Abstract
Understanding the molecular mechanisms of bacterial antibiotic resistance will help prepare against further emergence of multi-drug resistant strains. MacQ is an enzyme responsible for the multi-drug resistance of Acidovorax sp. strain MR-S7. MacQ has acylase activity against both N-acylhomoserine lactones (AHLs), a class of signalling compounds involved in quorum sensing, and β-lactam antibiotics. Thus, MacQ is crucial as a quencher of quorum sensing as well as in conferring antibiotic resistance in Acidovorax. Here, we report the X-ray structures of MacQ in ligand-free and reaction product complexes. MacQ forms a 170-kDa capsule-shaped molecule via face-to-face interaction with two heterodimers consisting of an α-chain and a β-chain, generated by the self-cleaving activity of a precursor polypeptide. The electron density of the spacer polypeptide in the hollow of the molecule revealed the close orientation of the peptide-bond atoms of Val20SP-Gly21SP to the active-site, implying a role of the residues in substrate binding. In mutational analyses, uncleaved MacQ retained degradation activity against both AHLs and penicillin G. These results provide novel insights into the mechanism of self-cleaving maturation and enzymatic function of N-terminal nucleophile hydrolases.
Similar content being viewed by others
Introduction
Acidovorax sp. strain MR-S7 is a gram-negative bacterium that was isolated from activated sludge in a treatment system for penicillin G-polluted wastewater1,2,3. We previously reported that Acidovorax sp. strain MR-S7 is resistant to various β-lactam antibiotics and is able to degrade a broad range of N-acylhomoserine lactones (AHLs)3. AHLs are a class of signalling compounds involved in the bacterial intercellular communication known as quorum sensing4. Quorum sensing regulates many bacterial behaviours, including luminescence5, biofilm formation2, 6,7,8,9,10,11, signal turnover12, pigment production13, 14, antibiotic production14, 15, swarming16, and virulence17,18,19,20,21. AHL-degrading enzymes interfere with the AHL-mediated quorum sensing system in a process termed quorum quenching22. Different pathogens that are serious health threats, particularly in nosocomial infections, produce virulence factors under the control of a quorum-sensing system. Thus, quorum quenching could be a key strategy to interfere with bacterial infectious diseases23,24,25,26. AHL-acylase is one of the enzymes capable of inactivating AHLs via hydrolysis of the amide bond of the acyl side-chain to produce homoserine lactone (HSL) and fatty acid3, 22. Several enzymes exhibiting AHL-acylase activity have been reported. They include AiiD from Ralstonia sp. XJ12B27, PvdQ and QuiP from Pseudomonas aeruginosa PAO128,29,30, AhlM from Streptomyces sp. M66431, AiiC from Anabaena sp. PCC712032, Aac from Shewanella sp. MIB01533, and HacA and HacB from P. syringae B728a34.
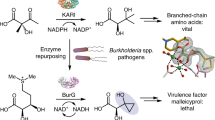
We recently identified a gene (macQ) responsible for β-lactam antibiotic resistance of Acidovorax sp. strain MR-S7. The recombinant MacQ protein exhibited bifunctional acylase activity against various ranges of AHLs and against multiple β-lactam antibiotics, including penicillin G, ampicillin, and amoxicillin (Fig. 1)3. The MacQ protein is comprised of 806 amino acid residues, including an N-terminal signal peptide (Fig. 2a), and belongs to the N-terminal nucleophile (Ntn) hydrolase superfamily. Ntn hydrolases are synthesized as inactive precursor single polypeptides, which become activated by self-cleaving activity, generating the mature hetero-multimeric enzyme composed of α- and β-chains35. The penicillin G/cephalosporin/AHL-acylases are more homologous to MacQ among the Ntn hydrolases. The precursor polypeptide of these homologues harbours two distinct cleavage sites that enable the formation of a mature heterodimer composed of α- and β-chains, with the release of a spacer polypeptide (SP)30, 36,37,38,39. The N-terminal Ser/Thr/Cys of the β-chain is the catalytically essential residue, which acts as a nucleophile to initiate amide bond cleavage of various substrate compounds40,41,42,43. Extensive functional and structural analyses have been reported for Ntn hydrolase enzymes. Two enzymes, AhlM and KcPGA, exhibit acylase activity against both AHLs and penicillin G31, 44, similar to MacQ. However, the structures and substrate adaptation mechanisms of those bifunctional acylases are unknown.
Amidohydrolysis of AHL (C10-HSL) and penicillin G catalyzed by MacQ. As shown in Table 1, MacQ is capable of degrading wide varieties of AHLs as well as penicillin G.
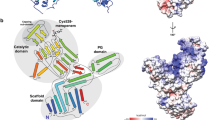
Heterodimeric structure of MacQ. (a) Schematic representation of the single MacQ precursor polypeptide and its self-cleaving maturation products. The N-terminal sequencing and MALDI-TOF/MS analysis indicated that the α-chain, SP, and β-chain correspond to residues 5α-182α (29–206), 1SP-27SP (207–233), and 1β-581β (234–814), respectively. The parentheses refer to the residue number of precursor polypeptide of MacQ including signal peptide sequence. The signal peptide (residue 1–24) was predicted with the SignalP 4.1 program (http://www.cbs.dtu.dk/services/SignalP/). (b) Stereo-view ribbon diagram of MacQ heterodimer with the SP generated by self-cleaving activity of a MacQ precursor single polypeptide chain. The α-chain, SP, and β-chain are coloured in orange, magenta, and green, respectively. The bound PAA and the catalytic Ser1β are also shown as stick models in yellow and blue, respectively, and are labelled.
