Abstract
To examine the pathogenic role of α-synuclein (αS) in Parkinson’s Disease, we have generated induced Pluripotent Stem Cell lines from early onset Parkinson’s Disease patients with SNCA A53T and SNCA Triplication mutations, and in this study have differentiated them to PSC-macrophages (pMac), which recapitulate many features of their brain-resident cousins, microglia. We show that SNCA Triplication pMac, but not A53T pMac, have significantly increased intracellular αS versus controls and release significantly more αS to the medium. SNCA Triplication pMac, but not A53T pMac, show significantly reduced phagocytosis capability and this can be phenocopied by adding monomeric αS to the cell culture medium of control pMac. Fibrillar αS is taken up by pMac by actin-rearrangement-dependent pathways, and monomeric αS by actin-independent pathways. Finally, pMac degrade αS and this can be arrested by blocking lysosomal and proteasomal pathways. Together, these results show that macrophages are capable of clearing αS, but that high levels of exogenous or endogenous αS compromise this ability, likely a vicious cycle scenario faced by microglia in Parkinson’s disease.
Similar content being viewed by others
Introduction
α-synuclein (αS) is a small, 14.5 kD, usually monomeric protein, that is highly expressed in neurons, where it can make up to 1 % of cytosolic protein, localising notably to presynaptic termini1. In Parkinson’s Disease (PD) it forms oligomers and fibrils, a major component of Lewy bodies and a hallmark of PD2. Overexpression or mutations in the SNCA gene, which encodes αS, can cause relatively early onset familial PD3,4,5. PD is characterised by loss of dopaminergic neurons in the Substantia nigra, therefore most studies focus on cell-autonomous pathological processes within dopaminergic neurons. However, non-neuronal cell processes likely also play a role in the progression of PD, with astrocytes and microglia being implicated through their expression of several key PD-related genes, including GBA, LRRK2 and SNCA 6.
Microglia are brain-resident macrophages, and are professional phagocytes, responsible for the homeostatic clearance of cellular debris, dying cells, incompetent synapses and aggregation-prone proteins. However, they can be provoked into a damaging, reactive state by inflammatory stimuli, triggering cytokine release (especially TNFα), potentially exacerbating neuronal damage and creating a vicious cycle of cytokine production and neuronal destruction (reviewed by refs 7 and 8). It is clearly important to examine the role of macrophages/microglia in clearing αS, and conversely, to understand whether the function of macrophages/microglia is affected by the presence of excess or mutant forms of the protein, as found in PD. Here, we examine this for the first time using human induced Pluripotent Stem Cells (iPSC) generated from PD patients with SNCA A53T mutation or the extremely rare SNCA Triplication. We differentiate the iPSC to adherent macrophages (pMac) via non-adherent macrophage precursors (pMacpre), following our previously published protocol9. These pMac represent yolk-sac-derived, tissue-resident macrophages10, 11 and therefore share the same ontogeny as microglia, which migrate into the fetal brain from the yolk-sac before the formation of the blood-brain barrier. Moreover, they can be skewed towards a microglial phenotype by co-culture with iPSC-neurons12. They are, therefore, a better model for microglia-related research questions than patient blood-derived monocytes, which derive from adult bone-marrow-hematopoiesis. pMac also overcome the limited availability of patient blood samples, and the complicating effects of medication and co-morbidities on immune cellular phenotypes.
We observe increased intracellular and extracellular αS in Triplication pMac, but not in A53T pMac. We report decreased phagocytosis by SNCA Triplication pMac, but not A53T pMac, and this decrease is phenocopied in healthy control pMac by addition of monomeric αS. pMac take up monomeric and fibrillar αS, degrading αS via lysosomal and proteasomal pathways. Macrophages therefore clear αS, but are easily intoxicated by higher than physiologically normal levels.
Results
SNCA Triplication, but not SNCA A53T mutation, causes elevated intracellular and extracellular αS protein levels in pMacpre and pMac
Multiple iPSC lines generated from 3 A53T patients, 1 Triplication patient and 4 normal control donors, all differentiated successfully to pMacpre and pMac (Table 1, Fig. 1A, Table S1, Figs S1–3). SNCA gene expression levels in pMac from control donors was not significantly different to levels in blood-derived monocytes and macrophages (Fig. S4). In controls and A53T pMac, αS positive puncta were found distributed throughout the cytosol by confocal microscopy, with a proportion of the signal apparently in the nucleus, as described previously for neurons13. Triplication pMac had many more αS puncta (Fig. 1B). Relative quantification of intracellular αS levels by flow cytometry, showed no significant difference in A53T pMacpre and pMac versus controls, but Triplication pMacpre and pMac had a significant (3-fold) increase in intracellular αS (Fig. 1C–E), and were significantly more granular as assessed by flow cytometry, whilst their size was not significantly different from controls (Figure S3B,C).
