Abstract
2,4,6-trinitrotoluene (TNT) has been reported to cause numerous adverse effects. However, the detailed molecular mechanisms underlying TNT-induced liver toxicity need to be elucidated. In this study, we used HepG2 (p53wt) and Hep3B (p53null) cell lines to investigate the cytotoxic effects of TNT. At first, we found that TNT significantly decreased cell viability and induced DNA damage. Thereafter, through transcriptomic analysis, we observed that the diverse biological functions affected included mitochondrial dysfunction and endoplasmic reticulum (ER) stress. Mitochondrial dysfunction was evidenced by the loss of mitochondrial membrane potential, increased expression of cleaved-caspase-9&-3 and increased caspase-3/7 activity, indicating that apoptosis had occurred. In addition, the expressions of some ER stress-related proteins had increased. Next, we investigated the role of reactive oxygen species (ROS) in TNT-induced cellular toxicity. The levels of DNA damage, mitochondrial dysfunction, ER stress and apoptosis were alleviated when the cells were pretreated with N-acetyl-cysteine (NAC). These results indicated that TNT caused the ROS dependent apoptosis via ER stress and mitochondrial dysfunction. Finally, the cells transfected with CHOP siRNA significantly reversed the TNT-induced apoptosis, which indicated that ER stress led to apoptosis. Overall, we examined TNT-induced apoptosis via ROS dependent mitochondrial dysfunction and ER stress in HepG2 and Hep3B cells.
Similar content being viewed by others
Introduction
2,4,6-trinitrotoluene (TNT) has been commonly used as an explosive throughout the world, and it is one of the most serious environmental contaminants in military sites where munitions were manufactured1. TNT has been shown to be highly toxic, mutagenic, and carcinogenic in some bacterial and animal tests2,3,4,5. In addition, TNT could lead to numerous adverse effects, including upper respiratory problems, gastrointestinal complaints, anemia, liver function abnormalities, and aplastic anemia6, 7. In China, a survey study of male workers from 8 Chinese military factories who were exposed to TNT for more than a year confirmed that TNT could increase the relative risk of 80%, especially liver cancer8. More recently, multiple studies have indicated that TNT-induced stress, including endoplasmic reticulum (ER) stress and oxidative stress, may lead to liver injury7, 9. However, the molecular mechanisms involved in stress-induced hepatotoxicity are still unclear, although some studies have shown that ER stress and the apoptotic pathway are involved in TNT-induced hepatic toxicity7, 9, 10. Noticeably, the role of reactive oxygen species (ROS) in mediating ER and mitochondrial stress needs to be fully investigated.
ROS profoundly impact a number of cellular responses such as DNA damage, cell cycle progression, and apoptotic cell death11,12,13. In eukaryotic cells, the mitochondrial electron transport is the main source of ROS during normal metabolism12. Excessive or sustained ROS can cause damage to proteins and DNA via diverse mechanisms, thereby activating or inhibiting the related signaling pathway14.
The ER plays an important role in chemical toxicant-induced apoptosis15. The ER is an organelle that maintains intracellular calcium homeostasis, protein synthesis, post-translational modification and proper protein folding16. A disturbance of ER Ca2+ homeostasis or the protein process can lead to ER stress, which in turn induces the production of ROS in the ER and mitochondria17. High ROS generation within mitochondria induces the opening of the mitochondrial permeability transition pore (mPTP)17. Subsequently, a number of proteins that regulate apoptosis become involved, contributing to cell death.
To determine the possibility of ROS involvement in apoptosis as described above, we detected ROS generation in cells by activating the mitochondrial and ER stress pathways. Further investigations into the links between ROS increase, DNA damage and apoptosis induced by ROS were also conducted. In this study, we investigated the detailed mechanisms underlying TNT toxicity in HepG2 cells. Furthermore, we investigated the effects of TNT toxicity in Hep3B cells and aimed to understand if the mechanisms of TNT toxicity in different human hepatoma cells were different based on the presence of p53 in HepG2 cells but not in Hep3B cells.
Results
Effects of TNT on cell viability, DNA damage and the activation of caspase-3/7 in HepG2 and Hep3B cells
To investigate the extent of the effect of TNT on HepG2 and Hep3B cells, we performed dose response or time course analysis of TNT-mediated proliferation inhibition, DNA damage and the activation of caspase-3/7 in HepG2 and Hep3B cells. We performed a CCK-8 assay to detect the level of cytotoxicity in TNT treated cells. The results show that TNT exhibited the cytotoxicity against the growth of cells in terms of dose response and time. Cell viability was reduced to about 50% after the cells were treated with TNT (80 μM) for 24 h in HepG2, and treated with TNT (60 μM) for 24 h in Hep3B (Fig. 1A).
TNT-induced cytotoxicity, DNA damage and apoptosis in HepG2 cells in a dose- and-time dependent manner. (A) HepG2 and Hep3B cells were treated with TNT (0–100 μM) for 24 h and 48 h, and cell viability was analyzed by CCK-8 assay. (B) DNA damage was determined by comet assay in HepG2 cells treated with TNT for 24 h. (C) Effects of TNT on γ-H2AX protein expressions in HepG2 and Hep3B cells. The cells were exposed to TNT for 24 h. The protein expression of γ-H2AX was analyzed by Western blot. (D) Effects of TNT on p53 protein expressions in HepG2 cells. The cells were exposed to TNT for 24 h. The protein expression of p53 was analyzed by Western blot. (E) Effects of TNT on protein expression of pro-caspase-9 and -3 and cleaved-caspase-9 and -3 in HepG2 cells. The cells were exposed to TNT (0, 30, and 80 μM) for 24 h. The protein expression was analyzed by Western blot. (F) The caspase-3/7 activity was assayed after the HepG2 and Hep3B cells were treated with TNT and with or without NAC for 24 h via microplate reader. The data are presented as mean ± SD for four independent experiments with triplicate determinations. *P < 0.05; **P < 0.01 as compared with control.
