encoded by codons with higher GC contents. Highly expressed coding sequences had higher codon usage bias, GC content, and expression breadth. Additionally, expression breadth was positively correlated with polypeptide length, but there was no correlation between gene expression and polypeptide length. Inferred selective pressure was also negatively correlated with both gene expression and expression breadth in all one-to-one orthologs, while positively but non-significantly correlated with gene expression in sequences with signatures of positive selection. Gene expression levels and expression breadth were significantly higher for single-copy genes than for multiple-copy genes. Similarly, the gene expression and expression breadth in sequences with signatures of purifying selection were higher than those of sequences with positive selective signatures. These results indicated that gene expression differed between single-copy and multiple-copy genes as well as sequences with signatures of positive and purifying selection.
Similar content being viewed by others
Introduction
Molecular biology and evolution research in the later 20th century revealed that homologous genes can be divided into paralogs and orthologs1. Paralogous genes, or paralogs, are derived from sequence duplication events within a single lineage2. Paralogs are often free to evolve novel functions because the functional redundancy provided by gene duplicates frees one of the copies from the selective constraint maintaining its function prior to duplication3. In contrast, orthologous genes, or orthologs, are distributed among different species that diverged from a single ancestral gene at a speciation event2. Accordingly, orthologs typically perform equivalent functions across different species. Therefore, these genes can be used to construct phylogenetic relationships and provide insight into the processes of molecular evolution2. For example, data from β-tubulin and translation elongation factor sequences suggest that Epichloë species likely originated in Eurasia4,5. Similar types of molecular data from orthologs have been used to make comparisons of the evolutionary rates between gymnosperms and angiosperms, revealing lower evolutionary rates among gymnosperms6. Yue, et al.7 found that annual plant species (i.e., Arabidopsis thaliana and Medicago truncatula) have evolved at higher rates than perennial plant species (i.e., Populus trichocarpa and Vitis vinifera) using both nuclear and chloroplast genome loci.
Other evolutionary patterns have also revealed themselves in the study of orthologs. For example, highly expressed genes have potentially undergone stronger purifying selection based on their important functional roles that require high transcript levels2. The levels and patterns of expression are not only the major determinants that explain nonsynonymous rate variation among genes but also a crucial determinant of gene retention rates after duplication8. Recently, Grusz, et al.9 demonstrated that the faster evolutionary rates characteristic of vittarioid fern lineages are likely not driven by positive selection, nor are they limited to any particular type of nucleotide substitution. Hodgins, et al.10 found that genes with low expression levels have a lot of neutral substitutions, but rapidly diverging genes tend to have higher expression divergence in conifers. In brief, orthologs are well suited to revealing phylogenetic relationships and understanding the mechanisms shaping gene expression.
Generally, gene expression can be changed by many factors. For example, tissue-specific expression, DNA dosage, tRNA abundance and external environment factors can affect gene expression patterns11,12,13,14. A growing body of studies has revealed that several gene sequence architecture features, including synonymous codon usage, amino acid composition, coding sequences (CDSs) length, GC content, and intron size, are correlated with expression levels in prokaryotes and eukaryotes15,16,17. Similarly, highly expressed genes are biased towards using optimal codons in the Chilodonella uncinata genome18. Camiolo, et al.19 found short and GC-rich CDSs were positively correlated with expression and optimal usage bias in four monocot, fifteen dicot, and two moss species. In Silene latifolia, gene expression was positively correlated with third codon position GC content (GC3), but strongly and negatively correlated with intronic GC content20. Moreover, gene expression has been shown to be correlated with the size of gene families. Larger multiple-copy gene families exhibit both lower expression levels and breadth than genes in single-copy gene families17. In addition, tissue-specific expression was more often observed among genes in multiple-copy gene families than genes in single-copy gene families17.
Peanut is a major oil and protein crop. Cultivated peanut (Arachis hypogaea) is an allotetraploid species with an AABB genome21. Its ancestral species are most likely Arachis duranensis and Arachis ipaënsis, which contributed the A and B genomes, respectively22,23,24,25. Recently, the genome sequences of A. duranensis and A. ipaënsis have been sequenced, assembled, and released26. A comprehensive tissue-specific transcriptome of cultivated peanut has also been released26,27. Collectively, this research can be used to understand the relationship between evolutionary trends and gene expression. In this study, we identified orthologs from A. duranensis and A. ipaënsis, and used codon usage bias, polypeptide length, GC content, substitution rate, expression breadth, and gene expression level as explanatory variables to evaluate patterns among orthologs in A. duranensis and A. ipaënsis. Our study had two key aims: (1) to examine the codon usage pattern and relationship between codon usage bias, expression level, and substitution rate in these one-to-one orthologs and (2) to characterize differences in the evolutionary trends and expression patterns between genes in single-copy gene families and those in multiple-copy gene families from these orthologs. This study provides insight into the evolution and expression of gene families in Arachis.
Results
One-to-one orthologs
A total of 7,435 one-to-one ortholog pairs were used in this study after the filtering criteria (see Materials and Methods) were applied to the A. duranensis and A. ipaënsis genomes (Table S1). These selected gene sequences can be classified into two groups based on the number of genes in their gene families, single-copy gene families (with one gene) and multiple-copy gene families (with more than one gene). We identified 925 single-copy and 6,510 multiple-copy genes in both genomes. Bertioli, et al.26 found approximately 9,236 sequences belonged to multiple-copy gene families in A. ipaënsis or A. duranensis, while 6,357 and 7,253 sequences belonged to single-copy gene families in A. ipaënsis and A. duranensis, respectively. Many single-copy genes were detected in A. duranensis and A. ipaënsis, indicating these genes originated before the divergence of A. duranensis and A. ipaënsis approximately 2.16 million years ago26. Although wild-type peanut underwent a whole genome duplication (WGD) event28, these single-copy genes have exhibited no changes in number.