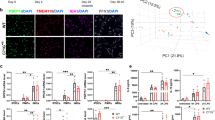
pMacpre and pMac expression of αS. (A) iPSC differentiation to macrophages (scale bar = 200 µm). (B) Intracellular staining for αS in pMac, representative z projected confocal images (scale bar = 20 µm). (C,D,E) Intracellular levels of αS by flow cytometry: (C) Representative αS FACs plots of pMacpre; (D) pMacpre αS levels (geometric mean fluorescence intensity, MFI); (E) pMac αS levels. (F) αS levels in 7-day supernatant from pMac. Values normalized to WT mean for each independent experiment. Also see Figs S1–3 Statistical analyses, one way ANOVA with Dunnett’s multiple comparisons test.
αS levels in 7-day tissue culture medium were not significantly different for A53T pMac (1001 pg/ml ± 261; mean ± SEM, n = 13) versus control (506 pg/ml ± 81; n = 12), but were significantly higher with Triplication (3473 pg/ml ± 572; n = 7) (Fig. 1F). αS levels in XVIVO15 medium alone were at the lower detection limit of the assay (58 pg/ml ± 1.8; n = 4). These αS levels secreted by pMac were comparable to those secreted by iPSC-dopaminergic neuronal cultures (previously published14, measured using the same assay platform), which ranged from ~100–400 pg/ml in 2-day supernatant from controls and ~200–800 pg/ml for lines from patients harbouring N370S mutations in the Parkinson’s disease-associated gene for glucocerebrosidase, GBA.
Cytokines in supernatants from unstimulated pMac were not significantly different from controls for Triplication or A53T for the majority of cytokines measured, including for the key proinflammatory cytokine TNFα (34-plex, Fig. 2 and Table S2). However, the chemokine CXCL1 (GRO-α), and the proinflammatory cytokines IL-18 and IL-22 were significantly constitutively upregulated in Triplication pMac versus controls (3, 2 and 1.5-fold respectively), suggesting a modest, specific dysregulation of cytokine production in macrophages overexpressing SNCA.
Cytokine and chemokine release by healthy control and SNCA mutant pMacs. Supernatants from pMac were assayed with Cytokine & Chemokine 34-Plex Human ProcartaPlex. Selected results are shown here, further cytokine results are shown in Table S2. Statistical analyses one way ANOVA with Dunnett’s multiple comparisons test.
Together, these results show that pMac express αS and release it to the medium, and that this is significantly increased in Triplication mutants, suggesting that their brain resident cousins, microglia, may contribute to accumulation of αS in the brain and possibly to the spreading of αS species.
Excess αS levels, but not A53T mutation, reduces phagocytosis in pMac
To investigate whether phagocytosis, endocytosis or pinocytosis pathways are perturbed in αS mutant pMac, particle uptake by macrophages was assayed with Alexafluor488-conjugated killed yeast particles (zymosan)9, 10 kD dextran or transferrin15, respectively. A53T pMac had a very modest increase in zymosan uptake (118% ± 7; n = 24). Triplication lines, in contrast, showed a highly significant reduction in zymosan uptake (51% ± 4; n = 17) versus controls (n = 33) (Fig. 3A–C). Meanwhile, only very modest or no differences were detected in dextran and transferrin uptake in SNCA mutant lines versus controls (Figure S5A–C). Therefore, excess endogenous αS compromises phagocytic but not endocytic or pinocytic ability in pMac.
High levels of αS reduce phagocytosis in pMac. Phagocytosis was measured by uptake of fluorescent zymosan by pMac. (A) Z-projection of confocal images (scale bar = 20 µm). (B) representative FACS plots of zymosan uptake (black line, untreated with cytochalasin D; gray, cytochalasin D-treated). (C) Quantification of (B) (3 independent experiments). (D) Incubation with monomeric αS for different lengths of time, then challenge with zymosan (one line). (E) Incubation with different concentrations of monomeric αS (24 hr), then challenge with zymosan. Values normalized to untreated mean for each of 3 independent experiments (C,E). Statistical analyses, one way ANOVA with Dunnett’s multiple comparisons test. Also see Figure S4A–C.
To explore whether increased levels of endogenously expressed αS was responsible for this phagocytic defect phenotype, we applied exogenous monomeric αS to the pMac. Phagocytosis of zymosan negatively correlated with monomeric αS dose and exposure time–2 µg/ml for 2 hrs showed an effect (Fig. 3D), and 10 µg/ml or 50 µg/ml for 24 hrs gave a highly significant reduction (Fig. 3E). Therefore, excess exogenous αS has the same detrimental effect as excess endogenously-derived αS on phagocytic ability of pMac, confirming that the effect seen with the Triplication lines is not due to another genetic defect in these lines.