To evaluate the ability of TNT to trigger genotoxic damage in hepatocytes, the HepG2 cells were first treated with different concentrations of TNT (in the range of 0 to 30 μM) for 24 h, and they were subsequently analyzed using an alkaline comet assay (Fig. 1B); the presence of a DNA-forming tail-like structure in a concentration-dependent manner was observed in cells treated with TNT. Second, HepG2 and Hep3B cells were treated with different concentrations of TNT (HepG2: 0–30 μM; Hep3B: 0–50 μM) for 24 h to determine the induction of protein related to the DNA damage marker: γ-H2AX18. We found that TNT induces γ-H2AX in a concentration-dependent manner in HepG2 and Hep3B cells (Fig. 1C). Third, we investigated the effect of TNT on p53 protein levels. We found that TNT significantly increases the expression of p53 as compared to HepG2 cells without added TNT (Fig. 1D).
As caspase cascade activation is a key event in apoptosis pathways, we found that TNT (30 μM and 80 μM) treatment with HepG2 reduced the expression of pro-caspase-3 and -9 and increased cleaved caspase-3 and -9, respectively, indicating that the treated cells induced apoptosis (Fig. 1E). This assumption was further substantiated by caspase-3/7 activity, which is an indicator of cell death (Fig. 1F). Our results demonstrate that TNT exposure induces significant DNA damage and apoptosis in HepG2 and Hep3B cells.
Transcriptome analysis of TNT-induced impaired biological function and/or pathways
In order to systematically understand the transcriptional response from HepG2 cells treated with 30 μM and 80 μM TNT for 24 h, we first used RNA-seq, a high throughput method, to detect the differential genes (> or <2-fold, P < 0.05) in cells before and after 30 μM and 80 μM TNT treatments. These differential genes detected in 30 μM or 80 μM TNT are listed in Supplementary Tables S1 and S2. To validate the RNA-seq results in this study, we selected several differential expressed genes (DEGs) of particular relevance to oxidative phosphorylation/protein processing in the endoplasmic reticulum during the 80 μM TNT treatment and sought to corroborate the RNA-seq data using quantitative RT-PCR (Supplementary Table S3). The results showed a strong correlation between the RNA-seq and qRT-PCR data, which strongly confirmed that the RNA-seq results in this study are reliable. In order to better understand the functions of differentially expressed genes, we subjected the data to analysis for relationships as revealed by the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway, Using KEGG analysis, differential genes were identified with biological pathways/functions that were related to oxidative phosphorylation, the metabolic pathway and protein processing in the endoplasmic reticulum. In addition, Alzheimer’s disease, Huntington’s disease, and Parkinson’s disease were detected irrespective of the 30 μM and 80 μM TNT treatments (Supplementary Tables S4 and S5).
TNT-induced ER stress in HepG2 cells
The altered redox homeostasis in the cell could cause ER stress, which in turn could induce the production of ROS in the ER and mitochondria12. To delineate the induction of ER stress by TNT in HepG2 cells, we investigated the induction of proteins related to ER stress after the 30 μM and 80 μM TNT treatments in the HepG2 cells. As shown in Fig. 2A and B, the expression of binding immunoglobulin protein (BiP) and protein disulfide isomerase (PDI) increased after the TNT treatment, but no differences were found in calnexin expression. The up-regulation of ER stress transducers inositol-requiring protein 1 alpha (IRE1-α) and PKR-like ER protein kinase (PERK) was observed. The dose response and time of C/EBP homologous protein (CHOP), transcriptional target (ER oxidase 1-like α [Erol-Lα]), phosphorylated eukaryotic initiation factor 2 (eIF2α), phosphorylated IRE1, and phosphorylated PERK also increased after the TNT treatment. We observed similar results in the Hep3B cells (Supplementary Fig. S1). These results indicated that TNT is capable of inducing ER stress in human hepatoma cells.
TNT-induced ER stress in HepG2 cells. The effects of (A) 30 μM TNT and (B) 80 μM TNT on the expression of ER stress related proteins. HepG2cells were treated with TNT at the indicated concentrations for 0, 6, 12, 18, and 24 h. Whole-cell lysates were obtained and subjected to Western blot analysis using antibodies against calnexin, PDI, BIP, IRE1-α, PERK, Ero1-Lα, CHOP, phosphorylated eIF2α, phosphorylated IRE1, phosphorylated PERK and β-actin. The bands were excised from different gels that were run under the same electrophoresis condition.
