Abstract
Niche-based and neutrality-based theories are two major classes of theories explaining the assembly mechanisms of local communities. Both theories have been frequently used to explain species diversity and composition in local communities but their relative importance remains unclear. Here, we analyzed 57 assemblages of angiosperm trees in 0.1-ha forest plots across China to examine the effects of environmental heterogeneity (relevant to niche-based processes) and spatial contingency (relevant to neutrality-based processes) on phylogenetic structure of angiosperm tree assemblages distributed across a wide range of environment and space. Phylogenetic structure was quantified with six phylogenetic metrics (i.e., phylogenetic diversity, mean pairwise distance, mean nearest taxon distance, and the standardized effect sizes of these three metrics), which emphasize on different depths of evolutionary histories and account for different degrees of species richness effects. Our results showed that the variation in phylogenetic metrics explained independently by environmental variables was on average much greater than that explained independently by spatial structure, and the vast majority of the variation in phylogenetic metrics was explained by spatially structured environmental variables. We conclude that niche-based processes have played a more important role than neutrality-based processes in driving phylogenetic structure of angiosperm tree species in forest communities in China.
Similar content being viewed by others
Introduction
Species in a local community are a subset of those in the species pool of the region where the local community is located. Which species of the regional pool are assembled into the local community depends on the dispersal ability of species in the regional species pool to reach the local community and the degree to which the arrived species can tolerate abiotic and biotic conditions in the local community1. Two major classes of theories have been proposed to explain the assembly mechanisms of local communities. Niche-based theories suggest that deterministic processes (such as environmental filtering and species interactions) determine species composition of local communities2, 3, whereas neutrality-based theories suggest that stochastic processes (such as historical processes that influence the species pool and dispersal limitations) determine the assembly of local communities4. Over the past decades, there has been increasing evidence showing that both deterministic and stochastic processes have played roles in regulating species composition of local communities but the relative importance of these processes remains unclear, partly because these processes are not mutually exclusive and may shift in importance at different spatial scales or in different study contexts1, 5,6,7. The balance between deterministic and stochastic processes in governing species assemblages is influenced by prevailing environmental conditions8. Deterministic processes are expected to be strong drivers of community structure in harsh environments whereas stochastic processes may dominate in local communities with benign environmental conditions8.
On one hand, distributions of species along environmental gradients reflect such traits as cold or drought tolerance. Species–environment interactions are mediated by phenotypic traits, which have evolved to reflect adaptive tradeoffs9, 10. Increasing evidence has shown that some species traits are phylogenetically conserved10, 11. As a result, phylogenetically closely related species are expected to share more similar phenotypic traits with each other than with more distantly related species12. Because interactions between species and their environments are mediated by phenotypic tradeoffs9, 10, one would expect that the maintenance of ancestral traits in some phylogenetic clades and the emergence of derived traits in other phylogenetic clades limit the potential habitat types that can be occupied by different clades13. Processes of habitat filtering would likely exert a strong effect on species assembly and result in patterns of phylogenetic structure in communities. Because ancestral angiosperm clades originated in tropical environment14 and because closely related species are expected to have similar niches15, the phylogenetic niche conservatism hypothesis predicts that angiosperm species in community assemblages would be more phylogenetically related (clustered) in harsher (e.g., colder or dryer) environments, as commonly observed in empirical studies16,17,18.
On the other hand, spatial contingency can cause dispersal limitation of some clades across a region due to neutral dynamics or historical factors19, 20. Empirical studies have found that spatial effects are important in regulating phylogenetic structure21 because dispersal limitations alone can cause closely related species to occupy nearby sites, and environmental variables tend to have a strong spatial structure22.