pMac take up fibrillar and monomeric αS
We next examined whether and how αS is taken up by control pMac. Phagocytic uptake of fluorescently labelled αS fibrils was readily observable by time-lapse microscopy (Fig. 4A, Video S1). Adding exogenous monomeric αS increased intracellular levels of αS in a dose-dependent manner (Fig. 4B). 10 µg/ml for 2 hrs gave levels similar to SNCA Triplication, and thus, was chosen for further analyses. Uptake of monomeric αS was not affected by blocking actin polymerisation with cytochalasin D (Fig. 4C), implying uptake by mechanisms that are actin-rearrangement-independent, so not phagocytosis or macropinocytosis or actin-dependent endocytosis16. Further, uptake was not inhibited by Dynasore (Figure S5G), so not dynamin-dependent endocytosis, so uptake of monomeric αS could include direct traverse across the plasma membrane17. In contrast, uptake of fibrillar αS was significantly reduced by blocking actin polymerisation, so likely to employ phagocytosis (or possibly macropinocytosis or actin-dependent endocytosis for smaller aggregates) (Fig. 4D,E).
pMac take up fibrillar and monomeric αS. (A) 10 µg/ml Oregon Green 488-labelled fibrillar αS was added to pMac (imaged every 5 mins, 18 hrs). Frames capture one selected phagocytosis event (scale bar = 50 µm). (B) Intracellular αS levels following incubation with monomeric αS (FACs), normalized to mean endogenous αS levels in untreated pMac. (C) pMac incubated with monomeric αS, with or without Cytocholasin D (FACs as in (B)). (D) pMac incubated with fluorescent αS fibrils, with and without Cytocholasin D (FACs), MFI fluorescent αS divided by MFI medium only. (E) as per (D) but lower dose and shorter time. (F) TNFα levels in pMac supernatants measured 24 hrs after monomeric or fibrillar αS or LPS. Statistical analyses: (C,D,E) two-tailed t-test; (B,F) one way ANOVA with Dunnett’s multiple comparisons test. Also see Figure S4D–F.
Lipopolysaccharide (LPS) triggered strong TNFα release in pMac (3082 ± 34; n = 3), but monomeric αS and fibrillar αS did not (29 pg/ml ± 3; n = 3; 38 pg/ml ± 14 respectively) (Fig. 4F). LPS stimulation of SNCA mutant pMac triggered the same level of TNFα release as control pMac (Supplementary Figure 5D).
Together, this shows that pMac readily take up monomeric αS by actin-rearrangement-independent pathways, and fibrillar αS by actin-dependent pathways, and are not provoked under these baseline conditions into releasing TNFα in response to αS, whether exogenous, endogenous, monomeric, fibrillar or mutant. This also shows that the phagocytic defect shown in the presence of excess αS is not the result of autocrine or paracrine TNFα signalling.
pMac degrade αS by both lysosomal and proteasomal pathways
To investigate whether pMac were capable of degrading exogenous αS, pMac were incubated with monomeric αS, then collected, washed and replated onto fresh plates (to completely remove any remaining extracellular αS, which readily sticks to plastic). Endogenous αS staining localised to small puncta, mainly in the nucleus, with a few puncta in the cytosol, and little co-localisation of αS with the lysosome marker LAMP1 (Figs 1A, 5A). After 2 hrs of exposure to monomeric αS, larger αS puncta were visible in the cytosol of a subset of cells, partly co-localising with LAMP1 (Fig. 5B), as well as accumulating near the cell surface. After washing, and replating for 4 hrs, αS no longer localised to the cell surface but showed increased co-localisation with LAMP1 (Fig. 5C).
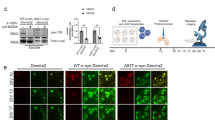
pMac degrade exogenous αS. (A) Immunocytochemistry of pMac: αS (green); lysosomal marker LAMP1 (red); nuclei (DAPI, blue; scale bar = 20 µm), region within white square is magnified below. (B) As (A) but pMac treated for 2 hrs with 10 µg/ml αS. (C) As (B), but pMac then washed and replated (4 hrs). (D) pMac incubated for 2 hrs with 10 µg/ml monomeric αS, washed and replated for the indicated number of hrs before assaying for intracellular αS by FACs; MFI relative to endogenous αS in untreated pMac. (E) Release of αS to supernatant of same experiment as (D). (F) As (D), with the addition of degradation pathway-specific drugs upon replating pMac after αS exposure and incubation for 8 hrs. (G) Pathway-specific drugs were added to cells (without exogenous αS exposure) for 2 and 8 hrs and assayed for intracellular accumulation of endogenous αS by FACs. (F,G) 1 line, 3 independent differentiations. Statistical analyses: (E) two-tailed t-test; (D,F,G) one way ANOVA with Dunnett’s multiple comparisons test.
The timecourse of depletion of monomeric αS from pMac was assessed by flow cytometry. αS levels in the pMac after αS-challenge and replating were ~4-fold higher than endogenous levels, indicating substantial uptake of αS, but by 22 hrs levels had dropped back to near-endogenous levels (Fig. 5D), implying that they had either expelled or degraded the excess.
