Abstract
Streptomyces are of great biological and industrial significance due to their complex morphological development and ability to produce numerous secondary metabolites. However, the intrinsic biochemical mechanisms underlying morphogenesis and secondary metabolism are rarely revealed, partially because of the limited availability of the biochemical tools in Streptomyces. Here we provided series of integrative vectors with various affinity tags, including single tags 3×FLAG, 3×HA, 3×Strep-tag II, 18×His, 13×Myc, and dual tags, all of which were driven from a strong constitutive promoter ermEp*. Using a sigma factor SigT from S. coelicolor as a model, we successfully expressed and immuno-detected SigT fused with all tags. Moreover, after SigT was N-terminally tagged with 3×FLAG and C-terminally tagged with 18×His, we isolated SigT-interactive proteins from the S. coelicolor lysate based on the tandem affinity purification (TAP). Particularly, among the proteins purified, the SigT cognate anti-sigma factor RstA ranked the top with the most total independent spectra. These data suggested the feasibility of these affinity tags in Streptomyces, which will be widely employed to explore the biochemical mechanisms to further understand the dynamic and elaborate regulation in this genus.
Similar content being viewed by others
Introduction
Streptomyces, the soil-dwelling filamentous bacteria, possess a periodic morphological development with a progression of cell types from vegetative mycelia to aerial mycelia, and to the production of spores, and have a complex secondary metabolism to produce invaluable antibiotics, immuno-suppressors, anti-tumor drugs, etc1. Due to their biological, industrial and clinical importance, numerous efforts have been made to reveal the regulatory pathways and their cross-talks, mainly at the genetic and transcriptional levels, all of which will be the basis to engineer the bacteria in this genus by systems biology, synthetic biology and metabolic engineering, etc2,3,4,5,6,7.
Abundant genetic tools, including transposons, linear/circular and high/low-copy vectors, have been developed initially to elucidate the gene functions and genomic information of Streptomyces 8. Complicated regulatory mechanisms have also been dissected based on the well-developed biochemical assays, such as in vitro transcription, electrophoretic mobility shift assay (EMSA) combined with in vivo assays, such as chromatin immunoprecipitation (ChIP). Particularly, the bld genes, whi genes, and several sigma factors essential for aerial hyphae development and sporulation have been discovered and the underlying genetic circuits were depicted9, 10. The highly conserved signaling pathways triggered by γ-butyrolactones but mediated by the pleiotropic regulator AdpA have been shown to globally regulate morphological transition and secondary metabolism9, 11, 12. Moreover, the nutrient-sensing pathways, such as the PhoRP two-component system and the orphan regulator GlnR, are also found in regulation of secondary metabolite production independently or in cross-talks13,14,15,16.
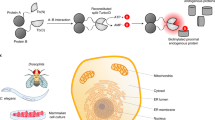
However, the regulatory complexity of Streptomyces, which essentially results from the in vivo dynamic protein turn-over, transient participation of particular proteins in different complexes, and competition or coordination of multiple regulatory factors on the regulons in a developmental phase-dependent manner, has been scarcely reported to date. Exploration of these mechanisms raises the requirement to develop protein-specific antibodies, or alternatively to label proteins with small tags without interference with their authentic functions. Along with commercial availability of high quality antibodies, several small affinity tags have been extensively used, especially in eukaryotic cells, to study the dynamic protein-protein and protein-DNA interaction, map the interactome, and reveal the protein complexes in different organelles, which has significantly helped us to understand their biology in nature and related diseases, and dramatically saved time and labors simultaneously17,18,19.
In Streptomyces, limited tags have been successfully used, such as FLAG in ChIP assays10, 20, eGFP for protein degradation monitoring21,22,23, and both tags for protein-protein interaction assays22. eGFP and mCherry have tremendously facilitated visualization of dynamic localization of morphogenesis-related proteins, revealing the subtle morphological changes and authentic protein functions24,25,26. However, these bulky fluorescent proteins might cause steric hindrance to impede the appropriate protein folding27 and block the interaction with others28.
Here we provided series of Streptomyces vectors containing various codon-optimized small single tag or dual tags, including 3×FLAG, 3×HA, 13×c-Myc, 18×His and 3×Strep-tag II. All these tags have been successfully expressed in fusion with the sigma factor SigT in S. coelicolor, and tandem affinity purification (TAP) was demonstrated for the first time in Streptomyces to show the feasibility of these tags without functional disruption of the fused proteins.
