Abstract
Local anesthetics (LAs) inhibit endoplasmic reticulum-associated protein degradation, however the mechanisms remain elusive. Here, we show that the clinically used LAs pilsicainide and lidocaine bind directly to the 20S proteasome and inhibit its activity. Molecular dynamic calculation indicated that these LAs were bound to the β5 subunit of the 20S proteasome, and not to the other active subunits, β1 and β2. Consistently, pilsicainide inhibited only chymotrypsin-like activity, whereas it did not inhibit the caspase-like and trypsin-like activities. In addition, we confirmed that the aromatic ring of these LAs was critical for inhibiting the proteasome. These LAs stabilized p53 and suppressed proliferation of p53-positive but not of p53-negative cancer cells.
Similar content being viewed by others
Introduction
Local anesthetics (LAs) are clinically important for treatment of arrhythmias, control of pain, and suppression of seizures1,2,3. They work by blocking voltage-gated Na+ channels responsible for initiating action potentials in excitable cells to cause the electrical conduction block of the heart and neurons as their acute effect4,5,6,7. LAs reach the binding site within the channel pore between the selective filter and the channel gates to modify the channel conformation through interactions of their aromatic ring with the aromatic group of amino acid residues located in the Na+ channel gate8, 9.
Recently, LAs such as quinidine, mexiletine and aprindine have been reported to stabilize Na+ channels (Nav1.5)10, 11 as well as human ether-go-go-related gene (hERG) K+ channels12, and Kir6.2 proteins13, when administered for a long time. Channel proteins with a short half-life time such as Nav1.514, cystic fibrosis transmembrane regulator15, gap junction channel connexin 4316, 17, Kv1.518, Kir6.213, and hERG12 are degraded through the ubiquitin-proteasome system (UPS). LAs have been suggested to have some effects on the UPS based on the results of microarray analyses showing up-regulation of channel proteins with a short half-life time by these agents19.
P53 is the key molecule for regulating cell proliferation, which is degraded through UPS20, and its accumulation is well-known to induce cellular apoptosis21. Among the anticancer properties of various medicines, it is reported that they modulate the expression of p53 and induce apoptosis, thus LAs-induced modification of the expression of p53 may contribute to developing the new anticancer agents. On the other hand, an increased expression of p53 may explain their harmful effects on the heart. An increase in p53 induces left ventricular dilatation and dysfunction in mice defficient in MDM2, an E3 ligase for p53. Furthermore, it has been demonstrated that p53 is critically involved in pressure overload–induced cardiac dysfunction22. Indeed, in clinical settings, such as in the Cardiac Arrhythmia Suppression Trial (CAST study)23, for preventing arrhythmias, long term treatment with Na+ channel blockers was reported deterioration of the cardiac function accompanied by death of patients with myocardial infarction, suggesting Na+ channel blockers may show deleterious effects on the heart via affecting proteasome activity. Thus, aiming at the development of new anticancer agents as well as at the attenuation of the harmful action of LAs, we explored the action of LAs on the proteasome.
LAs commonly have an aromatic ring with an amido bond. LAs can share this common structure with authentic proteasome inhibitors targeting the chymotrypsine-like site of the 20S proteasome such as MG132 and PS-34124, suggesting that the aromatic ring of the LAs may interact not only with Na+ channels but also with the chymotrypsine-like site of the 20S proteasome. Thus, LAs can access the chymotrypsine-like site of the 20S proteasome to inhibit it. The 20S proteasome consists of four heptameric rings (two outer α rings and two inner β rings), which are made up of seven structurally related α and β subunits, respectively, displaying an α1–7β1–7β1–7α1–7 organization25, 26. Three β1, β2, and β5 subunits of the inner β-rings contain catalytically active threonine residues at their N termini, in which the β1, β2, and β5 subunits are associated with caspase-like, trypsin-like, and chymotrypsin-like activities, respectively. Two pairs of these three active sites face the interior of the cylinder and reside in a chamber formed by the two β rings.
The initial purpose of the present study was to determine whether and how the LAs inhibit the UPS to stabilize the short-lived proteins degraded via proteasomes. We identified the structure-based mechanism of LAs binding to the chymotrypsine-like site of the β5 subunit of the 20S proteasome along with the common structure to exert its Na+ channel blocking action. We also demonstrated that LAs suppress the proliferation of MKN45 cells.
