Abstract
Because of ubiquity of thioesters, thioesterases play a critical role in metabolism, membrane biosynthesis, signal transduction, and gene regulation. In many bacteria, YbgC is such an enzyme, whose coding gene mostly resides in the tol-pal cluster. Although all other proteins encoded in the tol-pal cluster are clearly involved in maintaining cell envelope integrity and cell division, little is known about the physiological role of YbgC. In this study, we identify in Shewanella oneidensis, a γ-proteobacterium used as a research model for environmental microbes, YbgC as a motility regulator. The loss of YbgC results in enhanced motility, which is likely due to the increased rotation rate of the flagellum. The regulatory function of YbgC requires its thioesterase activity but could not be replaced by YbgC homologues of other bacteria. We further show that the regulation of YbgC is mediated by the second message c-di-GMP.
Similar content being viewed by others
Introduction
Gram-negative bacteria are characterized by the presence of an inner membrane (IM) and a distinct outer membrane (OM), separated by a semi-aqueous compartment termed the periplasm where the peptidoglycan (PG) layer resides1. These structures all together constitute the cell envelope, which is essential for survival and proliferation in environments. Integrity and function of the cell envelope in part relies on membrane-associated protein complexes that transport substrate and energy between the environment and the cytoplasm2,3,4. One such system is the Tol-Pal system; its loss causes a variety of phenotypes tied to cell envelope integrity and cell division4,5,6,7,8. Moreover, the systems are essential for the polar localization of some proteins, including chemoreceptors in Escherichia coli and polar localization factor TipN in Caulobacter crescentus 4, 9.
The Tol-Pal system consists of six established members, IM proteins TolA, TolQ, and TolR, periplasmic proteins TolB and YbgF, and OM protein Pal10,11,12. However, the tol-pal gene cluster commonly comprises seven open reading frames, ybgC-tolQ-tolR-tolA-tolB-pal-ybgF, although exceptions (either lacking ybgC or both ybgC and ybgF) are found (Fig. 1)13. To date, there is no evidence to support a role of YbgC in the Tol-Pal system. Operon organizations for these genes differ depending on species, for examples, two operons, ybgC-tolQ-tolR-tolA/tolB-pal-ybgF and ybgC-tolQ-tolR-tolA-tolB/pal-ybgF in E. coli and Pseudomonas putida, respectively14, 15. YbgC proteins, found only in bacteria, belong to the hot-dog thioesterase superfamily16. Consistently, esterase/thioesterase activity (acyl-CoA hydrolase) for YbgC proteins in E. coli, Haemophilus influenzae, and Helicobacter pylori has been demonstrated; as a consequence, it has been proposed that YbgC proteins might be involved in the biosynthesis of species-specific phospholipids17,18,19.
Organization of the tol-pal cluster in representative bacteria. Genes flanking the tol-pal cluster vary. Operon structures for the tol-pal cluster have been determined only in E. coli and P. putida, with promoters shown by arrows. In some bacteria, ybgC is missing in the tol-pal cluster and in extremely rare cases, both ybgC and ybgF are missing. Genes are drawn to scale. BLASTp E-values of S. oneidensis counterparts to E. coli YbgC, TolQ, TolR, TolA, TolB, Pal, and YbgF, are 5e-40, 7e-98, 8e-16, 2e-10, 9e-133, 8e-56, and 1e-22, respectively.
Bacteria have evolved complex mechanisms to control transition between the motile planktonic and sedentary biofilm-associated forms of life in response to both extra- and intra-cellular cues20. A key player in the decision is second messenger bis-(3′-5′)-cyclic dimeric guanosine monophosphate (c-di-GMP), which inhibits flagellar assembly and/or movement while enhancing biosynthesis of extracellular polymeric substance (EPS) required for biofilm formation21. A link between thioesterase and c-di-GMP has been established via diffusible signal factor (DSF) in certain bacteria22, 23.
Shewanella oneidensis, a Gram-negative γ-proteobacterium renowned for its respiratory versatility and enormous potential in bioremediation and microbial fuel cells, is now considered a research model organism in bacterial physiology24, 25. The bacterium is highly motile by virtue of its single polar flagellum, whose assembly has been studied extensively in recent years26,27,28,29,30,31,32. In our previous investigation into spatial and numerical control of flagellar biosynthesis, we revealed many novel features of FlhF, a protein proposed to be a determinant of polar flagellar assembly in polarly flagellated bacteria32. However, mechanisms underlying polar localization of FlhF and the flagellum, remain unknown.