Results
Construction of Streptomyces vectors with affinity tags
Antibodies are powerful tools in investigating protein functions nowadays. Both protein-specific polyclonal and monoclonal antibodies against endogenously expressed PhoP and ScbR2 have been reported in Streptomyces ChIP assays16, 29. However, though polyclonal antibodies can be readily raised, quality control for affinity and specificity is problematic, and preparation of monoclonal antibodies will be time-consuming and risky30, 31. One of resolutions to overcome these problems is to express the target proteins in fusion with commonly used small epitopes, such as 6×Histidine, 8-amino acid FLAG tag (DYKDDDDK), 9-amino acid influenza hemagglutinin (HA) tag (YPYDVPDYA), 10-amino acid c-Myc tag (EQKLISEEDL), etc32. The 8-amino acid Strep-tag II (WSHPQFEK) is a recently developed epitope exhibiting intrinsic affinity toward streptavidin, and can be used to efficiently purify fusion proteins by Strep-Tactin conjugated beads33. Fusion of one of these small tags to the target proteins has been proven to have the minimal influence on their native conformation and functions34. Moreover, there are commercially available monoclonal antibodies against these affinity tags for various purposes, such as immunoblot, immunoprecipitation (IP), immunohistochemistry (IHC) and ChIP, with the advantages of predominantly high affinity, specificity, and time- and labor-saving.
To expand their applications and increase their antigenicity in Streptomyces, multiple tandems of these small epitopes were generated after codon-optimization. The 22-amino acid 3×FLAG (DYKDHDGDYKDHDIDYKDDDDK), and tandemly arrayed small tags including 3×HA, 13×Myc, 18×His, 3× Strep-tag II were all expressed under the control of a strong constitutive promoter ermEp*22. Eight vectors (pSN1-pSN8) (Fig. 1) were the derives of the integrative promoter-probing vehicle pIJ866035, where ermEp* was inserted between two BglII sites to remove the multiple cloning sites (MCS) on the parental vector, and the egfp reporter gene was replaced by the synthetic DNA fragments encoding the tandem epitopes together with the redesigned MCS. To meet the demand of labeling proteins at alternative ends, the 18×His tag was positioned at the C-terminus of the fusion proteins, while others were N-terminally tagged. Meanwhile, for the purpose of tandem affinity purification (TAP), two vectors were constructed by combination of N-terminal 3×FLAG or 3× Strep-tag II with C-terminal 18×His, given that 18×His-tagged fusion proteins can be purified both at native and denatured conditions, and the purification matrix Ni2+-NTA is relatively inexpensive, while anti-FLAG beads and Strep-Tactin beads are in excellent quality control in protein purification and immunoprecipitation10, 33. Moreover, to avoid possible steric hindrance from the tandem epitopes, a linker sequence 10×Gly was placed between 3×HA, 3×FLAG, or 3×Strep-tag II and the target protein, while a linker sequence GGSGGGGGG was placed between the target protein and 18×His (Fig. 1).
Expression of fusion proteins with affinity tags
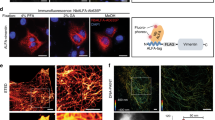
To validate these affinity tags in Streptomyces, an extracytoplasmic functions (ECF) sigma factor SigT from S. coelicolor was expressed from all above vectors. SigT can be physically protected by its cognate anti-sigma factor RstA from degradation, and is involved in dual positive feedback regulation of morphogenesis and secondary metabolism of S. coelicolor. We have previously tagged it with 3×FLAG and eGFP for immuno-detection and immunoprecipitation22, 23. However, a 10-amino acid flexible linker rich in Pro and Gly was essential for the proper expression of SigT-GFP22, suggesting that GFP, which is 236-amino acid long, might interfere with the proper conformation of SigT.
sigT coding sequence was cloned in all constructs in fusion with tags, and expressed in S. coelicolor. Immuno-blot assays showed SigT fusion proteins were successfully expressed in Streptomyces with α-FLAG, HA, Myc, His and Strep-tag II antibodies, respectively (Fig. 2). All the antibodies specifically recognized the corresponding epitopes, since only the SigT fusion protein bands were clearly observed in the immuno-blots. This highly reduced cross-reactivity will guarantee the exclusion of the false positive in downstream assays, especially such as ChIP. All the data suggested that these tags could be readily expressed and fused with Streptomyces proteins.