Codon usage
The GC contents of each of the three codon positions were assessed across all identified CDSs as GC1, GC2, and GC3, which correspond to the GC content of the first, second, and third codon positions, respectively (Table S2). The average GC1 was 50.37%, followed by GC3 at 42.95% and GC2 at 40.44% in A. duranensis CDSs. Similarly, A. ipaënsis exhibited average GC1, GC3, and GC2 values of 50.37%, 43.01%, and 40.44%, respectively. The average GC contents across the three codons positions of CDSs were 44.59% in A. duranensis and 44.61% in A. ipaënsis, indicating that two wild peanuts have high a AT content (i.e., 55.41% in A. duranensis and 55.39% in A. ipaënsis) in CDSs. These results are consistent with previous research in eudicots, which found that GC1 content exceeded GC2 content, GC3 content exceeded GC2 content, and AT content exceeded GC content overall29. The average frequency of optimal codons (Fop) value was 0.39 in both A. duranensis and A. ipaënsis. Fop was negatively correlated with polypeptide length but positively correlated with GC content in A. duranensis and A. ipaënsis (Table 1). These results showed that codon usage bias in A. duranensis and A. ipaënsis one-to-one orthologs tended towards shorter polypeptides having higher GC contents.
Gene expression
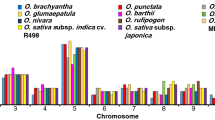
Although gene expression patterns in most tissues were similar between A. duranensis and A. ipaënsis one-to-one orthologs, gene expression patterns in main stem leaf, lateral leaf, Pattee 1 (i.e., stage 1) pod, Pattee 3 pod, Pattee 5 pericarp, Pattee 6 pericarp, Pattee 5 seed, and Pattee 6 seed tissues were biased (Fig. 1 and Table S3). For example, gene expression patterns in A. duranensis and A. ipaënsis perianth tissue were similar, while gene expression patterns in A. duranensis main stem leaves and lateral leaves differed from those in A. ipaënsis (Fig. 1 and Table S3). In addition, expression breadth of 1,873 ortholog pairs differed between A. duranensis and A. ipaënsis (Table S4); for example, Aradu.NIH31 and Araip.I19HU were an ortholog pair, while their expression breadth values were 1 and 22, respectively. These results revealed biases in gene expression patterns and expression breadth between A. duranensis and A. ipaënsis. Moreover, these results are consistent with those described by Clevenger, et al.27, who found that 76.8–98.1% of expressed gene pairs exhibited balanced expression, but striking differences in expression bias were exhibited in a tissue-specific context in the two cultivated peanut subgenomes.
Gene expression level of one-to-one orthologs in 22 tissues in Arachis duranensis and Arachis ipaënsis based on RNA-seq data. (A1 and B1) seedling leaf 10 d post emergence; (A2 and B2) main stem leaf; (A3 and B3) lateral leaf; (A4 and B4) vegetative shoot tip from the main stem; (A5 and B5) reproductive shoot tip from the first lateral leaf; (A6 and B6) 10 d roots; (A7 and B7) 25 d nodules; (A8 and B8) perianth; (A9 and B9) gynoecium; (A10 and B10) androecium; (A11 and B11) aerial gynophore tip; (A12 and B12) subterranean gynophore tip; (A13 and B13) Pattee 1 pod; (A14 and B14) Pattee 1 stalk; (A15 and B15) Pattee 3 pod; (A16 and B16) Pattee 5 pericarp; (A17 and B17) Pattee 5 seed; (A18 and B18) Pattee 6 pericarp; (A19 and B19) Pattee 6 seed; (A20 and B20) Pattee 7 seed; (A21 and B21) Pattee 8 seed; (A22 and B22) Pattee 10 seed. The FPKM value for each gene in these various tissues was normalized using a log2-transformation.
There was a positive correlation between Fop, GC content, expression breadth, and gene expression level, but no correlation between polypeptide length and gene expression level except for negative correlations between gene expression level and both seedling leaf 10 d post emergence and perianth tissues in A. duranensis and A. ipaënsis one-to-one orthologs (Fig. 2 and Table S5). Furthermore, we identified correlations between Fop, polypeptide length, GC content, expression breadth, and average gene expression levels among 22 tissues (Fig. 2 and Table S5). Average expression level was positively correlated with Fop, GC content, and expression breadth (Fig. 2 and Table S5). However, there was also no correlation between polypeptide length and average gene expression level. These results indicated that highly expressed one-to-one orthologs with higher GC content have codon usage bias (as demonstrated by Fop) and broad expression breadth in one-to-one orthologs. In addition, expression breadth was positively correlated with polypeptide length, GC1, GC2, and overall GC content in one-to-one orthologs (Table 2). A given gene with broader expression breadth tended to have longer polypeptide lengths and higher GC content, including GC1, GC2, and overall GC content in one-to-one orthologs.