In our continued efforts to determine factors that influence the polar localization of the flagellum in S. oneidensis, we initiated this study by searching for S. oneidensis homologues to polar localization factors established in other bacteria, including the Tol-Pal system. During the investigation, we found that YbgC is involved in motility by regulating c-di-GMP turnover but unlikely to be associated with DSF molecules.
Methods
Bacterial strains, plasmids and culture conditions
All bacterial strains and plasmids used in this study were listed in Table 1. Information for primers used in this study was available upon request. For genetic manipulation, E. coli and S. oneidensis were grown in Lysogeny broth (LB, Difco, Detroit, MI) under aerobic conditions at 37 and 30 °C, respectively. When appropriate, the growth medium was supplemented with chemicals at the following final concentrations: 2, 6-diaminopimelic acid (DAP), 0.3 mM; ampicillin, 50 μg/ml; kanamycin, 50 μg/ml; gentamycin, 15 μg/ml; and streptomycin, 100 μg/ml.
In-frame mutant construction and complementation
In-frame deletion strains for S. oneidensis were constructed using the att-based fusion PCR method as described previously33. In brief, two fragments flanking the gene of interest were amplified by PCR, which were linked by the second round of PCR. The fusion fragments were introduced into plasmid pHGM01 by using Gateway BP clonase II enzyme mix (Invitrogen) according to the manufacturer’s instruction. Verified mutagenesis vectors were maintained in E. coli WM3064, which was used as the donor for subsequent conjugation, resulting in vector transfer into S. oneidensis. Integration of the mutagenesis constructs into the chromosome was selected by resistance to gentamycin and confirmed by PCR. Verified transconjugants were grown in LB broth in the absence of NaCl and plated on LB supplemented with 10% sucrose. Gentamycin-sensitive and sucrose-resistant colonies were screened by PCR for deletion of the target gene. Mutants were verified by sequencing the site for intended mutation.
Mutants used in previous studies have been verified by genetic complementation (Table 1). For newly constructed mutants, plasmid pHG102 was used for genetic complementation27. The coding sequence of the target genes was amplified and inserted into multiple cloning site of pHG102 under the control of the S. oneidensis arcA promoter, which is constitutively active34. For inducible gene expression, gene of interest generated by PCR was introduced into pHGE-Ptac under the control of isopropyl-β-D-1-thiogalactopyranoside (IPTG)-inducible promoter Ptac 35. After sequencing verification, resulting vectors were transferred into the relevant strains via conjugation for complementation and/or expression.
Physiological characterization of S. oneidensis strains
Growth of S. oneidensis strains under aerobic or anaerobic conditions was determined by recording the optical density of cultures at 600 nm (OD600) and by visualizing colonies on plates. MS defined medium containing 0.02% (w/v) of vitamin free Casamino Acids was used as previously described, with 30 mM lactate as electron donor36. For aerobic growth, mid-log cultures were inoculated into fresh medium to an OD600 of ∼0.01 and shaken at 200 rpm at 30 °C. For anaerobic growth, cultures were purged with nitrogen and inoculated into fresh media prepared anaerobically to an OD600 of ∼0.01. The electron acceptor used in this study was fumarate at 20 mM.
Motility testing (swimming) was carried out by spotting 0.5 µl of mid-log phase (∼0.3 of OD600) liquid culture of S. oneidensis strains on LB plates with an agar concentration of 0.25% (w/v). To facilitate comparison, the wild-type strain and/or flagellin-free mutant (FFM) which is nonmotile27, were always included on the same plate. Photograph was taken 16 h after incubation at 30 °C unless otherwise noted. For microscopic analysis, swimming cells were scraped from the leading edges of each swarm, stained for flagellar filaments, and visualized on a glass slide with a Motic BA310 phase-contrast microscope28. Determination of the swimming speed of the cells in liquid media was carried out essentially as described elsewhere37. In brief, after ∼20 µl silicone was dropped onto a microscope slide, the cover slide (60 × 24 mm) was immediately placed on top and evenly pushed down. Slides were dried at room termperature for at least 4 h prior to use. For cell preparation, an aliquot (∼400 µl) of mid-log cultures grown in LB was placed under the cover slide of the microscopic slides and immediately analyzed microscopically. Micrographs were captured with a Moticam 2306 charged-coupled-device camera and Motic Images Advanced 3.2 software.
