Abstract
Ability to distinguish between closely related Wolbachia strains is crucial for understanding the evolution of Wolbachia-host interactions and the diversity of Wolbachia-induced phenotypes. A useful model to tackle these issues is the Drosophila suzukii – Wolbachia association. D. suzukii, a destructive insect pest, harbor a non-CI inducing Wolbachia ‘wSuz’ closely related to the strong CI-inducing wRi strain. Multi locus sequence typing (MLST) suggests presence of genetic homogeneity across wSuz strains infecting European and American D. suzukii populations, although different Wolbachia infection frequencies and host fecundity levels have been observed in both populations. Currently, it is not clear if these differences are due to cryptic wSuz polymorphism, host background, geographical factors or a combination of all of them. Here, we have identified geographical diversity in wSuz in D. suzukii populations from different continents using a highly diagnostic set of markers based on insertion sequence (IS) site polymorphism and genomic rearrangements (GR). We further identified inter-strain diversity between Wolbachia infecting D. suzukii and its sister species D. subpulchrella (wSpc). Based on our results, we speculate that discernible wSuz variants may associate with different observed host phenotypes, a hypothesis that demands future investigation. More generally, our results demonstrate the utility of IS and GRs in discriminating closely related Wolbachia strains.
Similar content being viewed by others
Introduction
Wolbachia are obligate-intracellular bacteria infecting more than half of the arthropod species1. Although they are typically maternally inherited by cladogenic transmission or introgression events, horizontal transmission can also occur between closely or distantly related species2. Wolbachia can spread and maintain themselves in the host by manipulating host reproductive biology3. The most studied manipulating strategy is cytoplasmic incompatibility (CI) that favors infected females to enhance rapid bacterial spread throughout the population4. In the absence of or in combination with CI, Wolbachia may beneficially affect their hosts’ fitness, for example by providing essential nutrients5, increasing stem cell proliferation6 and protecting against pathogenic RNA viruses7,8,9,10. Various studies indicate the presence of multiple Wolbachia strains in the same host or of different strains in several populations of the same host, inducing various phenotypes11,12,13,14. Such a large variety of phenotypes caused by Wolbachia within the same or different hosts indicate a complex mechanism behind distinct host-Wolbachia interactions. The correct typing of Wolbachia strain diversity is, therefore, a prerequisite to correctly understand their biology in a given host.
Various molecular tools based on multi-locus sequence typing (MLST) genes together with the hyper-variable Wolbachia surface protein (wsp) gene15,16,17,18 have been successfully used for Wolbachia strain typing. Wolbachia has been classified in distinct types or strains that can be grouped into at least 16 supergroups (named A–F and H–Q)19. It is, however, challenging to distinguish among very closely related bacterial strains using single gene phylogenetic or the MLST system alone due to their limited resolution15,20,21,22,23. For example, the MLST system was insufficient to discriminate closely related Wolbachia strains infecting natural populations of D. melanogaster 13,15,18,24. Moreover, MLST failed to differentiate between wRi, wSuz and wSpc Wolbachia strains harbored by their natural hosts D. simulans, D. suzukii and D. subpulchrella (sister species of D. suzukii), respectively25,26,27. However, comparison of wRi (complete genome) and wSuz (draft genome) revealed several differences such as Insertion sequence (IS) presence/absence polymorphism and genomic rearrangements (GRs)25. Whole genome sequencing (WGS), indeed, maximizes the chances of finding informative characters that are less likely to occur in the few genes sampled by MLST and provides enough information to effectively discriminate between indistinguishable strains28. For example, a population genomics study allowed the identification of previously uncharacterized wMel diversity within several D. melanogaster wild populations29. However, WGS can be time consuming and expensive for large-scale population genetic studies.
Using a different approach, Riegler and colleagues13,30 applied a set of hyper-variable markers based on site polymorphism of IS elements, variable number tandem repeat (VNTR) loci, and chromosomal inversions to discriminate closely related A-supergroup Wolbachia strains. IS elements are bacterial class-II transposons of discrete DNA segments that can replicate and spread in the genome through a cut-and-paste mechanism as reviewed in31. The majority of IS elements are bound by short terminal inverted repeat (TIRs) sequences of variable lengths that are repeated in opposite orientations at the 5’ and 3’ ends of these elements. ISs are classified into about 20 families on the basis of several conserved features within families, such as structure, insertion site preference, sequence organization, and similar TIRs31,32. Together with TIRs, these elements can also undergo ectopic (non-allelic homologous) recombination events resulting in GRs. The genomes of Wolbachia, in particular, display a very high number of IS elements representing about 10% of the bacterial genome33. These elements can exhibit a large amount of variability in their genomic content and have thus been proven very useful for discriminating very closely related bacterial strains13,33,34,35,36,37.
According to MLST, different populations of D. suzukii harbor the same wSuz strain, which in turn is indistinguishable from the new strain (wSpc) harbored by D. subpulchrella 26,27. Contrary to their closely related wRi strain that causes strong CI in D. simulans, wSuz and wSpc have been characterized by either very low or a complete lack of CI-inducing capability26,27. We have previously detected differences in wSuz prevalence (and to a lesser extent its CI inducibility) in different D. suzukii populations. European (EU) wSuz infection frequencies are three times significantly higher compared to American (US) ones27. Both populations have been reported inducing no considerable CI26,27, but EU (French) D. suzukii reportedly showed a lower, although statistically insignificant, hatch rate in the CI cross27. If D. suzukii actually harbors a single strain of ‘wSuz’, we should assume that observed differences in their natural infection prevalence and CI levels are either dependent on the host genetic background or caused by other environmental factors such as temperature or exposure to insecticides27. Alternatively, there may exist slightly different cryptic variants of wSuz in nature affecting variable levels of their persistence ability in various D. suzukii populations, but have not yet been distinguished based on standard MLST typing method. Unsuccessful determination of hidden wSuz diversity may, therefore, under-estimate the actual biological complexities behind wSuz-D. suzukii interactions.