Correlation analyses of gene expression level and various attributes of Arachis duranensis and Arachis ipaënsis one-to-one orthologs. The analyzed sequence attributes include frequency of optimal codons (Fop), polypeptide length, GC1 content, GC2 content, GC3 content, overall GC content, expression breadth, Ka, Ks, and Ka/Ks. (A and B) Arachis duranensis and Arachis ipaënsis, respectively. (1) seedling leaf 10 d post emergence; (2) main stem leaf; (3) lateral leaf; (4) vegetative shoot tip from the main stem; (5) reproductive shoot tip from the first lateral leaf; (6) 10 d roots; (7) 25 d nodules; (8) perianth; (9) gynoecium; (10) androecium; (11) aerial gynophore tip; (12) subterranean gynophore tip; (13) Pattee 1 pod; (14) Pattee 1 stalk; (15) Pattee 3 pod; (16) Pattee 5 pericarp; (17) Pattee 5 seed; (18) Pattee 6 pericarp; (19) Pattee 6 seed; (20) Pattee 7 seed; (21) Pattee 8 seed; (22) Pattee 10 seed; (23) average gene expression level. The heat map was constructed based on Pearson correlation coefficients in Table S5. **Indicates significance at P < 0.01.
Substitution rate
For a total of 6,732 one-to-one orthologs, Ka (nonsynonymous per site substitution rate) and Ks (synonymous per site substitution rate) values were calculated and filtering criteria were applied (Table S6). The average Ka, Ks, and Ka/Ks values were 0.05, 0.02, and 0.30, respectively. There were 6,598 Ka/Ks ratio values (nonsynonymous to synonymous per site substitution rate ratios) less than 1, indicating these one-to-one orthologs underwent purifying selection. However, there were 134 Ka/Ks values greater than 1 (Table S7), indicating positive selection shaped these one-to-one orthologs. The average synonymous substitution rate of the 6,732 one-to-one orthologs was 11.57 × 10−9 Ks/year. The average synonymous substitution rate for Arachis genes was previously estimated at 8.12 × 10−9 Ks/year26. The present results thus indicated that the synonymous substitution rate was elevated in 6,732 one-to-one orthologs. Moreover, Ka/Ks values were negatively correlated with gene expression level (Fig. 2 and Table S5), expression breadth, GC1, GC3, and overall GC content (Table 3). Moreover, Ka and Ks values were not correlated with Fop, polypeptide length, GC content, gene expression level, and expression breadth (Fig. 2, Table 3, and Table S5).
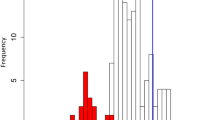
Gene expression level and expression breadth of gene sequences that experienced purifying selection were significantly higher than those in gene sequences that have been shaped by positive selection (Mann–Whitney U test, P < 0.01; Fig. 3). In the two groups, Fop was negatively correlated with polypeptide length, but positively correlated with GC content (Table 1). However, there were inconsistent correlations between gene expression level from 22 tissues and variables including Fop, polypeptide length, GC content, expression breadth, Ka, Ks, and Ka/Ks; the average gene expression level was positively correlated with Fop, GC1 content, GC3 content, overall GC content, and expression breadth, while no correlation was observed between average gene expression level and both Ka and Ks values in the two groups (Fig. 4 and Table S8). It should be noted that Ka/Ks was positively and non-significantly correlated with average gene expression level (r = 0.16, P > 0.05) in genes under positive selection, but negatively and significantly correlated with average gene expression level (r = −0.22, P < 0.01) in genes under purifying selection (Fig. 4 and Table S8). Moreover, expression breadth was positively correlated with polypeptide length in positive and negative groups (Table 2). However, expression breadth was positively correlated with Fop and GC3 content in genes under positive selection, and positively correlated with GC1 content, GC2 content, and overall GC content in genes under purifying selection (Table 2).
Comparative of gene expression level and expression breadth. (A) Gene expression level in sequences that have experienced purifying and positive selection. (B) Gene expression level in single-copy and multiple-copy genes. (C) Expression breadth in sequences that have experienced purifying and positive selection, and single-copy and multiple-copy genes. Blue (A), red (B), purple (C), and green (C) colors indicate sequences that have experienced purifying selection, sequences that have experienced positive selection, single-copy genes and multiple-copy genes, respectively. (1) seedling leaf 10 d post emergence; (2) main stem leaf; (3) lateral leaf; (4) vegetative shoot tip from the main stem; (5) reproductive shoot tip from the first lateral leaf; (6) 10 d root; (7) 25 d nodules; (8) perianth; (9) gynoecium; (10) androecium; (11) aerial gynophore tip; (12) subterranean gynophore tip; (13) Pattee 1 pod; (14) Pattee 1 stalk; (15) Pattee 3 pod; (16) Pattee 5 pericarp; (17) Pattee 5 seed; (18) Pattee 6 pericarp; (19) Pattee 6 seed; (20) Pattee 7 seed; (21) Pattee 8 seed; (22) Pattee 10 seed; (23) average gene expression level.
Correlation analyses of gene expression level and various attributes in sequences that have experienced positive and purifying selection. The analyzed sequence attributes include frequency of optimal codons (Fop), polypeptide length, GC1 content, GC2 content, GC3 content, overall GC content, expression breadth, Ka, Ks, and Ka/Ks. (U and O) Sequences that have experienced positive and purifying selection, respectively. (1) seedling leaf 10 d post emergence; (2) main stem leaf; (3) lateral leaf; (4) vegetative shoot tip from the main stem; (5) reproductive shoot tip from the first lateral leaf; (6) 10 d roots; (7) 25 d nodules; (8) perianth; (9) gynoecium; (10) androecium; (11) aerial gynophore tip; (12) subterranean gynophore tip; (13) Pattee 1 pod; (14) Pattee 1 stalk; (15) Pattee 3 pod; (16) Pattee 5 pericarp; (17) Pattee 5 seed; (18) Pattee 6 pericarp; (19) Pattee 6 seed; (20) Pattee 7 seed; (21) Pattee 8 seed; (22) Pattee 10 seed; (23) average gene expression level. The heat map was constructed based on Pearson correlation coefficients in Table S7. *Indicates significance at P < 0.05; **Indicates significance at P < 0.01.