Results
Inhibition of the 20S Proteasome Activity
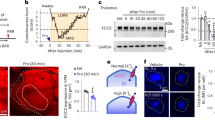
In an attempt to clarify the mechanism of action of LAs, we examined whether LAs inhibited the activity of the 20S proteasome in vitro. LAs with an aromatic ring, such as pilsicainide and lidocaine, inhibited 20S proteasome activity although the extent of inhibition was significantly lower than that attained with MG132 or bortezomib (Fig. 1a). IC50 values were 4.1 ± 2.4 nM, 1.1 ± 1.8 nM, 3.5 ± 1.5 μM, 23 ± 0.2 μM, 20 ± 2.5 μM, and 14 ± 6.3 μM for MG132, bortezomib, pilsicainide, lidocaine, 4-F, and mexiletine, respectively (Explanation of 4-F is described later and supplementary information). Neither the Ca2+ channel blocker verapamil which contains two aromatic rings, nor the K+ channel blocker E-4031, affected the activity of the 20S proteasome (Fig. 1a). Tetrodotoxin (TTX), a Na+ channel blocker without an aromatic ring, did not inhibit 20S proteasome activity, either (Fig. 1a and Supplementary Fig. S1), suggesting that the single aromatic ring of LAs might play a role in the blockade of proteasome activity. Actually, quinidine, mexiletine and aprindine have a single aromatic ring similar to lidocaine and pilsicainide, and both aprindine and mexiletine inhibit the 20S proteasome, although we previously reported that bepridil (Supplementary Fig. S1), a Na+ channel blocker with two aromatic rings, did not show the inhibitory action on the proteasome27. To substantiate this idea, we examined whether modification of the aromatic ring of LAs could also affect 20S proteasome activity. Fluorinated derivatives 2-F, 3-F, 4-F, 2,4-F and 2,6-F with altered electronic properties of the compound28, 29 (Supplementary Table S1 and Fig. S1) partially inhibited the chymotrypsin-like activity of the 20S proteasome (Fig. 1a,b), while M-3 did not (Fig. 1a,b). IC50 values for 2-F, 3-F, 4-F, 2,4-F and 2,6-F were 21.5 ± 46.8 μM, 20.5 ± 21.8 μM, 23 ± 0.2 μM, 23 ± 22 μM, and 28 ± 33 μM, respectively. We also conducted a [3H]-pilsicainide binding assay in the absence and presence of pilsicainide, lidocaine, 4-F as well as M-3. The [3H]-pilsicainide binding assay revealed binding of pilsicainide, lidocaine, and 4-F to the 20S proteasome with K d = 1.0 ± 2.8 μM, K d = 21.2 ± 3.5 μM and K d = 20 ± 3.2 μM, respectively, whereas no binding capacity of M-3 was observed (Fig. 1c).
Effects of LAs on 20S proteasome activity and binding to the 20S proteasome. (a) Effects of LAs and other related chemical agents (■: pilsicainide; ▼: lidocaine; ∆: MG132; ♦: bortezomib; □: verapamil; ○: E-4031; ◊: TTX; ▲: 4-F; ●: M-3) on 20S proteasome activity in vitro. Each point represents the mean ± SEM of more than 7 determinations. 20S proteasome activities were expressed relative to their values in the absence of drugs (100%). The calculated IC50 values were 4.1 ± 2.4 nM, 1.1 ± 1.8 nM, 3.5 ± 1.5 μM, 23 ± 0.2 μM, 20 ± 2.5 μM, 14 ± 6.3 μM for MG132, bortezomib, pilsicainide, lidocaine, 4F, and mexiletine, respectively. (b) Inhibitory action of pilsicainide, 2-F, 3-F, 4-F, 2,4-F, 2,6-F and M-3 at 10 μM (left) and 100 μM (right) on the chymotrypsin-like activity of the proteasome. (c) Inhibition of binding of [3H]-pilsicainide to the 20S proteasome by cold pilsicainide, 4-F, M-3 and lidocaine in vitro. Each point represents the mean ± SEM of more than 7 determinations. The binding fraction of [3H]-pilsicainide to the 20S proteasome was expressed relative to the values in the absence of cold pilsicainide, 4-F, M-3 or lidocaine (100%). K d values for pilsicainide, lidocaine, and 4-F are1.0 ± 2.8 μM, 21.2 ± 3.5 μM, and 4-F, respectively, whereas no binding capacity of M-3 was observed.