To assess expulsion of αS by pMac, extracellular αS levels in supernatants from the above experiment were measured (Fig. 5E). 30 mins after replating, extracellular αS levels were similar to 4-day supernatant from untreated pMac (838 pg/ml ± 106; n = 3, and 882 pg/ml ± 188; n = 3). By 22h, levels of secreted αS nearly doubled (1407 pg/ml ± 313; n = 3), but were still about 7000 fold below input, and were below the levels measured in SNCA Triplication pMac supernatants.
To establish whether αS taken up by pMac is degraded within the cells, and if so by what mechanism, degradation pathway-specific drugs were added into the assay at the point of replating. Leupeptin (broad-spectrum protease inhibitor), MG132 (26S proteasome inhibitor) and Bafilomycin A1 (prevents vesicle acidification, thereby inhibiting lysosomal function) were each able to inhibit degradation of αS, as measured after 8 hrs, by 2–3-fold (Fig. 5F). Applying these pathway inhibitors to pMac not treated with exogenous αS did not significantly alter endogenous αS levels (Fig. 5G).
Together, these results show that endogenous αS is relatively stable over the timeframe of these assays, and that pMac can expel exogenously acquired αS and also degrade it by both lysosomal and proteasomal pathways.
Discussion
This study demonstrates that αS can compromise the ability of professional phagocytes to conduct their normal homeostatic phagocytic functions, which includes clearance of αS, and likely contributes to the build-up of αS in PD patients. Our results are the first to use iPSC-derived macrophages from PD patients harbouring SNCA mutations and controls to study αS, enabling expression at the correct gene dosage in this highly relevant, authentic human cell type.
SNCA Triplication is extremely rare4. The availability of iPSC from these patients has so far been limited to samples from two individuals, a 48 year old male18,19,20 and a 55 year old female21, 22, the latter being the same individual from which we have derived the iPSC used in this study, and which are the first SNCA Triplication iPSC to be derived with non-integrating reprogramming vectors. Because of this rarity, we were not able to extend our results to iPSC-macrophages from additional SNCA Triplication patients. However, the observed phenotype was reproduced across all three clones from this patient. Moreover, we have shown for the first time that exogenous αS phenocopies the phagocytic defect displayed by SNCA Triplication pMac, indicating that excess αS affects phagocytosis, whether endogenous or exogenously derived.
Our results showing the negative impact of excess endogenous αS on phagocytosis concur with and extend the results of others using mice over-expressing human SNCA from a bacterial artificial chromosome23. That system relies on expression of human αS on a mouse background, and αS levels are not as precisely comparable to the SNCA Triplication patient as in our pMac system. Our results also concur with and extend results using SNCA triplication patient monocytes23. pMac, being entirely in vitro, are not exposed to any confounding or compounding in vivo or ex vivo factors, such as chronic inflammation, so results obtained with pMac likely reflect a primary defect, rather than a secondary effect. Patient monocytes may be compromised by the effects of PD drug regimens and the patients’ physiological status, whereas pMac have never been exposed to patient cytokines, drugs, etc. Consistent with this, pMac do not produce TNFα in response to monomeric or fibrillar αS or SNCA mutation, presumably because they have not been exposed to any priming cytokines, whereas increased TNFα shown to be released from mouse microglia overexpressing SNCA is likely due to chronic activation in the in vivo system23, 24.
In our system A53T SNCA did not lead to a significant increase in αS levels (Fig. 1B–E), and did not affect phagocytosis (Fig. 2A–C), in contrast to the increased αS and decreased phagocytosis seen with SNCA Triplication. This was perhaps surprising, given that in neurons A53T is associated with accumulation of αS, but our results suggest that pMac, which are professional clearers of unwanted material, are better able to process mutant, misfolded A53T αS than neurons and do not accumulate it so readily.
αS uptake by other cell types has been described previously, including free passage through the plasma membrane17, micropinocytosis25 and dynamin-dependent endocytosis26. Our results show for the first time that actin dynamics are involved in the uptake of fibrillar but not monomeric forms of αS in pMac, and time-lapse video of fibrillary uptake supports this. In Alzheimer’s Disease, microglia have been shown to congregate around amyloid beta plaques, yet are not able to phagocytose the plaques and instead become chronically activated27. We have previously shown that dopaminergic neurons harbouring PD-associated GBA mutations secrete increased levels of αS versus controls14. We postulate that macrophages and microglia take up αS as part of their homeostatic, non-inflammatory portfolio, but are similarly rendered less phagocytically competent by the locally very high levels of αS found in PD brains.