Identification of SigT-interactive proteins by tandem affinity purification (TAP)
Previous report has shown that an artificial extension of target proteins with 3×FLAG tag will not disturb the functionality in vivo by ChIP assays in studying protein-DNA interaction20. Here we further validated the feasibility of these affinity tags in the tandem affinity purification (TAP), which is a powerful tool in exploring the protein-protein interaction in vivo 36.
SigT has been shown subject to protein degradation during the secondary metabolism of S. coelicolor 23, but its degradation was blocked in the proteasome-deficient ΔprcB/A mutant21, suggesting that SigT remain intact while interacting with other proteins involved in its degradation in this mutant. Therefore the TAP assay was demonstrated with SigT as a model in the ΔprcB/A mutant to explore its potential interactome (Fig. 3). SigT was expressed in pSN7 with dual tags 3×FLAG at its N-terminus and 18×His at C-terminus. 3FLAG-SigT-18His could be immuno-detected in the ΔprcB/A mutant (Fig. 3, lane 1). Though some SigT fusion protein flowed away after loading on Ni2+-NTA, initial affinity binding to nickel could capture most SigT protein (Fig. 3, lane 2–7). Then the eluent was reloaded on the anti-FLAG M2 monoclonal antibody-conjugated agarose. After careful washes to a low background, SigT together with its interactive-proteins dissociated from the beads by heat denature (Fig. 3, lane 8–12). After buffer exchange and trypsin digestion, the final eluent was subject to HPLC-MS/MS for peptide sequencing and identification.
Immuno-blot assays of samples during tandem affinity purification (TAP) with anti-FLAG antibody. Total protein from S. coelicolor ΔprcB/A expressing 3FLAG-SigT-18His was subject to sequential nickel-based and anti-FLAG antibody-based affinity purification. Samples were taken during TAP as described in Materials and Methods. L: total lysate. FT: flow through. W: wash. E: elution.
Totally 28 proteins were identified in this TAP assay (Table 1) (see Supplementary data for the detailed information). Among them, SigT ranked No.2 with 100% group probability and showed 3 independent sequenced peptides, consistent with this protein as a bait in TAP. Interestingly, the anti-sigma factor RstA (SCO3891) had the most total independent spectra, and similarly, three peptides were independently sequenced as SigT, while all other 26 proteins only had one matched peptide. These data were consistent with our previous immunoprecipitation assay that SigT could physically interact with RstA22. Moreover, other proteins potentially involved in biosynthesis, regulation, transportation, etc, together with several proteins with unknown functions were also identified, suggesting that SigT might participate in diverse physiological processes by protein-protein interaction. All these data suggested that our dual tag strategy could be applied in TAP to study protein-protein interaction in Streptomyces.
Discussion
The filamentous bacteria Streptomyces are becoming the focus for its industrial values to produce numerous secondary metabolites in medicinal and agricultural applications. High through-put genome sequencing also showed a large biosynthetic gene cluster pool in this genus, suggesting a huge capacity for natural product production. Many genetic tools and in vitro assays have been established to reveal the genetic circuits and pathway cross-talks controlling the physiology of Streptomyces. Here we provided biochemical approaches with several affinity tags for in vivo protein expression and detection, and also for the first time established tandem affinity purification (TAP) in Streptomyces to study protein-protein interaction with SigT as a bait. These in vivo biochemical tools can also be expanded to explore the real-time dynamics of protein-DNA interaction (such as ChIP), to establish the protein-protein interaction network (or interactome), and to examine protein-protein competition or coordination in regulating gene expression on the promoters of regulons, etc. All these mechanisms in Streptomyces will be the basis of building up biological models of their morphological development and rational designing of biosynthetic and regulatory circuits to exploit their potentialities in secondary metabolite production to the utmost.