GFP fusions, visualization and Western blot
To validate protein production, constructs expressing GFP fused to the C-terminal of target proteins were prepared as before35. After verification by sequencing, the vectors were moved into relevant S. oneidensis strains by conjugation. Quantitation of GFP signals was also performed38. In brief, mid-log phase cultures were collected, washed with phosphate-buffered saline containing 0.05% Tween 20, and resuspended in the wash buffer to an OD600 of ∼0.1. One hundred μl of the cell suspensions were transferred into black 96-well plates at various time intervals and fluorescence was measured using a fluorescence microplate reader (M200 Pro Tecan) with excitation at 485 nm and detection of emission at 515 nm. To determine localization of FlhF proteins, FlhF-GFP fusion proteins were used essentially the same as previously described32. For visualization of GFP fusions, cells were prepared with the protocol as described previously35. Slides were stored at 4 °C, and images were collected using a Zeiss ISM710 spectral two-photon confocal microscope.
Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and Western blotting analysis were performed as as previously described35. In brief, The mid-log phase cells were harvested, washed with phosphate buffered saline (PBS), resuspended in the same buffer, and subjected to SDS-PAGE (12%). After membrane transfer for 2 h at 60 V using a Criterion blotter (Bio-Rad), the blotting membrane was probed with the primary antibody Mouse Anti-eGFP-tag Monoclonal Antibody (GenScript) and then the second antibody Goat anti-Mouse IgG-HRP (Horse Radish Peroxidase) (Roche Diagnostics). Detection was performed using a chemiluminescence Western blotting kit (Roche Diagnostics) in accordance with the manufacturer’s instructions and images were visualized with the UVP Imaging System. Protein concentrations of cell lysates were determined using a Bradford assay with bovine serum albumin (BSA) as a standard (Bio-Rad).
Flagellin extraction, purification and analysis
Flagellar filament isolation and purification was performed essentially the same as previously described28. In brief, 250 ml bacterial batch culture was centrifuged at 5000 g for 10 min at 4 °C. The cell pellet was resuspended in 10 ml PBS buffer, pH 7.0 and vortexed for 20 min to shear off flagella. The cell debris was removed by centrifugation at 10,000 g for 30 min at 4 °C and the supernatant containing flagellar filaments was filtered through a 0.45 μm-pore filter. The filtrate was centrifuged at 100,000 g for 2 h and the pellet containing concentrated filaments was resuspended in double distilled H2O. Protein concentrations of each sample were determined by Bradford assay using BCA as standard using the Pierce BCA protein assay kit (Thermo). Protein samples were separated by using SDS-PAGE (10%) and stained with Coomassie brilliant blue as before28. To determine the ratio of flagellins FlaA to FlaB, LC/MS/MS analysis of flagellins was carried out as previously described28, 29.
Expression and purification of the recombinant proteins
The YbgC recombinant proteins were produced and purified to homogeneity as described before32. Briefly, the ybgC gene of S. oneidensis was amplified by PCR with the high-fidelity DNA polymerase Pyrobest (Takara), cloned into pET28a, in which an N-terminal six-His-tag encoding sequence was fused, and verified by sequencing. E. coli BL21(DE3) carrying the vector of interest was grwon in LB at 37 °C to an OD600 of ∼0.5 and induced by the addition of 0.1 mM IPTG for 3 to 4 h at 20 °C. The cells were harvested by centrifugation and disrupted by using a precooled French press (Constant cell disruption system, One Shot model; United Kingdom) at 18,000 lb/in2 for one cycle. After removal of the cell debris, the supernatant containing the His6-tagged recombinant proteins was purified by nickel-nitrilotriacetic acid (Ni2+-NTA)–agarose affinity chromatography using the purification buffers (wash buffer [50 mM NaHPO4, 300 mM NaCl, 40 mM imidazole, 10% glycerol] and elution buffer [50 mM NaHPO4, 300 mM NaCl, 250 mM imidazole, 10% glycerol]) according to the manufacturer’s instructions (GE healthcare). Imidazole and salts were then removed from the eluted fractions by overnight dialysis against 20 mM sodium phosphate buffer (pH 7.5).