Here, we describe the X-ray crystal structures of the bifunctional quorum-quenching and β-lactam antibiotic-degrading Ntn hydrolase MacQ in ligand-free form and in complex with reaction products, fatty acid, or phenyl acetic acid. The structures revealed that the three distinct polypeptide chains generated by self-cleaving activity assemble into a capsule-like higher oligomeric complex.
Results and Discussion
Recombinant expression of MacQ by Escherichia coli
The expression of macQ isolated from Acidovorax sp. strain MR-S7 was performed using Escherichia coli cells, as previously described3. The precursors of AHL-acylases and related enzymes are cleaved into three polypeptide chains (α-chain, internal SP, and β-chain) and the catalytically active mature enzymes are generated as a heterodimer consisting of α- and β-chains, with the SP being released30. As expected, SDS-PAGE analysis confirmed two bands at approximately 20 kDa and 62 kDa, which corresponded to the α- and β-chains of MacQ, respectively (Fig. 2a). Although we created an expression construct to produce recombinant MacQ with His6-tags at both the N- and C-termini (Fig. 2a), N-terminal sequence analysis for the 20 kDa band revealed that the origin of the α-chain begins at Ser29 and a truncated N-terminal His6-tag. MALDI-TOF/MS analysis predicted a subunit weight for the α-chain of 19.23 kDa. On the basis of these data, we presume that the α-chain of the purified sample consists of 178 amino acid residues from Ser29 (Ser5α) to Ala206 (Ala182α), with a theoretical molecular weight of 19.21 kDa (Fig. 2a). The cleavage site at Ala206 agreed well with that of the AHL-acylase homologue PvdQ from P. aeruginosa strain PAO1, as previously reported30. MALDI TOF/MS analysis also detected minor peaks between 19.2–19.6 kDa, suggesting that the cleavage position between the α-chain and SP might not be specific. The other cleavage site was at Ser234, which defines the Ntn residue of the β-chain (Ser1β).
MacQ forms a capsule-shaped dimer of heterodimer containing SP chains
The crystal structures of MacQ, both ligand-free and in complex with the reaction products of decanoic acid (C10) and phenyl acetic acid (PAA), were determined at 2.6 Å, 2.2 Å, and 1.75 Å resolution, respectively. The initial atomic model of ligand-free MacQ was built by the molecular replacement method using the PvdQ heterodimer as a search model (PDB code, 2wyb; 34% amino-acid sequence identity)36. The C10 complex crystallized in the P1 space group, similar to that of ligand-free MacQ, while the PAA complex crystallized in the P21 space group. The structure of the PAA complex was solved by the molecular replacement method using the ligand-free MacQ as a search model.
The MacQ structure clearly revealed a typical Ntn-hydrolase fold comprised of a four-layered α-β-β-α motif45 mainly composed of two polypeptide chains, α- and β-chains, generated by self-cleaving activity. The N-terminal nucleophile is Ser1β, which corresponds to Ser234 in the precursor sequence (Fig. 2). The DALI structure similarity search46 demonstrated the closest structural homology of MacQ to PvdQ [PDB codes, 3L94 (α-chain)47/2WYC (β-chain)36; Z score, 21.9/49.7; rmsd, 1.2/2.2 Å for 163/546 Cα atoms; sequence identity for the fit region, 38%/35%]. However, two novel findings in the structure of MacQ compared to the structures of PvdQ and other related enzymes were evident. First, electron density was observed for the SP lying in the vicinity of the active-site pocket (Figs 2b and 3c). This was quite unexpected, because the SP was not found in any previously reported structures of penicillin G/cephalosporin/AHL-acylases and is believed to fall away in the course of the autoproteolytic maturation. The SP atoms were refined with full occupancy, while the B-factor values of the SP were relatively high compared to those of α- and β-chain, indicating increased conformational heterogeneity or a lower occupancy of the SP as compared to the α- and β-chains. Second, the crystal structure analysis revealed that MacQ forms a high oligomeric quaternary structure. The two heterodimers consisting of the α- and β-chain are arranged face-to-face, resulting in the formation of an approximately 170-kDa capsule-shaped molecule (dimer of heterodimer) containing two SP chains (Fig. 3). The unit-cell of the P1 lattice contains two capsule-shaped molecules, which are identical and can be superimposed with a root mean squared (RMS) difference of 0.47 Å for 1512 Cα atoms. The asymmetric unit of the P21 lattice also contains two PAA-bound molecules, which superimposed well on those of ligand-free and C10 complexes of MacQ in the P1 lattice, with an RMS difference of ~1.0 Å for 1505 Cα atoms. These results suggest that the lattice forces and the inter-molecular crystal contacts do not affect the overall oligomeric structure of MacQ. Analytical gel filtration also showed that the MacQ predominantly exists as a dimer of heterodimer in solution, although a relatively small amount of heterodimer was also observed (Supplementary Fig. S1).
Capsule-shaped assembly of MacQ. (a) Ribbon diagram of molecular structure of MacQ formed face-to-face by two heterodimers shown in Fig. 2b. One heterodimer is shown as the same colour scheme to that in Fig. 2b, and the other heterodimer is shown in gray. Two perpendicular views are shown. (b) Cross section of the molecule in surface representation showing the interior space of the capsule-like molecule of MacQ and the substrate-binding pocket indicated by the red arrow. The bound PAA is also shown as a yellow stick model. (c) mFo-DFc omit map for the observed SP region contoured at 2.6σ level. The final refined model of the SP is shown as stick model and the residues are labelled. The bound PAA and the Ser1β are also shown in stick model. As shown here, the model for the residues Gly1SP-Ala7SP and Glu23SP-Gly27SP are not built due to the poor electron density.