Our previous comparison of wRi and wSuz genomes have provided a putative diagnostic set of markers based on IS site polymorphism and genomic rearrangements25. In this study, we validated these diagnostic markers using PCR and Sanger sequencing and revealed an a) intra-strain diversity within wSuz from different D. suzukii populations worldwide and b) inter-strain Wolbachia diversity between previously (MLST-based) indistinguishable wSuz and wSpc strains. These findings will aid in our understanding of Wolbachia diversity and infection dynamics within and between D. suzukii populations and related species. We also discuss the potential implications of wSuz geographical diversity in symbiont-based pest management programs.
Results
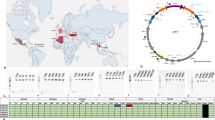
We selected 32 polymorphic insertion sequence (IS) loci and two large-scale genomic rearrangements (GRs) based on the comparison of wRi and wSuz genomes25 (Fig. 1). Of the 32 IS-associated loci, eight belonged to ISWpi1 group from the IS5 family, 23 to ISWpi5 group from the IS66 family, and one belonged to ISWpi7 of the IS110 transposon family (listed in Supplementary Table S1). We designed 34 sets of primers and verified these diagnostic markers by PCR amplification (and Sanger-sequencing, when necessary) using genomic DNA extracted from D. simulans, D. subpulchrella and two individuals each from thirteen D. suzukii populations (Table 1). The cumulative results of IS presence-absence polymorphism and the GR based diagnostic PCRs from different Wolbachia strains are shown in Table 2.
Genome comparison of wSuz and wRi for candidate marker loci selection. wSuz contigs have been oriented according to wRi genome. Annotated CDSs in either plus or minus strand are represented with brown and cream colored boxes respectively. The grey twisted ribbons represent the two genomic rearrangements detected in wSuz relative to wRi. The orientation of the primers used for validating GR1 and GR2 in both genomes are represented with black and red arrowheads respectively. The inner circle in wRi genome represents annotated IS elements, color-coded based on their group affiliation (red: ISWpi1, green:ISWpi2, orange:ISWpi4, blue: ISWpi5 and purple: ISWpi7). Colored lines linking wRi to wSuz genome represent the 32 polymorphic IS loci used in the present study. The graph was designed with Circos software81.
IS insertion site polymorphism and genomic rearrangements differentiate wSuz, wSpc and wRi Wolbachia strains
Out of the 32 polymorphic IS loci, 27 were specific of wRi (IS2-IS13, IS15-IS21, IS23-IS30), two were specific of wSuz (IS31 and IS32), and three were shared between wSpc and wRi (IS1, IS14 and IS22). The latter were demonstrated by the similar amplicon sizes in wSpc and wRi (2,576bps, 2,000bps and 1,820bps respectively) compared to wSuz (1,600bps, 512bps and 330bps respectively) (Fig. 2a–c). Sequence analysis, however, revealed the presence of IS target-site variations at all these three loci (Fig. 2a–c). At IS1 locus, an ISWpi1 (wRi_003610) element was shared, but reversely orientated in wSpc and wRi (Fig. 2a). At IS14 and IS22 loci, two ISWpi5 elements (wRi_p03000 and wRi_002290, respectively) were shared among wSpc and wRi, but the exact insertion sites differed between the two strains: at IS14, the ISWpi5 element in wSpc was inserted 84bp upstream relative to wRi, whilst for the IS22 locus, the insertion in wSpc was 8bp downstream to that of wRi (Fig. 2b & c).
Inter-strain polymorphism between closely related wSuz, wSpc and wRi Wolbachia strains. Arrows with different shades of green represent different ORFs while the blue arrows represent transposase genes. Red arrowheads correspond to terminal inverted repeats (TIRs). A–C. wRi-specific IS loci (a) IS element present at locus IS1 belonging to ISWpi1 group shows inversion between wRi and wSpc, and absence from wSuz strain, (b) and (c) Two IS elements (loci ID- IS14 and IS22) show independent insertion events between wRi and wSpc however completely absent from wSuz genome. (d) wSuz-specific IS element (locus ID- IS31) belonging to ISWpi1 group shows insertion only in wSuz from all populations producing 1292 bps long amplicon, but absent from wRi and wSpc with 398 bps amplicon size. (e) Genomic rearrangement (GR1) showing amplification in wSuz only, absent in wRi and wSpc. Lanes from left: wRi, wSpc, wSuz_ITA, wSuz_FRA, wSuz_JPN, wSuz_CHN, wSuz_USA, wSuz_CAN and STC-Wolbachia negative control. The full-length gel pictures are presented in Supplementary Figure S2.
Two large-scale genome rearrangements (GR1 and GR2) further discriminated wSuz, wSpc and wRi (Table 2). Primers flanking both GR regions in wSuz (Fig. 1, Supplementary Table S1) were used to confirm the rearrangements using PCR: GR1 was confirmed as a genomic inversion in all wSuz variants compared to wSpc and wRi (Fig. 2e); GR2 was inverted in both wSuz and wSpc, but not in wRi (Fig. 3b).
