Abstract
The Hongshan chicken is a Chinese indigenous breed that has two distinctly different tail types. Some chickens have stunted tails as compared to the normal phenotype, and they are termed rumpless. Rumplessness in other chicken breeds was caused by a reduction in the number of coccygeal vertebrae. However, X-ray examination showed that rumpless Hongshan chickens possess the normal number of coccygeal vertebrae. Our analyses of the main tail feathers and tissue sections led us to speculate that their stunted tail appearance may be the result of abnormal feather development. To investigate the genetic mechanism underlying rumplessness in Hongshan chickens, we analyzed the results of various crosses. The results indicated that rumplessness is a Z-linked dominant character. In addition, we chose some normal and rumpless individuals for pool-sequencing. Nucleotide diversity and Fst were calculated, and a selective sweep was detected on the Z chromosome. These analyses allowed us to reduce the search area to 71.8–72 Mb on the Z chromosome (galGal5.0). A pseudogene LOC431648 located in this region appeared a strong candidate involving in Wnt/β-catenin signaling pathway to regulate feather development in chickens.
Similar content being viewed by others
Introduction
Beautiful feathers are powerful tools for attracting mates for male birds. Initiation and development of chicken feather provides a useful model for studies on feather growth. Several molecular pathways are involved in feather development1. Establishment of feather tracts is the first step in feather formation. Noggin, sonic hedgehog bone morphogenetic protein 2 (BMP2), Wnt, and β-catenin were shown to play a role in this step2,3,4,5. Thereafter, feather bud formation starts. Wnt-7a, β-catenin, L-fringe, neural cell adhesion molecule (NCAM), Gremlin, and Wnt-11 are involved with a restrictive expression pattern4,5,6,7,8,9,10. For feather pattern formation, both the activators, fibroblast growth factors (FGF), such as FGF2 and FGF4, and inhibitors of BMPs are necessary11,12,13.
There are several indigenous breeds of chicken with distinct phenotypic traits in China. For instance, Silkies chicken are characterized by dark blue flesh, viscera, and bones and silky feather and Dongxiang blue-shell chicken lays eggs with blue shells. These traits have been investigated in recent years14, 15. Indigenous breeds are excellent models for researching the genetic basis of phenotypic diversity, and the sex-linked characters that are useful for studying the Z chromosome evolution or some related issues.
Hongshan chicken is an indigenous dual-purpose breed in Hubei Province, China. The birds are characterized with yellow beaks, shanks, and feathers, but have two distinctly different types of tails16. Some chickens have cocked tails, as in other chicken breeds, whereas others have pendulous tails, a condition termed rumplessness (Fig. 1). Roosters with normal tails possess a long sickle feather, and both normal roosters and hens have a greater number of main tail feathers than rumpless chickens have.
Rumplessness phenotypes have been investigated in other chicken breeds and animal species. In some cases, abnormality of coccygeal vertebrae has been identified17. A genome-wide association study (GWAS) in Araucana chickens suggested that the rumpless (Rp) gene was located on chromosome 2, proximal to the Iroquois homeobox genes IRX1 and IRX2 18, 19. A recessive type of rumplessness was shown to be controlled by the gene rp-2, homozygosity for rp-2 resulted in a similar phenotype as that of the dominant Rp gene20. Another type of dominant rumplessness caused by mutation of the Brachyury (T) gene has been identified21 and shown to be characterized by a reduction in the number coccygeal vertebrae. Moreover, mutation of the T gene was found to be related to taillessness in mice22 and dogs23. To the best of our knowledge, the physiological and genetic mechanisms of rumplessness in Hongshan chicken have not been studied. Therefore, the gene or mutation causing the tail malformation in Hongshan chicken is unknown.
Determining the basis of rumplessness in Hongshan chicken may be important for protecting, developing, and utilizing this chicken breed. Therefore, in the present study, we investigated the anatomical changes associated with rumplessness using X-ray imaging and microscopy of tissue sections in order to determine the phenotypic characteristics. Further, we carried out a series of crossing experiments to identify the genetic mechanism of rumplessness. With the rapid development of next-generation sequencing (NGS) technologies over the last few years, whole genome sequencing has become a powerful method for gene mapping24. Using NGS, we estimated some population genetic indices to identify the mutation causing rumplessness.