Docking Simulations of LAs binding to the 20S proteasome
Automated docking simulations were performed to find the binding mode of pilsicainide to the active site region of bovine proteasomes. Hetényi and van der Spoel have reported a considerable reliability automated docking simulations with the program AutoDock 3.0 in predicting the binding mode of a ligand molecule to a protein without prior knowledge about its binding site30. In this study, we have adopted a two-step method to search a plausible binding site of pilsicainide: firstly, binding results were classified into clusters roughly by their positions and secondly, a cluster was selected that was significantly populated and contained a binding conformation of low energy. Docking simulations of pilsicainide against two segments of the β ring (β1-β2-β3 section and β4-β5-β6 section) showed a significantly low docked energy in a conformational cluster in β4-β5-β6 but not in β1-β2-β3 (Fig. 2a). The energetically most stable and most populated cluster was found in the pocket-like structure of the β5 subunit (Fig. 2b). For pilsicainide binding, one sufficient cluster was found in a grid map around the β5 subunit and no sufficient clusters were found for a grid map around β4 and β6. The cluster found in a grid map around β5 contained 17-binding conformations, including a binding conformation with the lowest energy. The binding mode and the binding conformation with the lowest energy are shown in Fig. 2b. Through interactions of its aromatic ring with the aromatic residue of Tyr136 and of its amido group with both Tyr113 and Gly129, pilsicainide occupied a groove (S1′ pocket) formed by three β-strands at the subunit interface of β4 and β5; i.e., Val26-Met28 of β4, Arg120-Ser122 and Thr125-Val128 of β5, which is located in the vicinity of the nucleophilic attacking residue Thr1 of the subunit (S1 pocket). The K I value (dissociation constant of inhibitor) estimated from the binding energy was determined to be 1 μM, which was a little lower than that from the binding experiment. In contrast to pilsicainide, which showed a well populated and low binding-energy cluster, other analogs tested did not showed such a significant binding clusters. The 4-F derivative was energetically favored but showed less populated clusters against the β4-β5-β6 section compared to pilsicainide, while lidocaine showed only less energetically favored and less populated clusters compared to pilsicainide (Fig. 2c). The conformation of the lowest binding energy in the cluster that lays in the binding pocket showed binding-free energy and a K I value for lidocaine of −7 kcal/mol and 20 μM (Fig. 2a and c), respectively. This suggests that lidocaine has less affinity to the binding site of pilsicainide, which was consistent with the results from their inhibitory action on 20S proteasome activities.
Predicted binding mode of LAs to the proteasome. (a) Docking simulations of pilsicainide against two parts of the β-subunit ring. (b) The binding mode of pilsicainide in the β5 subunit. The approximate position of S1 site pocket is designated. CPK model of pilsicainide and the surrounding protein matrix surface (left panel). Subunits β4 and β5 are colored yellow and orange, respectively. Amino acid residues surrounding the pilsicainide molecule and the nucleophilic active site residue of Thr1 are represented by the stick model and named (right panel). Green broken lines denote putative hydrogen bonds between the ligand and the protein. (c) Docking simulations of pilsicainide and its derivative 4-F, and lidocaine against β4-β5-β6 subunits.
Trypsin-like and caspase-like activities of the wild-type 20S proteasome were not inhibited by pilsicainide (Supplementary Fig. S2). These results were in accordance with the finding that pilsicainide binds only to the β5 catalytic subunit of the 20S proteasome.
Stabilization of Proteins by LAs and MG132
Inhibition of the 20S proteasome results in stabilization of proteins degraded through the UPS. The tumor suppressor p53 is well known to be degraded through the UPS. Like MG132, both pilsicainide and lidocaine increased the expression of p53-FLAG in COS7 cells together with a concomitant accumulation of its ubiquitinated form (Supplementary Fig. S3a). This p53 is ubiquitinated primarily by E3 ligase Mdm231 and its C438A mutant works as a dominant negative form. Co-expression of the C438A mutant of Mdm2 with p53-FLAG abolished the effects of pilsicainide, lidocaine and MG132, providing further evidence for proteasome inhibition by these LAs (Supplementary Fig. S3b). 4-F partially increased the protein expression of p53-FLAG, while M-3 failed to affect it (Fig. 3a,b).
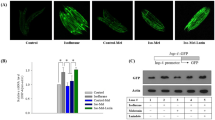
Pilsicainide and lidocaine increase transfected and endogenous p53. (a), (b) Dose-dependent effects of MG132 and LAs on ubiquitinated p53-FLAG protein. Cell lysates were analyzed by Western blot analysis (a). Quantitative analysis of p53-FLAG stability using densitometry is shown in (b). Each point was expressed relative to the values in the absence of drugs (100%). (c), (d) Dose-dependent effects of LAs on endogenous p53 expression in MKN45 cells. MKN45 cells were cultured for 24 h in the presence of pilsicainide, lidocaine, 4-F, M-3 or MG132. Cell lysates were analyzed by Western blot analysis (c). Quantitative analysis of the protein level of endogenous p53 was conducted using densitometry (d).
Given these results, we examined the effects of LAs on the proliferation of MKN45 gastric cancer cells that express p53. Both pilsicainide and lidocaine increased the p53 protein level in MKN45 cells in a dose-dependent manner. 4-F increased p53 a little and M-3 failed to increase it (Fig. 3c,d).