Active-site structure and presumed substrate-binding mechanism
To gain insights into the substrate-binding mechanism of MacQ, we attempted to determine the structures of MacQ in complex with substrates (C10-HSL and penicillin G). In accordance with a previously reported procedure36, the crystals were soaked in saturated substrate solution with a lower pH value (pH 5.5) before X-ray diffraction. The reaction product, C10/PAA, was successfully trapped at the hydrophobic active-site pocket, while the electron density for the hydrolyzed counterpart, HSL/6-aminopenicillanic acid (6-APA), was not observed. The results suggest that the more hydrophilic HSL/6-APA diffused into the solvent immediately after the enzymatic amidohydrolyzing reaction.
The MacQ homodimer contains two active sites. The C10 was found in a different binding state in each active-site pocket. In one active site, the carboxyl group of C10 hydrogen-bonded with the side-chain hydroxyl and amino group of the catalytic residue Ser1β, while in the other active site the hydroxyl oxygen (Oγ) of Ser1β was covalently linked to the carbonyl carbon of the C10 molecule (Fig. 4a and b). These distinct binding states have also been observed in the structural analyses of PvdQ in complex with dodecanoic acid36. Despite the different binding states of C10 at the active site pocket, the two heterodimers superimposed well with an RMS difference of ~0.3 Å for 756 Cα atoms. No side-chain conformational changes were detected around the active site, and therefore the factors that cause the different binding modes of the substrate in the different active sites remain unclear. In contrast, PAA was bound in the same state in each active-site pocket without covalent bonds. The atomic models of PAA in two different conformations fit well to the observed electron density and were refined with reasonable B-factor values in all active sites (Fig. 4c and d). The C10/PAA was accommodated in the highly hydrophobic active-site pocket created by the side-chains of Trp24β, Phe32β, Phe50β, Gln57β, Ile58β, His68β, Val70β, Trp165β, Trp189β, and Val190β (Fig. 4a and c). Amino-acid sequence alignments of MacQ, AhlM, and PvdQ showed that these residues are not completely identical, but are highly conserved (Supplementary Fig. S2). Interestingly, although the induced-fit mechanism for the binding of dodecanoic acid was reported in the structural studies of PvdQ36, the active-site of MacQ was determined to be motionless and the side-chain conformations of all the aforementioned residues superimposed well among ligand-free, C10 complex, and PAA complex structures, with a constant pocket volume of approximately 170 Å3. These observations might partially account for the broad substrate specificity of MacQ.
Active site structure of MacQ. The colour scheme is similar to that in Fig. 2b. (a) Bound C10 fatty acid covalently linked to the Ser1β. The side-chains of nearby residues created the hydrophobic C10-binding pocket are also shown and labelled. The expected binding position of HSL moiety is roughly indicated as a red dotted circle. The residues of the SP near the active site are also shown as stick models and labelled. (b) mFo-DFc omit map for the bound C10 and the Ser1β contoured at 2.5σ level. Two binding states of C10 were observed: the covalently linked C10 to Oγ of Ser1β (upper) and the noncovalently bound C10 (lower). (c) Bound PAA. The expected binding position of 6-APA moiety is roughly indicated as a red dotted circle. (d) mFo-DFc omit map for the bound PAA and the Ser1β contoured at 2.5σ level. Two PAA molecules modelled with slightly different positions and conformations fit well onto the observed electron density map.
The remaining SP in the hollow surface of the molecule is probably serendipitous, but we presume that the SP could be involved in MacQ function for the following reasons. First, the SP is observed in all heterodimeric units of MacQ in the same conformation, both in the presence and absence of ligands (Fig. 5e). Second, the residues from Gln19SP to Gly22SP form a type-II β-turn structure together with ordered solvent molecules, and the main-chain atoms that form a peptide linkage between the Val20SP and Gly21SP lie at the entrance of the hydrophobic pocket, where the residues can interact with the HSL/6-APA moiety of the bound substrate. The distance between Oγ of the Ser1β and the peptide-bond atoms of Val20SP-Gly21SP is ~7 Å. It is considered that the loose binding of C10/PAA without an induced-fit mechanism might be compensated for by the presumed interactions between SP and HSL/6-APA. To investigate whether the SP is important for enzymatic function of MacQ, we created an SP-shortened mutant and examined its enzymatic activity level.
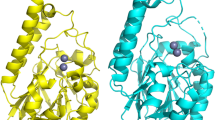
Mutational analyses of MacQ. (a) Cleavage site of MacQ and PvdQ and the amino acid sequence of S210A and SPΔ7 mutants around SP region. Deletion or substitutions are highlighted in yellow. The Gly21SP (Gly203 in the uncleaved form), the nearest SP residue to the active site, is indicated by the red asterisk. The seven-residue deletion is the minimum requirements to delete the Gly21SP (Gly203) in the cleaved form of MacQ with keeping the cleavage sequence Gly209-Ser210. (b) SDS-PAGE analysis of wild-type MacQ and mutants S210A and SPΔ7 under reducing (β-mercaptoethanol (βME) + ) and non-reducing (βME −) conditions. The results clearly show that both S210A and SPΔ7 mutants were not cleaved. (c) Overall structure of MacQ SPΔ7. The structure shows the same capsule-like molecule as that of wild-type MacQ, but consists of two uncleaved monomers. The residues 183–202, which correspond to the SP of the wild-type enzyme, are coloured in dark magenta. (d) Structure superimposition of wild-type MacQ heterodimer and SPΔ7 monomer. The α-chain, SP, and β-chain of wild-type MacQ are coloured in orange, magenta, and green, respectively. The SPΔ7 is coloured in gray, while the Gly183-Gly202 is coloured in dark magenta, as shown in (a). The conformation of the SP region of wild-type MacQ and the corresponding Gly183-Gly202 of SPΔ7 is completely different. (e) Structure superimposition of the SP/Ser1β of wild-type MacQ in ligand-free and C10/PAA complex and the Gly183-Ser203 of the SPΔ7. A total of 12 chains for wild-type MacQ and 4 chains for SPΔ7 are shown. The SPΔ7 residues of 193–197 for chain B, 178–200 for chain C and 179–200 for chain D could not be built due to poor electron density. The carbon atoms are coloured in magenta for the SP of wild-type enzyme, and in gray for chain A, cyan for chain B, yellow for chain C, and light blue for chain D of the SPΔ7.