Among the genes under positive selection, Ka/Ks values were negatively correlated with Fop, overall GC content, and expression breadth, but positively correlated with polypeptide length, GC1 content, and GC2 content (Table 3). However, Ka/Ks values were negatively correlated with GC1 content, GC3 content, overall GC content, and expression breadth among genes under purifying selection (Table 3). The Ka and Ks values were negatively correlated with polypeptide length, but positively correlated with GC1 content and expression breadth in the positive selection group, respectively. Moreover, positive correlations were exhibited between the Ks value and GC1 content, overall GC content, and expression breadth among the genes under positive selection (Table 3). However, Ka and Ks values were only negatively correlated with expression breadth in the group of genes under purifying selection (Table 3). Overall, correlations differed both between genes under positive and purifying selection in a pattern that was potentially consistent with differences in synonymous and nonsynonymous substitution rates.
Comparison of single-copy and multiple-copy gene families
To understand differences in gene expression levels and evolutionary trends between single-copy and multiple-copy gene families, we first compared gene expression levels and expression breadth between single-copy and multiple-copy gene families in A. duranensis and A. ipaënsis. We found that gene expression levels and expression breadth were significantly higher in single-copy gene families than those in multiple-copy gene families (Mann–Whitney U test, P < 0.01; Fig. 3). Further, Fop was negatively correlated with polypeptide length in multiple-copy gene families, and positively correlated with GC content in single-copy and multiple-copy gene families (Table 1).
Although the gene expression level of some tissues was not correlated with Fop and GC content, average gene expression level was positively correlated with Fop and GC content in single-copy and multiple-copy gene families (Fig. 5 and Table S9). In addition, the gene expression level among 22 tissues was positively correlated with expression breadth in single-copy and multiple-copy gene families, but negatively correlated with Ka/Ks value in multiple-copy gene families. There was no correlation between polypeptide length, Ka, Ks, and gene expression level in single-copy and multiple-copy gene families. Expression breadth was positively correlated with polypeptide length, overall GC content, and GC1 content among the single-copy and multiple-copy gene families (Table 2). However, expression breadth was positively correlated with Fop and GC2 content in multiple-copy gene families (Table 2). Moreover, Ks was negatively correlated with expression breadth, and Ka/Ks was negatively correlated with GC2 content in single-copy gene families (Table 3). However, Ka/Ks values were negatively correlated with GC1, GC3, overall GC content, and expression breadth in multiple-copy gene families (Table 3).
Correlation analyses of gene expression level and various attributes of single-copy and multiple-copy genes. The analyzed sequence attributes include frequency of optimal codons (Fop), polypeptide length, GC1 content, GC2 content, GC3 content, overall GC content, expression breadth, Ka, Ks, and Ka/Ks. (S and M) Single-copy and multiple-copy genes, respectively. (1) seedling leaf 10 d post emergence; (2) main stem leaf; (3) lateral leaf; (4) vegetative shoot tip from the main stem; (5) reproductive shoot tip from the first lateral leaf; (6) 10 d roots; (7) 25 d nodules; (8) perianth; (9) gynoecium; (10) androecium; (11) aerial gynophore tip; (12) subterranean gynophore tip; (13) Pattee 1 pod; (14) Pattee 1 stalk; (15) Pattee 3 pod; (16) Pattee 5 pericarp; (17) Pattee 5 seed; (18) Pattee 6 pericarp; (19) Pattee 6 seed; (20) Pattee 7 seed; (21) Pattee 8 seed; (22) Pattee 10 seed; (23) average gene expression level. The heat map was constructed based on Pearson correlation coefficients in Table S8. **Indicates significance at P < 0.01.
Discussion
The A. duranensis and A. ipaënsis genome sequences were released in 2015. Bertioli, et al.26 found most genes had a one-to-one correspondence between the two species using both full-length and partial sequence alignment methods. In cultivated peanut, Clevenger, et al.27 identified 8,816 full-length homologs using reciprocal BLAST. In the present study, we report that 7,435 full-length sequences are one-to-one ortholog pairs in A. duranensis and A. ipaënsis using local BLAST. The number of one-to-one orthologs determined by a previous study was larger than that identified in our study because we excluded partial sequences and genes with unknown functions. Accordingly, many one-to-one ortholog pairs were possibly excluded in this study based on our screening strategies. Although it is popular to use OrthoMCL to identify orthologs, misinterpretation of the data is possible, and the number of orthologs can therefore be controlled by setting an inflation parameter30,31. Future studies may improve upon this method to identify more one-to-one orthologs.
Previous studies have shown that orthologs that experienced positive selective pressure are involved in abiotic or biotic stress resistance17,32. However, we found most one-to-one orthologs that experienced positive selective pressures play a role in binding, photosynthesis, and other pathways, rather than resistance to stresses (Table S7). Nevertheless, four Ka/Ks values from toll/mammalian interleukin-1 receptor (TIR)–nucleotide-binding site–leucine-rich repeat (NBS–LRR) (TNL) genes exceeded 1. TNL belongs to the NBS–LRR gene family, which is associated with disease resistance33. Song, et al.34 determined that paralogous genes mainly underwent purifying selection in A. duranensis and A. ipaënsis. These results suggested that the biological function of NBS-LRR differed between A. duranensis and A. ipaënsis. Recently, Michelotto, et al.35 demonstrated that A-type wild peanut is more resistant to disease than B-type wild peanut. Similarly, Pandey, et al.36 found that A-type wild peanut had more resistance genes than B-type wild peanut based on the number of quantitative trait loci.