Suppression of Proliferation of Cancer Cells
Furthermore, pilsicainide and lidocaine suppressed proliferation of MKN45 cells in a dose-dependent manner, whereas M-3 did not inhibit their proliferation (Fig. 4a). These effects of pilsicainide and lidocaine were not observed in Kato III gastric cancer cells that do not express p53 (Fig. 4b). To test the inhibitory action of LAs on the proliferation of MKN45 cells in vivo, MKN45 cells were pretreated with lidocaine and then transplanted into the intestine of SOD SKID mice. Lidocaine treatment resulted in suppression of tumor growth (Fig. 4c).
Effects of LAs on cancer cells. (a) Effects of LAs on the proliferation of MKN45 cells. MKN45 cells were cultured for 24 h in the presence of MG132, pilsicainide, lidocaine, 4-F or M-3. Proliferation of MKN43 was estimated by MTT assays. (b) Effects of LAs on the proliferation of MKN45 and Kato III cells cultured for 24 h in the presence of MG132 (10 nM), pilsicainide (1 μM) and lidocaine (20 μM). (c) In the in vivo experiments, lidocaine significantly decreased the proliferation of MKN45 cells dispersed among mouse mesenchymal cells to a similar extent as MG132.
Discussion
In this work, we demonstrated that LAs stabilized the short-lived protein degraded in the cytosol by proteasomes, and LAs with an aromatic ring, such as pilsicainide, bound directly to the proteasome β5 subunit and inhibited its chymotrypsin-like activity, although the affinity to S1′ pocket differed between pilsicainide and lidocaine. Modification of the aromatic ring of pilsicainide attenuated its inhibitory action on the 20S proteasome.
The eukaryotic 20S proteasome has at least three different types of the specificity against substrate polypeptides, i.e., caspase-like (for acidic amino acid residues), trypsin-like (for basic amino acid residues), and chymotrypsin-like (for hydrophobic and aromatic amino acid residues) specificities32. These three specificities have been assigned to three active subunits β1, β2, and β5, respectively33, determined by using synthesized fluorogenic peptides in vitro. The subunit β5 has the chymotrypsin-like activity and mutations in the active site in this subunit were shown to be lethal or to disturb cell growth33. The predicted binding site of pilsicainide was apart from the nucleophilic attacking residue Thr1 of the subunit. From crystallographic25, 34 and modeling studies35, it was reported that inhibitors specific for the proteasome mainly targeted the S1 site.
In our estimated model, however, pilsicainide bound to the groove that is located in the vicinity of S1 site, as shown in Fig. 2. This groove is presumably acting as the S1′ site of proteinase as an analogy of chymotrypsin, and the inhibitor is disturbing the substrate binding. Thus, this S1′ site could be another target for designing a new inhibitor for the proteasome. The inhibitory action of LAs on the chymotrypsin-like activity of the 20S proteasome may come from their common structure. LAs essentially bear an aromatic ring and an amido group in their structure not only for conferring lipophilicity to a drug that allows it to pass through membranes, but also for CH-π or π-stacked binding to the aromatic group of amino acid residues to block Na+ channels activities. Our estimated model demonstrated that pilsicainide bound to the S1′ site via interaction of its aromatic ring with Tyr136 (CH-π or π-stacked binding) and/or interaction with the amido group of Tyr113 and Gly129 (hydrogen bonding). It is well-known that fluoride closely mimics the steric requirement of hydrogen at the enzyme receptor site, but its strong electronegativity significantly alters the reactivity of neighboring centers. Therefore, fluorinated derivatives have been a tool for the investigation of molecular interactions of the chemicals with amino acid residues. Since aromatic residues are capable of forming π-electron interactions with the aromatic drug portion, which are known to be crucial for high-affinity LA block of Na+ channel, aromaticity is crucial for Na+ channel function and pharmacology. Fluorinated derivatives of pilsicainide revealed a decrease in electron-density of their aromatic ring, which perturb not only the interaction with the aromatic group of Tyr136, but also interaction between the amido groups of Tyr113 and Gly129 (Fig. 2b), leading to less potent inhibition of the 20S proteasome activity (Fig. 1b) in comparison with pilsicainide. Thus, either the M-3 compound analogue of pilsicainide or their fluorinated derivatives attenuated both their bind to and block of the 20S proteasome (Fig. 1c), as well as their stabilization of p53-FLAG (Fig. 3). Taken together, the aromatic ring of LAs played a pivotal role by occupying the groove that is located in the vicinity of S1 site to interfere with the access of hydrophobic and aromatic amino acid residues of the targeted peptide to this site.