Spacer-shortened mutant MacQ is an enzymatically active single polypeptide
The self-cleaving maturation mechanism has been extensively studied using cephalosporin acylase (CA)37,38,39, 48. Various SP-shortened mutants of CA were created48. They retained self-cleaving activity similar to wild-type CA. On the basis of the previous reports regarding CA, we created an SP-shortened MacQ by deleting seven residues from Val20SP to Val26SP (from Val203 to Val209; Fig. 5a). We expected that the SP-shortened mutant, termed SPΔ7, would provide important clues for the enzymatic function of the C-terminal region of the SP. The recombinant SPΔ7 was expressed and purified by the same procedure as the wild-type enzyme. Unexpectedly, SDS-PAGE analysis showed that the purified SPΔ7 was uncleaved and was present as a single polypeptide chain with a subunit weight of approximately 80 kDa (Figs. 5a and b), suggesting that the seven-residue deletion of the SP inactivated the self-cleaving maturation, despite the fact that SPΔ7 possesses the catalytic serine (Ser1β in the cleaved form). We also investigated whether SPΔ7 retains AHL/penicillin G acylase activity. Interestingly, SPΔ7 exhibited unambiguous degrading activity against almost all of the substrates tested for wild-type MacQ (Table 1, Supplementary Figs S3 and S4)3. These results indicate that the self-cleaving maturation is not a requisite process for the enzymatic activation of MacQ. In light of this finding, we were interested in determining the crystal structure of SPΔ7 to understand how the uncleaved polypeptide chains fold and form the active site.
Structure analysis of MacQ SPΔ7 mutant
We successfully determined the crystal structure of MacQ SPΔ7 in ligand-free form. SPΔ7 crystals were obtained only using a different crystallization solution containing tacsimate buffer (pH 6.0), but the crystals displayed unit-cell parameters with an identical space group similar to those of the PAA complex of the wild-type MacQ. Unfortunately, attempts to determine the structure of ligand-bound SPΔ7 were unsuccessful because of the low reproducibility and fragility of SPΔ7 crystals. The asymmetric unit contains two capsule-like molecules that form the same structure as the wild-type MacQ. However, each molecule was determined to be a homodimer composed of two uncleaved SPΔ7 polypeptides (Fig. 5b,c, and d). The overall structure of SPΔ7 superimposed very well on that of the wild-type enzyme without its SP region (Fig. 5d and e). We could build the continuous atomic model for chain A (Supplementary Fig. S5), while several residues corresponding to the SP region of the wild-type enzyme were disordered for the chain B, and most of the residues of the SP region (residue 179–200) were disordered for chains C and D. The conformation of the SP region of the SPΔ7 (residue 183–202) was found to be completely different from that of the SP in the wild-type enzyme (Fig. 5e).
The structure of SPΔ7 indicates a possible mechanism of the autoproteolytic cleavage between Gly202 and Ser203 (Supplementary Fig. S6). It is likely that the water molecule that hydrogen-bonds with the main-chain O of Gln201 and His225 (Wat X in Supplementary Fig. S6) forms the pseudo-tetrahedral geometry and deprotonates the side-chain hydroxyl of Ser203. The activated hydroxyl of Ser203 then carries out the nucleophilic attack on the main-chain C of the Gly202. The oxyanionic intermediate is likely stabilized by the main-chain amide of Val272, thereby lowering the activation energy for the peptide bond cleavage. In the SPΔ7 structure, the ~3.4 Å distance between the main-chain O of Gly202 and the main-chain amide of Val272 would be too distant to form a hydrogen bond. The seven-residue deletion may affect the main-chain conformation of the Gly202-Ser203, leading to the destabilization of the intermediate structure. The SPΔ7 structure also revealed that the side-chain N of Asn480 approaches the main-chain O of Gly202 in a direction nearly vertical to the peptide plane. The conformation of the “Gly-Ser” in the wild-type MacQ precursor might have a more favourable inter-atomic geometry to access the oxyanion hole created by the main-chain N of Val272 and possibly the side-chain N of Asn480 (Supplementary Fig. S6).
Because the uncleaved SPΔ7 is enzymatically active for both AHLs and penicillin G, the question arises as to how the SPΔ7 recognizes the substrate at the active site. The hydrophobic active-site pocket where the C10/PAA is bound in the wild-type enzyme is also present in the structure of SPΔ7. However, the pocket is insulated from the solvent because of the presence of the uncleaved polypeptide chain (residues Leu201-Ser203) (Fig. 6). Thus, it is likely that certain local conformational changes are required for the binding of the substrate and for catalytic reaction. To check whether the Ser210 (corresponding to Ser203 of SPΔ7 or Ser1β of wild-type enzyme) is really essential for autoproteolytic activation and AHL/penicillin G-degrading function31, 49, 50, we created a S210A mutant and examined its enzymatic activity.