This article has revealed the physiological change in rumpless Hongshan chicken, which was caused by abnormal development of feather. This was a newfound sex-linked mutation in chickens. We provided a suspicious region on Z chromosome by aid of NGS, and a pseudogene in this region probably involving in feather development was reported to be a candidate gene.
Materials and Methods
Animals and ethics statement
The birds used in this study were derived from the “Hongshan Chicken Purification and Rejuvenation” breeding base in Hubei Province, where the Hongshan chicken breed were maintained and initiated from 2003. In this breeding base, rumpless and normal chickens were reared and bred separately. The progenies were sequentially classified to two populations after completion of tail development.
The approval for performing the experiments was obtained from the Animal Care and Use Committee of China Agricultural University (Approval ID: XXCB-20090209), and maintenance and housing of the birds conformed to their required standards.
Body weight
The body weights of Hongshan chickens were measured at different ages. Different groups of chickens were weighed at each age. The frequency of rumplessness was low in our experimental population, therefore, the number of rumpless chickens was lower than that of normal chickens at each age. At the age of 18 weeks, 13 rumpless and 31 normal females were weighed; at 21 weeks, nine rumpless and 30 normal females were weighed; and at 35 weeks, nine rumpless and 21 normal males were weighed.
Two types of Hongshan chicken were raised under same conditions, where an environmentally controlled house with conventional cage system. The temperature was about 20 °C, 2 chickens in each cage, automatic water and free food intake. The diet of each developmental stage was provided by Hubei Tongxing Agricultural Company Limited (Suizhou, China).
Morphological evaluation
We compared the number of coccygeal vertebrae in normal and rumpless Hongshan chicken using X-ray analysis of 15 rumpless chickens (two females and 13 males) and three normal chickens (two females and one male). Skin samples from around the main tail feather of five rumpless and five normal males were collected and fixed in 4% paraformaldehyde overnight, embedded in paraffin wax, and used to prepare 5-µm sections, which were stained using hematoxylin and eosin (HE), as described previously25.
Inheritance of rumplessness
First, we sought to determine whether rumplessness is a recessive or a dominant trait. We performed four crosses: rumpless × rumpless (cross 1), normal × normal (cross 2), normal × rumpless (cross 3), and rumpless × normal (cross 4); the male parent is listed first.
Moreover, we carried out two crosses using roosters of other breeds and Hongshan rumpless females to confirm the results of the former crosses: sex-linked dwarfism × rumpless (cross 5) and sex-linked green shank × rumpless (cross 6). Both the sex-linked dwarf and green-shanked breeds possess normal tails, and the two traits are Z chromosome linked and are recessive26,27,28,29.
Mapping the rumpless allele by whole genomic data
Wing vein blood was obtained from rumpless roosters, normal roosters, and rumpless hens of the same Hongshan population. DNA was isolated from the blood samples using phenol-chloroform protocols. The DNA concentration was determined using a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific Inc.); three pools (24 rumpless males, 24 normal males, and 27 rumpless female) of DNA were prepared with an equal amount of DNA from each sample. DNA libraries were constructed with approximately 500 bp insert size using TruSeq DNA PCR-Free Sample Prep Kit (Illumina), and 2 × 100 bp paired-end reads were sequenced by the Illumina Hiseq. 2000 protocols (BGI, Shenzhen, China). We obtained abundant clean data. After filtering with NGS QC Toolkit (v2.3)30 with default parameters, 191.1 × 106, 187.5 × 106, and 354.9 × 106 high quality read pairs were obtained from rumpless males, normal males, and rumpless females, respectively.
The sequencing data were first mapped to the chicken reference genome (Gallus_gallus-5.0, http://hgdownload.soe.ucsc.edu/downloads.html#chicken) with the Burrows-Wheeler Aligner (BWA)31. The BAM files were sorted and duplicate reads removed using Picard toolkit (https://github.com/broadinstitute/picard). The Genome Analysis Toolkit (GAKT)32 was used for single-nucleotide polymorphism (SNP) calling.