In this study, we found that LAs with an aromatic ring such as pilsicainide and lidocaine inhibit the proteasome activity. The 20S proteasome is a potential target in cancer therapy36, 37. However, these LAs are not very good inhibitors because their binding affinities were relatively low. Although based on the structure of pilsicainide, it will be possible to design anti-cancer drugs, that are hardly realized. Na+ channel blockers are used widely in the world for treating arrhythmias in patients with heart disease. According to the CAST study23, it is well-recognized that Na+ channel blockers significantly increase the mortality rate in patients with myocardial infarction, while its underlying mechanism remains unknown. Accumulation of p53 in the myocardium has been reported to be associated with chronic heart failure38, 39. Taken together, the present results that Na+ channel blockers, at clinical concentrations, increase p53 via inhibition of the proteasome, Na+ channel blockers could increase the mortality rate of patients with heart failure, which may explain in part the side effects observed in the CAST study. Therefore, the present study may contribute to the development of new selective Na+ channel blockers without inhibitory actions on the proteasome via modification of the aromatic ring, and safer for patients with heart failure. The clinical implications of this novel action of pilsicainide should be a subject of future investigations.
Methods
Plasmids and Expression
pcDNA3/p53-FLAG was a kind gift from Dr. Nakayama (Kyushu University). pcDNA3/Mdm2 (C438A) was a kind gift from Dr. Yasuda (Tokyo Pharmaceutical College). pCR/CMV/hH1 was a kind gift from Dr. Makita (Hokkaido University). COS7 cells were maintained in Dulbecco’s modified Eagle medium (Gibco BRL)/10% fetal bovine serum at 37 °C in a 5% CO2 incubator. Cells were transfected using lipofectamine (Gibco BRL) according to the instructions provided by the manufacturer. Forty-eight hours after transfection, cells were assayed. Proteasome inhibitors or LAs were applied 36 hours after transfection. All drugs except for MG132 were dissolved in culture medium or distilled water. MG132 was dissolved in dimethyl sulfoxide (DMSO). The final concentration of DMSO in the culture or reaction medium was equal to or less than 0.01% v/v. Pilsicainide was kindly provided by Daiichi Suntory Pharmaceutical Co., Ltd. (Osaka, Japan).
Western Blot Analysis and Immunoprecipitation
The same amounts (10 μg) of protein were separated on SDS-PAGE and electro-transferred to a polyvinylidene difluoride membrane. Membranes were probed with antibodies against FLAG (1:1000, Cosmo Bio), GFP (1:1000, Molecular Probes), ubiquitin (1:1000, MBL) or β-actin (1:1000, Oncogene) and were developed using an ECL detection system. Immunoprecipitation was carried out in PBS/1% Triton X-100, 0.5% SDS, 0.25% sodium deoxycholate, 1 mM ethylenediaminetetraacetic acid (EDTA), and 10 μg/ml aprotinin, 10 μg/ml leupeptine, 10 μg/ml pepstatin and 1 mM phenylmethylsulfonylfluoride for 2 h at 4 °C. Immune complexes were collected with protein G agarose (GE Healthcare Life Science) and bound proteins were analyzed by SDS-PAGE followed by immunoblotting.
Measurement of 20S Proteasome Activity
20S Proteasome activity was assessed by fluorescence of free AMC (7-Amino-4-methylcoumarin, excitation: 380 nm, emission: 460 nm) liberated from a substrate peptide (Suc-Leu-Leu-Val-Tyr-MCA). The reaction mixture contained the 20S proteasome (500 μg/ml), the substrate peptide (10 μM) and the indicated drugs in 25 mM HEPES, 0.5 mM EDTA, 0.03% SDS, pH 7.6. The mixture was incubated at 37 °C for 1 h. AMC fluorescence liberated in the absence of drugs was taken as the basal value (100%). The fluorescence liberated in the absence of SDS was taken as the background value (0%)13. Most of the activity assays were conducted using an assay kit (Calbiochem), but some were conducted in hand-made buffer solutions.