Comparison of molecular surface and active site cavities of wild-type MacQ and SPΔ7 mutant. (a) Top view and (b) side view molecular surface representation of heterodimeric wild-type MacQ and monomeric (uncleaved) SPΔ7. The colour scheme corresponds to Fig. 5d. The Ser1β of wild-type MacQ is located at the depth of the active-site cleft and is accessible from the solvent, while the corresponding Ser203 in SPΔ7 is buried inside due to the presence of uncleaved chain. (c) Active site cavities of the heterodimeric wild-type MacQ and monomeric (uncleaved) SPΔ7. The Ser1β/203 is represented as a stick model and coloured in red. C10/PAA binding pocket and the putative HSL/6-APA binding position are indicated by asterisk in red and black, respectively. The red arrow represents the putative releasing route of the HSL/6-APA from the pocket in the vicinity of Ser1β. The corresponding pocket is not formed in the structure of SPΔ7 due to the presence of uncleaved polypeptide chain.
Uncleaved S210A mutant is also active for AHLs
The recombinant S210A MacQ mutant was expressed and purified by the same procedure as the wild-type enzyme and SPΔ7. SDS-PAGE analysis of the purified sample showed that the S210A mutant is also present as an uncleaved form, similar to the SPΔ7 mutant (Fig. 5b). Ntn is also essential for self-cleaving maturation as well as enzymatic activity39,40,41,42,43. Thus, it is reasonable that the S210A mutant was uncleaved. Next, we assessed whether the S210A mutant is enzymatically active. The mutant could not degrade penicillin G, but was enzymatically active for various AHLs (Table 1, Supplementary Figs S3 and S4). Although the mechanisms of substrate binding and catalytic reaction of the S210A mutant remain unclear, it is evident that neither self-cleaving maturation nor Ser210 are essential for AHL acylase activity of MacQ, except for the C6-HSL, OC6-HSL and OC8-HSL short-chain AHLs.
Conclusions
MacQ is an enzyme belonging to the Ntn hydrolase superfamily. It catalyzes the amidohydrolase reaction for various ranges of AHLs and for β-lactam antibiotics. We determined the crystal structures of MacQ in ligand-free form and complexed with the C10 and PAA reaction products. MacQ forms a previously unreported 170-kDa capsule-like molecule consisting of a self-cleaved α-chain, SP, and β-chain. The SP showed increased conformational heterogeneity compared to the α- and β-chain, while the peptide-bond atoms of Val20SP-Gly21SP, which show the relatively low B-factor values in the structure of the SP, lie in the vicinity of the active-site pocket, likely accessible to the hydrophilic HSL/6-APA moiety of the bound substrate. In addition, both SP-shortened SPΔ7 and S210A mutants of MacQ result in the uncleaved, immature form of the protein, but retain amidohydrolyzing activity against AHLs, except for the short-chain AHLs, C6-HSL and OC6-HSL. These results suggest that MacQ is a structurally and enzymatically unique protein that does not conform to the common theory of autoproteolytic activation of Ntn-hydrolases and exemplifies the versatility of highly diversified Ntn hydrolase superfamily45. Future work will aim at investigating the underlying enzyme mechanism of MacQ by further mutational and structural analyses.
Methods
Recombinant protein expression and purification
The expression construct for wild-type MacQ was previously described3. The SPΔ7 and S210A mutants were created by the inverse PCR method51. Overexpression and purification of recombinant MacQ was conducted as previously described3. Briefly, overexpression of MacQ and its mutants with His-tags at both the N- and C-termini were induced by addition of 0.1 mM isopropyl-β-d-thiogalactopyranoside (IPTG) for 18 hr at 18 °C, using E. coli strain OrigamiTM 2 (DE3) (Novagen). After cell disruption by sonication, the supernatant was applied to a Ni affinity chromatography column (HIS-SELECT; Sigma-Aldrich), and the His-tagged sample was eluted according to the manufacturer’s instruction. The molecular weights of the subunit components of purified MacQ were confirmed with a Voyager DE Pro matrix-assisted laser desorption ionization time-of-flight (MALDI-TOF) mass spectrometer (Applied Biosystems). The α-chain of MacQ was separated by SDS-PAGE and transferred onto polyvinylidine difluoride (PVDF) membranes. The Coomassie Blue R250-stained bands on PVDF membranes were excised and applied to the N-terminal sequence analyses (Procise 492HT peptide sequencer; Applied Biosystems). The protein concentration was determined using Bradford protein assay kit (BioRad) with bovine serum albumin as a standard. The purified sample was dialyzed against the buffer consisting of 20 mM Tris-HCl pH 7.5, 50 mM NaCl, and 1 mM dithiothreitol (DTT) and was concentrated to 8 mg/mL for crystallization, using a centrifugal filter (molecular weight cut-off, 100 kDa; Millipore).
Analytical gel-filtration chromatography
The purified wild-type MacQ (cleaved form) and the S210A mutant (uncleaved form) were loaded on a Superose 6 10/300 GL column (GE Healthcare) equilibrated with 50 mM sodium phosphate pH 8.0 and 200 mM NaCl, at a flow rate of 0.4 mL/min. Ferritin (440 kDa), catalase (232 kDa), aldolase (158 kDa), ovalbumin (43 kDa) and ribonuclease A (13.7 kDa) were used as molecular-weight standards to calibrate the column. The molecular weight standard proteins were obtained from GE Healthcare.