Three population genetic indices were estimated from the sequence data. First, the nucleotide diversity (π) was used to measure the degree of polymorphism within the population33. The second index, fixation index (Fst), measures population differentiation due to genetic structure34. Third, selective sweep refers to reduction or elimination of variation among nucleotides near a mutation under strong positive selection35. In our analyses, π was calculated by PoPoolation36, with parameter set: window size of 10 K, step size of 5 K, minimum allele count of 2, a minimum base quality of 20, a minimum coverage of 10 and a maximum coverage of 200. Fst was calculated by PoPoolation237, with same parameter set as π. SweeD38 was used to detect selective sweeps, with default parameters.
Verification experiments
We collected tail feather follicles from four rumpless and four normal Hongshan chickens three weeks after their tail feathers were removed. Total RNAs were extracted from the tail feather follicles using TRIzol, and reverse transcribed to cDNA using EasyScript One-step gDNA Removal and cDNA Synthesis SuperMix (TransGen Biotech). The expression levels of two genes, DTWD2 and LOC431648, on the Z chromosome were analyzed by quantitative polymerase chain reaction (q-PCR) on an ABI 7500 system (Applied Biosystems, Foster City, CA), with Power SYBR Green PCR Master Mix (Applied Biosystems). All reactions were run in triplicate. Relative gene expression was calculated by the 2−ΔΔCt method39. Primer Premier 540 was used to design the q-PCR primers, the product length was set as 100–250 bp. Because LOC431648 is a pseudogene with high sequence similarity with the mRNA sequence of OCRL on chromosome 4, our primers targeted the amplification of the inconsistent regions between LOC431648 and OCRL. We sequenced the amplification products of LOC431648 to determine whether they were derived from chromosome 4 or Z.
We amplified the whole sequence of the gene LOC431648, and extended the sequence to 1500 bp upstream and downstream to search for potential SNPs or structural variations in 12 rumpless and 12 normal Hongshan chickens (six male and six female).
Data Availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Results
Body weights of the two types of Hongshan chicken
No significant differences in body weight were found between rumpless and normal females at 18 or 21 weeks, or in males at 35 weeks (one-way ANOVA) (Fig. 2). At 18, 21, and 35 weeks, the mean body weights of rumpless Hongshan chickens were 1.10, 1.23, and 1.73 kg, respectively, and that of normal Hongshan chicken 1.03, 1.19, and 1.73 kg, respectively.
Coccygeal vertebral and feather development in rumpless chickens
The X-ray analysis showed that rumpless chickens have a normal coccygeal vertebral structure (Fig. 3A), indicating that rumplessness was not caused by variations in skeletal structure. In addition, visual inspection revealed the presence of a normal oil gland on their tails. The altered tail phenotype of rumpless chickens appeared to be the result of changes in the morphology of tail feathers. The main tail feather of rumpless chickens was more slender and frizzy than normal feathers (Fig. 3B,C).
Morphological observations of two types of Hongshan chicken. (A) X-ray image of normal (top) and rumpless (bottom) rooster. (B) A picture of the main tail feather of normal (left) and rumpless (right) roosters. (C) Section of a feather in the feather follicle at 100 × (left) and 400 × magnification (right); normal rooster on top and rumpless rooster at the bottom.
Rumplessness in Hongshan chicken is a sex-linked dominant character
From the phenotypes of cross 1–4, we found that rumplessness was a Z-linked dominant trait (Table 1). However, there was a lack of male offspring in cross 1–4. Roosters of two other breeds with normal tails in cross 5 and 6 confirmed this result. One of the breeds used in cross 5 and 6 suffered from sex-linked dwarfism, which is a recessive character controlled by the gene GHR on the Z chromosome41. We obtained 40 offspring, 22 of which were dwarf females with normal tails, 16 rumpless males with normal body size, and the two others were males with normal tail and normal body size. The second breed was a Jianghan chicken that had green shanks, which is a recessive trait determined by an unknown gene on the Z chromosome14, 42. We obtained 39 offspring, 17 of which were rumpless and yellow-shanked males, 19 were normal tailed and green-shanked females, two were rumpless and green-shanked males, and one was a normal tailed and yellow-shanked male. Excluding some outliers, our results were similar to the crosses using only Hongshan chickens, which confirmed that rumplessness was a Z-linked dominant trait.