We explored whether pMac degrade and/or expel αS, and by what mechanisms, because if they expel substantial amounts, their microglial cousins could be potential contributors to the spread of αS in the brain. Microtubule-associated protein Tau, which forms tangles in Alzheimer’s Disease, is capable of being spread by release from murine microglia, and subsequent uptake by nearby neurons28. We found that pMac expelled αS both constitutively and following uptake, but lysosomal and proteasomal degradation of αS was also evident in pMac. In other cell types, αS has been shown to be ubiquitinated by Nedd4 before degradation by these routes29, 30. Since pMac, and by extension microglia, are capable of uptake, excretion and degradation of αS, manipulating the balance could be therapeutically useful. The use of anti-αS antibodies to promote αS uptake by microglia has already been investigated in mice31, 32, and humanised versions are starting to be trialled in PD patients. The pMac used here, as close cousins of microglia, offer a simple monoculture system to investigate the role of genes involved in neurodegeneration in the myeloid lineage. pMicroglia, developed from pMac recently by co-culture of pMac with iPSC-neurons by ourselves12 and by others along broadly similar routes33,34,35,36, offer an even more physiologically relevant in vitro system and will be used to extend the observations made here in pMac. These models will enable us to understand and be able to fine-tune the balance between successful phagocytosis/destruction of opsonised αS by microglia, and destructive microglial over-activation and/or propagation of αS species.
Methods
Reagents were from ThermoFisher (Invitrogen) unless stated otherwise.
Reprogramming of patient fibroblasts to iPSC and differentiation to macrophages
As previously described9, 14, see Supplemental Information for methodological details.
All lines were derived from dermal fibroblasts from healthy donors or Parkinson’s disease patients, through StemBANCC (SF180, SF828, SF829, SF830, SF831), or the Oxford Parkinson’s Disease Centre (SF840, SF841, SF856, SF854, AH016): participants were recruited to this study having given signed informed consent, which included derivation of hiPSC lines from skin biopsies (Ethics Committee that specifically approved this part of the study: for control donors, National Health Service, Health Research Authority, NRES Committee South Central, Berkshire, UK, REC 10/H0505/71, and for SNCA patients REC 07/H0720/161); all experiments were performed in accordance with UK guidelines and regulations and as set out in the REC.
Determination of αS levels
pMacpre/pMac were fixed (4% PFA, 10 mins), washed (PBS), blocked/permeabilized (flow cytometry buffer: PBS, 10 µg/mL human IgG [Sigma], 1% FCS [Hyclone], 0.01% sodium azide, 0.1% Saponin [Sigma], 30 mins), incubated with rabbit anti-αS antibody (MJFR1, Abcam) or rabbit IgG1 (ab27478, Abcam) (1:250, 45 mins), washed x3 (flow cytometry buffer), incubated with donkey anti-rabbit IgG-Alexa647 (1:500), washed/resuspended (PBS) and assayed by flow cytometer (Calibur, BD) to obtain intracellular αS levels. To compare results of replicate experiments carried out on different occasions, data was normalized to the mean of healthy control lines of the corresponding experiment.
7 day supernatant from pMac cultures was centrifuged (5 mins, 400 g) to remove cell debris, transferred to a new tube and stored at −80 °C. Supernatants were applied undiluted to a human αS detection kit (MesoScale Discovery, MSD).
Particle uptake
Uptake of dead yeast particles (zymosan) was assayed as described previously for pMac9, 37. Briefly, fluorescent zymosan particles were applied (2 particles/cell, 30 mins, 37 °C) in pMac differentiation medium (as a negative control for zymosan uptake, pMac were pre-treated (one hr) with 10 µM Cytochalasin D to inhibit actin polymerisation), or fluorescent dextran (D-22910) or fluorescent transferrin were applied at 50 µg/ml, fluorescence of all extracellular particles blocked (0.025% Trypan blue [Sigma] in PBS), washed (PBS), released (TrypLE Express, 10 mins), collected gently with a cell scraper, fixed (4% PFA) and analysed by flow cytometer (Calibur, BD).
Recombinant αS
Recombinant human monomeric was purified as described previously38, 39. Endotoxin in αS preparations was removed using high capacity endotoxin spin columns (Pierce 88276) prior to labelling and/or aggregation into fibrils. A limulus amebocyte lysate (LAL) chromogenic kit (Pierce 88282) was used to measure endotoxin levels before and after treatment. Preparations containing endotoxin levels were >1 EU per mg of αS were used in experiments at a final dilution ≤10 µg/ml (i.e., ≤0.01 EU/ml). Monomeric αS was labeled with Oregon Green 488 succinimidyl-ester (Invitrogen) as per manufacturer’s instructions. Assembled fibrils contained a molar ratio of ≤1:10 (dye:protein). For αS uptake and degradation experiments, pMac were differentiated in ultra-low attachment surface plates (Corning) to prevent αS attachment, and to allow resuspension of pMac without the use of a cell scraper. Before and during αS stimulation pMac were treated with Cytochalasin D (Sigma, 10 µM, to inhibit actin polymerisation) or Dynasore (Cayman Chemical, 80 µM, inhibits dynamin). After αS incubation, cells were lifted by pipetting, washed 3x with PBS and either used directly for determination of αS levels or replated in fresh medium to assay αS degradation. When replating, the following drugs were used to block specific degradation pathways, as described previously40, 41: Leupeptin (Alfa Aesar, 100 µM, broad protease inhibitor); MG132 (Sigma, 25 µM, blocks 26S proteasome); Bafilomycin A1 (InvivoGen, 400 nM, inhibits vacuolar-type H+ ATPases, preventing vesicle acidification and lysosomal function).