Enzyme assay
The activities of MacQ and the generated mutants against various ranges of AHLs were examined by an AHL-inactivation assay using the green fluorescence protein (GFP)-based biosensor strains E. coli MT102 (harbouring plasmid pJBA132) and P. putida F117 (harbouring plasmid pKR-C12)52, 53, as previously described3. The amidohydrolyzing activity against penicillin G was examined by analyzing the reaction products by gas chromatography-mass spectrometry (GC-MS), as previously described3.
Crystallization
All crystallization screenings were carried out at 20 °C by the vapor-diffusion technique. Initial crystallization screening was performed using commercially available sparse matrix screening kits (Hampton Research and Emerald BioStructures), by the sitting-drop vapor-diffusion method in 96-well plates. Drops were formed by mixing 0.5 μL of sample and an equal volume of reservoir solution, and were equilibrated against 70 μL reservoir solution. Subsequent optimization of the hit conditions was performed using hanging-drop vapor-diffusion using 24-well plates. The crystals of wild-type MacQ were obtained using the reservoir solution consisting of 100 mM Tris-HCl pH 7.5–8.5, 0.1–0.2 M calcium acetate, and 14–18% polyethylene glycol (PEG) 3350. The SPΔ7 mutant was also crystallized using the reservoir solution consisting of 8% tacsimate pH 6.0 and 16% PEG 3350.
Structure determination and model refinement
Prior to X-ray diffraction, crystals were soaked into the cryoprotectant solution consisting of crystallization mother liquor supplemented with 20% glycerol, and were flash-cooled with a nitrogen gas stream at 100 K. For the data collection of the ligand complex, the crystals were briefly soaked in a freshly prepared solution consisting of 100 mM bis-Tris pH 5.5, 0.2 M calcium acetate, 18% PEG 3350, 20% glycerol, and saturated substrate (C10-HSL/penicillin G). X-ray diffraction data were collected with synchrotron radiation at the Photon Factory (Tsukuba, Japan). The raw data were processed using the HKL2000 program package54. The crystals of ligand-free and C10-bound MacQ belong to the space group P1 with unit-cell parameters approximately a = 86, b = 90, c = 123 Å, α = 103, β = 105, and γ = 106°. The crystals of wild-type MacQ with PAA complex and the SPΔ7 mutant belong to the space group P21 with unit-cell parameters approximately a = 102, b = 139, c = 122 Å, α = 90, β = 111 and γ = 90°. The structure of ligand-free MacQ was determined by the molecular replacement method with the program PHASER55 using the PvdQ heterodimer (PDB code, 2WYB) as a search model. After density modification by the program DM56, manual model building and iterative model fitting were performed with the graphics program COOT57. The structures of PAA-bound MacQ and the SPΔ7 mutant in space group P21 were determined by molecular replacement method using the ligand-free MacQ as a search model. Model refinement was performed using the program REFMAC558. The stereochemical quality of the final refined model was performed using the program PROCHECK59. Molecular drawings were prepared using the program PyMOL (http://www.pymol.org/). Crystallographic parameters and refinement statistics are summarized in Supplementary Table S1.
Data availability statements
The protein structures reported in this study have been deposited in the RCSB Protein Data Bank (http://www.rcsb.org) under accession codes 4YF9, 4YFA, 4YFB and 5C9I. Other data generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Miura, T. et al. Genome sequence of the multiple-β-lactam-antibiotic-resistant bacterium Acidovorax sp. strain MR-S7. Genome Announc. 1, e00412–13, doi:10.1128/genomeA.00412-13 (2014).
Kusada, H., Hanada, S., Kamagata, Y. & Kimura, N. The effects of N-acylhomoserine lactones, β-lactam antibiotics and adenosine on biofilm formation in the multi-β-lactam antibiotic-resistant bacterium Acidovorax sp. strain MR-S7. J. Biosci. Bioeng. 118, 14–19 (2014).
Kusada, H. et al. A novel quorum-quenching N-acylhomoserine lactone acylase from Acidovorax sp. strain MR-S7 mediates antibiotic resistance. Appl. Environ. Microbiol. 83, e00080–17 (2017).
Waters, C. M. & Bassler, B. L. Quorum sensing: cell-to-cell communication in bacteria. Annu. Rev. Cell Dev. Biol. 21, 319–346 (2005).
Dunlap, P. V. Quorum regulation of luminescence in Vibrio fischeri. J. Mol. Microbiol. Biotechnol. 1, 5–12 (1999).
Davies, D. G. et al. The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science 280, 295–298 (1998).
Fuqua, W. C., Winans, S. C. & Greenberg, E. P. Quorum sensing in bacteria: the LuxR-LuxI family of cell density-responsive transcriptional regulators. J. Bacteriol. 176, 269–275 (1994).
Hammer, B. K. & Bassler, B. L. Quorum sensing controls biofilm formation in Vibrio cholerae. Mol. Microbiol. 50, 101–104 (2003).
Tseng, B. S. et al. Quorum sensing influences Burkholderia thailandensis biofilm development and matrix production. J. Bacteriol. 198, 2643–2650 (2016).
Remuzgo-Martínez, S. et al. Biofilm formation and quorum-sensing-molecule production by clinical isolates of Serratia liquefaciens. Appl. Environ. Microbiol. 81, 3306–3315 (2015).
De Kievit, T. R. Quorum sensing in Pseudomonas aeruginosa biofilms. Environ. Microbiol. 11, 279–288 (2009).