Previous studies have demonstrated that synonymous substitution rates in herbaceous lineages are higher than those in woody relatives7,37. One of the major factors underlying this difference is the shorter generation times of herbaceous lineages. Annual plants reach their first flowering more quickly than perennials, and thus they, on average, experience more frequent cell divisions per unit time prior to reproduction7. The average synonymous substitution rate in Arachis genes was similar to that in Medicago, though higher than that in Lotus, Glycine, and Phaseolus26. Here, the average synonymous substitution rate of 6,732 one-to-one orthologs (11.57 × 10−9 Ks/year) was higher than that among Arachis genes (8.12 × 10−9 Ks/year26), indicating one-to-one orthologs play a crucial role in sustaining biological functions. Arachis species originated in the high elevations of South America26, where UV radiation is relatively strong. Accordingly, plants under these conditions have higher synonymous substitution rates, consistent with an elevated rate of damage repair from pyrimidine dimers. Therefore, the elevated synonymous substitution rates of orthologs in Arachis may actually be an outcome of adaptive evolution. In addition, a higher substitution rate is considered a stronger and more important overall evolutionary force than positive selection in CDSs38,39.
Selection appeared to increase the efficiency and accuracy of transcription and translation, as supported by the positive correlation between codon usage bias (Fop) and gene expression level in A. duranensis and A. ipaënsis among one-to-one orthologs. This positive correlation between codon usage bias and gene expression has been reported in some plants previously, including Populus tremula40, Silene latifolia20, Picea spp.17, and Cardamine spp.41. On the other hand, codon usage bias (as demonstrated by Fop) was negatively correlated with polypeptide length, but positively correlated with GC content in this study. Similar results were derived from the genomes of four monocots, fifteen dicots, and two mosses described by Camiolo, et al.19, confirming that short and higher-GC DNA sequences exhibit relatively high levels of expression and optimal usage bias. Similarly, Ingvarsson40 showed Fop values were negatively correlated with protein lengths, but strongly and positively correlated with GC3 content in Populus tremula. In rice, Wang and Hickey42 found that codon usage bias was negatively correlated with gene length, and short genes contained high GC content compared to long genes.
In this study, highly expressed genes were subjected to stronger selective constraint than genes with low expression levels based on the negative correlation between selection pressure and both gene expression and expression breadth in A. duranensis and A. ipaënsis one-to-one orthologs. These results strongly support an expression-rate of sequence evolution anticorrelation model (E-R anticorrelation)43. This model can be explained by at least four hypotheses, including the expression cost hypothesis, the protein misfolding avoidance hypothesis, the protein misinteraction avoidance hypothesis, and the mRNA folding requirement hypothesis44. The expression cost hypothesis proposes that the optimal expression level of a gene corresponds to a trade-off between the benefits and costs associated with its expression45,46. The protein misfolding avoidance hypothesis asserts that protein misfolding is cytotoxic and thus reduces fitness47. However, Yang, et al.48 proposed that natural selection on traits other than misfolding avoidance against protein–protein misinteraction, which wastes functional molecules and is potentially toxic, constrains the evolution of surface residues. Park, et al.49 considered that selection for mRNA folding can impact the nonsynonymous-to-synonymous nucleotide substitution rate ratio, requiring a revision of the current interpretation of this ratio as a measure of protein-level selection. However, there is a positive but nonsignificant correlation between selective pressure and average gene expression in sequences experiencing positive pressure. Although the current study does not immediately suggest a reasonable explanation, the fitness of plants may increase more through the production of new biological functions than by avoiding misfolded proteins. Accordingly, subfunctionalization and neofunctionalization may lead to higher levels of gene expression that increase fitness.
The gene expression and expression breadth of single-copy gene family sequences were significantly higher than those in multiple-copy gene family sequences. The same result was observed in conifers10,17 and flowering plants8,31. Single-copy genes and their expression patterns have been shown to evolve more slowly than genes in multiple-copy genes31,50. These previous results were consistence with our result showing that gene expression levels and expression breadth in sequences that experienced purifying selection exceeded those in sequences experiencing positive selection.
Our results clarify the relationship between gene expression level and molecular evolution in A. duranensis and A. ipaënsis one-to-one orthologs using transcriptome data. The same codon usage bias was detected among all one-to-one orthologs. Additionally, gene expression and expression breadth were significantly higher in single-copy gene families than in multiple-copy gene families. Similarly, the gene expression and expression breadth in sequences that had experienced purifying selection were higher than those in sequences that had experienced positive selection. In addition, our results demonstrated that selective pressure was negatively correlated with gene expression and expression breadth in all one-to-one orthologs, while positively but not significantly correlated with gene expression in sequences that had experienced positive selection. This study provides the foundation for further research on gene expression and evolution in Arachis.
Materials and Methods
Sequence retrieval and expression data collection
The A. duranensis and A. ipaënsis CDSs were downloaded from PeanutBase (http://peanutbase.org/download)26. To avoid including partial sequences in these analyses, the following evaluation criteria were adopted: (1) CDSs were required to start with an ATG codon and end in TAA, TAG, or TGA codons and (2) CDSs were required to lack premature termination codons or ambiguous codons. Functional annotation of each gene was described by Bertioli, et al.26. Genes with unknown functions were excluded in the present study. Gene families were classified as either single-copy gene families (i.e., consisting of one gene) or multiple-copy gene families (i.e., consisting of more than one gene) based on gene number.