Statistical analysis
GraphPad Prism was used for statistical analysis. One way ANOVA and Dunnett’s multiple comparisons test was used for all analyses, except where indicated (for single comparisons), where Student’s 2-tailed t-test was used. n.s. = not significant, * = p < 0.05, ** = p < 0.01, *** = p < 0.001, **** = p < 0.0001. Numbers given in parentheses in the text are in the form (mean ± SEM, n).
Accession numbers
SNP datasets and Illumina HT12v4 expression array datasets for previously unpublished iPSC lines have been deposited in GEO, under Accession number GSE89886.
References
McLean, P. J., Kawamata, H., Ribich, S. & Hyman, B. T. Membrane association and protein conformation of alpha-synuclein in intact neurons. Effect of Parkinson’s disease-linked mutations. The Journal of biological chemistry 275, 8812–8816 (2000).
Spillantini, M. G. et al. Alpha-synuclein in Lewy bodies. Nature 388, 839–840, doi:10.1038/42166 (1997).
Polymeropoulos, M. H. et al. Mapping of a gene for Parkinson’s disease to chromosome 4q21-q23. Science 274, 1197–1199 (1996).
Singleton, A. B. et al. alpha-Synuclein locus triplication causes Parkinson’s disease. Science 302, 841, doi:10.1126/science.1090278 (2003).
Ibanez, P. et al. Causal relation between alpha-synuclein gene duplication and familial Parkinson’s disease. Lancet 364, 1169–1171, doi:10.1016/S0140-6736(04)17104-3 (2004).
Dzamko, N., Geczy, C. L. & Halliday, G. M. Inflammation is genetically implicated in Parkinson’s disease. Neuroscience 302, 89–102, doi:10.1016/j.neuroscience.2014.10.028 (2015).
Amor, S., Puentes, F., Baker, D. & van der Valk, P. Inflammation in neurodegenerative diseases. Immunology 129, 154–169, doi:10.1111/j.1365-2567.2009.03225.x (2010).
Hirsch, E. C., Vyas, S. & Hunot, S. Neuroinflammation in Parkinson’s disease. Parkinsonism & related disorders 18(Suppl 1), S210–212, doi:10.1016/S1353-8020(11)70065-7 (2012).
van Wilgenburg, B., Browne, C., Vowles, J. & Cowley, S. A. Efficient, long term production of monocyte-derived macrophages from human pluripotent stem cells under partly-defined and fully-defined conditions. PloS one 8, e71098, doi:10.1371/journal.pone.0071098 (2013).
Buchrieser, J., James, W. & Moore, M. D. Human Induced Pluripotent Stem Cell-Derived Macrophages Share Ontogeny with MYB-Independent Tissue-Resident Macrophages. Stem cell reports 8, 334–345, doi:10.1016/j.stemcr.2016.12.020 (2017).
Vanhee, S. et al. In vitro human embryonic stem cell hematopoiesis mimics MYB-independent yolk sac hematopoiesis. Haematologica 100, 157–166, doi:10.3324/haematol.2014.112144 (2015).
Haenseler, W. et al. A Highly Efficient Human Pluripotent Stem Cell Microglia Model Displays a Neuronal-Co-culture-Specific Expression Profile and Inflammatory Response. Stem cell reports 8, 1727–1742 (2017).
Yu, S. et al. Extensive nuclear localization of alpha-synuclein in normal rat brain neurons revealed by a novel monoclonal antibody. Neuroscience 145, 539–555, doi:10.1016/j.neuroscience.2006.12.028 (2007).
Fernandes, H. J. et al. ER Stress and Autophagic Perturbations Lead to Elevated Extracellular alpha-Synuclein in GBA-N370S Parkinson’s iPSC-Derived Dopamine Neurons. Stem cell reports 6, 342–356, doi:10.1016/j.stemcr.2016.01.013 (2016).
Carter, G. C., Bernstone, L., Baskaran, D. & James, W. HIV-1 infects macrophages by exploiting an endocytic route dependent on dynamin, Rac1 and Pak1. Virology 409, 234–250, doi:10.1016/j.virol.2010.10.018 (2011).
Ozdener, G., Bais, M. & Trackman, P. Determination of cell uptake pathways for tumor inhibitor lysyl oxidase propeptide. Molecular Oncology 10, 1–23, doi:10.1016/j.molonc.2015.07.005 (2016).
Ahn, K. J., Paik, S. R., Chung, K. C. & Kim, J. Amino acid sequence motifs and mechanistic features of the membrane translocation of alpha-synuclein. Journal of neurochemistry 97, 265–279, doi:10.1111/j.1471-4159.2006.03731.x (2006).