A candidate region on the Z chromosome detected by sequencing data analysis
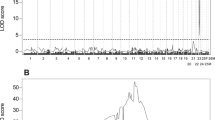
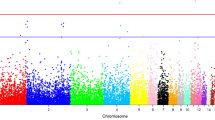
We obtained three pools of sequencing data, namely, rumpless roosters, normal roosters, and rumpless hens. Some population genetic indices were calculated (Fig. 4, Table S1). The π value of 71.8–72 Mb on the Z chromosome in rumpless females was lower than that in normal males; there was a selective sweep at the same region in rumpless chickens. However, the Fst peak was at 79 Mb, and there was a weak signal near 71.8–72 Mb. Two genes were present in this region, DTW domain containing 2 (DTWD2; Z, 71760525–71835352) and inositol polyphosphate 5-phosphatase OCRL-1-like (LOC431648; Z, 72020597–72022434).
Analysis of three pools of sequencing data (rumpless females, rumpless males, and normal males). Nucleotide diversity on Z chromosome (−log10-transformed) of (A) rumpless females, and (B) normal males. Relative differences between rumpless females and normal males for −log10π on (C) Z chromosome, and (D) genome. Likelihood of selective sweeps on Z chromosome of (E) rumpless females and rumpless males, and (F) rumpless males and normal males. Fst between rumpless females and normal males on (G) Z chromosome and (H) genome.
No significant differences were detected between rumpless and normal Hongshan chickens for the expression level of the DTWD2 and LOC431648 genes.
Based on sequence variation between LOC431648 and OCRL, our analysis confirmed that the amplification product was derived from LOC431648. The complete DNA of LOC431648 was sequenced, but no fixed SNP was identified within this pseudogene.
Discussion
Our analyses demonstrate that rumplessness in Hongshan chicken is not caused by changes in coccygeal vertebral structure, and no significant difference was found on body weight, therefore, it is probably not a true example of rumplessness as defined by Dunn (1925). However, as the appearance of this type of Hongshan chicken is similar to rumplessness in other chicken breeds18, we shall continue to refer to it as rumpless in the rest of this study.
Because rumplessness in Hongshan chicken is a qualitative trait, we assumed that it was controlled by a single gene. The results of our crossing experiments supported our assumption, and revealed that rumplessness in Hongshan chicken is a Z-linked dominant trait. However, the crosses produced some outliers (Table 1), which might be explained in two possible ways. Firstly, an error in assessment might have led to a normal individual being scored as rumpless because of developmental delay or feather shedding. Secondly, intermediate type existed in the population, most heterozygous individuals were classified as rumpless, but a few of them might be counted as normal ones by mistake.
We sequenced the pools of DNA from rumpless males and normal males at first. Since we found that rumplessness is a sex-linked dominant phenotype, we added a pool of DNA from rumpless females to reduce the effect of heterozygotes in the sequencing analysis, and our analysis focused on the Z chromosome.
The three population genetics indices used in our sequencing data analysis have been proved to be effective in other similar researches. The π has previously been used in analysis of adaptation to high-altitude hypoxia in dogs43, and to regulatory mutations that disrupt asymmetric hair pigmentation in horses44. Fst has been used to select for sheep without horns (poll)45, to analyze head crests in rock pigeon46, and to investigate the evolution of vision in chicken47. Selective sweep is an efficient method for pool-seq data analysis48 and has been used in a study of rabbit domestication49.
In our researches, both the π and selective sweep analyses identified a region at 71.8–72 Mb on the Z chromosome, and the Fst analysis identified a closely linked region. Two genes, DTWD2 and LOC431648, close to this region were selected as possible candidates causing rumplessness.
A previous study on genome-wide association showed that several SNPs near the human homolog of DTWD2 were associated with the maximum number of alcoholic drinks consumed in 24 hours50, which may be irrelevant to rumplessness in chickens. Therefore, we focused on the LOC431648 gene. The LOC431648 is a processed pseudogene, and its sequence is very similar to the mRNA sequence of OCRL on chromosome 4. Although pseudogenes tend to have lost some functionality compared to the intact gene51, some are functional and can perform regulatory activities similar to those of noncoding DNA52. Our sequencing results showed that LOC431648 was expressed.
The OCRL gene encodes an inositol polyphosphate-5-phosphatase and is involved in inositol phosphate metabolism. Intriguingly, inositol metabolism has a role in the Wnt/β-catenin signaling pathway53 that participates in the induction of feather primordia and affects the shape of feather buds in chickens10.