RNA-seq data derived from various tissues in cultivated peanut have also been released in PeanutBase27. We specifically collected RNA-seq data generated from leaf, shoot, root, nodule, perianth, gynoecium, androecium, gynophores, pod, pericarp, and seed tissues. All details of the sequencing, de novo transcriptome assembly, and expression level evaluation were described by Clevenger, et al.27. Briefly, cultivated peanuts (cultivar ‘Tifrunner’) were grown in a greenhouse (maintained at 24–30 °C). All tissues were harvested at 14:00 except for flower samples, which were collected at 8:30. Three biological replicates of each tissue were sampled from three different plants. The total RNA of each pooled tissue sample was extracted. TruSeq RNA Sample Preparation v2 kits were used for library construction, and paired-end 2 × 100 bp sequencing was conducted using an Illumina HiSeq. 2500 instrument with a total of 209 cycles of TruSeq Rapid SBS Kit v1 (Illumina, San Diego, CA, USA) chemistry. Second, an in silico amphidiploid genome was created by simply disregarding scaffolds and concatenating the A. duranensis genome assembly with the A. ipaënsis genome assembly and labeling each simply as corresponding to the “A” and “B” genomes, respectively. Once the reads were mapped, the SAM file was run through the genome-guided pipeline. Third, total reads were mapped to the transcript assembly from 58 libraries (consisting of samples from 22 distinct tissue types and developmental stages including vegetative and seed stages) using Bowtie, allowing two mismatches within any particular 25-bp seed. Fragments per kilobase per million reads mapped (FPKM) were estimated using RSEM51 for each library. When reads mapped to multiple transcripts, RSEM fractionates the read count among the transcripts so read counts are not integers. Transcripts that had less than 1 FPKM in all 58 libraries were filtered out using the Trinity package, because they were deemed to lack sufficient minimum read coverage. The FPKM values for each gene were distinguished for both the A (A. duranensis) and B (A. ipaënsis) genomes from cultivated peanut. The orthologs were identified between cultivated peanut and its diploid ancestors (A. duranensis [A genome] and A. ipaënsis [B genome] available from http://peanutbase.org/gene_expression/atlas). The FPKM value for each gene in these various tissues was normalized using a log2-transformation for both the A and B genomes. The heat maps in Figs 1, 2, 4 and 5 were generated in R using the heatmap.2 function available in the gplots CRAN library package. Expression breadth, defined as the number of tissues in which a gene is expressed (FPKM value > 0), was estimated in A. duranensis and A. ipaënsis, respectively.
Identification of orthologs
We used A. duranensis CDSs as a query for comparison with A. ipaënsis CDSs using local BLAST, and vice versa. The following evaluation criteria were used as thresholds prior to inclusion of CDSs in subsequent analyses27: (1) alignment exceeding 80% of the length of the longer sequence, (2) identity > 80%, and (3) E-value ≤ 10−10. These pairs were also excluded if one CDS matched more than one hit or was annotated to have an unknown function.
MAFFT52 was used to align ortholog pairs. PAL2NAL53 was used to convert protein sequences into their corresponding nucleotide sequences. PAML 4.0 54 was used to calculate the Ka/Ks (nonsynonymous to synonymous per site substitution rates) ratio. Ortholog pairs with Ks < 0.01 were excluded because low sequence divergence could result in unreliable estimates. In addition, as Ka approaches 0, the Ka/Ks value becomes essentially a constant. Hence, we excluded ortholog pairs with Ka/Ks < 0.001. Generally, Ka/Ks = 1, Ka/Ks > 1, and Ka/Ks < 1 indicated neutral, positive, and purifying selection, respectively. Absolute rates of substitution at synonymous sites, u, were calculated in pairwise comparisons using the formula u = d/2 T, where d is the synonymous substitution rate and T is 2.16 million years, corresponding to the estimated divergence time between A. duranensis and A. ipaënsis26.
Calculation of codon usage bias
Codon bias, measured as the frequency of optimal codons (Fop), and polypeptide length were estimated using CodonW (version 1.4, http://codonw.sourceforge.net). For a gene with extreme codon bias, Fop equals 1, while for a gene with random codon usage, Fop equals 0 55. GC content was also calculated using an in-house Perl script.
Statistical analysis
A Mann–Whitney U test was used to make comparisons of all variables between single-copy and multiple-copy genes as well as between sequences in the positive and purifying groups. Correlations were assessed among all variables estimated from genes, including Fop, GC content, polypeptide length, substitution rate, gene expression level, and expression breadth. A one-way ANOVA test was performed, and P-values of less than 0.05 were considered significant in the correlation analyses. All analyses were conducted in JMP 9.0 (SAS Institute, Inc., Cary, NC, USA). Pearson correlation coefficients indicated the strength of the correlation between two variables, with the strongest relationships having the highest correlation coefficients. However, the strengths of these correlation coefficients are typically low in molecular biology data sets because of the high number of factors influencing these large data sets. In this study, we inferred there was no correlation if the correlation coefficient was less than 0.1 based on findings from previous studies40,56.
Change history
27 February 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Fitch, W. M. Homology: a personal view on some of the problems. Trends Genet 16, 227–231 (2000).
Kuzniar, A., van Ham, R. C. H. J., Pongor, S. & Leunissen, J. A. M. The quest for orthologs: finding the corresponding gene across genomes. Trends Genet 24, 539–551 (2008).
Lynch, M., O’Hely, M., Walsh, B. & Force, A. The probability of preservation of a newly arisen gene duplicate. Genetics 159, 1789–1804 (2001).