TNT-induced apoptosis through mitochondrial dysfunction pathway
ROS formation from mitochondria dysfunction would cause the mitochondrial membrane potential (MMP) to collapse completely19, 20. To investigate the effect of TNT on mitochondrial function, we first measured MMP using JC-1 dye. This fluorescent dye can spread selectively within the mitochondria depending on the membrane potential12. The exposure of HepG2 cells (30 μM or 80 μM) and Hep3B cells (50 μM or 60 μM) to TNT for 24 h caused disruption of the MMP, which was detected by a decrease in the red/green ratio of JC-1 fluorescence, as shown in the results of the cells treated with 100 μM H2O2 (the positive control; Fig. 3A). The loss of MMP was always accompanied by the mitochondrially mediated apoptotic pathway in human cells, as reported by Zhou et al.12. Additionally, MMP is a highly regulated process that is primarily controlled through interactions between pro- and anti-apoptotic members of the B-cell lymphoma 2 (Bcl-2) family21. Accordingly, we measured the expression of Bcl-2-associated X protein (Bax) and Bcl-2 using Western blot analysis (Fig. 3B). The results showed a dose-dependent suppression of Bcl-2 expression, but the expression of Bax exhibited the opposite results. These results suggest that the mitochondrially mediated apoptotic pathway was stimulated by TNT.
TNT triggered the mitochondrial dysfunction and apoptosis pathways. (A) TNT (0, 30, and 80 μM) caused a disruption of MMP in HepG2 cells, and TNT (0, 50, and 60 μM) caused a disruption of MMP in Hep3B cells after treatment for 24 h, as evidenced by an increased proportion of cells with green fluorescent light and a decrease proportion of cells with a higher red (JC-1 aggregates)/green (JC-1 monomers) ratio of JC-1 fluorescence. (B) TNT-induced mitochondrial dysfunction, as evidenced by Western blot analysis using antibodies against Bax, Bcl-2, and β-actin. (C) The cells were treated with different concentrations of TNT for 90 min in the presence or absence of 10 mM NAC, and the dissipation of MMP was measured. (the positive control: 100 μM H2O2)
The role of ROS formation in inducing DNA damage, ER stress, mitochondrial dysfunction, apoptosis, and cell death
There has been much evidence to support the important role of ROS in initiating cascades of cell death22. We proved that TNT indeed generates ROS (Fig. 4A and B). To assess the effect of TNT on oxidative DNA damage, NAC—the antioxidant and ROS scavenger— was used for the functional role in TNT-induced DNA damage. The results revealed that the level of DNA damage was reduced in HepG2 and Hep3B cells after the addition of NAC, as compared to the control group (Fig. 1B–D). We indicated that TNT could result in oxidative DNA damage. We also tested the effect of NAC on ER stress, mitochondrial dysfunction, apoptosis, and cell viability. The pretreatment of cells with NAC decreased the ratio of expressed Bax to expressed Bcl-2 (Fig. 3B) Furthermore, NAC significantly blocked TNT-induced MMP loss (Fig. 3C), indicating that the process of mitochondrial-related apoptosis was inhibited (Fig. 1F). Lastly, we showed that NAC attenuated TNT-induced ER stress and cytotoxicity (Fig. 5A,B and Supplementary Fig. S1). These findings strongly suggest that TNT induces oxidative stress and generating oxidative relation lesions. Additionally, CHOP was the best characterized apoptosis factor related to the ER stress pathway. We decreased the expression of CHOP protein using the siRNA method (80% knockdown efficiency; Supplementary Fig. S2) to investigate whether ER stress induced apoptosis. The results revealed that apoptosis was decreased when the expression of CHOP protein was inhibited in HepG2 cells (Fig. 6). Overall, ROS formation elicited apoptosis through ER stress and mitochondrial dysfunction.
TNT induced the generation of ROS. (A) ROS expression levels of HepG2 and Hep3B cells exposed to different doses of TNT for 0 to 90 min. (B) The HepG2 cells were treated with 80 μM TNT, and the Hep3B cells were treated with 60 μM TNT, both for 90 min in the presence or absence of 10 mM NAC. The values represent the mean ± SD and were derived from at least three independent experiments. Triplicate measurements were performed for each experiment. *P < 0.05; **P < 0.01 (vs. control cells).
ROS-dependent ER stress was involved in the TNT-mediated ROS production. (A) HepG2 cells were treated with 80 μM TNT for 24 h in the presence or absence of 10 mM NAC, given the expression of ER stress-related proteins. The bands were excised from different gels that were run under the same electrophoresis condition. (B) The cells were treated with 0–100 μM TNT for 24 h in the presence or absence of 10 mM NAC, and then the cell viability was determined. The values represent the mean ± SD and were derived from at least three independent experiments. Triplicate measurements were performed for each experiment. **P < 0.01 (vs. control cells).
Effects of CHOP siRNA on TNT-induced ER stress in HepG2 cells. The cells were transfected with or without CHOP siRNA and treated with or without TNT (80 μM) and with or without NAC for 24 h. Caspase-3/7 activity was detected via microplate reader. The data are presented as mean ± SD for four independent experiments with triplicate determinations. *P < 0.05; **P < 0.01 as compared with control.
Discussion
Cultures of HepG2 (p53wt) and Hep3B (p53null) cells are frequently used in in vitro models for human biotransformation in the human liver and is of great importance for toxicological and pharmaceutical studies23. Qiu et al. reported that p53 may not be the crucial factor to determine the differences in biological responses between HepG2 and Hep3B cell lines in confronting toxicants24. HepG2 cells of human origin retained active xenobiotic metabolizing enzymes and genotoxic sensitivity. Furthermore, HepG2 cells with the enzyme activity, and gene expression were similar to normal human liver cells25, 26. The use of HepG2 cells as the target for detecting the toxicity of pesticides, including TNT and dinitrotoluenes (byproducts of TNT), has recently been reported9, 27,28,29,30. We utilized this cell line to detect the genotoxicity of dicrotophos and to address the molecular mechanism of 3,3′-dichlorobenzidine-mediated toxicity30, 31. Recently, we proved the role of miRNA in inhibiting DNA repair proteins in HepG2 cells- treated with 4-aminobiphenyl, a well-known carcinogen that induces bladder cancer32. In this study, we have shown that TNT has a significant inhibitory effect on the growth of HepG2 and Hep3B cells. HepG2 and Hep3B cells are of great relevance for detecting cytotoxic and genotoxic substances23; therefore, these cell lines were utilized to elucidate the molecular mechanism of TNT-induced apoptosis through an ROS-dependent destructive cycle involving DNA damage, ER stress and mitochondrial dysfunction.