The relative importance of niche-based deterministic processes (represented by climatic variation) and neutrality-based stochastic processes (represented by spatial variation) in generating phylogenetic community structure can be assessed by variation partitioning approaches21. Specifically, variation in phylogenetic structure can be partitioned into its purely environmental and spatial effects and their joint effects; this should allow one to assess the degree to which the environment and space each govern patterns of phylogenetic structure in community assemblages. If environmental filtering processes play a dominant role in structuring species assemblages, one would expect that the proportion of variation in community structure explained by environmental variables to be greater than that of spatial variables. Alternatively, if spatial contingency is a more important determinant of phylogenetic structure of the species assemblages, one would expect that the proportion of variation explained by spatial variables to be greater than that explained by environmental variables.
In this paper, we assess the relative importance of environmental heterogeneity and spatial contingency in driving phylogenetic structure of angiosperm tree assemblages in China, which covers a wide range of geographical extent and possesses a wide range of variation in climate (e.g., from tropical rain forests northward to boreal forests). Trees are large and thus tree stems and buds are easier to be damaged by cold climate, compared with shrubs and herbaceous plants that may be protected from cold temperature by understory vegetation and/or snow cover during winter; thus, trees are more sensitive to extreme climatic conditions than shrubs and herbs. Adaptation to cold tolerance may be difficult for tropical trees because it requires complex modifications of biochemistry, physiology and morphology to protect stems and buds from cold damage13, 23. Thus, angiosperm trees are an ideal taxonomic system for assessing the relative importance of environmental and spatial variables in shaping phylogenetic structure of community assemblages.
Results
The number of angiosperm tree species per plot varied greatly among the forest plots examined, ranging from 2 to 66 with an average of 21 species per plot. Similarly, environment also varied greatly among the forest plots (Table S1). For example, annual mean temperature ranged from −4.7 to 23.2 °C, and annual precipitation ranged from 442 to 2032 mm (Table S1).
A phylogeny for 187 angiosperm tree genera in the studied forest plots (for illustrative purposes; analyses are based on a species-level phylogeny, see Methods). Genera within a cluster of the same color belong to the same family. The number of species in each genus was indicated by four different symbols: open triangles, 1 species; open circles, 2–5 species; filled triangles, 6–10 species; filled circles, > 10 species.
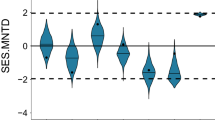
Phylogenetic structure of the forest communities varied across the spatial extent of the study at a varying degree, depending on the particular phylogenetic metric. PD was positively correlated with MPD and negatively correlated with MNTD (P < 0.05 in both cases; Fig. 1); there was no significant relationship between MPD and MNTD (Fig. 1). When the standardized effect sizes of these three metrics were considered, they were positively correlated to each other (P < 0.05 in all cases, Fig. 1).
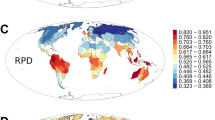
PD, MPD, and MNTD each were significantly correlated with each of the eight environmental variables examined (P < 0.05 in all cases; Table 1). PDses was significantly correlated with seven of the eight environmental variables while MPDses and MNTDses were significantly correlated with only three and one environmental variables, respectively (Table 1). Compared with the three metrics of the standardized effect size, on average, their counterparts were more strongly correlated with environmental variables (Table 1; the mean of absolute values of the 24 correlation coefficients was 0.661 for PD, MPD and MNTD and 0.301 for PDses, MPDses and MNTDses; t-test, P < 0.001).
Of the six best-fit models for the relationships between phylogenetic metrics and environmental variables (Table 2), BIO6 (minimum temperature of the coldest month) was retained in five models and was the most important explanatory variable in four of the five models, as indicated by its standardized regression coefficients. Of the six phylogenetic metrics, environmental variables explained the highest amount (86.4%) of the variation in PD and the lowest amount (28.9%) of the variation in MNTDses (Table 2).
Spatial variables explained less amount of the variation in each phylogenetic metric, compared with environmental variables (compare Table 2 with Table 3). Of the six phylogenetic metrics, spatial variables explained the highest amount (85.8%) of the variation in PD and the lowest amount (14.8%) of the variation in MNTDses (Table 3).