Materials and Methods
Strains and media
Streptomyces coelicolor wild type M145 and the ΔprcB/A mutant21 were used for protein expression in vivo. Escherichia coli strain TG1 was a host for the routine plasmid sub-cloning. E. coli strain ET12456 containing pUZ8002 was used for conjugation of plasmids from E. coli to Streptomyces 8.
All E. coli cells were cultured in LB medium. The inter-species conjugations were demonstrated according to the protocol described on the SFM medium8. For SigT fusion protein expression, all Streptomyces strains were cultured in TSB supplemented with 5% PEG6000 (TSBP) to the logarithmic stage. For the TAP assay, the S. coelicolor ΔprcB/A mutant expressing 3FLAG-SigT-18His was initially cultured in TSBP for 1 day, and then transferred to the liquid R5- medium for the secondary metabolism development after continuous culture for 2 days21.
Plasmid construction
All the plasmids and primers in this study were listed in Supplementary data Tables S1 and S2, respectively. Fragments I, II, III and IV containing NdeI-3HA-linker-MCS-TAA-NotI, NdeI-3FLAG-linker-MCS-linker-18His-TGA-NotI, NdeI-3Strep-tag II-linker-MCS-TAA-NotI and NdeI-MCS-13Myc-TAA-NotI (herein TAA and TGA encode the stop codons, and MCS is multiple cloning sites), respectively, were synthesized with optimized-codons for Streptomyces in Qinglan company, China (see Supplementary data for detailed sequences). ermEp* was amplified with primers 1 and 2 from pIJ8630-ermEp*22, digested with BamHI, and inserted into the BglII site of pIJ866035 to create the vector pIJ8660-ermEp*. Then the egfp gene was replaced with fragments I-IV at the NdeI/NotI site to give rise to plasmids pSN2, 7, 4 and 5, respectively. BglII/EcoRI fragments from pSN2 and pSN7 were cloned into BglII/EcoRI-digested pSN5 to create plasmids pSN1 and pSN6, respectively. Then pSN1 was digested by BglII and EcoRI, and the smaller DNA fragment was gel-recovered and inserted into BglII/EcoRI-digested pSN7 to make plasmid pSN3. BglII/EcoRI fragment of pSN4 were replaced with BglII/EcoRI fragment from pSN7 to produce plasmid pSN8. sigT was amplified from the genomic DNA of M145 with primers 3 and 4, digested with BglII and XbaI, and ligated into the BglII/XbaI site of pSN2 to pSN8 to create sigT expression plasmids as listed in Table S1.
Western blot
Streptomyces total protein was prepared from the mycelia and Western blot was demonstrated as described previously with antibodies against HA (Tiangen, China), Myc (Tiangen, China), FLAG (Sigma, USA), His (Abmart), Strep-tag II (IBA, Germany)21. The primary antibodies were diluted in 1:2000, while the anti-mouse HRP-conjugated secondary antibody was diluted in 1:5000.
Tandem affinity purification (TAP) and protein identification
The total protein was prepared from mycelia of the ΔprcB/A mutant expressing 3FLAG-SigT-18His in buffer A (50 mM Tris-HCl pH 8.0, 100 mM NaCl, 10 mM imidazole). Initial nickel-based purification was described by the manufacture (Merck). Briefly, the lysate was loaded on the buffer A-equilibrated Ni2+-NTA (the flow-through as FT), and incubated for 2 hours at 4 °C. After two rounds of washing with 10-column volumes of buffer A (W1, W2), the binding proteins were eluted with buffer B (50 mM Tris-HCl pH 8.0, 100 mM NaCl, 5% glycerol, 500 mM imidazole) for 6 times, each with 500 μl of buffer B (E1-E6). All the eluents were combined and EDTA was added to a final concentration of 1 mM. Then the protein mixture was incubated with 250 μl of anti-FLAG M2 agarose gel (Sigma) equilibrated with buffer C (50 mM Tris-HCl, pH 7.5, 150 mM NaCl) for 2 hours at 4 °C. After a short centrifugation (500 rpm, 3 min), the supernatant (FT) was discarded and the pellet was washed 3 times with 1 ml of buffer C (W1). After centrifugation at 500 rpm for 3 min, the supernatant was discarded as much as possible. 500 μl of 20 mM Tris-HCl (pH 8.0) buffer was added to the pellet and boiled for 5 min for protein elution, and this process was repeated for 2 times (E1-E3). All the eluents were combined and the purified proteins were digested with trypsin and identified by HPLC/MS/MS as described before37.