RNA-seq provides a far more precise measurement of levels of transcripts than other traditional methods33. With this method, we successfully measured the change of transcriptome responses in zebrafish embryos before and after triadimefon34. Accordingly, in this study, RNA-seq was utilized to assess the transcriptome responses in cells under TNT stress. KEGG analysis revealed that oxidative phosphorylation was one of the enriched transcriptional responses. Many genes related to oxidative phosphorylation were dysregulated after the TNT treatment (Supplementary Tables S4 and S5), suggesting that mitochondrial function could be affected by TNT. Mitochondrial dysfunction in the form of oxidative stress and mutations can contribute to the pathogenesis of various neurodegenerative diseases such as Parkinson’s disease (PD), Alzheimer’s disease (AD), and Huntingon’s disease (HD)35. When the cells were treated with 80 μM TNT, the most significant response was the dysregulation of genes related to AD and HD, which could have resulted from mitochondrial dysfunction, as suggested by Arun et al.35, who reported on the role of mitochondrial dysfunction in the pathogenesis of these diseases.
Several recent studies have shown that the generation of ROS by diverse cell-death stimuli not only turns on the cell death signals but also leads directly to DNA damage36. In this study, the comet assay and γ-H2AX protein expressions results clearly showed that TNT could cause DNA damage in HepG2 cells, and DNA damage was alleviated when the cells were pretreated with NAC. p53 is a major cellular stress sensor that is activated in response to DNA damage such as telomere dysfunction and other adverse stimuli such as ROS37. Chung et al. reported that DNA damage induced by drugs such as sinularin can initiate p53-dependent and p53-independent pathways38. In this study, a significant increase in p53 protein levels was observed in the HepG2 cells. We also observed DNA damage, ER stress, mitochondrial dysfunction, and apoptosis in Hep3B cells, a p53-null cell line, when the cells were treated with TNT. These results suggest that TNT-induced cell apoptosis occurs independently of p53 in Hep3B cells. Given that DNA damage usually comes with oxidative stress and that ROS could be involved in leading to mitochondrial dysfunction38, we used JC-1 dye to stain viable mitochondria and flow cytometry to measure mitochondrial membrane potential. We indeed showed that TNT can disrupt mitochondrial membrane potential. Two sites in the mitochondria, complex I and complex III, have been suggested as the major sites for ROS production39. Zhou et al. showed that ROS production increased in human leukemia cell lines when complex III activity was decreased by miltirone12. Similarly, we detected the dysregulation of genes related to complex I and complex III in cells after TNT treatment, although the activity of these complexes was not measured. Therefore, we reasonably suggested that ROS mainly originated from mitochondria when the levels of mitochondrial dysfunction were rescued in cells pretreated with NAC.
In this study, we showed that TNT could induce a pronounced increase of ROS and lead to apoptosis. A critical question arises regarding the apoptotic mechanism by which ROS elevation occurs during TNT treatment. Woo et al. has been demonstrated that apoptosis is related to MMP depolarization and that the permeabilization of outer mitochondrial membranes is a critical step in apoptosis40. Our results demonstrated that TNT induced MMP depolarization. Furthermore, the loss of MMP was attenuated when cells were pretreated with NAC, indicating that ROS participates in mitochondrial dysfunction. Bcl-2, which belongs to the anti-apoptotic family, can bind to Bax and prevent the permeabilization of the outer mitochondrial membrane41. TNT decreased the expression of Bcl-2 and increased the expression of Bax, leading to an increase in the Bax/Bcl-2 ratio. In addition, TNT significantly decreased the expression of pro-caspase-9 and -3 and increase cleaved caspase-9 and -3 respectively, and increased the activity of caspase-3/7. These results indicate that TNT could induce the mitochondria-related apoptotic pathway. However, the above results were reversed when these cells were pretreated with NAC, indicating that ROS could play an important role in apoptosis.
The activation of unfolded protein response (UPR) plays a protective role in cells under ER stress. Physiological processes that demand a high rate of protein synthesis and secretion must sustain activation of the UPR’s adaptive programs without triggering cell death pathways. However, the activation of UPR by excessive ER stress can convert its role to cytotoxic by activation of multiple apoptotic pathways in mammalian cells cells42. IRE1-α, eIF2α and PERK constitute the core stress regulators of the UPR and transduce signals from the ER to the cytoplasm and nucleus after ER stress42. PDI is an ER chaperon, that is responsible for the formation of disulfide bonds in proteins43. BiP, which is an intracellular chaperon, regulates the unfolded protein response during ER stress44. Ero1-Lα is an upstream signal of ER stress and has been conjectured to hyperoxidize the ER lumen and cause cytotoxic ROS production, leading to cell death45. Another mechanism was associated with the role of Bax in CHOP-induced apoptosis. Additionally, CHOP, a transcription factor, can decrease Bcl-2 transcription and promote ROS production46, as our results in the present study have shown. These results implicated the involvement of ER stress in TNT-induced apoptosis.