Intra-strain polymorphism of wSuz within different D. suzukii populations. (a) wSuz specific IS element (Locus ID- IS32) showing 1,794 bps amplicon size polymorphism in European wSuz (wSuz_ITA and wSuz_FRA) strains in comparison to other wSuz strains showing amplification of 304 bp size, similar to wRi and wSpc. (b) Genomic rearrangement (GR2) showing size polymorphism in American (USA) and Canadian (CAN) D. suzukii only. Upper panel - schematic diagram of Inverted translocation (IT) shown in wRi and wSuz genome. The full-length gel pictures are presented in Supplementary Figure S2.
Polymorphism in wSuz strains from different D. suzukii host populations
We detected intra-strain polymorphism within wSuz strains from different D. suzukii populations listed in Table 1. Hereafter, several D. suzukii populations from different countries, but of the same continent, have been referred by their continental names. A wSuz-specific IS element at locus IS32 was exclusively found in European samples (wSuz_ITA, wSuz_FRA), and not in American (wSuz_USA, wSuz_CAN) and Asian (wSuz_CHN, wSuz_JPN) populations (Table 2, Fig. 3a). Sequence analysis further confirmed that IS32 belongs to the ISWpi5 group and is inserted six nucleotides upstream to the stop codon of a gene homologous to wRi_002820. The wRi_002820 homologues in wRi, wSpc and non-European wSuz strain variants remained intact and coded for a hypothetical protein33, however, we detected low similarities to the SMC (Structural Maintenance of Chromosomes) protein family. The ISWpi5 insertion in European wSuz variant at the same locus resulted in 9 extra amino acids addition at the C-terminus of the protein due to in-frame position of left TIR of the IS element (Supplementary Fig. S1).
Sequence comparison of GR2 showed that this inverted region, spanning more than 40Kb in size, is flanked by two nearly identical ISWpi7 elements (wRi_p07230, wRi_007660) and results in the truncation of an ankyrin (ANK) gene represented by two pseudogenes wRi_p07220 and wRi_p07650 flanking the inversion in wRi genome (Fig. 3b). In contrast, the ANK gene was intact in wSpc and all of the wSuz variants, except for those infecting American D. suzukii (wSuz_USA and wSuz_CAN), where a similar ISWpi7 element truncated the ANK gene causing no inversion as confirmed by PCR and Sanger sequencing (Fig. 3b). Overall, IS- and GR- based diagnostic markers revealed the existence of at least three different wSuz genotypes infecting D. suzukii populations from American, Asian and European continents.
Phylogenetic analyses recapitulate genomic differences
On the basis of our IS and GR strain typing patterns, we constructed a character-state matrix (Supplementary Table S2) and performed phylogenetic analysis. Maximum parsimony and Bayesian analysis resulted in identical tree topologies (Fig. 4). wSuz strains were found clearly monophyletic: European and American wSuz genotypes originated independently from a more ancestral Asian infection although with weak support values due to relatively few synapomorphic characters available to compute phylogeny.
Phylogeny from all polymorphic loci. Cladogram of wRi, wSpc and wSuz Wolbachia strains inferred from the 34 character- state matrix. Support values for each node are placed, on the left is the Bayesian posterior probability and right is the percentage bootstrap support from TNT based parsimony analysis. Black stars on the map represent each sampled population of D. suzukii used in this study. The map was modified from d-maps.com (http://d-maps.com/m/world/centreeurope/centreeurope22.gif).
Discussion
Identification and discrimination of “cryptic” (not yet discovered and very closely related) Wolbachia genotypes is essential to understand the biology and the evolution of host-Wolbachia associations. Previous screenings based on MLST failed to discriminate between wSuz (harbored by D. suzukii), wSpc (D. subpulchrella) and wRi (D. simulans) Wolbachia strains, suggesting the presence of a monomorphic Wolbachia infecting different host species26,27. The same studies suggested the absence of genetic polymorphism in Wolbachia infecting different D. suzukii populations. Indeed, whole genome comparison of wRi and wSuz strains revealed extensive sequence similarity between the two Wolbachia strains25,38,39 indicating that wRi and wSuz are very closely related and diverged very recently. Moreover, the newly released draft genome of wSpc strain indicated a closer relationship between wSuz and wSpc40 (pre-print, https://doi.org/10.1101/135475). Despite the high level of similarity, wRi and wSuz differed substantially in terms of their insertion sequence (IS) site polymorphism and genomic rearrangements (GRs). In this study, we have shown the utility of these polymorphic markers to distinguish wSpc from wRi and wSuz, as well as to identify intra-strain wSuz diversity among different continental populations of D. suzukii (from America, Asia and Europe).
We first detected target site variations as well as sequence inversion of IS elements at the three loci (IS1, IS14 and IS22) shared between wRi and wSpc. IS element inversions have previously been reported in Wolbachia and attributed to the effect of ectopic recombination between the TIRs of IS elements41. In case of the IS1 locus, ectopic recombination has presumably resulted in the complete inversion of the insertion element including the asymmetric TIRs in wRi and wSpc. Furthermore, target site polymorphism was detected in case of IS14 and IS22 loci in wSpc compared to wRi. Both cases involved the insertion of an ISWpi5 element, a member of the IS66 family. Shared insertions of the same IS element at slightly different sites suggests possible independent insertion events in the two strains; however, it is not clear whether IS elements of the IS66 family exhibit sequence-specific or region-specific target preference42,43. An alternate parsimonious scenario would be that the observed target site polymorphism is the result of IS excision and local re-integration in either wRi or wSpc genomes after their divergence from a common ancestral genotype. Our results have practical implications for improving IS polymorphism-based Wolbachia strain typing methodologies. Many of the previous studies focus on simple PCR amplicon size polymorphism detection (presence/absence patterns) by gel electrophoresis13,30,35. We, however, advocate that for obtaining higher resolution strain typing, sequencing of the IS element as well as the respective insertion site is also important to uncover orientation- or target site-based variations, which otherwise can be neglected due to the similar PCR amplicon size obtained.