Docking Simulation of Compounds of the Bovine 20S Proteasome
Docking simulations were performed using the AutoDock program package, version 3.0.540. The crystal structure coordinates of the bovine proteasome were obtained from Brookhaven Protein Databank [PDB ID: 1IRU26]. The structure coordinates of the fourteen β subunits located in the inner rings of the proteasome were extracted from the data file for the docking simulation. Missing residues in the protein structure were reconstructed using the program Swiss-PDB Viewer41. Polar hydrogen atoms were reconstructed using the program ‘protonate’ supplied with AutoDock distribution. Atomic charges and atomic solvation parameters in the protein were assigned using the programs ‘q.kollua’ and ‘addsol’, respectively. The structure and Mulliken atomic charges of pilsicainide were calculated using Gaussian98 (Gaussian Inc., USA). In the ligand structure used for docking, freedom of torsion at the peptide bond was frozen and all non-polar hydrogen atoms were merged to their bonded carbon atom. Two grid maps were generated to cover three proteinase active subunits; β1, β2, and β5 (designated as H, I, and L in the PDB file, respectively). Grid maps were generated with 0.375 Å spacing within a rectangular volume of 38.6 Å × 47.6 Å × 47.6 Å. Default values were used for parameters of the simulation, except for the ‘maximum number of energy evaluations’ and ‘maximum number of generations’, both of which were set to 107. The Lamarckian genetic algorithm with the pseudo Solis and Wets search was used for the calculations. The results of 100 individual trials were divided into clusters of conformations with an rms tolerance of 1.5 Å. These clusters were aligned with the docked energy of each cluster, which was determined as the lowest docked energy within the cluster. The cluster with a significant population and the lowest docking energy was selected as the set containing the most probable binding conformations. The conformation with the lowest docking energy within this set was selected as the most probable binding conformation.
Ligand Binding Assay
Commercially available 20S proteasome was purchased from Calbiochem. LAs and their derivatives at increasing concentrations (10–8 to 10–2 M) were incubated with [3H]-pilsicainide for 1 h at room temperature in buffer containing 50 mM HEPES supplemented with 1% bovine serum albumin (pH 7.4). Following the reaction termination by addition of 100% tricarboxylic acid, the bound and unbound ligands were separated by centrifugation. Specific binding was defined as the difference between binding capacities in the pellet and supernatant.
Analysis of Data on Dose-Dependent Inhibition of the 20S proteasome
Concentration dependence of the effects of each drug on the stability of the short-lived protein and the 20S proteasome activity was fitted using the following equation:
where, B% represents the changes in expression of the short-lived protein and the 20S proteasome activity induced by drugs at a given concentration of the [drug], and Bmax represents the maximum attainable reduction. IC50 and n are the half-maximal inhibitory concentration and Hill coefficient, respectively.
One-way ANOVA test and Student-t test were carried out to compare the groups. All data are expressed as the mean ±SEM. The level of statistical significance was set at p < 0.05.
References
Vaughan Williams, E. M. A classification of antiarrhythmic actions reassessed after a decade of new drugs. J Clin Pharmacol 24, 129–147, doi:10.1002/j.1552-4604.1984.tb01822.x (1984).
Catterall, W. A. & Mackie, K. The Pharmacological Basis of Therapeutics. Hardman, J. G., Limbird, L. E., Molinoff, P. B., Ruddon, R. W., Gilman, A. G., editors. New York, pp. 331–347 (1996).
Roden, D. M. The Pharmacological Basis of Therapeutics. Hardman J. G., Limbird L. E., Molinoff P. B., Ruddon R. W., Gilman A. G.., editors. New York, pp. 839–874 (1996).
Courtney, K. R. Mechanism of frequency-dependent inhibition of sodium currents in frog myelinated nerve by the lidocaine derivative GEA. J Pharmacol Exp Ther 195, 225–236, PMID:1081138 (1975).
Hille, B. Local anesthetics: hydrophilic and hydrophobic pathways for the drug-receptor reaction. J Gen Physiol 69, 497–515, doi:10.1085/jgp.69.4.497 (1977).
Hondeghem, L. M. & Katzung, B. G. Time- and voltage-dependent interactions of antiarrhythmic drugs with cardiac sodium channels. Biochim Biophys Acta 472, 373–398, doi:10.1016/0304-4157(77)90003-X (1977).
Bean, B. P., Cohen, C. J. & Tsien, R. W. Lidocaine block of cardiac sodium channels. J Gen Physiol 81, 613–642, doi:10.1085/jgp.81.5.613 (1983).
Tsang, S. Y., Tsushima, R. G., Tomaselli, G. F., Li, R. A. & Backx, P. H. A multifunctional aromatic residue in the external pore vestibule of Na+ channels contributes to the local anesthetic receptor. Mol Pharmacol 67, 424–434, doi:10.1124/mol.67.2. (2005).
Kuroda, Y. et al. Interactions between local anesthetics and Na+ channel inactivation gate peptides in phosphatidylserine suspensions as studied by 1H-NMR spectroscopy. Chem Pharm Bull (Tokyo) 48, 1293–1298, doi:10.1248/cpb.48.1293 (2000).
Valdivia, C. R. et al. A novel SCN5A arrhythmia mutation, M1766L, with expression defect rescued by mexiletine. Cardiovasc Res 55, 279–289, doi:10.1016/S0008-6363(02)00445-5 (2002).
Valdivia, C. R. et al. A trafficking defective, Brugada syndrome-causing SCN5A mutation rescued by drugs. Cardiovasc Res 62, 53–62, doi:10.1016/j.cardiores.2004.01.022 (2004).