When each phylogenetic metric was simultaneously regressed on the environmental and spatial variables retained in the two best-fit models for the phylogenetic metric (Tables 2 and 3), of the six phylogenetic metrics, environmental and spatial variables explained the highest amount (91.8%) of the variation in PD and the lowest amount (27.9%) of the variation in MNTDses (Fig. 2). Variation partitioning analyses showed that the variation in a phylogenetic metric explained independently by environmental variation was, on average, much larger than that explained independently by spatial structure (14% versus 4%; Fig. 2), and that of the variation in a phylogenetic metric that was explained by the environmental and spatial variables, the vast majority was explained by spatially structured environmental variation (i.e., the amount of the variation in a phylogenetic metric that was explained by the environmental and spatial variables jointly; Fig. 2). For example, environmental and spatial variables jointly explained 80.4% of the variation in PD (Fig. 2).
Histogram of each phylogenetic metric and relationships between phylogenetic metrics. An ellipse includes 75% of the sample. The center of each ellipse is the sample mean of the x and y variables in the plot, and its axes are determined by the standard deviations of x and y. Values are Pearson’s correlation coefficients (*Indicating P < 0.05).
Discussion
This study is one of the few that assess the relative importance of environmental and spatial factors in driving patterns of phylogenetic structure of species assemblages. Species assemblages examined here were distributed across wide spans in both environmental gradients and spatial extents (e.g., a long latitudinal gradient). These ample extents of environment and space are ideal for examining the relative importance of the two types of driving factors on the formation of phylogenetic structure.
Our study showed that among the eight environmental variables examined, mean annual temperature (BIO1) was most strongly correlated with phylogenetic structure (Table 1). However, when all environmental variables were simultaneously considered in a regression analysis, it is minimum temperature (BIO6) that was retained in the vast majority of the best-fit models. This suggests that minimum temperature is the most important driver of phylogenetic structure patterns for angiosperm tree species, at least in China. This is consistent with the climatic extreme hypothesis23, 24.
We found that MPDses was significantly and positively correlated with temperature, regardless of whether mean annual temperature or minimum temperature was considered. This indicates that angiosperm tree species in a forest community are phylogenetically more closely related with decreasing temperature. This pattern is consistent with the phylogenetic niche conservatism hypothesis, which predicts more phylogenetical clustering in colder environments. We also found that MNTDses was not correlated with any temperature variables (P > 0.05; Table 1). Because MPDses measures phylogenetic relatedness among taxa at both deep and shallow levels within a phylogenetic tree and emphasizes phylogenetic relatedness for major clades (e.g., orders and families) branching at deep nodes whereas MNTDses measures phylogenetic relatedness at a shallower level within the phylogenetic tree among taxa descending from superficial nodes, the fact that temperature is correlated with MPDses but not with MNTDses suggests that cold tolerance evolved at deep divergences in angiosperm evolutionary history.
Our variation partitioning analyses showed that the variation in phylogenetic metrics explained independently by environmental variation was much larger than that explained independently by spatial structure. Our environmental variables included only climate variables. If our environmental variables also included soil variables, which are not available for our forest plots, we believe environmental variables would have even stronger effects on phylogenetic metrics than did spatial variables, enhancing our conclusion. Our finding is consistent with that of Cisneros et al. (2016)25 who found that ∼12% and ∼1% of the variation in phylogenetic structure of bat communities in Costa Rica are independently explained by environmental and spatial variables, respectively. Duarte et al. (2012)26 also demonstrated that the phylogenetic structure of Araucaria forests in Brazil is better explained by environmental factors than by space. In a subtropical forest in China, Liu et al. (2013)27 also found that environmental variables explained more variation than did spatial variables for several tree traits (e.g. maximum height). Taken together, these findings suggest that niche-based processes (e.g., niche availability, species–habitat associations, and physiological limitations to environmental conditions) have played a more important role in driving phylogenetic structure patterns in local communities than neutrality-based processes (e.g., ecological drift, dispersal limitation, differential colonization or extinction dynamics). However, these findings are inconsistent with those of Gavilanez & Stevens (2013)21 who found that purely spatial processes play a stronger role in structuring primate communities than niche mechanisms in tropical America. More studies are needed to determine whether it is general that environment plays a more important role than space in driving phylogenetic structure of plant and animal communities.