We further detected intra-strain Wolbachia polymorphism in wSuz strain from different geographical populations of D. suzukii host. Historically originating from Asia, D. suzukii has recently invaded Europe and America44,45. Population studies suggested that the two continents were invaded independently from two distinct Asian regions46,47. The presence of geographical diversity in wSuz Wolbachia strains (Fig. 4) is in agreement with this scenario, suggesting that founding D. suzukii individuals carried different wSuz variants in each of the two continents. We cannot exclude, however, the effects of environmental constraints that may have triggered rapid genomic changes in Wolbachia either due to adaptation and/or relaxed selection in a new environment. For example, a rapid adaptive evolution of wMel-Pop strain of D. melanogaster has been previously reported after its artificial transfer to Aedes aegypti mosquito cell lines48. Another study showed altered behavior of Wolbachia when passaged for several generations through heterozygous mutant lines of D. melanogaster 49. Concurrent with this, it has been suggested that the IS mobility is able to promote the evolutionary adaptation of their hosts50,51. However, a different study pointed out that cryptic and low-titer Wolbachia infections within or between host populations can shift in prevalence during strong bottleneck events, for example during artificial host transfers52. Under these scenarios, an alternative explanation for the geographical diversity of wSuz could be that the different wSuz genotypes may initially coexisted in the native Asian populations of D. suzukii as low-titer or rare variants within or between populations and during its colonization of America and Europe, D. suzukii might have experienced a mixture of bottlenecks46,47 and differential selective pressures in the two continents to evolve into new genotypes. More D. suzukii samples from other Asian populations and at more time points will be needed in order to test this hypothesis.
The presence of wSuz variants among different D. suzukii populations raises another interesting question as to what extent this genetic diversity could be associated with phenotypic variations in the host. Earlier studies on Wolbachia from European and American D. suzukii populations revealed that wSuz does not induce significant levels of CI and is imperfectly maternally transmitted from the mother to the progeny26,27. To maintain its infection status in the wild, CI is often considered as the driving force for Wolbachia-mediated sweeps in insect host populations53; the persistence of wSuz despite inducing no apparent CI under laboratory conditions in both continental populations points towards some positive fitness effects. Indeed, experimental data show wSuz-mediated high fecundity54 and strong protection against RNA viruses55 in D. suzukii. However, these fitness advantages are not conserved among different populations of D. suzukii: first, wSuz infection is higher prevalent and provides with more fecundity in European populations than in American ones26,27,54; second, we detected higher wSuz anti-viral protection ability in the American population than in the European one (Kaur R., Martinez J., Jiggins F., Rota-Stabelli O., Miller W.J., Tissue-specificity of Wolbachia in Drosophila vary in their interactions towards Drosophila C Virus and Flock House Virus, manuscript in preparation). Because the symbiont strain rather than the host genetic background has been demonstrated to determine the degree of Wolbachia-mediated antiviral protection effect56, we speculate that the observed differences in the antiviral protection (and perhaps fecundity and infection frequency) may be, at least partially, attributed to the different wSuz genotypes we have detected. It is important to stress that Wolbachia-induced phenotypes depend not only on the Wolbachia genetic background but also on the genetic background of the host57 and, more importantly, on the host-Wolbachia associations58,59,60. Indeed, population studies indicated a certain degree of genetic diversity between European and American D. suzukii 46,47,61. It is therefore highly plausible that a rather complex series of interactions took place in the European and American D. suzukii-wSuz systems, leading to observed differences.
One of the most interesting genomic events we have found is at the IS32 locus where the insertion element is exclusively present in European wSuz variant (Fig. 3a), making it a highly diagnostic marker for characterizing wSuz intra-strain diversity. This insertion terminally disrupts the ORF of a Wolbachia gene named wRi_002820, likely encoding a protein involved in tRNA synthesis, DNA repair and chromosomal segregation in wAu Wolbachia strain62. Another interesting event is the large-scale genomic rearrangement - GR2, flanked by two nearly identical inverted repeat elements in wRi genome. Similar genomic events associated with flanking inverted or direct repeats have previously been detected in other Wolbachia strains, e.g. wMelPop, giving rise to large-scale inversions48,63 or extensively amplifying Octomom locus64 respectively, and differentiating it from closely related wMel strain. GR2 is, therefore, another diagnostic marker for screening wSuz genotypes since the 5’-flanking inverted IS element resulting in GR2 is found in American wSuz only. This IS element, similar to wRi, results in truncation of an Ankyrin (ANK) repeat domain protein, but without causing an inversion (Fig. 3b), suggesting that this chromosomal inversion event is specific to wRi only. Furthermore, it is known that such insertion/truncation events may cause gene inactivation or alter gene regulation and expression50,65 resulting in potential phenotypic changes. Proteins with eukaryotic domains such as ANK repeats are considered primary candidates for mediating host-Wolbachia interactions; variability in ANK repeat structure and number could affect the affinity, specificity, localization, expression and function of these ANK proteins66,67. Thus, we prudently hypothesize that the structural variability of these proteins in wSuz variants might be associated with different observed inter-continental phenotypes and host-Wolbachia associations in D. suzukii. Life trait experiments involving American-European D. suzukii cross infections should be performed to verify our working hypothesis.