Much evidence has shown that ROS can disturb ER protein folding and induce ER stress. In our results, we found that levels of ER stress can be attenuated by NAC. Hence, ER stress may be associated with TNT-induced oxidative stress. To investigate the possible role of ER stress in TNT-induced apoptosis, the knockdown of CHOP was used to alleviate the ER stress. These treated cells significantly attenuated the TNT-mediated apoptosis.
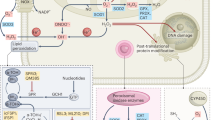
In summary, the present findings demonstrate that TNT effectively induced apoptosis through ROS-dependent DNA damage, ER stress and mitochondrial dysfunction. We provided the detailed mechanistic basis for TNT-induced cell death under oxidative stress. Besides, NAC could significantly increase the viability of TNT-treated cells (Fig. 5). Finally, this study presented the mechanisms of TNT-induced HepG2 cell apoptosis through ROS-dependent mitochondria and ER stress pathways (Fig. 7).
Materials and Methods
Materials
The TNT (99.9% purity) used in the present study was provided by the Taiwan Explosive Chemical Laboratory, Combined Logistics Forces, Taiwan. The N-acetylcysteine (NAC) and dimethyl sulfoxide (DMSO) came from Sigma-Aldrich (St. Louis, MO, USA).
Cell culture
HepG2 and Hep3B cells were cultured in Dulbecco’s modified Eagle medium (DMEM, Lonza, BE12-614, Barcelona, Spain) containing 10% (v/v) fetal bovine serum (Lonza), 2 mM l-glutamine, 50 U/mL penicillin, and 0.1 mg/mL streptomycin (Sigma) in a humidified incubator with 5% CO2 at 37 °C.
Cell treatment
The cells were cultured in 10-cm2 dishes until 70% confluence was reached. The medium was then replaced with a fresh serum-free medium containing the indicated TNT stock solution. Different concentrations of TNT (0–100 μg/mL) were dissolved in 0.5% DMSO. The negative controls were exposed to 0.5% DMSO. The treated cells were exposed to the TNT stock solution for a total of 24 h or 48 h.
Cell viability
The cell viability of HepG2 and Hep3B cells was detected using Cell Counting Kit-8 (CCK-8 Donjindo Molecular Technologies, Inc., MD, USA) according to the manufacturer’s instruction. Cells in a 96-well plate (5 × 104 cells/well) were incubated with or without various concentrations of TNT for 12 h or 24 h. Thereafter, 10 μL of CCK-8 solution was added to each well, and the cells were incubated at 37 °C for 1–4 h. The absorbance was measured at 450 nm using a microplate spectrofluorometer (Mark; Bio-Rad, Hercules, CA, USA).
Comet assay
The comet assay was performed under alkaline conditions using our previously described methods47. At least 300 images were randomly selected from each sample and analyzed for DNA damage with the Comet IV computer software (Perceptive Instruments, UK). The tail moment comet parameter (mean ± SD) was used as an indicator of DNA damage.
Total RNA isolation
The total RNA from the HepG2 cells was obtained using TRIzol Reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions. Briefly, after the cells had been treated cells for 24 h, they were washed with 1 mL TRIzol Reagent and 200 μL of chloroform during a short incubation. After mixing vigorously, the solution was centrifuged at 13,000 × g for 20 min. The RNA was then precipitated in isopropyl alcohol, washed with 75% ethanol, pelleted, and resolubilized in nuclease-free water. RNA quantity and purity were measured spectrophotometrically (BioPhotometer, Eppendorf). The samples were considered suitable for further processing if the A260/A280 ratios were between 1.8 and 2.0. RNA integrity was determined with 1.8% agarose electrophoresis gel.
Gene expression profiles: The RNA sequencing
The RNA sequencing (RNA-seq) was performed according to our previously described methods34. The total extracted RNA was first treated with DNase I to degrade any possible DNA contamination. Poly-A mRNA was isolated using oligo (dT) magnetic beads and then fragmented. The cleaved RNA fragments were transcribed into first-strand cDNA using reverse transcriptase and random hexamer primers, which was followed by second-strand cDNA synthesis. The double -stranded cDNA was further subjected to end-repair, phosphorylation, 3′-adenylation, and adaptor ligation in sequence. Adaptor ligated fragments were selected according to the size (250–350 bp), and cDNA fragments in the desired range were extracted from the gel. The fragments were enriched by PCR amplification. After qualitative and quantitative analysis by an Agilent 2100 Bioanalyzer and the ABI StepOnePlus Real-Time PCR System, respectively, the cDNA libraries were subjected to RNA-seq via Illumina HiSeq. 2000 (BGI, Shenzhen, China).
Analysis of differential gene expression
The original image data generated by the sequencer were converted into sequences. After the filtration of the low-quality reads, the raw reads were cleaned up by removing adapter sequences based on the illumine pipeline. Gene expression profiling was measured by mapping clean reads onto assembled sequences using SOAP software. Gene expression levels were calculated by using the reads per kilobase of transcript sequence per million mapped reads (RPKM)48. This study followed a modified version of Audic’s method screen for the differentially expressed genes (DEGs)49. The false discovery rate (FDR) method was used to determine the threshold of the p-value. We used an FDR of ≦0.001 and an absolute value of log2 ratio ≧1 as the threshold to judge the significance of gene expression difference.
Pathway analysis
All genes must cooperate with each other to exercise their biological functions. We mapped all DEGs onto terms from the Kyoto Encyclopedia of Genes and Genomes (KEGG) database (http://www.genome.jp/kegg/pathway.html), where a q-value of <0.05 indicated significantly enriched terms in DEGs.