Byers, B. et al. SNCA Triplication Parkinson’s Patient’s iPSC-derived DA Neurons Accumulate α-Synuclein and Are Susceptible to Oxidative Stress. PLOS ONE 6, e26159, doi:10.1371/journal.pone.0026159 (2011).
Flierl, A. et al. Higher Vulnerability and Stress Sensitivity of Neuronal Precursor Cells Carrying an Alpha-Synuclein Gene Triplication. PLOS ONE 9, e112413, doi:10.1371/journal.pone.0112413 (2014).
Oliveira, L. M. et al. Elevated α-synuclein caused by SNCA gene triplication impairs neuronal differentiation and maturation in Parkinson’s patient-derived induced pluripotent stem cells. Cell death & disease 6.
Devine, M. et al. Parkinson’s disease induced pluripotent stem cells with triplication of the α-synuclein locus. Nature communications 2, 440, doi:10.1038/ncomms1453 (2011).
Mazzulli, J., Zunke, F., Isacson, O., Studer, L. & Krainc, D. α-Synuclein-induced lysosomal dysfunction occurs through disruptions in protein trafficking in human midbrain synucleinopathy models. Proceedings of the National Academy of Sciences of the United States of America 113, 1931–1936, doi:10.1073/pnas.1520335113 (2016).
Gardai, S. J. et al. Elevated alpha-synuclein impairs innate immune cell function and provides a potential peripheral biomarker for Parkinson’s disease. PloS one 8, e71634, doi:10.1371/journal.pone.0071634 (2013).
Thome, A. D., Harms, A. S., Volpicelli-Daley, L. A. & Standaert, D. G. microRNA-155 Regulates Alpha-Synuclein-Induced Inflammatory Responses in Models of Parkinson Disease. The Journal of neuroscience: the official journal of the Society for Neuroscience 36, 2383–2390, doi:10.1523/JNEUROSCI.3900-15.2016 (2016).
Lee, H. J. et al. Assembly-dependent endocytosis and clearance of extracellular alpha-synuclein. The international journal of biochemistry & cell biology 40, 1835–1849, doi:10.1016/j.biocel.2008.01.017 (2008).
Chai, Y. J. et al. The secreted oligomeric form of alpha-synuclein affects multiple steps of membrane trafficking. FEBS letters 587, 452–459, doi:10.1016/j.febslet.2013.01.008 (2013).
Krabbe, G. et al. Functional impairment of microglia coincides with Beta-amyloid deposition in mice with Alzheimer-like pathology. PloS one 8, e60921, doi:10.1371/journal.pone.0060921 (2013).
Asai, H. et al. Depletion of microglia and inhibition of exosome synthesis halt tau propagation. Nature neuroscience 18, 1584–1593, doi:10.1038/nn.4132 (2015).
Tofaris, G. K. et al. Ubiquitin ligase Nedd4 promotes alpha-synuclein degradation by the endosomal-lysosomal pathway. Proceedings of the National Academy of Sciences of the United States of America 108, 17004–17009, doi:10.1073/pnas.1109356108 (2011).
Sugeno, N. et al. Lys-63-linked ubiquitination by E3 ubiquitin ligase Nedd4-1 facilitates endosomal sequestration of internalized alpha-synuclein. The Journal of biological chemistry 289, 18137–18151, doi:10.1074/jbc.M113.529461 (2014).
Bae, E. J. et al. Antibody-aided clearance of extracellular alpha-synuclein prevents cell-to-cell aggregate transmission. The Journal of neuroscience: the official journal of the Society for Neuroscience 32, 13454–13469, doi:10.1523/JNEUROSCI.1292-12.2012 (2012).
Games, D. et al. Reducing C-terminal-truncated alpha-synuclein by immunotherapy attenuates neurodegeneration and propagation in Parkinson’s disease-like models. The Journal of neuroscience: the official journal of the Society for Neuroscience 34, 9441–9454, doi:10.1523/JNEUROSCI.5314-13.2014 (2014).
Muffat, J. et al. Efficient derivation of microglia-like cells from human pluripotent stem cells. Nature medicine (2016).
Pandya, H. et al. Differentiation of human and murine induced pluripotent stem cells to microglia-like cells. Nature neuroscience (2017).
Abud, E. et al. iPSC-Derived Human Microglia-like Cells to Study Neurological Diseases. Neuron 94 (2017).
Douvaras, P. et al. Directed Differentiation of Human Pluripotent Stem Cells to Microglia. Stem cell reports 8, 1516–1524 (2017).
Jiang, Y. et al. Derivation and functional analysis of patient-specific induced pluripotent stem cells as an in vitro model of chronic granulomatous disease. Stem cells 30, 599–611, doi:10.1002/stem.1053 (2012).