We finally discuss the potential implications of genetic diversity found in D. suzukii (and D. subpulchrella) for Wolbachia-based pest management programs. Wolbachia is a promising tool for developing control strategies of arthropod pest populations based on the CI phenotype68,69. Previous studies have shown no CI inducing capability in Italian, French, East and West US coast D. suzukii populations26,27. In addition wSpc, similar to wSuz, does not induce CI in its native host D. subpulchrella 26. However, the aforementioned closely related wSuz, wSpc and wRi strains could have quite different effects on the host biology, if transfected or introgressed in a different host system. Various experiments have been carried out successfully to test this cross-compatibility hypothesis, with artificial transinfection of CI-inducing Wolbachia among several Drosophila species both intra-14 and inter-specifically70,71. Future experiments involving artificial transinfection or introgression of D. suzukii with closely related Wolbachia strains such as wSpc or wRi can be performed in order to assess their modification and rescue capabilities to aid the development of bi-directional CI-based pest control programs72,73. Moreover, a correlation between IS-distinctive wPip Wolbachia genetic variants and CI crossing types has been shown in Culex pipiens mosquito populations35,74,75. We propose that different geographical D. suzukii populations harboring wSuz variants should be inter-crossed to better explore the host-Wolbachia genetic background effects on CI-induction.
Methods
Fly strains and rearing
Details of different Drosophila-Wolbachia associations assayed in this study as well as their sources and origin are listed in Table 1. All live flies were maintained on standard fly food in vials at a constant temperature of 22°C with a 12:12 light:dark cycle.
Candidate marker loci selection
We previously detected several structural variations such as insertion sequence (IS) site polymorphism and genomic rearrangements (GRs) separating wSuz from the close-related wRi strain25. A total of 34 candidate markers including 32 IS site polymorphic loci together with two large-scale GRs were chosen to study previously uncharacterized inter- and intra-strain Wolbachia polymorphism (Fig. 1). Primers were designed on their respective 5′ and 3′ flanking regions using Primer 376 as implemented in Geneious software version 7.0.6 (Biomatters, New Zealand). Primer sequences are listed in Table S1. Conserved protein domains of diagnostic IS target genes were identified using the NCBI’s conserved domain database in conjunction with BlastP and also independently verified using EMBL-EBI’s InterProScan77 and Pfam78. BlastP analysis was conducted using the NCBI BlastP program.
PCR amplification and sequencing
Host genomic DNAs were extracted using DNeasy tissue kit (Qiagen) according to the manufacturer’s instructions. Diagnostic PCR assays were performed in 20 μl reaction mixtures containing 1x GoTaq reaction buffer, 3.0 mM MgCl2, 0.5 μM of forward and reverse primer, 35 μM dNTPs, 1U of Taq Polymerase (Promega) and 30–50ng of DNA template. PCR amplification was performed on a BioRad Thermal Cycler using the following thermal profiles: 1 cycle (94°C for 3 min), 35 cycles (94°C for 30 sec, 60°C for 30 sec, 72°C for 1 min) and 1 cycle (72°C for 8 mins). Amplicons were examined using gel-electrophoresis on 1% Agarose gel stained with ethidium bromide. Gel images were visualized using an ultraviolet gel documentation system (iNTAS, Goettingen, Germany). Images were cropped to remove extraneous gel area. The Qiagen® Nucleotide Removal Kit was used to purify the reaction products, followed by Sanger sequencing analysis. All sequences have been deposited in Genbank under accession numbers MF034744 – MF034749.
Phylogenetic analysis
We conducted Parsimony and Bayesian analyses on a character state matrix in which each genomic locus listed in Table S1 was considered as an independent character. The presence/absence pattern of the characters was deduced directly from the amplified PCR bands of two individuals from each population. Presence of insertion sequence was designated with 1, and absence with 0. Whenever an IS element at a defined insertion locus was of a different size than expected, it was designated with a number higher than 1. Parsimony analysis was performed in TNT (Tree analysis using New Technology) program v1.579 by implementing traditional TBR (tree bisection reconnection) heuristic search algorithm, using 1000 replicates, saving 10 trees per replicate and replacing existing trees. To assess confidence in the resulting phylogenetic estimate, the data were subjected to a bootstrap using symmetric resampling79 and a search with 33% change probability (100 replicates), and jackknife analysis using a traditional search with a 36% removal probability replicated 5,000 times. Bayesian phylogenetic analysis was performed with MrBayes v3.2.580 using the Mk model of Lewis (2001) with the assumption that only characters that varied among taxa were included (i.e. coding = variable). Two simultaneous iterations of the Bayesian analysis were run using four simultaneous Monte Carlo Markov Chains (MCMC) for 1,000,000 generations. Trees were sampled every 100 generations. Posterior probabilities representing a measure of clade credibility were generated from the majority-rule tree composed from trees sampled from both runs, after excluding the first 25% of trees as burn-in.
Data Availability
All data generated or analyzed during this study are included in this article (and related Supplementary information files).
References
Weinert, L. A., Araujo-Jnr, E. V., Ahmed, M. Z. & Welch, J. J. The incidence of bacterial endosymbionts in terrestrial arthropods. Proc. R. Soc. B Biol. Sci 282, 20150249–20150249 (2015).
Raychoudhury, R., Baldo, L., Oliveira, D. C. S. G. & Werren, J. H. Modes of acquisition of Wolbachia: Horizontal transfer, hybrid introgression, and codivergence in the Nasonia species complex. Evolution (N. Y). 63, 165–183 (2009).