Our results showed that the vast majority of the total variation in phylogenetic structure that was explained by environmental and spatial variables was explained jointly by the two types of variables, regardless of which phylogenetic metric was considered. For example, of 91.8% of the variation in PD that was explained by environmental and spatial variables, 80.4% was explained jointly by the two types of variables. Similar results have been found in other studies. For example, in Cisneros et al.’s (2016)25 study for bat communities in Costa Rica, on average, approximately 84% of the variation in phylogenetic structure is explained jointly by environmental and spatial variables. This overlapping effect by environment and space may represent a dispersal effect that is correlated with topography or the joint effect of multiple environmental factors that have a similar spatial structure (i.e., spatially structured environmental variation). It is difficult to tease apart the variation jointly explained by environment and space28, 29.
Methods
Study sites and data collection
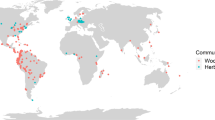
Forest plots used in this study were sampled in 15 protected areas across China (see Fig. S1 in Qian et al. 201618). These areas were distributed across a wide range of geographical space, covering a latitudinal gradient of 35° from tropical rain forests to boreal forests. We sampled four forest plots in each area. Each forest plot was 20 by 50 m (0.1 ha). Location (latitude, longitude, and elevation) of each plot was recorded. Woody individuals with diameter at breast height being 3 cm or larger were identified to species. All species included in this study are native tree species. Three forest plots each included less than two angiosperm tree species and were excluded because some phylogenetic metrics used in this study require at least two species in a forest plot. As a result, 57 forest plots were included in this study. A species list of angiosperm trees was compiled for each forest plot. Botanical nomenclature at the species level was standardized according to the Flora of China30. The 57 plots contained 462 angiosperm tree species in 187 genera and 63 families (Fig. 3).
Results of partial regression analyses partitioning the variation in each phylogenetic metric into four portions: (a) uniquely accounted for by environmental variables, (b) accounted for jointly by environmental and spatial variables, (c) uniquely accounted for by spatial variables, and (d) unexplained variance. For each phylogenetic metric, environmental and spatial variables included in the partial regression analyses are shown in Tables 2 and 3.
Phylogeny reconstruction
The phylogeny used in this study for the 462 species is the same phylogeny as in Qian et al. (2016)18. This phylogeny was generated using an updated version of Zanne et al.’s (2014)31 phylogeny (i.e., PhytoPhylo; Qian & Jin, 2016; available at https://github.com/jinyizju/S.PhyloMaker) as a backbone, which was in turn built based on seven gene regions (i.e., ribosomal cluster regions: 18 S rDNA, 26 S rDNA, ITS, chloroplast genes: matK, rbcL, atpB, and the chloroplast trnL-F intron) and time-calibrated based on fossil data.