Caspase-3/7 activity assay
Cells from each treatment group were harvested after 24 h and caspase activity was detected by a Caspase-Glo 3/7 assay kit (Promega, Taipei, Taiwan). Briefly, 100 μL of Caspase-Glo 3/7 reagent was added to each well of the plates, followed by mixing for 1 min. Subsequently, these plates were incubated at room temperature for 2 h. Absorbance values were measured with a microplate reader at 405 nm (Synergy HTX Multi-Mode Reader; Biotek, Taipei, Taiwan).
Detection of intracellular ROS
The production of intracellular ROS was detected using fluorescent dye 2′,7′- dichlorodihydrofluorescein diacetate (H2DCFDA, Sigma, USA). Briefly, 5 × 105 cells/well cultured in a 6-well plate were incubated with or without different concentrations (HepG2: 0, 30, or 80 μM; Hep3B:0, 50, 60 μM) of TNT for the indicated time (with 100 μM H2O2 as a positive control). The cells were stained with 10 μM H2DCFDA for 30 min. Then the cells were examined under a fluorescence microscope, and the fluorescence intensity generated in the cells was detected using a fluorescence plate reader at an excitation wavelength of 488 nm and an emission wavelength of 525 nm.
Western blot analysis
Western blot analysis was performed as previously described32. The culture medium was replaced with a new medium when the HepG2 cells were 70% confluent, then, the cells were exposed to various concentrations of TNT for 24 h. Subsequently, the cells were washed twice with PBS and then lysed. From each sample, 50 μg protein was subjected to 10% SDS polyacrylamide gel electrophoresis, transferred to PVDF membranes, and blocked with 5% skim milk at room temperature for 1 h. After blocking, the membrane was incubated with antibodies against pro-caspase-9, pro-caspase-3, cleaved-caspase-9, cleaved-caspase-3, β-actin, calnexin, PDI, BIP, IRE1-α, PERK, Ero1-Lα, CHOP, Bax, Bcl-2, phosphorylation of IRE1, phosphorylation of PERK and phosphorylation of eIF2α (Cell Signaling, USA) for 1 h. Then, the membranes were washed with 0.1% PBST (PBS and 0.05% Tween 20) and incubated with secondary antibodies conjugated to horseradish peroxidase for 1 h. Bands were detected after chemiluminescent HRP (Immunobilon Western, Millipore) was added. The band density was measured with ImageQuant-TL7.0 software (GE Healthcare).
Mitochondrial Membrane Potential (MMP) measurement
The disruption of MMP was measured using fluorochrome dye JC-1 by flow cytometry as reported by Zhou et al.12. After the cells were treated with different concentration of TNT, the cells were harvested and washed twice with PBS. Then, the cells were treated with JC-1 for 30 min and analyzed in a flow cytometer. Each group acquired more than 10,000 individual cells.
Validation by reverse transcription quantitative PCR
To confirm the expression results of the mRNAs obtained from RNA-seq, SYBR Green I® Quantitative Real-Time PCR (qRT-PCR) was performed with an IQ-5 Real-Time PCR System (BIO RAD). All of the primer sequences used for qRT-PCR are listed in Supporting Information Table S6. In brief, a total of 10 ng RNA for each sample was used to perform qRT-PCR. cDNA was synthesized from the total RNA using a qScript cDNA Synthesis Kit (Quanta BioSciences, Gaithersburg, MD, USA) according to the manufacturer’s instructions. Subsequently, amplifications were carried out under the following conditions: 95 °C for 10 min (initial denaturation), 35 combined cycles at 95 °C for 30 s (denaturation), 57 °C for 45 s (annealing), and 72 °C for 1 min (extension). Fluorescence was measured in real time. The cycle threshold (Ct) values were calculated using the LightCycler3 data analysis software. The fold expression or repression of the target gene relative to the internal control gene in each sample was calculated by following the ΔΔCt model. All the samples were analyzed in triplicate, and the mean value of these triplicate measurements was used to calculate the mRNA transcriptions.
The siRNA transfection
The cells were seeded in 6-well plates at a density of 5 × 105 cells/mL and allowed to reach approximately 70% confluence on the day of transfection. The small interfering RNA (siRNA) against CHOP was purchased commercially from Sigma, USA. The cells were transfected with CHOP siRNA (100 nM) using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. After 24 h, the cells were treated with TNT for 24 h and examined by Western blot.
References
Chien, C. C., Kao, C. M., Chen, D. Y., Chen, S. C. & Chen, C. C. Biotransformation of trinitrotoluene (TNT) by Pseudomonas spp. isolated from a TNT-contaminated environment. Environ Toxicol Chem 33, 1059–1063 (2014).
Berthe-Corti, L., Jacobi, H., Kleihauer, S. & Witte, I. Cytotoxicity and mutagenicity of a 2,4,6-trinitrotoluene (TNT) and hexogen contaminated soil in S. typhimurium and mammalian cells. Chemosphere 37, 209–218 (1998).
Honeycutt, M. E., Jarvis, A. S. & McFarland, V. A. Cytotoxicity and mutagenicity of 2,4,6-trinitrotoluene and its metabolites. Ecotoxicol Environ Saf 35, 282–287 (1996).
Ross, R. H. & Hartley, W. R. Comparison of water quality criteria and health advisories for 2,4,6-trinitrotoluene. Regul Toxicol Pharmacol 11, 114–117 (1990).