Werren, J. H., Baldo, L. & Clark, M. E. Wolbachia: master manipulators of invertebrate biology. Nat. Rev. Microbiol. 6, 741–51 (2008).
Stouthamer, R., Breeuwer, J. A. J. & Hurst, G. D. D. Wolbachia Pipientis : Microbial manipulator of arthropod reproduction. Annu. Rev. Microbiol. 53, 71–102 (1999).
Hosokawa, T., Koga, R., Kikuchi, Y., Meng, X.-Y. & Fukatsu, T. Wolbachia as a bacteriocyte-associated nutritional mutualist. Proc. Natl. Acad. Sci. 107, 769–774 (2010).
Fast, E. M. et al. Wolbachia enhance Drosophila stem cell proliferation and target the germline stem cell niche. Science 334, 990–992 (2011).
Teixeira, L., Ferreira, Á. & Ashburner, M. The bacterial symbiont Wolbachia induces resistance to RNA viral infections in Drosophila melanogaster. PLoS Biol. 6, 2753–2763 (2008).
Hedges, L. M., Brownlie, J. C., O’Neill, S. L. & Johnson, K. N. Wolbachia and virus protection in insects. Science 322, 702–702 (2008).
Aliota, M. T., Peinado, S. A., Velez, I. D. & Osorio, J. E. The wMel strain of Wolbachia reduces transmission of Zika virus by Aedes aegypti. Nat. Publ. Gr. 1–7 doi:https://doi.org/10.1038/srep28792 (2016).
Mains, J. W., Brelsfoard, C. L., Rose, R. I. & Dobson, S. L. Female adult Aedes albopictus suppression by Wolbachia-infected male mosquitoes. Sci. Rep. 6, 33846 (2016).
Bourtzis, K., Nirgianaki, A., Markakis, G. & Savakis, C. Wolbachia infection and cytoplasmic incompatibility in Drosophila species. Genetics 144, 1063–1073 (1996).
Merçot, H. & Charlat, S. Wolbachia infections in Drosophila melanogaster and D. simulans: Polymorphism and levels of cytoplasmic incompatibility. in. Genetica 120, 51–59 (2004).
Riegler, M., Sidhu, M., Miller, W. J. & O’Neill, S. L. Evidence for a global Wolbachia replacement in Drosophila melanogaster. Curr. Biol. 15, 1428–1433 (2005).
Rousset, F. & Destordeur, E. Properties of Drosophila simulans strains experimentally infected by different clones of the bacterium Wolbachia. Heredity (Edinb). 72, 325–331 (1994).
Baldo, L. et al. Multilocus sequence typing system for the endosymbiont Wolbachia pipientis. Appl. Environ. Microbiol. 72, 7098–7110 (2006).
Baldo, L. & Werren, J. H. Revisiting Wolbachia supergroup typing based on WSP: Spurious lineages and discordance with MLST. Curr. Microbiol. 55, 81–87 (2007).
Zhou, W., Rousset, F. & O’Neill, S. Phylogeny and PCR-based classification of Wolbachia strains using wsp gene sequences. Proc. R. Soc. B Biol. Sci. 265, 509–515 (1998).
Paraskevopoulos, C., Bordenstein, S. R., Wernegreen, J. J., Werren, J. H. & Bourtzis, K. Toward a Wolbachia multilocus sequence typing system: Discrimination of Wolbachia strains present in Drosophila species. Curr. Microbiol. 53, 388–395 (2006).
Glowska, E., Dragun-Damian, A., Dabert, M. & Gerth, M. New Wolbachia supergroups detected in quill mites (Acari: Syringophilidae). Infect. Genet. Evol. 30, 140–146 (2015).
Baldo, L., Lo, N. & Werren, J. H. Mosaic nature of the Wolbachia surface protein. J. Bacteriol. 187, 5406–5418 (2005).
Bordenstein, S. R. & Wernegreen, J. J. Bacteriophage flux in endosymbionts (Wolbachia): Infection frequency, lateral transfer, and recombination rates. Mol. Biol. Evol. 21, 1981–1991 (2004).
Jiggins, F. M., Bentley, J. K., Majerus, M. E. N. & Hurst, G. D. D. How many species are infected with Wolbachia? Cryptic sex ratio distorters revealed to be common by intensive sampling. Proc. R. Soc. B Biol. Sci. 268, 1123–1126 (2001).
Werren, J. H. & Bartos, J. D. Recombination in Wolbachia. Curr. Biol. 11, 431–435 (2001).
Ishmael, N. et al. Extensive genomic diversity of closely related Wolbachia strains. Microbiology 155, 2211–2222 (2009).
Siozios, S. et al. Draft genome sequence of the Wolbachia endosymbiont of Drosophila suzukii. Genome Announc. 1, e00032-13–e00032-13 (2013).
Hamm, C. A. et al. Wolbachia do not live by reproductive manipulation alone: Infection polymorphism in Drosophila suzukii and D. Subpulchrella. Mol. Ecol. 23, 4871–4885 (2014).
Cattel, J. et al. Wolbachia in European populations of the invasive pest Drosophila suzukii: Regional variation in infection frequencies. PLoS One 11, (2016).
Miller, J. M. Whole-genome mapping: A new paradigm in strain-typing technology. Journal of Clinical Microbiology 51, 1066–1070 (2013).
Richardson, M. F. et al. Population genomics of the Wolbachia endosymbiont in Drosophila melanogaster. PLoS Genet. 8, (2012).