Phylogenetic metrics
Faith’s phylogenetic diversity (PD) and Webb’s mean pairwise distance (MPD) and mean nearest taxon distance (MNTD) are commonly used metrics to quantify phylogenetic diversity32,33,34. Accordingly, we used them to quantify phylogenetic diversity in the angiosperm tree communities of this study. All these phylogenetic metrics are based on branch lengths of the phylogeny but they emphasize different depths of evolutionary histories across a phylogeny. PD measures the total amount of phylogenetic distance among species in a community. We used PD metric32 to quantify the phylogenetic diversity of each forest plot as the total phylogenetic branch length joining the basal node (i.e., the basal node of angiosperms in our case) to the tips of all the species in the forest plot. MPD measures the mean phylogenetic distance separating all assemblage members from each other (i.e., a tree-wide assessment of relatedness among co-occurring members) and thus quantifies the overall relatedness of the assemblage members. MNTD measures the average distance to the closest relative for each taxon (i.e., an assessment of terminal relatedness among co-occurring taxa). These three metrics are affected by species richness in community assemblages. To account for species richness, we calculated the standardized effect size (ses) of each phylogenetic diversity metric using the following formula: X ses = (X observed – X randomized )/SD randomized, where X ses represents the standardized effect size of a phylogenetic diversity metric (i.e., PDses, MPDses, or MNTDses), X observed represents the observed value of the metric, X randomized represents the mean of randomized values, and SD randomized represents the standard deviation of the randomized values. The null model that we used to calculate the metrics shuffled the names of taxa across the tips of the phylogeny 999 times. PDses, MPDses and MNTDses are, respectively, PD, MPD and MNTD standardized with respect to a regional species pool; they reflect whether species in an assemblage are overdispersed or clustered with respect to the species pool. For each of PDses, MPDses and MNTDses, a positive value indicates phylogenetic evenness or overdispersion and a negative value indicates phylogenetic clustering. Note that MPDses and MNTDses are, respectively, equal to −1 NRI (net relatedness index) and −1 NTI (nearest taxon index)34. All phylogenetic metrics were calculated using Picante35.
Environmental and spatial variables
We selected eight environmental variables to examine the relationship between environment and phylogenetic structure of the angiosperm tree communities. The environmental variables, which were extracted from the WorldClim database (http://www.worldclim.org)36, are annual mean temperature (BIO1), temperature seasonality (BIO4), maximum temperature of the warmest month (BIO5), minimum temperature of the coldest month (BIO6), annual precipitation (BIO12), precipitation seasonality (BIO15), precipitation of the driest quarter (BIO17), and precipitation of the warmest quarter (BIO18). Previous studies have shown that these environmental variables are among major determinants of animal and plant distributions37.
Spatial variables were represented by eigenvectors derived from principal coordinates of neighbour matrices (PCNM) based on the geographic coordinates of communities22, 38. PCNM allows assessing spatial effects on phylogenetic structure of local communities at multiple scales. PCNM variables were generated by performing a principal coordinate analysis (PCoA) on a truncated distance matrix connecting the forest plots. PCNM vectors were calculated with geographic distances between plots being truncated at the maximum distance connecting all sites (809.7 km), which was determined based on a minimum spanning tree criterion39. Seven PCNM variables resulted from the analysis for the forest plots. We used SAM 4.0 software39 to conduct the PCNM analysis.
Data analysis
We conducted correlation analyses to assess the relationship of each of the six phylogenetic metrics with other five phylogenetic metrics and with the eight environmental variables. For each phylogenetic metric, we built models with all possible combinations of the eight environmental variables and used the corrected Akaike information criterion (AICc) to evaluate performance of each model and to select the model with the lowest AICc as the best-fit model40. Similarly, we built models with all possible combinations of the seven spatial variables (PCNM vectors) and used AICc to select the best-fit model.
For each phylogenetic metric, we conducted a series of partial regressions41 to partition the variance of the phylogenetic metric. We first regressed each phylogenetic metric on the set of environmental variables and the set of spatial variables retained in the two best-fit models separately, and then regressed each phylogenetic metric on the two sets of variables simultaneously. We conducted variation partitioning analyses41 to partition the variation in each phylogenetic metric into four fractions: (1) variation accounted for by environmental variation alone, (2) variation accounted for by environmental and spatial variations jointly, (3) variation accounted for by spatial variation alone, and (4) variation accounted for by neither environmental nor spatial variation.
We used SYSTAT42 to conduct all statistical analyses.