Ficker, E., Obejero-Paz, C. A., Zhao, S. & Brown, A. M. The binding site for channel blockers that rescue misprocessed human long QT syndrome type 2 ether-a-gogo-related gene (HERG) mutations. J Biol Chem 277, 4989–4998, doi:10.1074/jbc.M107345200 (2002).
Tanaka, H. et al. Proteasomal degradation of Kir6.2 channel protein and its inhibition by a Na+ channel blocker aprindine. Biochem Biophys Res Commun 331, 1001–1006, doi:10.1016/j.bbrc.2005.04.011 (2005).
Staub, O. et al. Regulation of stability and function of the epithelial Na+ channel (ENaC) by ubiquitination. EMBO J 16, 6325–6336, doi:10.1093/emboj/16.21.6325 (1997).
Ward, C. L., Omura, S. & Kopito, R. R. Degradation of CFTR by the ubiquitin-proteasome pathway. Cell 83, 121–127, doi:10.1016/0092-8674(95)90240-6 (1995).
Laing, J. G. & Beyer, E. C. The gap junction protein connexin43 is degraded via the ubiquitin proteasome pathway. J Biol Chem 270, 26399–26403, doi:10.1074/jbc.270.44.26399 (1995).
Beardslee, M. A., Laing, J. G., Beyer, E. C. & Saffitz, J. E. Rapid turnover of connexin43 in the adult rat heart. Circ Res 83, 629–635, doi:10.1161/01.RES.83.6.629 (1998).
Kato, M. et al. Evidence for proteasomal degradation of Kv1.5 channel protein. Biochem Biophys Res Commun 337, 343–348, doi:10.1016/j.bbrc.2005.09.053 (2005).
Raghavendra Prasad, H. S., Qi, Z., Srinivasan, K. N. & Gopalakrishnakone, P. Potential effects of tetrodotoxin exposure to human glial cells postulated using microarray approach. Toxicon 44, 597–608, doi:10.1016/j.toxicon.2004.07.018 (2004).
Suh, K. S. et al. The role of the ubiquitin proteasome system in lymphoma. Crit Rev Oncol Hematol 87, 306–322, doi:10.1016/j.critrevonc.2013.02.005 (2013).
Yu, J. & Zhang, L. The transcriptional targets of p53 in apoptosis control. Biochem Biophys Res Commun 331, 851–858, doi:10.1016/j.bbrc.2005.03.189 (2005).
Toko, H. et al. Ca2+/calmodulin-dependent kinase IIδ causes heart failure by accumulation of p53 in dilated cardiomyopathy. Circulation 122, 891–899, doi:10.1161/CIRCULATIONAHA.109.935296 (2010).
Echt, D. S. et al. Mortality and morbidity in patients receiving encainide, flecainide, or placebo. The Cardiac Arrhythmia Suppression Trial. N Engl J Med 324, 781–788, doi:10.1056/NEJM199103213241201 (1991).
Adams, J. Proteasome inhibition: a novel approach to cancer therapy. Trends Mol Med 8, S49–54, doi:10.1016/S1471-4914(02)02315-8 (2002).
Groll, M. et al. Structure of 20S proteasome from yeast at 2.4 A resolution. Nature 386, 463–471, doi:10.1038/386463a0 (1997).
Unno, M. et al. The structure of the mammalian 20S proteasome at 2.75 Å resolution. Structure 10, 609–618, doi:10.1016/S0969-2126(02)00748-7 (2002).
Suzuki, S. et al. Stabilization of Kv1.5 channel protein by bepridil through its action as a chemical chaperone. Eur J Pharmacol 696, 28–34, doi:10.1016/j.ejphar.2012.09.025 (2012).
Renou, M. & Guerrero, A. Insect parapheromones in olfaction research and semiochemical-based pest control strategies. Annu Rev Entomol 45, 605–630, doi:10.1146/annurev.ento.45.1.605 (2000).
Schlosser, M. Enantiocontrolled Synthesis of Fluoorganic Compounds: Stereochemical Challenges and Biomedicinal Targets, Ed. Soloshonok, V. A. Wiley, Chichester, pp. 653–659 (1999).
Hetényi, C. & van der Spoel, D. Efficient docking of peptides to proteins without prior knowledge of the binding site. Protein Sci 11, 1729–1737, doi:10.1110/ps.0202302 (2002).
Yogosawa, S., Miyauchi, Y., Honda, R., Tanaka, H. & Yasuda, H. Mammalian Numb is a target protein of Mdm2, ubiquitin ligase. Biochem Biophys Res Commun 302, 869–872, doi:10.1016/S0006-291X(03)00282-1 (2003).