Thioesterase activity assay
The thioesterase activity of YbgC was determined by the difference in UV-visible light absorption between the substrate and the hydrolytic product as described previously19. In brief, hydrolysis reactions of the aryl-CoA substrates (Sigma-Aldrich) were monitored at 25 °C by recording the absorbance of 5-thio-2-nitrobenzoate at 412 nm, which was formed by 5,5′-dithio-bis-2-nitrobenzoic acid (DTNB) with CoASH released after acyl-CoA hydrolysis for 20 min. All kinetic measurements were carried out in 200 mM sodium phosphate buffer (pH 7.0) in triplicate at 25 °C. The concentration of the enzyme was adjusted to ensure that consumption of the substrate was less than 5% within the first 3 min of the reaction, during which the initial velocity (v) was measured. Kinetic data were collected by a Synergy 2 Multi-Detection microplate reader (M200 Pro, Tecan) and processed by GraphPad Prism. The kinetic parameters of maximum velocity (v max) and K m were determined using a nonlinear regression fitting from the initial velocity data according to the Michaelis-Menten equation and the k cat value was calculated from the ratio of v max and the concentration of the thioesterase monomer.
DNA synthesis and site-directed mutagenesis
The ybgC genes of H. influenzae and H. pylori as well as the wspR WT and wspR C385T (WspRR129C) genes of Pseudomonas fluorescens SBW25 were synthesized39. For site-directed mutagenesis of YbgC, residues of interest were replaced by intended ones according to the method used before29. Plasmid pHGE-Ptac-ybgC was used as the template with a QuikChange II XL site-directed mutagenesis kit (Stratagene). Mutated PCR products were generated, subsequently digested by DpnI, and transformed into E. coli WM3064. After sequencing verification, the resulting vectors were transferred into the relevant S. oneidensis strains by conjugation.
Expression analysis
Expression of genes of interest was assessed using a single-copy integrative lacZ reporter system and quantitative reverse transcription PCR (qRT-PCR) as described previously31, 40. A fragment covering the sequence upstream of each operon tested from −400 to +1 was then amplified and cloned into the reporter vector pHGEI01, verified by sequencing, and the correct plasmid was then transferred into relevant S. oneidensis strains by conjugation. Once transferred into S. oneidensis strains, pHGEI01 containing promoter of interest integrates into the chromosome and the antibiotic marker is then removed by an established approach41. Cells grown to the mid-log phase under experimental settings were collected and β-galactosidase activity was measured with an assay kit and recorded with a Synergy 2 Multi-Detection microplate reader (M200 Pro, Tecan) as described previously40.
For qRT-PCR, total RNA was isolated from relevant S. oneidensis cells of mid-exponential phase using a combination of Trizon (invitrogen) with the RNeasy Mini Kit (Qiagen) and qRT-PCR analyses were carried out with an ABI7300 96-well system (Applied Biosystems) as described previously42. The expression of each gene was determined from three replicas in a single real-time qRT-PCR experiment. The Cycle threshold (C T ) values for each gene of interest were averaged and normalized against the C T value of the arcA gene, whose abundance was constant under experimental conditions34. Relative abundance (RA) of each gene was standardized to the C T values of both the arcA gene using the equation RA = 2−∆CT, yielding similar fold differences.
Determination of intracellular c-di-GMP levels
Intracellular levels of c-di-GMP were determined with LC-MS using a previously reported procedure with slight modifications43. Swimming cells were scraped from the leading edges of each swarm on semi-solid MS plates, suspended in lysis buffer (40% acetonitrile, 40% methanol, 0.1% formic acid), followed by 15 min of incubation on ice. Insoluble material was removed by 30,000 g for min at 4 °C. The resulting supernatant was collected and analyzed by using liquid chromatography-tandem mass spectrometry on an Exactive hybrid quadrupol-Orbitrap mass spectrometer (Thermo Scientific), coupled with a Thermo Accela UHPLC system. Ions were detected using multiple-reaction monitoring mode. Peaks were integrated manually in Thermo Xcalibur QualBrowser, and relative concentrations of c-di-GMP in all mutants were calculated by normalized to the average level in the wild-type.