Riegler, M., Iturbe-Ormaetxe, I., Woolfit, M., Miller, W. J. & O’Neill, S. L. Tandem repeat markers as novel diagnostic tools for high resolution fingerprinting of Wolbachia. BMC Microbiol. 12, S12 (2012).
Chandler, M. & Mahillon, J. Insertion sequences revisited. Mob. DNA II 62, 305–366 (2002).
Siguier, P., Filée, J. & Chandler, M. Insertion sequences in prokaryotic genomes. Current Opinion in Microbiology 9, 526–531 (2006).
Klasson, L. et al. The mosaic genome structure of the Wolbachia wRi strain infecting Drosophila simulans. Proc. Natl. Acad. Sci. 106, 5725–5730 (2009).
Bordenstein, S. R. & Reznikoff, W. S. Mobile DNA in obligate intracellular bacteria. Nat. Rev. Microbiol. 3, 688–699 (2005).
Duron, O. et al. Transposable element polymorphism of Wolbachia in the mosquito Culex pipiens: Evidence of genetic diversity, superinfection and recombination. Mol. Ecol. 14, 1561–1573 (2005).
Salzberg, S. L., Puiu, D., Sommer, D. D., Nene, V. & Lee, N. H. Genome sequence of the Wolbachia endosymbiont of Culex quinquefasciatus JHB. J. Bacteriol. 191, 1725 (2009).
Wu, M. et al. Phylogenomics of the reproductive parasite Wolbachia pipientis wMel: A streamlined genome overrun by mobile genetic elements. PLoS Biol. 2, (2004).
Comandatore, F. et al. Phylogenomics and analysis of shared genes suggest a single transition to mutualism in Wolbachia of nematodes. Genome Biol. Evol. 5, 1668–1674 (2013).
Gerth, M., Gansauge, M.-T., Weigert, A. & Bleidorn, C. Phylogenomic analyses uncover origin and spread of the Wolbachia pandemic. Nat. Commun. 5, 5117 (2014).
Conner, W. R. et al. Genome comparisons indicate recent transfer of wRi-like Wolbachia between sister species Drosophila suzukii and D. subpulchrella. bioRxiv (2017).
Ling, A. & Cordaux, R. Insertion sequence inversions mediated by ectopic recombination between terminal inverted repeats. PLoS One 5, (2010).
Siguier, P., Gourbeyre, E. & Chandler, M. Bacterial insertion sequences: Their genomic impact and diversity. FEMS Microbiol. Rev. 38, 865–891 (2014).
Siguier, P., Gourbeyre, E., Varani, A., Ton-hoang, B. A. O. & Chandler, M. Everyman’s Guide to bacterial insertion sequences. Microbiol. Spectr. 3, 1–35 (2015).
Rota-Stabelli, O., Blaxter, M. & Anfora, G. Drosophila suzukii:. Current Biology 23, (2013).
Cini, A., Ioriatti, C. & Anfora, G. A review of the invasion of Drosophila suzukii: in Europe and a draft research agenda for integrated pest management. Bulletin of Insectology 65, 149–160 (2012).
Adrion, J. R. et al. Drosophila suzukii: The genetic footprint of a recent, worldwide invasion. Mol. Biol. Evol. 31, 3148–3163 (2014).
Fraimout, A. et al. Deciphering the routes of invasion of Drosophila suzukii by means of ABC random forest. Mol. Biol. Evol. msx050 doi:https://doi.org/10.1093/molbev/msx050 (2017).
Woolfit, M. et al. Genomic evolution of the pathogenic Wolbachia strain, wMelPop. Genome Biol. Evol. 5, 2189–2204 (2013).
Newton, I. L. G. & Sheehan, K. B. Passage of Wolbachia pipientis through mutant Drosophila melanogaster induces phenotypic and genomic changes. Appl. Environ. Microbiol. 81, 1032–1037 (2015).
Vandecraen, J., Chandler, M., Aertsen, A. & Van Houdt, R. The impact of insertion sequences on bacterial genome plasticity and adaptability. Crit. Rev. Microbiol. 1–22 doi:https://doi.org/10.1080/1040841X.2017.1303661 (2017).
Schneider, D. & Lenski, R. E. Dynamics of insertion sequence elements during experimental evolution of bacteria. Research in Microbiology 155, 319–327 (2004).
Schneider, D. I. et al. Uncovering Wolbachia diversity upon artificial host transfer. PLoS One 8, (2013).
Hoffmann, A. A., Hercus, M. & Dagher, H. Population dynamics of the Wolbachia infection causing cytoplasmic incompatibility in Drosophila melanogaster. Genetics 148, 221–231 (1998).
Mazzetto, F., Gonella, E. & Alma, A. Wolbachia infection affects female fecundity in Drosophila suzukii. Bull. Insectology 68, 153–157 (2015).
Cattel, J., Martinez, J., Jiggins, F., Mouton, L. & Gibert, P. Wolbachia-mediated protection against viruses in the invasive pest Drosophila suzukii. Insect Mol. Biol. 25, 595–603 (2016).
Martinez, J. et al. Symbiont strain is the main determinant of variation in Wolbachia -mediated protection against viruses across Drosophila species. Mol. Ecol. doi:https://doi.org/10.1111/mec.14164 (2017).
Dean, M. D. A Wolbachia-associated fitness benefit depends on genetic background in Drosophila simulans. Proc. R. Soc. B Biol. Sci. 273, 1415–1420 (2006).
Sakamoto, H. et al. Transinfection reveals the crucial importance of Wolbachia genotypes in determining the type of reproductive alteration in the host. Genet. Res. 85, 205–210 (2005).