References
Hopwood, D.A. Streptomyces in Nature and Medicine: The Antibiotic Makers (Oxford University Press; 1 edition) (2007).
Niu, G., Chater, K. F., Tian, Y., Zhang, J. & Tan, H. Specialised metabolites regulating antibiotic biosynthesis in Streptomyces spp. FEMS Microbiol Rev 40, 554–73 (2016).
Bai, C. et al. Exploiting a precise design of universal synthetic modular regulatory elements to unlock the microbial natural products in Streptomyces. Proc Natl Acad Sci USA 112, 12181–6 (2015).
Flardh, K. & Buttner, M. J. Streptomyces morphogenetics: dissecting differentiation in a filamentous bacterium. Nat Rev Microbiol 7, 36–49 (2009).
Chaudhary, A. K., Dhakal, D. & Sohng, J. K. An insight into the “-omics” based engineering of streptomycetes for secondary metabolite overproduction. Biomed Res Int 2013, 968518 (2013).
Liu, G., Chater, K. F., Chandra, G., Niu, G. & Tan, H. Molecular regulation of antibiotic biosynthesis in Streptomyces. Microbiol Mol Biol Rev 77, 112–43 (2013).
Martin, J. F. & Liras, P. Engineering of regulatory cascades and networks controlling antibiotic biosynthesis in Streptomyces. Curr Opin Microbiol 13, 263–73 (2010).
Kieser, T., Bibb, M.J., Butter, M.J., Chater, K.F. & Hopwood, D.A. Practical Streptomyces Genetics, (The John Innes Foundation, 2000).
McCormick, J. R. & Flardh, K. Signals and regulators that govern Streptomyces development. FEMS Microbiol Rev 36, 206–31 (2012).
Bush, M. J., Chandra, G., Bibb, M. J., Findlay, K. C. & Buttner, M. J. Genome-Wide Chromatin Immunoprecipitation Sequencing Analysis Shows that WhiB Is a Transcription Factor That Cocontrols Its Regulon with WhiA To Initiate Developmental Cell Division in Streptomyces. mBio 7, e00523–16 (2016).
Biarnes-Carrera, M., Breitling, R. & Takano, E. Butyrolactone signalling circuits for synthetic biology. Curr Opin Chem Biol 28, 91–8 (2015).
Higo, A., Hara, H., Horinouchi, S. & Ohnishi, Y. Genome-wide distribution of AdpA, a global regulator for secondary metabolism and morphological differentiation in Streptomyces, revealed the extent and complexity of the AdpA regulatory network. DNA Res 19, 259–73 (2012).
Urem, M., Swiatek-Polatynska, M. A., Rigali, S. & van Wezel, G. P. Intertwining nutrient-sensory networks and the control of antibiotic production in Streptomyces. Mol Microbiol 102, 183–95 (2016).
Sola-Landa, A., Rodriguez-Garcia, A., Amin, R., Wohlleben, W. & Martin, J. F. Competition between the GlnR and PhoP regulators for the glnA and amtB promoters in Streptomyces coelicolor. Nucleic Acids Res 41, 1767–82 (2013).
Martin, J. F. et al. Cross-talk of global nutritional regulators in the control of primary and secondary metabolism in Streptomyces. Microb Biotechnol 4, 165–74 (2011).
Allenby, N. E., Laing, E., Bucca, G., Kierzek, A. M. & Smith, C. P. Diverse control of metabolism and other cellular processes in Streptomyces coelicolor by the PhoP transcription factor: genome-wide identification of in vivo targets. Nucleic Acids Res 40, 9543–56 (2012).
Vo, T. V. et al. A Proteome-wide Fission Yeast Interactome Reveals Network Evolution Principles from Yeasts to Human. Cell 164, 310–23 (2016).
Bard-Chapeau, E. A. et al. EVI1 oncoprotein interacts with a large and complex network of proteins and integrates signals through protein phosphorylation. Proc Natl Acad Sci USA 110, E2885–94 (2013).