Song, H. & Nan, Z. Origin, divergence, and phylogeny of asexual Epichloë endophyte in Elymus species from western China. PLoS ONE 10, e0127096 (2015).
Song, H. et al. Advances in research on Epichloë endophytes in Chinese native grasses. Front Microbiol 7, 1399 (2016).
Chen, J. et al. Sequencing of the needle transcriptome from Norway spruce (Picea abies Karst L.) reveals lower substitution rates, but similar selective constraints in gymnosperms in gymnosperms. BMC Genomics 13, 589 (2012).
Yue, J. X. et al. Genome-wide investigation reveals high evolutionary rates in annual model plants. BMC Plant Biol 10, 242 (2010).
Yang, L. & Gaut, B. S. Factors that contribute to variation in evolutionary rate among Arabidopsis genes. Mol Bio Evol 28, 2359–2369 (2011).
Grusz, A., Rothfels, C. J. & Schuettpelz, E. Transcriptome sequencing reveals genome-wide variation in molecular evolutionary rate among ferns. BMC Genomics 17, 692 (2016).
Hodgins, K. A., Yeaman, S., Nurkowski, K. A., Rieseberg, L. H. & Aitken, S. N. Expression divergence is correlated with sequence evolution but not positive selection in conifers. Mol Bio Evol 33, 1502–1516 (2016).
Song, H. et al. Global analysis of WRKY genes and their response to dehydration and salt stress in soybean. Front Plant Sci 7, 9 (2016).
Song, H. et al. Genome-wide identification and characterization of WRKY gene family in peanut. Front Plant Sci 7, 534 (2016).
Yao, H., Dogra Gray, A., Auger, D. L. & Birchler, J. A. Genomic dosage effects on heterosis in triploid maize. Proc Natl Acad Sci USA 10, 2665–2669 (2013).
Thoma, S. et al. Tissuse-specific expression of a gene encoding a cell wall-localized lipid transfer protein from Arabidopsis. Plant Physiol 105, 35–45 (1994).
Williford, A. & Demuth, J. P. Gene expression levels are correlated with synonymous codon usage, amino acid composition, and gene architecture in the red flour beetle, Tribolium castaneum. Mol Bio Evol 29, 3755–3766 (2012).
Arunkumar, R., Josephs, E. B., Williamson, R. J. & Wright, S. I. Pollen-specific, but not sperm-specific, genes show stronger purifying selection and higher rates of positive selection than sporophytic genes in Capsella grandiflora. Mol Bio Evol 30, 2475–2486 (2013).
De La Torre, A. R., Lin, Y. C., Van de Peer, Y. & Ingvarsson, P. K. Genome-wide analysis reveals diverged pattern of codon bias, gene expression, and rates of sequence evolution in Picea gene families. Genome Biol Evol 7, 1002–1015 (2015).
Maurer-Alcalá, X. X. & Katz, L. A. Nuclear architecture and patterns of molecular evolution are correlated in the Chilodonella uncinata. Genome Biol Evol 8, 1634–1642 (2016).
Camiolo, S., Melito, S. & Porceddu, A. New insights into the interplay between codon bias determinants in plants. DNA Res, 1–9 (2015).
Qiu, S., Bergero, R., Zeng, K. & Charlesworth, D. Patterns of codon usage bias in Silene latifolia. Mol Bio Evol 28, 771–780 (2011).
Bertioli, D. J. et al. An overview of peanut and its wild relatives. Plant Genet Resour-C 9, 134–149 (2011).
Kochert, G. et al. RFLP and cytogenetic evidence on the origin and evolution of allotetraploid domesticated peanut, Arachis hypogaea (Leguminosae). Am J Bot 83, 1282–1291 (1996).
Seijo, J. et al. Physical mapping of the 5S and 18S-25S rRNA genes by FISH as evidence that Arachis duranensis and A. ipaënsis are the wild diploid progenitors of A. hypogaea (Leguminosae). Am J Bot 91, 1294–1303 (2004).
Seijo, G. et al. Genomic relationships between the cultivated peanut (Arachis hypogaea, Leguminosae) and its close relatives revealed by double GISH. Am J Bot 94, 1963–1971 (2007).
Ramos, M. et al. Chromosomal and phylogenetic context for conglutin genes in Arachis based on genomic sequence. Mol Genet Genomics 275, 578–592 (2006).
Bertioli, D. J. et al. The genome sequences of Arachis duranensis and Arachis ipaensis, the diploid ancestors of cultivated peanut. Nat Genet 48, 438–446 (2016).
Clevenger, J., Chu, Y., Scheffler, B. & Ozias-Akins, P. A developmental transcriptome map for allotetraploid Arachis hypogaea. Front Plant Sci 7, 1446 (2016).
Chen, X. et al. Draft genome of the peanut A-genome progenitor (Arachis duranensis) provides insights into geocarpy, oil biosynthesis, and allergens. Proc Natl Acad Sci USA 113, 6785–6790 (2016).
Li, N., Li, Y., Zheng, C., Huang, J. & Zhang, S. Genome-wide comparative analysis of the codon usage patterns in plants. Genes Genom 38, 723–731 (2016).
Trachana, K. et al. Orthology prediction methods: A quality assessment using curated protein families. Bioessays 33, 769–780 (2011).
De Smet, R. et al. Convergent gene loss following gene and genome duplications creates single-copy families in flowering plants. Proc Natl Acad Sci USA 110, 2898–2903 (2013).