Luk, K. C. et al. Exogenous alpha-synuclein fibrils seed the formation of Lewy body-like intracellular inclusions in cultured cells. Proceedings of the National Academy of Sciences of the United States of America 106, 20051–20056, doi:10.1073/pnas.0908005106 (2009).
Luk, K., Hyde, E., Trojanowski, J. & Lee, V. Sensitive fluorescence polarization technique for rapid screening of alpha-synuclein oligomerization/fibrillization inhibitors. Biochemistry 46, 12522–12529, doi:10.1021/bi701128c (2007).
Raposo, R. A. et al. Protein kinase C and NF-kappaB-dependent CD4 downregulation in macrophages induced by T cell-derived soluble factors: consequences for HIV-1 infection. Journal of immunology 187, 748–759, doi:10.4049/jimmunol.1003678 (2011).
Luhr, K. M. et al. Scrapie protein degradation by cysteine proteases in CD11c+ dendritic cells and GT1-1 neuronal cells. Journal of virology 78, 4776–4782 (2004).
Dafinca, R. et al. C9orf72 Hexanucleotide Expansions Are Associated with Altered Endoplasmic Reticulum Calcium Homeostasis and Stress Granule Formation in Induced Pluripotent Stem Cell-Derived Neurons from Patients with Amyotrophic Lateral Sclerosis and Frontotemporal Dementia. Stem cells 34, 2063–2078, doi:10.1002/stem.2388 (2016).
Sandor, C. et al. Transcriptomic profiling of purified patient-derived dopamine neurons identifies convergent perturbations and therapeutics for Parkinson’s disease. Human molecular genetics. doi:10.1093/hmg/ddw412 (2017).
Devine, M. J. et al. Parkinson’s disease induced pluripotent stem cells with triplication of the alpha-synuclein locus. Nature communications 2, 440, doi:10.1038/ncomms1453 (2011).
Acknowledgements
Financial support: The Wellcome Trust WTISSF121302 and the Oxford Martin School LC0910-004 (James Martin Stem Cell Facility Oxford, W.H., S.A.C.); the MRC Dementias Platform UK Stem Cell Network Capital Equipment MC_EX_MR/N50192X/1, Partnership MR/N013255/1 (W.H., S.A.C., R.W.-M.) and Momentum MC_PC_16034 (W.H., S.A.C.) Awards; the Swiss National Foundation Early Postdoc Mobility, 148607, and ARUK Oxford pilot grant (W.H.). The research leading to these results has received support from the Innovative Medicines Initiative Joint Undertaking under grant agreement n° 115439, resources of which are composed of financial contribution from the European Union’s Seventh Framework Programme (FP7/2007–2013) and EFPIA companies’ in kind contribution. This publication reflects only the author’s views and neither the IMI JU nor EFPIA nor the European Commission are liable for any use that may be made of the information contained therein. J.V. is supported by the Monument Trust Discovery Award from Parkinson’s UK. We thank the High-Throughput Genomics Group at the Wellcome Trust Centre for Human Genetics, Oxford (Funded by Wellcome Trust grant reference 090532/Z/09/Z and MRC Hub grant G0900747 91070) for the generation of Illumina genotyping and transcriptome data. S.W. is supported by an Alzheimer’s Research UK Senior Fellowship and the NIHR Queen Square Dementia Biomedical Research Unit. K.G. is an employee of the Federal Government, USA.
Author information
Authors and Affiliations
Contributions
Conceptualization, S.A.C., W.S.J. and W.H.; Methodology, W.H., S.A.C. and H.L.; Investigation, W.H., F.Z. and H.L.; Formal Analysis, W.H.; Writing, W.H. and S.A.C.; Original Draft Writing, W.H., Review & Editing, S.A.C., R.W.-M. and W.S.J.; Funding Acquisition, S.A.C., R.W.-M., W.H. and W.S.J.; Resources, K.C.L., S.W., J.V., G.D., H.H., K.G. and F.R.; Supervision S.A.C., R.W.-M. and W.S.J.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Haenseler, W., Zambon, F., Lee, H. et al. Excess α-synuclein compromises phagocytosis in iPSC-derived macrophages. Sci Rep 7, 9003 (2017). https://doi.org/10.1038/s41598-017-09362-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-09362-3
This article is cited by
-
Tau depletion in human neurons mitigates Aβ-driven toxicity
Molecular Psychiatry (2024)
-
Lysosomal acidification dysfunction in microglia: an emerging pathogenic mechanism of neuroinflammation and neurodegeneration
Journal of Neuroinflammation (2023)
-
CRISPR/Cas9 assisted stem cell therapy in Parkinson's disease
Biomaterials Research (2023)
-
Characterization of the pathogenic α-Synuclein Variant V15A in Parkinson´s disease
npj Parkinson's Disease (2023)
-
Toll-like receptors and NLRP3 inflammasome-dependent pathways in Parkinson’s disease: mechanisms and therapeutic implications
Journal of Neurology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.