Snider, J. et al. Mapping the functional yeast ABC transporter interactome. Nat Chem Biol 9, 565–72 (2013).
den Hengst, C. D. et al. Genes essential for morphological development and antibiotic production in Streptomyces coelicolor are targets of BldD during vegetative growth. Mol Microbiol 78, 361–79 (2010).
Mao, X. M. et al. Proteasome involvement in a complex cascade mediating SigT degradation during differentiation of Streptomyces coelicolor. FEBS Lett 588, 608–13 (2014).
Mao, X. M. et al. Involvement of SigT and RstA in the differentiation of Streptomyces coelicolor. FEBS Lett 583, 3145–50 (2009).
Mao, X. M. et al. Dual positive feedback regulation of protein degradation of an extra-cytoplasmic function sigma factor for cell differentiation in Streptomyces coelicolor. J Biol Chem 288, 31217–28 (2013).
Donczew, M. et al. ParA and ParB coordinate chromosome segregation with cell elongation and division during Streptomyces sporulation. Open Biol 6, 150263 (2016).
Fuchino, K. et al. Dynamic gradients of an intermediate filament-like cytoskeleton are recruited by a polarity landmark during apical growth. Proc Natl Acad Sci USA 110, E1889–97 (2013).
Richards, D. M., Hempel, A. M., Flardh, K., Buttner, M. J. & Howard, M. Mechanistic basis of branch-site selection in filamentous bacteria. PLoS Comput Biol 8, e1002423 (2012).
Sokolovski, M., Bhattacherjee, A., Kessler, N., Levy, Y. & Horovitz, A. Thermodynamic Protein Destabilization by GFP Tagging: A Case of Interdomain Allostery. Biophys J 109, 1157–62 (2015).
Zhang, F. et al. Probing the impact of GFP tagging on Robo1-heparin interaction. Glycoconj J 31, 299–307 (2014).
Wang, W. et al. Angucyclines as signals modulate the behaviors of Streptomyces coelicolor. Proc Natl Acad Sci USA 111, 5688–93 (2014).
Cooper, H.M. & Paterson, Y. Production of polyclonal antisera. Curr Protoc Cell Biol Chapter 16, Unit 16 2 (2001).
Cooper, H.M. & Paterson, Y. Production of polyclonal antisera. Curr Protoc Immunol Chapter 2, Unit 2 4 (2001).
Zhao, X., Li, G. & Liang, S. Several affinity tags commonly used in chromatographic purification. J Anal Methods Chem 2013, 581093 (2013).
Schmidt, T. G. & Skerra, A. The Strep-tag system for one-step purification and high-affinity detection or capturing of proteins. Nat Protoc 2, 1528–35 (2007).
Terpe, K. Overview of tag protein fusions: from molecular and biochemical fundamentals to commercial systems. Appl Microbiol Biotechnol 60, 523–33 (2003).
Sun, J., Kelemen, G. H., Fernandez-Abalos, J. M. & Bibb, M. J. Green fluorescent protein as a reporter for spatial and temporal gene expression in Streptomyces coelicolor A3(2). Microbiology 145(Pt 9), 2221–7 (1999).
Gerace, E. & Moazed, D. Affinity Purification of Protein Complexes Using TAP Tags. Methods Enzymol 559, 37–52 (2015).
Mao, X. M. et al. Transcriptional Regulation of the Daptomycin Gene Cluster in Streptomyces roseosporus by an Autoregulator, AtrA. J Biol Chem 290, 7992–8001 (2015).
Acknowledgements
We gratefully thank Prof. Mark J. Buttner and Mervyn J. Bibb in the John Innes Centre, Norwich, United Kingdom, for the parental plasmids pIJ8630 and pIJ8660. This work was supported financially by the National Key Research and Development Program (2016YFD0400805), Natural Science Foundation of China (31370103, 31571284) to Xu-Ming Mao, and Natural Science Foundation of China (31520103901, 31470212) to Yong-Quan Li.
Author information
Authors and Affiliations
Contributions
X.-M.M., N.S. and Y.-Q.L. conceived and designed the project, X.-M.M., N.S. and Y.Z. performed the experiments. All the authors analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mao, XM., Sun, N., Zheng, Y. et al. Development of Series of Affinity Tags in Streptomyces . Sci Rep 7, 6854 (2017). https://doi.org/10.1038/s41598-017-07377-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-07377-4
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.