We propose that variations in (or near) the LOC431648 gene cause rumplessness in Hongshan chicken. No differential expression of the gene was detected by q-PCR and no fixed variations in DNA sequence were identified. The feather follicle was collected three weeks after removing the tail feather, when the feather bud had already formed. Therefore, the LOC431648 expression might have reduced to moderate levels owing to restrictive expression. Besides, the difference in gene expression occurred in a small location of the feather follicle, but the RNA extracted from a mixture of feather follicle and some skin around it might affect the q-PCR results.
Considering that no differential expression of the gene was detected by q-PCR, more powerful tools need to be adopted, like fluorescence in situ hybridization (FISH). Furthermore, we cannot exclude the possibility of epigenetic modifications. To investigate these possibilities, further research will be necessary.
References
Chen, C. F. et al. Development, regeneration, and evolution of feathers. Annual review of animal biosciences 3, 169–195, doi:10.1146/annurev-animal-022513-114127 (2015).
Fliniaux, I., Viallet, J. P. & Dhouailly, D. Signaling dynamics of feather tract formation from the chick somatopleure. Development 131, 3955–3966, doi:10.1242/dev.01263 (2004).
Scaal, M. et al. BMPs induce dermal markers and ectopic feather tracts. Mechanisms of development 110, 51–60 (2002).
Widelitz, R. B. et al. Wnt-7a in feather morphogenesis: involvement of anterior-posterior asymmetry and proximal-distal elongation demonstrated with an in vitro reconstitution model. Development 126, 2577–2587 (1999).
Widelitz, R. B., Jiang, T. X., Lu, J. & Chuong, C. M. beta-catenin in epithelial morphogenesis: conversion of part of avian foot scales into feather buds with a mutated beta-catenin. Developmental biology 219, 98–114, doi:10.1006/dbio.1999.9580 (2000).
Chen, C. W. & Chuong, C. M. Dynamic expression of lunatic fringe during feather morphogenesis: a switch from medial-lateral to anterior-posterior asymmetry. Mechanisms of development 91, 351–354 (2000).
Jiang, T. X., Jung, H. S., Widelitz, R. B. & Chuong, C. M. Self-organization of periodic patterns by dissociated feather mesenchymal cells and the regulation of size, number and spacing of primordia. Development 126, 4997–5009 (1999).
Bardot, B. et al. Drm/Gremlin, a BMP antagonist, defines the interbud region during feather development. The International journal of developmental biology 48, 149–156, doi:10.1387/ijdb.041804bb (2004).
Ohyama, A., Saito, F., Ohuchi, H. & Noji, S. Differential expression of two BMP antagonists, gremlin and Follistatin, during development of the chick feather bud. Mechanisms of development 100, 331–333 (2001).
Chang, C. H. et al. Distinct Wnt members regulate the hierarchical morphogenesis of skin regions (spinal tract) and individual feathers. Mechanisms of development 121, 157–171, doi:10.1016/j.mod.2003.12.004 (2004).
Song, H. K., Lee, S. H. & Goetinck, P. F. FGF-2 signaling is sufficient to induce dermal condensations during feather development. Developmental dynamics: an official publication of the American Association of Anatomists 231, 741–749, doi:10.1002/dvdy.20243 (2004).
Jung, H. S. et al. Local inhibitory action of BMPs and their relationships with activators in feather formation: implications for periodic patterning. Developmental biology 196, 11–23, doi:10.1006/dbio.1998.8850 (1998).
Noramly, S. & Morgan, B. A. BMPs mediate lateral inhibition at successive stages in feather tract development. Development 125, 3775–3787 (1998).
Dorshorst, B., Okimoto, R. & Ashwell, C. Genomic regions associated with dermal hyperpigmentation, polydactyly and other morphological traits in the Silkie chicken. The Journal of heredity 101, 339–350, doi:10.1093/jhered/esp120 (2010).
Wang, Z. et al. An EAV-HP insertion in 5′ Flanking region of SLCO1B3 causes blue eggshell in the chicken. PLoS genetics 9, e1003183, doi:10.1371/journal.pgen.1003183 (2013).
Liang, Z. et al. Purification and Rejuvenation Breeding for Hongshan Chicken. Hubei Agricultural Sciences 48, 667–670 (2009).