References
Willis, C. G. et al. Phylogenetic community structure in Minnesota oak savanna is influenced by spatial extent and environmental variation. Ecography 33, 565–577 (2010).
Hutchinson, G. E. Concluding remarks. Cold Spring Harb Sym. 22, 415–427 (1957).
Tokeshi, M. Niche apportionment or random assortment—species abundance patterns revisited. J. Anim. Ecol. 59, 1129–1146 (1990).
Hubbell, S.P. The Unified Neutral Theory of Biodiversity and Biogeography. Princeton University Press, Princeton (2001).
Allen, T. F. H. & Starr, T. B. Hierarchy: perspectives for ecological complexity. University of Chicago Press, Chicago (1982).
Weiher, E. & Keddy, P. Ecological assembly rules: perspectives, advances, retreats. Cambridge University Press (1999).
Leibold, M. A. & McPeek, M. A. Coexistence of the niche and neutral perspectives in community ecology. Ecology 87, 1399–1410 (2006).
Chase, J. M. Drought mediates the importance of stochastic community assembly. P. Natl. Acad. Sci. USA 104, 17430–17434 (2007).
Ackerly, D. D. & Reich, P. B. Convergence and correlations among leaf size and function in seed plants: a comparative test using independent contrasts. Am. J. Bot. 86, 1272–1281 (1999).
Brodribb, T. J., Feild, T. S. & Jordan, G. J. Leaf maximum photosynthetic rate and venation are linked by hydraulics. Plant Physiol. 144, 1890–1898 (2007).
Feild, T. S. & Arens, N. C. The ecophysiology of early angiosperms. Plant cell and environment. Plant Cell Environ 30, 291–309 (2007).
Blomberg, S. P. & Garland, T. Tempo and mode in evolution: phylogenetic inertia, adaptation and comparative methods. J. Evolution. Biol 15, 899–910 (2002).
Donoghue, M. J. A phylogenetic perspective on the distribution of plant diversity. P. Natl. Acad. Sci. USA 105, 11549–11555 (2008).
Feild, T. S., Arens, N. C. & Dawson, T. E. The ancestral ecology of angiosperms: emerging perspectives from extant basal lineages. Int. J. Plant Sci. 164, S129–S142 (2003).
Harvey, P. H. & Pagel, M. S. The comparative method in evolutionary ecology. Oxford University Press, Oxford (1991).
Qian, H., Zhang, Y., Zhang, J. & Wang, X. Latitudinal gradients in phylogenetic relatedness of angiosperm trees in North America. Global Ecol. Biogeog 22, 1183–1191 (2013).
Qian, H., Hao, Z. & Zhang, J. Phylogenetic structure and phylogenetic diversity of angiosperm assemblages in forests along an elevational gradient in Changbaishan, China. J. Plant Ecol 7, 154–165 (2014).
Qian, H., Field, R., Zhang, J., Zhang, J. & Chen, S. Phylogenetic structure and ecological and evolutionary determinants of species richness for angiosperm trees in forest communities in China. J. Biogeog. 43, 603–615 (2016).
Cavender-Bares, J., Kozak, K. H., A. Fine, P. V. & Kembel, S. W. The merging of community ecology and phylogenetic biology. Ecol. Lett. 12, 693–715 (2009).
Parmentier, I. & Hardy, O. J. The impact of ecological differentiation and dispersal limitation on species turnover and phylogenetic structure of inselberg’s plant communities. Ecography 32, 613–622 (2009).
Gavilanez, M. M. & Stevens, R. D. Role of environmental, historical and spatial processes in the structure of Neotropical primate communities: contrasting taxonomic and phylogenetic perspectives. Global Ecology and Biogeography 22, 607–619 (2013).
Borcard, D. & Legendre, P. All-scale spatial analysis of ecological data by means of principal coordinates of neighbour matrices. Ecol. Model. 153, 51–68 (2002).