Bolt, H. M., Degen, G. H., Dorn, S. B., Plottner, S. & Harth, V. Genotoxicity and potential carcinogenicity of 2,4,6-TNT trinitrotoluene: structural and toxicological considerations. Rev Environ Health 21, 217–228 (2006).
Hathaway, J. A. Trinitrotoluene: A Review of Reported Dose-Related Effects Providing Documentation for a Workplace Standard. J Occup Med 19, 341–345 (1977).
Deng, Y. et al. Analysis of common and specific mechanisms of liver function affected by nitrotoluene compounds. PLoS One 6, e14662 (2011).
Yan, C. et al. The retrospective survey of malignant tumor in weapon workers exposed to 2,4,6-trinitrotoluene. Zhonghua Lao Dong Wei Sheng Zhi Ye Bing Za Zhi 20, 184–188 (2002).
Song, L. et al. Trinitrotoluene Induces Endoplasmic Reticulum Stress and Apoptosis in HepG2 Cells. Med Sci Monit 21, 3434–3441 (2015).
Glass, K. Y., Newsome, C. R. & Tchounwou, P. B. Cytotoxicity and expression of c-fos, HSP70, and GADD45/153 proteins in human liver carcinoma (HepG2) cells exposed to dinitrotoluenes. Int J Environ Res Public Health 2, 355–361 (2005).
Shi, Y. et al. ROS-dependent activation of JNK converts p53 into an efficient inhibitor of oncogenes leading to robust apoptosis. Cell Death Differ 21, 612–623 (2014).
Zhou, L. et al. Miltirone exhibits antileukemic activity by ROS-mediated endoplasmic reticulum stress and mitochondrial dysfunction pathways. Sci Rep 6, 20585 (2016).
Romanov, V., Whyard, T. C., Waltzer, W. C., Grollman, A. P. & Rosenquist, T. Aristolochic acid-induced apoptosis and G2 cell cycle arrest depends on ROS generation and MAP kinases activation. Arch Toxicol 89, 47–56 (2015).
Cooke, M. S., Evans, M. D., Dizdaroglu, M. & Lunec, J. Oxidative DNA damage: mechanisms, mutation, and disease. Faseb j 17, 1195–1214 (2003).
Chen, Y. W., Yang, Y. T., Hung, D. Z., Su, C. C. & Chen, K. L. Paraquat induces lung alveolar epithelial cell apoptosis via Nrf-2-regulated mitochondrial dysfunction and ER stress. Arch Toxicol 86, 1547–1558 (2012).
Soboloff, J. & Berger, S. A. Sustained ER Ca2+ depletion suppresses protein synthesis and induces activation-enhanced cell death in mast cells. J Biol Chem 277, 13812–13820 (2002).
Csordas, G., Thomas, A. P. & Hajnoczky, G. Quasi-synaptic calcium signal transmission between endoplasmic reticulum and mitochondria. Embo j 18, 96–108 (1999).
Zhao, M., Howard, E. W., Guo, Z., Parris, A. B. & Yang, X. p53 pathway determines the cellular response to alcohol-induced DNA damage in MCF-7 breast cancer cells. PLoS One 12, e0175121 (2017).
Ozgur, R., Turkan, I., Uzilday, B. & Sekmen, A. H. Endoplasmic reticulum stress triggers ROS signalling, changes the redox state, and regulates the antioxidant defence of Arabidopsis thaliana. J Exp Bot 65, 1377–1390 (2014).
Jacobson, J. & Duchen, M. R. Mitochondrial oxidative stress and cell death in astrocytes–requirement for stored Ca2+ and sustained opening of the permeability transition pore. J Cell Sci 115, 1175–1188 (2002).
Tait, S. W. & Green, D. R. Mitochondria and cell death: outer membrane permeabilization and beyond. Nat Rev Mol Cell Biol 11, 621–632 (2010).
Petrosillo, G., Ruggiero, F. M., Pistolese, M. & Paradies, G. Ca2+-induced reactive oxygen species production promotes cytochrome c release from rat liver mitochondria via mitochondrial permeability transition (MPT)-dependent and MPT-independent mechanisms: role of cardiolipin. J Biol Chem 279, 53103–53108 (2004).
Wilkening, S., Stahl, F. & Bader, A. Comparison of primary human hepatocytes and hepatoma cell line HepG2 with regard to their biotransformation properties. Drug Metab Dispos 31, 1035–1042 (2003).
Qiu, G. H. et al. Distinctive pharmacological differences between liver cancer cell lines HepG2 and Hep3B. Cytotechnology 67, 1–12 (2015).
Knasmuller, S. et al. Use of metabolically competent human hepatoma cells for the detection of mutagens and antimutagens. Mutat Res 402, 185–202 (1998).
Mersch-Sundermann, V., Knasmuller, S., Wu, X. J., Darroudi, F. & Kassie, F. Use of a human-derived liver cell line for the detection of cytoprotective, antigenotoxic and cogenotoxic agents. Toxicology 198, 329–340 (2004).
Pirozzi, A. V., Stellavato, A., La Gatta, A., Lamberti, M. & Schiraldi, C. Mancozeb, a fungicide routinely used in agriculture, worsens nonalcoholic fatty liver disease in the human HepG2 cell model. Toxicol Lett 249, 1–4 (2016).
Medina-Diaz, I. M. et al. Downregulation of human paraoxonase 1 (PON1) by organophosphate pesticides in HepG2 cells. Environ Toxicol 32, 490–500 (2017).