Dunn, L. C. The inheritance of rumplessness in the domestic fowl. Journal of Heredity 16, 127–134 (1925).
Noorai, R. E., Freese, N. H., Wright, L. M., Chapman, S. C. & Clark, L. A. Genome-wide association mapping and identification of candidate genes for the rumpless and ear-tufted traits of the Araucana chicken. PloS one 7, e40974, doi:10.1371/journal.pone.0040974 (2012).
Freese, N. H., Lam, B. A., Staton, M., Scott, A. & Chapman, S. C. A novel gain-of-function mutation of the proneural IRX1 and IRX2 genes disrupts axis elongation in the Araucana rumpless chicken. PloS one 9, e112364, doi:10.1371/journal.pone.0112364 (2014).
Landauer, W. Recessive Rumplessness of Fowl with Kyphoscoliosis and Supernumerary Ribs. Genetics 30, 403–428 (1945).
Zwilling, E. The Development of Dominant Rumplessness in Chick Embryos. Genetics 27, 641–656 (1942).
Dunn, L. C. & Gluecksohn-Schoenheimer, S. The Inheritance of Taillessness (Anury) in the House Mouse. II. Taillessness in a Second Balanced Lethal Line. Genetics 24, 587–609 (1939).
Indrebo, A. et al. A study of inherited short tail and taillessness in Pembroke Welsh corgi. The Journal of small animal practice 49, 220–224, doi:10.1111/j.1748-5827.2007.00435.x (2008).
Mascher, M. et al. Mapping-by-sequencing accelerates forward genetics in barley. Genome biology 15, R78, doi:10.1186/gb-2014-15-6-r78 (2014).
Jiang, T. X., Widelitz, S. S., Chuong, R. B. C. M. In Molecular Basis of Epithelial Appendage Morphogenesis . Austin (ed. Chuong CM) 395-408 (1998).
Leung, F. C., Styles, W. J., Rosenblum, C. I., Lilburn, M. S. & Marsh, J. A. Diminished hepatic growth hormone receptor binding in sex-linked dwarf broiler and leghorn chickens. Proceedings of the Society for Experimental Biology and Medicine. Society for Experimental Biology and Medicine 184, 234–238 (1987).
Burnside, J., Liou, S. S. & Cogburn, L. A. Molecular cloning of the chicken growth hormone receptor complementary deoxyribonucleic acid: mutation of the gene in sex-linked dwarf chickens. Endocrinology 128, 3183–3192, doi:10.1210/endo-128-6-3183 (1991).
Knox, C. W. The Inheritance of Shank Color in Chickens. Genetics 20, 529–544 (1935).
Bitgood, J. J. Linear relationship of the loci for barring, dermal melanin inhibitor, and recessive white skin on the chicken Z chromosome. Poultry science 67, 530–533 (1988).
Patel, R. K. & Jain, M. NGS QC Toolkit: a toolkit for quality control of next generation sequencing data. PloS one 7, e30619, doi:10.1371/journal.pone.0030619 (2012).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595, doi:10.1093/bioinformatics/btp698 (2010).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome research 20, 1297–1303, doi:10.1101/gr.107524.110 (2010).
Nei, M. & Li, W. H. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proceedings of the National Academy of Sciences of the United States of America 76, 5269–5273 (1979).
Holsinger, K. E. & Weir, B. S. Genetics in geographically structured populations: defining, estimating and interpreting F(ST). Nature reviews. Genetics 10, 639–650, doi:10.1038/nrg2611 (2009).
Smith, J. M. & Haigh, J. The hitch-hiking effect of a favourable gene. Genetical research 23, 23–35 (1974).
Kofler, R. et al. PoPoolation: a toolbox for population genetic analysis of next generation sequencing data from pooled individuals. PloS one 6, e15925, doi:10.1371/journal.pone.0015925 (2011).
Kofler, R., Pandey, R. V. & Schlotterer, C. PoPoolation2: identifying differentiation between populations using sequencing of pooled DNA samples (Pool-Seq). Bioinformatics 27, 3435–3436, doi:10.1093/bioinformatics/btr589 (2011).