Buschiazzo, E., Ritland, C., Bohlmann, J. & Ritland, K. Slow but not low: genomic comparisons reveal slower evolutionary reta and higher dN/dS in conifers compared to angiosperms. BMC Evol Biol 12, 8 (2012).
Meyers, B. C., Kozik, A., Griego, A., Kuang, H. H. & Michelmore, R. W. Genome-wide analysis of NBS-LRR-encoding genes in Arabidopsis. Plant Cell 15, 809–834 (2003).
Song, H. et al. Comparative analysis of NBS-LRR genes and their response to Aspergillus flavus In Arachis. PLoS ONE 12, e0171181 (2017).
Michelotto, M. D. et al. Identification of fungus resistant wild accessions and interspecific hybrids of the genus Arachis. PLoS ONE 10, e0128811 (2015).
Pandey, M. K. et al. Genetic dissection of novel QTLs for resistance to leaf spots and tomato spotted wilt virus in peanut (Arachis hypogaea L.). Front Plant Sci 8, 25 (2017).
Yang, Y. et al. Dissecting molecular evolution in the highly diverse plant clade caryophyllales using transcriptome sequencing. Mol Bio Evol 32, 2001–2014 (2015).
Gossmann, T. I. et al. Genome-wide analyses reveal little evidence for adaptive evolution in many plant species. Mol Bio Evol 27, 1822–1832 (2010).
Slotte, T. et al. Genomic determinants of protein evulotion and polymorphism in Arabidopsis. Genome Biol Evol 3, 1210–1219 (2011).
Ingvarsson, P. K. Gene expression and protein length influence codon usage and rates of sequence evolution in Populus tremula. Mol Bio Evol 24, 836–844 (2007).
Ometto, L., Li, M., Bresadola, L. & Varotto, C. Rates of evolution in stress-related genes are associated with habitat preference in two Cardamine lineages. BMC Evol Biol 12, 7 (2012).
Wang, H. C. & Hickey, D. A. Rapid divergence of codon usage patterns within the rice genome. BMC Evol Biol 7, S6 (2007).
Drummond, D. A., Bloom, J. D., Adami, C., Wilke, C. O. & Arnold, F. H. Why highly expressed proteins evolve slowly. Proc Natl Acad Sci USA 102, 14338–14343 (2005).
Zhang, J. & Yang, J. Determinants of the rate of protein sequence evolution. Nat Rev Genet 16, 409–420 (2015).
Cherry, J. L. Expression level, evolutionary rate, and the cost of expression. Genome Biol Evol 2, 757–769 (2010).
Gout, J. F., Kahn, D. & Duret, L. The relationship among gene expression, the evolution of gene dosage, and the rate of protein evolution. PLoS Genet 6, e1000944 (2010).
Geiler-Samerotte, K. A. et al. Misfolded proteins impose a dosage-dependent fitness cost and trigger a cytosolic unfolded protein response in yeast. Proc Natl Acad Sci USA 108, 680–685 (2011).
Yang, J. R., Liao, B. Y., Zhuang, S. M. & Zhang, J. Protein misinteraction avoidance causes highly expressed proteins to evolve slowly. Proc Natl Acad Sci USA 109, E831–E840 (2012).
Park, C., Chen, X., Yang, J. R. & Zhang, J. Differential requirements for mRNA folding partially explain why highly expressed proteins evolve slowly. Proc Natl Acad Sci USA 110, E678–E686 (2013).
Han, M. V., Demuth, J. P., McGrath, C. L., Casola, C. & M.W., H. Adaptive evolution of young gene duplication in mannals. Genome Res 19, 859–867 (2009).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Bio Evol 30, 772–780 (2013).
Suyama, M., Torrents, D. & Bork, P. PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34, 609–612 (2006).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol Bio Evol 24, 1586–1591 (2007).
Sharp, P. M. & Li, W. H. The codon adaption index-a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res 15, 1281–1295 (1987).
Whittle, C. A. & Extavour, C. G. Codon and amino acid usage are shaped by selection across divergent model organisms of the Pancrustacea. G3-Gene Genom Genet 5, 2307–2321 (2015).
Acknowledgements
This study was supported by the National Basic Research Program of China (2014CB138702) and the National Natural Science Foundation of China (31502001).
Author information
Authors and Affiliations
Contributions
H.S. and Z.N. conceived and designed this research. H.S. analyzed data and wrote the manuscript. H.G. and J.L. executed the statistical analyses. P.T. participated in the discussion of the results. Z.N. contributed to the evaluation and discussion of the results and manuscript revision.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Song, H., Gao, H., Liu, J. et al. Comprehensive analysis of correlations among codon usage bias, gene expression, and substitution rate in Arachis duranensis and Arachis ipaënsis orthologs. Sci Rep 7, 14853 (2017). https://doi.org/10.1038/s41598-017-13981-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-13981-1
This article is cited by
-
Genome-wide identification, evolutionary and expression analysis of the cyclin-dependent kinase gene family in peanut
BMC Plant Biology (2023)
-
Deciphering evolutionary dynamics of WRKY genes in Arachis species
BMC Genomics (2023)
-
Genome-wide characterization of Ficus carica MADS-box transcription factors with a focus on their roles during fruit development
Horticulture, Environment, and Biotechnology (2023)
-
Genome-Level Investigation of WRKY Transcription Factors and Their Potential Roles in Fruit Peel Ripening and Coloration in the Common Fig (Ficus carica L.)
Journal of Plant Growth Regulation (2023)
-
Chloroplast gene expression level is negatively correlated with evolutionary rates and selective pressure while positively with codon usage bias in Ophioglossum vulgatum L
BMC Plant Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.