Iturbe-Ormaetxe, I. & O’Neill, S. L. Wolbachia-host interactions: connecting phenotype to genotype. Current Opinion in Microbiology 10, 221–224 (2007).
Veneti, Z., Clark, M. E., Karr, T. L., Savakis, C. & Bourtzis, K. Heads or tails: Host-parasite interactions in the Drosophila-Wolbachia system. Appl. Environ. Microbiol. 70, 5366–5372 (2004).
Ramasamy, S. et al. The evolution of olfactory gene families in Drosophila and the genomic basis of chemical-ecological adaptation in Drosophila suzukii. Genome Biol. Evol. 8, 2297–2311 (2016).
Sutton, E. R., Harris, S. R., Parkhill, J. & Sinkins, S. P. Comparative genome analysis of Wolbachia strain wAu. BMC Genomics 15, 928 (2014).
Sun, L. V., Riegler, M. & O’Neill, S. L. Development of a physical and genetic map of the virulent Wolbachia strain wMelPop. J. Bacteriol. 185, 7077–7084 (2003).
Chrostek, E. & Teixeira, L. Mutualism breakdown by amplification of Wolbachia genes. PLoS Biol. 13, (2015).
Zhang, Z. & Saier, M. H. Jr A novel mechanism of transposon-mediated gene activation. PLOS Genet. 5, e1000689 (2009).
Iturbe-Ormaetxe, I., Burke, G. R., Riegler, M. & O’Neill, S. L. Distribution, expression, and motif variability of ankyrin domain genes in Wolbachia pipientis. J. Bacteriol. 187, 5136–5145 (2005).
Siozios, S. et al. The Diversity and evolution of Wolbachia ankyrin repeat domain genes. PLoS One 8, e55390 (2013).
Bourtzis, K. Transgenesis and the management of vector-borne disease. Transgenes. Manag. Vector-Borne Dis. 627, 104–113 (2008).
Brelsfoard, C. L. & Dobson, S. L. Wolbachia-based strategies to control insect pests and disease vectors. Asian Pacific J. Mol. Biol. Biotechnol. 17, 55–63 (2009).
Reynolds, K. T., Thomson, L. J. & Hoffmann, A. A. The effects of host age, host nuclear background and temperature on phenotypic effects of the virulent Wolbachia strain popcorn in Drosophila melanogaster. Genetics 164, 1027–1034 (2003).
Clancy, D. J. & Hoffmann, A. A. Behavior of Wolbachia endosymbionts from Drosophila Simulans in Drosophila Serrata, A Novel Host. Am. Nat. 149, 975–988 (1997).
Werren, J. H. Biology of Wolbachia. Annu. Rev. Entomol. 124, 587–609 (1997).
O’Neill, S. L. & Karr, T. L. Bidirectional incompatibility between conspecific populations of Drosophila simulans. Nature 348, 178–180 (1990).
Atyame, C. M. et al. Multiple Wolbachia determinants control the evolution of cytoplasmic incompatibilities in Culex pipiens mosquito populations. Mol. Ecol. 20, 286–298 (2011).
Atyame, C. M. et al. Wolbachia divergence and the evolution of cytoplasmic incompatibility in Culex pipiens. PLoS One 9, (2014).
Untergasser, A. et al. Primer3-new capabilities and interfaces. Nucleic Acids Res. 40, (2012).
Zdobnov, E. M. & Apweiler, R. InterProScan - an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17, 847–848 (2001).
Finn, R. D. et al. Pfam: The protein families database. Nucleic Acids Research 42, (2014).
Goloboff, P. A. et al. Improvements to resampling measures of group support. Cladistics 19, 324–332 (2003).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Krzywinski, M. et al. Circos: An information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Acknowledgements
We are thankful to Kevin Floate (Canada), Dong Chu (China) and Vaughn Walton (USA) for providing D. suzukii samples from the field. R.K. is funded by PhD fellowship from the FIRS>T (FEM International Research School-Trentino) programme at Fondazione Edmund Mach, Italy and research grant (P28255-B22) from the Austrian Science Fund (FWF), Austria. O.R.S. is supported by "Scopazzi" project financed by Associazione Produttori Ortofrutticoli Trentini and Fondazione Edmund Mach. We are thankful to Kirsty Mossmann (University of Manchester, UK) for reviewing the English language.
Author information
Authors and Affiliations
Contributions
R.K., S.S. and O.R.S. conceived the experiments. O.R.S. and W.J.M. provided the research material. R.K. performed the experiments. R.K., S.S. and O.R.S. analyzed the data. R.K. and S.S. prepared the figures. R.K. drafted the first version of the manuscript. All of the authors edited and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kaur, R., Siozios, S., Miller, W.J. et al. Insertion sequence polymorphism and genomic rearrangements uncover hidden Wolbachia diversity in Drosophila suzukii and D. subpulchrella . Sci Rep 7, 14815 (2017). https://doi.org/10.1038/s41598-017-13808-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-13808-z
This article is cited by
-
Infection effects of the new microsporidian species Tubulinosema suzukii on its host Drosophila suzukii
Scientific Reports (2021)
-
Endosymbiont diversity in natural populations of Tetranychus mites is rapidly lost under laboratory conditions
Heredity (2020)
-
Large scale genome reconstructions illuminate Wolbachia evolution
Nature Communications (2020)
-
Distinct genotypes and phenotypes in European and American strains of Drosophila suzukii: implications for biology and management of an invasive organism
Journal of Pest Science (2020)
-
Diversity and distribution of Wolbachia in relation to geography, host plant affiliation and life cycle of a heterogonic gall wasp
BMC Evolutionary Biology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.