Pavlidis, P., Zivkovic, D., Stamatakis, A. & Alachiotis, N. SweeD: likelihood-based detection of selective sweeps in thousands of genomes. Molecular biology and evolution 30, 2224–2234, doi:10.1093/molbev/mst112 (2013).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) Method. Methods 25, 402–408, doi:10.1006/meth.2001.1262 (2001).
Singh, V. K., Mangalam, A. K., Dwivedi, S. & Naik, S. Primer premier: program for design of degenerate primers from a protein sequence. BioTechniques 24, 318–319 (1998).
Ouyang, J. H. et al. Single nucleotide polymorphism (SNP) at the GHR gene and its associations with chicken growth and fat deposition traits. British poultry science 49, 87–95, doi:10.1080/00071660801938817 (2008).
Li, G. et al. A genome-wide association study identifies novel single nucleotide polymorphisms associated with dermal shank pigmentation in chickens. Poultry science 93, 2983–2987, doi:10.3382/ps.2014-04164 (2014).
Gou, X. et al. Whole-genome sequencing of six dog breeds from continuous altitudes reveals adaptation to high-altitude hypoxia. Genome research 24, 1308–1315, doi:10.1101/gr.171876.113 (2014).
Imsland, F. et al. Regulatory mutations in TBX3 disrupt asymmetric hair pigmentation that underlies Dun camouflage color in horses. Nature genetics 48, 152–158, doi:10.1038/ng.3475 (2016).
Kijas, J. W. et al. Genome-wide analysis of the world’s sheep breeds reveals high levels of historic mixture and strong recent selection. PLoS biology 10, e1001258, doi:10.1371/journal.pbio.1001258 (2012).
Shapiro, M. D. et al. Genomic diversity and evolution of the head crest in the rock pigeon. Science 339, 1063–1067, doi:10.1126/science.1230422 (2013).
Wang, M. S. et al. Positive selection rather than relaxation of functional constraint drives the evolution of vision during chicken domestication. Cell research 26, 556–573, doi:10.1038/cr.2016.44 (2016).
Boitard, S., Schlotterer, C., Nolte, V., Pandey, R. V. & Futschik, A. Detecting selective sweeps from pooled next-generation sequencing samples. Molecular biology and evolution 29, 2177–2186, doi:10.1093/molbev/mss090 (2012).
Carneiro, M. et al. Rabbit genome analysis reveals a polygenic basis for phenotypic change during domestication. Science 345, 1074–1079, doi:10.1126/science.1253714 (2014).
Pan, Y. et al. Genome-wide association studies of maximum number of drinks. Journal of psychiatric research 47, 1717–1724, doi:10.1016/j.jpsychires.2013.07.013 (2013).
Vanin, E. F. Processed pseudogenes: characteristics and evolution. Annual review of genetics 19, 253–272, doi:10.1146/annurev.ge.19.120185.001345 (1985).
Hirotsune, S. et al. An expressed pseudogene regulates the messenger-RNA stability of its homologous coding gene. Nature 423, 91–96, doi:10.1038/nature01535 (2003).
Gao, Y. & Wang, H. Y. Inositol pentakisphosphate mediates Wnt/beta-catenin signaling. The Journal of biological chemistry 282, 26490–26502, doi:10.1074/jbc.M702106200 (2007).
Acknowledgements
This study was supported by the earmarked fund for the National Scientific Supporting Projects of China (201504310411128), and Beijing Innovation Team of the Modern Agro-industry Technology Research System (BAIC04-2016). We thank our colleagues of the poultry team of National Engineering Laboratory for Animal Breeding of China Agricultural University, and Institute of Animal Husbandry and Veterinary Science of Hubei Academy of Agricultural Sciences for their assistance in sample collection and helpful comments on the manuscript.
Author information
Authors and Affiliations
Contributions
L.Q. conceived and designed the experiments; Q.W. wrote the main manuscript text, performed computational analyses, and carried out the experimental validations; J.P. kept and provided experimental samples, performed crossing experiments and X-ray inspection; A.P. and J.S. participated in collecting samples. Q.W. and J.P. contributed equally to this work. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Q., Pi, J., Pan, A. et al. A novel sex-linked mutant affecting tail formation in Hongshan chicken. Sci Rep 7, 10079 (2017). https://doi.org/10.1038/s41598-017-10943-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-10943-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.