Zhang, H. B., Wang, L. H. & Zhang, L. H. Genetic control of quorum-sensing signal turnover in Agrobacterium tumefaciens. Proc. Natl. Acad. Sci. USA 99, 4638–4643 (2002).
McClean, K. H. et al. Quorum sensing and Chromobacterium violaceum: exploitation of violacein production and inhibition for the detection of N-acylhomoserine lactones. Microbiology 143, 3703–3711 (1997).
Thomson, N. R. et al. Biosynthesis of carbapenem antibiotic and prodigiosin pigment in Serratia is under quorum sensing control. Mol. Microbiol. 36, 539–556 (2000).
Pierson, L. S. 3rd, Keppenne, V. D. & Wood, D. W. Phenazine antibiotic biosynthesis in Pseudomonas aureofaciens 30-84 is regulated by PhzR in response to cell density. J. Bacteriol. 176, 3966–3974 (1994).
Daniels, R., Vanderleyden, J. & Michiels, J. Quorum sensing and swarming migration in bacteria. FEMS Microbiol. Rev. 28, 261–289 (2004).
Jones, S. et al. The lux autoinducer regulates the production of exoenzyme virulence determinants in Erwinia carotovora and Pseudomonas aeruginosa. EMBO J. 12, 2477–2482 (1993).
Pirhonen, M., Flego, D., Heikinheimo, R. & Palva, E. T. A small diffusible signal molecule is responsible for the global control of virulence and exoenzyme production in the plant pathogen Erwinia carotovora. EMBO J. 12, 2467–2476 (1993).
Passador, L. et al. Expression of Pseudomonas aeruginosa virulence genes requires cell-to-cell communication. Science 260, 1127–1130 (1993).
Gao, R. et al. Genome-wide RNA sequencing analysis of quorum sensing-controlled regulons in the plant-associated Burkholderia glumae PG1 strain. Appl. Environ. Microbiol. 81, 7993–8007 (2015).
Cheng, F., Ma, A., Zhuang, X., He, X. & Zhuang, G. N-(3-oxo-hexanoyl)-homoserine lactone has a critical contribution to the quorum-sensing-dependent regulation in phytopathogen Pseudomonas syringae pv. tabaci 11528. FEMS Microbiol. Lett. 363, fnw265 (2016).
Dong, Y. H. & Zhang, L. H. Quorum sensing and quorum-quenching enzymes. J. Microbiol. 43, 101–109 (2005).
Chen, F. et al. Quorum quenching enzymes and their application in degrading signal molecules to block quorum sensing-dependent infection. Int. J. Mol. Sci. 14, 17477–17500 (2013).
Grandclément, C., Tannières, M., Moréra, S., Dessaux, Y. & Faure, D. Quorum quenching: role in nature and applied developments. FEMS Microbiol. Rev. 40, 86–116 (2016).
Tang, K. et al. MomL, a novel marine-derived N-acyl homoserine lactonase from Muricauda olearia. Appl. Environ. Microbiol. 81, 774–782 (2015).
Papaioannou, E. et al. Quorum-quenching acylase reduces the virulence of Pseudomonas aeruginosa in a Caenorhabditis elegans infection model. Antimicrob. Agents Chemother. 53, 4891–4897 (2009).
Lin, Y. H. et al. Acyl-homoserine lactone acylase from Ralstonia strain XJ12B represents a novel and potent class of quorum-quenching enzymes. Mol. Microbiol. 47, 849–860 (2003).
Huang, J. J., Petersen, A., Whiteley, M. & Leadbetter, J. R. Identification of QuiP, the product of gene PA1032, as the second acyl-homoserine lactone acylase of Pseudomonas aeruginosa PAO1. Appl. Environ. Microbiol. 72, 1190–1197 (2006).
Huang, J. J., Han, J. I., Zhang, L. H. & Leadbetter, J. R. Utilization of acyl-homoserine lactone quorum signals for growth by a soil pseudomonad and Pseudomonas aeruginosa PAO1. Appl. Environ. Microbiol. 69, 5941–5949 (2003).
Sio, C. F. et al. Quorum quenching by an N-acyl-homoserine lactone acylase from Pseudomonas aeruginosa PAO1. Infect. Immun. 74, 1673–1682 (2006).
Park, S. Y. et al. Identification of extracellular N-acylhomoserine lactone acylase from a Streptomyces sp. and its application to quorum quenching. Appl. Environ. Microbiol. 71, 2632–2641 (2005).
Romero, M. et al. Quorum quenching activity in Anabaena sp. PCC 7120: identification of AiiC, a novel AHL-acylase. FEMS Microbiol Lett 280, 73–80 (2008).
Morohoshi, T. et al. Identification and characterization of N-acylhomoserine lactone-acylase from the fish intestinal Shewanella sp. strain MIB015. Biosci. Biotechnol. Biochem. 72, 1887–1893 (2008).
Shepherd, R. W. & Lindow, S. E. Two dissimilar N-acyl-homoserine lactone acylases of Pseudomonas syringae influence colony and biofilm morphology. Appl. Environ. Microbiol. 75, 45–52 (2009).
Brannigan, J. A. et al. A protein catalytic framework with an N-terminal nucleophile is capable of self-activation. Nature 378, 416–419 (1995).
Bokhove, M., Nadal Jimenez, P., Quax, W. J. & Dijkstra, B. W. The quorum-quenching N-acyl homoserine lactone acylase PvdQ is an Ntn-hydrolase with an unusual substrate-binding pocket. Proc. Natl. Acad. Sci. USA 107, 686–691 (2010).