Groll, M. & Huber, R. Substrate access and processing by the 20S proteasome core particle. Int J Biochem Cell Biol 35, 606–616, doi:10.1016/S1357-2725(02)00390-4 (2003).
Heinemeyer, W., Fischer, M., Krimmer, T., Stachon, U. & Wolf, D. H. The active sites of the eukaryotic 20S proteasome and their involvement in subunit precursor processing. J Biol Chem 272, 25200–25209, doi:10.1074/jbc.272.40.25200 (1997).
Groll, M., Koguchi, Y., Huber, R. & Kohno, J. Crystal structure of the 20S proteasome:TMC-95A complex: a non-covalent proteasome inhibitor. J Mol Biol 311, 543–548, doi:10.1006/jmbi.2001.4869 (2001).
Smith, D. M. et al. Docking studies and model development of tea polyphenol proteasome inhibitors: applications to rational drug design. Proteins 54, 58–70, doi:10.1002/prot.10504 (2004).
Goldberg, A. L., Stein, R. & Adams, J. New insights into proteasome function: from archaebacteria to drug development. Chem Biol 2, 503–508, doi:10.1016/1074-5521(95)90182-5 (1995).
Dou, Q. P. & Li, B. Proteasome inhibitors as potential novel anticancer agents. Drug Resist Updat 2, 215–223, doi:10.1054/drup.1999.0095 (1999).
Altin, S. E. & Schulze, P. C. p53-upregulated modulator of apoptosis (PUMA): a novel proapoptotic molecule in the failing heart. Circulation 124, 7–8, doi:10.1161/CIRCULATIONAHA.111.036178 (2011).
Sano, M. et al. p53-induced inhibition of Hif-1 causes cardiac dysfunction during pressure overload. Nature 446, 444–448, doi:10.1038/nature05602 (2007).
Morris, G. M. et al. Automated Docking Using a Lamarckian Genetic Algorithm and an Empirical Binding Free Energy Function. J Comp Chem 19, 1639–1662, doi:10.1002/(SICI)1096-987X(19981115)19:14<1639::AID-JCC10>3.0.CO;2-B (1998).
Guex, N. & Peitsch, M. C. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18, 2714–2723, doi:10.1002/elps.1150181505 (1997).
Acknowledgements
This work was supported by grants-in-aid for Scientific Research [13670713(IH), 22570113(YM), 24112008(YS), 26000014(KT)], a grant-in-aid for Target Protein Research [101074100001 (YM, KT)] from the Ministry of Education, Science and Culture of Japan, a grant-in-aid for A-STEP of Japan Science and Technology of Japan [AS242Z02580Q (MU & YM)], a grant-in-aid from the Kato Memorial Bioscience Foundation (MU), a grant-in-aid from the Vehicle Racing Commemorative Foundation (IH), a grant-in-aid from Diponegoro University [SP DIPA-042.01.2.400898 (UB)], and by a Project Research Program under the approval 25P6-1 and 23P2-5 of the Research Reactor Institute, Kyoto University. Synchrotron radiation experiments were conducted under approvals 2011AB6647, 2011AB6650, 2012AB6749, 2012B6754, 2014A6959, 2016AB6649 of SPring-8 and 2011G518 and 2011G519 of PF. We would also like to thank Mr. Takuma Maekawa and members of SPring-8 BL44XU and PF for their help with X-ray intensity data collection, and to Professor Takamitsu Kohzuma of Ibaraki University, for his assistance to YO.
Author information
Authors and Affiliations
Contributions
U.B. purified the yeast 20S proteasome and assayed proteasome activity and wrote the manuscript. M.U. conducted purification of the yeast 20S proteasome, measured the activities of yeast 20S proteasome mutants, and helped to write the manuscript. K.N., A.K., T.M., Y.O. and R.S. performed isolation, purification, and enzyme activity assay. P.L. cultivated transfected cells, and performed western blotting. M.K. contributed to the enzymatic assay of the proteasome. M.I. contributed to the proliferation assay of cancer cells, transplantation of cancer cells in thr animal model. S.T. contributed to cell culture and proliferation of the cancer cells. Y.S. performed yeast cell mutagenesis. H.Y. contributed to the enzyme purification and structure analysis. I.S. and Y.K. executed auto-docking simulation of drugs and the mammalian 20S proteasome. T.I. contributed to drug analogues synthesis and NMR analysis. H.N. contributed to immunoprecipitation assay. I.H., Y.M. and K.T. directed and supervised the research.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bahrudin, U., Unno, M., Nishio, K. et al. Inhibitory effects of local anesthetics on the proteasome and their biological actions. Sci Rep 7, 5079 (2017). https://doi.org/10.1038/s41598-017-04652-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-04652-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.