Tchounwou, P. B., Wilson, B. A., Ishaque, A. B. & Schneider, J. Transcriptional activation of stress genes and cytotoxicity in human liver carcinoma cells (HepG2) exposed to 2,4,6-trinitrotoluene, 2,4-dinitrotoluene, and 2,6-dinitrotoluene. Environ Toxicol 16, 209–216 (2001).
Hseu, Y. C., Hsu, T. W., Lin, H. D., Chen, C. H. & Chen, S. C. Molecular mechanisms of discrotophos-induced toxicity in HepG2 cells: The role of CSA in oxidative stress. Food Chem Toxicol 103, 253–260 (2017).
Chen, L. C. et al. Molecular mechanisms of 3,3′-dichlorobenzidine-mediated toxicity in HepG2 cells. Environ Mol Mutagen 55, 407–420 (2014).
Huan, L. C. et al. MicroRNA regulation of DNA repair gene expression in 4-aminobiphenyl-treated HepG2 cells. Toxicology 322, 69–77 (2014).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10, 57–63 (2009).
Hsu, L. S., Chiou, B. H., Hsu, T. W., Wang, C. C. & Chen, S. C. The regulation of transcriptome responses in zebrafish embryo exposure to triadimefon. Environ Toxicol 32, 217–226 (2017).
Arun, S., Liu, L. & Donmez, G. Mitochondrial Biology and Neurological Diseases. Curr Neuropharmacol 14, 143–154 (2016).
Fang, J., Seki, T. & Maeda, H. Therapeutic strategies by modulating oxygen stress in cancer and inflammation. Adv Drug Deliv Rev 61, 290–302 (2009).
Sahin, E. & Depinho, R. A. Linking functional decline of telomeres, mitochondria and stem cells during ageing. Nature 464, 520–528 (2010).
Chung, T. W. et al. Sinularin induces DNA damage, G2/M phase arrest, and apoptosis in human hepatocellular carcinoma cells. BMC Complement Altern Med 17, 62 (2017).
Turrens, J. F. Superoxide production by the mitochondrial respiratory chain. Biosci Rep 17, 3–8 (1997).
Woo, I. S. et al. TMEM14A inhibits N-(4-hydroxyphenyl)retinamide-induced apoptosis through the stabilization of mitochondrial membrane potential. Cancer Lett 309, 190–198 (2011).
Fei, Q. & Ethell, D. W. Maneb potentiates paraquat neurotoxicity by inducing key Bcl-2 family members. J Neurochem 105, 2091–2097 (2008).
Ron, D. & Walter, P. Signal integration in the endoplasmic reticulum unfolded protein response. Nat Rev Mol Cell Biol 8, 519–529 (2007).
Perri, E. R., Thomas, C. J., Parakh, S., Spencer, D. M. & Atkin, J. D. The Unfolded Protein Response and the Role of Protein Disulfide Isomerase in Neurodegeneration. Front Cell Dev Biol 3, 80 (2015).
Panayi, G. S. & Corrigall, V. M. Immunoglobulin heavy-chain-binding protein (BiP): a stress protein that has the potential to be a novel therapy for rheumatoid arthritis. Biochem Soc Trans 42, 1752–1755 (2014).
Mera, K. et al. ER signaling is activated to protect human HaCaT keratinocytes from ER stress induced by environmental doses of UVB. Biochem Biophys Res Commun 397, 350–354 (2010).
McCullough, K. D., Martindale, J. L., Klotz, L. O., Aw, T. Y. & Holbrook, N. J. Gadd153 sensitizes cells to endoplasmic reticulum stress by down-regulating Bcl2 and perturbing the cellular redox state. Mol Cell Biol 21, 1249–1259 (2001).
Wu, J. C., Hseu, Y. C., Chen, C. H., Wang, S. H. & Chen, S. C. Comparative investigations of genotoxic activity of five nitriles in the comet assay and the Ames test. J Hazard Mater 169, 492–497 (2009).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5, 621–628 (2008).
Audic, S. & Claverie, J. M. The significance of digital gene expression profiles. Genome Res 7, 986–995 (1997).
Author information
Authors and Affiliations
Contributions
Ssu-Ching Chen conceived the experiments, Ssu-Ching Chen and Hung-Yu Liao wrote the main manuscript text, Chih-Ming Kao and Chao-Ling Yao analysed the results, Po-Wei Chiu and Chun-Chen Yao prepared Figures 1 and 5. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liao, HY., Kao, CM., Yao, CL. et al. 2,4,6-Trinitrotoluene Induces Apoptosis via ROS-Regulated Mitochondrial Dysfunction and Endoplasmic Reticulum Stress in HepG2 and Hep3B Cells. Sci Rep 7, 8148 (2017). https://doi.org/10.1038/s41598-017-08308-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-08308-z
This article is cited by
-
Obesity Promotes Experimental Colitis by Increasing Oxidative Stress and Mitochondrial Dysfunction in the Colon
Inflammation (2020)
-
The explosive trinitrotoluene (TNT) induces gene expression of carbonyl reductase in the blue mussel (Mytilus spp.): a new promising biomarker for sea dumped war relicts?
Archives of Toxicology (2020)
-
Connexin32 plays a crucial role in ROS-mediated endoplasmic reticulum stress apoptosis signaling pathway in ischemia reperfusion-induced acute kidney injury
Journal of Translational Medicine (2018)
-
Toxic effect and mechanism of four ionic liquids on seedling taproots of Arabidopsis thaliana
Environmental Science and Pollution Research (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.