Kim, Y., Kim, S., Earnest, T. N. & Hol, W. G. Precursor structure of cephalosporin acylase: insights into autoproteolytic activation in a new N-terminal hydrolase family. J. Biol. Chem. 277, 2823–2829 (2002).
Kim, J. K. et al. Crystal structures of glutaryl 7-aminocephalosporanic acid acylase: insight into autoproteolytic activation. Biochemistry 42, 4084–4093 (2003).
Kim, J. K. et al. Insight into autoproteolytic activation from the structure of cephalosporin acylase: A protein with two proteolytic chemistries. Proc. Natl. Acad. Sci. USA 103, 1732–1737 (2006).
Kim, J. H., Krahn, J. M., Tomchick, D. R., Smith, J. L. & Zalkin, H. Structure and function of the glutamine phosphoribosylpyrophosphate amidotransferase glutamine site and communication with the phosphoribosylpyrophosphate site. J. Biol. Chem. 271, 15549–15557 (1996).
Seemuller, E., Lupas, A., Stock, D. & Lowe, J. Proteasome from Thermoplasma acidophilum: a threonine protease. Science 268, 579–582 (1995).
Bokhove, M. et al. Structures of an isopenicillin N converting Ntn-hydrolase reveal different catalytic roles for the active site residues of precursor and mature enzyme. Structure 18, 301–308 (2010).
Duggleby, H. J., Tolley, S. P., Hill, C. P. & Dodson, E. J. Penicillin acylase has a single-amino-acid catalytic center. Nature 373, 264–268 (1995).
Mukherji, R. et al. A new role for penicillin acylases: degradation of acyl homoserine lactone quorum sensing signals by Kluyvera citrophila penicillin G acylase. Enzyme Microb. Technol. 56, 1–7 (2014).
Oinonen, C. & Rouvinen, J. Structural comparison of Ntn-hydrolases. Protein Sci. 9, 2329–2337 (2000).
Holm, L. & Rosenström, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–549 (2010).
Drake, E. J. & Gulick, A. M. Structural characterization and high-throughput screening of inhibitors of PvdQ, an NTN hydrolase involved in pyoverdine synthesis. ACS Chem. Biol. 6, 1277–86 (2011).
Yin, J., Deng, Z., Zhao, G. & Huang, X. The N-terminal nucleophile serine of cephalosporin acylase executes the second autoproteolytic cleavage and acylpeptide hydrolysis. J. Biol. Chem. 286, 24476–24486 (2011).
Lee, Y. S. & Park, S. S. Two-step autocatalytic processing of the glutaryl 7-aminocephalosporanic acid acylase from Pseudomonas sp. strain GK16. J. Bacteriol. 180, 4576–4582 (1998).
Rajendhran, J. & Gunasekaran, P. Molecular cloning and characterization of thermostable β-lactam acylase with broad substrate specificity from Bacillus badius. J. Biosci. Bioeng. 103, 457–463 (2007).
Hemsley, A. et al. A simple method for site-directed mutagenesis using the polymerase chain reaction. Nucleic Acids Res. 17, 6545–6551 (1989).
Andersen, J. B. et al. gfp-Based N-acyl homoserine-lactone sensor systems for detection of bacterial communication. Appl. Environ. Microbiol. 67, 575–85 (2001).
Steidle, A. et al. Visualization of N-acylhomoserine lactone-mediated cell-cell communication between bacteria colonizing the tomato rhizosphere. Appl. Environ. Microbiol. 67, 5761–5770 (2001).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–667 (2007).
Cowtan, K. D. & Zhang, K. Y. J. Density modification for macromolecular phase improvement. Prog. Biophys. Mol. Biol. 72, 245–270 (1999).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. Sect. D 60, 2126–2132 (2004).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. Sect. D 67, 355–367 (2011).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Acknowledgements
We would like to thank Mizuho Muramatsu at AIST for her assistance with the HPLC and GC-MS analyses. We are also grateful to Dr. Nozomi Asano at Hokkaido University for MALDI TOF-MS analysis. The authors also acknowledge the technical staff at Photon Factory (PF) for their kind support in the X-ray diffraction studies. This work was supported by an internal grant from AIST. Synchrotron radiation experiments were conducted under the approval number 2012G579 at PF.
Author information
Authors and Affiliations
Contributions
Y.Y., S.H., Y.K., T.T., and N.K. designed and conducted the research. Y.Y., H.K., T.E., and N.K. performed the protein expression, purification, and crystallization. H.K., T.E., and N.K. performed the enzyme assay. Y.Y. performed the crystallographic analysis. Y.Y., H.K., and N.K. wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yasutake, Y., Kusada, H., Ebuchi, T. et al. Bifunctional quorum-quenching and antibiotic-acylase MacQ forms a 170-kDa capsule-shaped molecule containing spacer polypeptides. Sci Rep 7, 8946 (2017). https://doi.org/10.1038/s41598-017-09399-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-09399-4
This article is cited by
-
Selection and Molecular Response of AHL-lactonase (aiiA) Producing Bacillus sp. Under Penicillin G-induced Conditions
The Protein Journal (2023)
-
Efficient side-chain deacylation of polymyxin B1 in recombinant Streptomyces strains
Biotechnology Letters (2022)
-
The Komagataeibacter europaeus GqqA is the prototype of a novel bifunctional N-Acyl-homoserine lactone acylase with prephenate dehydratase activity
Scientific Reports (2021)
-
Enhancing Catalytic Efficiency of an Actinoplanes utahensis Echinocandin B Deacylase through Random Mutagenesis and Site-Directed Mutagenesis
Applied Biochemistry and Biotechnology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.