Latham, R. E. & Ricklefs, R. E. Global patterns of tree species richness in moist forests: energy-diversity theory does not account for variation in species richness. Oikos 67, 325–333 (1993).
Grinnell, J. Barriers to distribution as regards birds and mammals. Am. Nat. 48, 248–254 (1914).
Cisneros, L. M., Fagan, M. E. & Willig, M. R. Environmental and spatial drivers of taxonomic, functional, and phylogenetic characteristics of bat communities in human-modified landscapes. PeerJ 4, e2551 (2016).
Duarte, L. D. S., Prieto, P. V. & Pillar, V. D. Assessing spatial and environmental drivers of phylogenetic structure in Brazilian Araucaria forests. Ecography 35, 952–960 (2012).
Liu, X., Swenson, N. G., Zhang, J. & Ma, K. The environment and space, not phylogeny, determine trait dispersion in a subtropical forest. Funct. Ecol. 27, 264–272 (2013).
Borcard, D., Legendre, P. & Drapeau, P. Partialling out the spatial component of ecological variation. Ecology 73, 1045–1055 (1992).
Cottenie, K. Integrating environmental and spatial processes in ecological community dynamics. Ecol. Lett 8, 1175–1182 (2005).
Wu, C.-Y., Raven, P.H. & Hong, D.-Y. Flora of China, vols. 2-25 (Science Press, Beijing and Missouri Botanical Garden Press, St. Louis, 1994–2013).
Zanne, A. E. et al. Three keys to the radiation of angiosperms into freezing environments. Nature 506, 89–92 (2014).
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992).
Webb, C. O. Exploring the phylogenetic structure of ecological communities: an example for rain forest trees. Am. Nat. 156, 145–155 (2000).
Webb, C. O., Ackerly, D. D., McPeek, M. A. & Donoghue, M. J. Phylogenies and community ecology. Annu. Rev. Ecol. Evol. S 33, 475–505 (2002).
Kembel, S. W. et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26, 1463–1464 (2010).
Hijmans, R. J., Cameron, S. E., Parra, J. L., Jones, P. G. & Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. Int. J. Climatol. 25, 1965–1978 (2005).
Davies, R. G. et al. Environmental predictors of global parrot (Aves: Psittaciformes) species richness and phylogenetic diversity. Global Ecol. Biogeog 16, 220–233 (2007).
Borcard, D., Legendre, P., Avois-Jacquet, C. & Tuomisto, H. Dissecting the spatial structure of ecological data at multiple scales. Ecology 85, 1826–1832 (2004).
Rangel, T. F. L. V. B., Diniz-Filho, J. A. F. & Bini, L. M. SAM: a comprehensive application for Spatial Analysis in Macroecology. Ecography 33, 46–50 (2010).
Burnham, K. P. & Anderson, D. R. Model selection and multimodel inference Springer, New York (2002).
Legendre, P. & Legendre, L. Numerical Ecology 2nd edn. Elsevier, Amsterdam (1998).
Wilkinson, L., Hill, M., Welna, J. P., & Birkenbeuel, G. K. SYSTAT for Windows: statistics SYSTAT Inc. Evanston (1992).
Acknowledgements
We thank two anonymous reviewers for their constructive comments, and the financial support of the Special Fund for Environmental Research in the Public Interest (201409055), National Natural Science Foundation of China (41501060) and National Basic Research Program (2009CB421105) to S. Chen.
Author information
Authors and Affiliations
Contributions
H. Qian conceived and designed the study, performed the analyses, and wrote the article. S. Chen contributed the forest plot data and generated Figure 3. J.-L. Zhang participated in calculating phylogenetic metrics and generated Figure 1. All authors participated in revising the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Qian, H., Chen, S. & Zhang, JL. Disentangling environmental and spatial effects on phylogenetic structure of angiosperm tree communities in China. Sci Rep 7, 5634 (2017). https://doi.org/10.1038/s41598-017-04679-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-04679-5
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.