Bioinformatics and statistical analyses
Homologues of proteins of interest were identified via a BLASTp search of the NCBI’s nonredundant protein database, using the amino acid sequence as the query. Pairwise and multiple amino acid sequence alignments were performed by using Clustal Omega program44. The relative intensity of specific protein signals on SDS-PAGE was measured using ImageJ45. Student’s t test was performed for pairwise comparisons. Statistical analysis tools integrated in Excel were used to determine correlation of data sets with coefficient of determination (R2) and polynomial regression. Values were presented as means +/− standard error (SE).
Results
S. oneidensis has a Tol-Pal but lacks a counterpart for PaPoc
In addition to the Tol-Pal system, two other systems recently have been proposed to play a critical role in regulation of flagellum polarity in polarly flagellated bacteria46, 47. One is HubP of Vibrio cholerae, which is polarly localized and functions to recruit other polarly posited proteins, including FlhF48. However, functional orthologs (SO_3069 and Sputcn32_2422) of VcHubP in Shewanella are dispensable for flagellar positioning at the pole although it is crucial for normal flagellar function49. The other is the Poc complex (TonB3-PocA-PocB) of Pseudomonas aeruginosa, which plays an essential role in coordinating both polarly located pilus and flagellum50. To test whether S. oneidensis possesses counterparts of E. coli Tol-Pal and PaPoc, we performed a BLASTp search against the S. oneidensis proteome. Clearly, S. oneidensis possesses a complete Tol-Pal system (Fig. 1). In the case of PaPoc, multiple putative homologues were found (Table S1). Based on E-value, protein size, and synteny, among the homologous proteins of PaPocA, it is apparent that SoTolQ is the most likely analogue. More importantly, SoTolR is the only protein that is homologous to PaPocB. Hence, we propose that S. oneidensis possesses a Tol-Pal complex, but may not have a counterpart of PaPoc.
SoYbgC has a role in motility
Tol-Pal appears to be only known system that may be involved in regulating the polar localization of the flagellum in S. oneidensis 4, 9. To test this, attempts were made to construct In-frame deletion mutants for each gene within the tol-pal cluster. Knock-out mutants for all genes except tolA were obtained, implying that TolA is probably essential to S. oneidensis. With respect to growth, all of resulting mutants but ∆SoybgC displayed significantly reduced rate compared with the wild-type (Fig. 2A). Depletion of each of three Tol proteins (TolQ, TolR, and TolB) had effects more substantial than the lack of Pal or YbgF. Importantly, the defects are due to the mutations because cells largely restored normal growth when the respective genes were expressed in trans (Fig. S1).
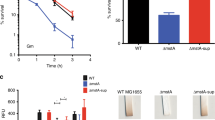
Characteristics of S. oneidensis mutants for genes in the tol-pal cluster. (A) Growth under aerobic conditions. Shown were wild-type (WT) and its isogenic mutants. The fresh medium was inoculated with mid-log phase cultures (∼0.3 of OD600) for each strain. Both LB (shown) and MS defined medium were used and similar results were obtained. ∆tol represents all tol mutants (∆tolQ, ∆tolR, and ∆tolB) because growth defects in these strains were similar. (B) Morphology of cells expressing FlhF-GFP. Cells of indicated strains grown to the mid-log phase in LB broth were examined for morphological phenotype and FlhF localization with a confocal microscope. Arrows highlight clear examples of blebs found in tol mutants but not other mutants. The scale bar represents 1 µm in all panels. (C) Motility on the semi-solid agar plates. Strains indicated were grown to the mid-log phase were spotted on plates along with nonmotile FFM (flagellin-free mutant) and incubated for 16 h. Complementation of the ∆ybgC strain by inducible expression from pHGE-Ptac was performed with IPTG ranging from 0 to 1 mM. Relative motility for each mutant was given by setting the motility of WT (the diameter) as 100%. Asterisks indicate statistically significant differences (*P < 0.05; **P < 0.01; ***P < 0.001; n ≥ 3). All experiments were performed at least three times with either representative results shown or with the standard error of the mean (SEM) presented as error bars.
A common phenotype caused by depletion of one or more components of the Tol-Pal system is that cells grow into cell-chain. Microscopic analysis of these S. oneidensis mutants revealed a similar phenotype, albeit varying in degree on individual mutations and tol mutants being more drastic (longer chain) (Fig. 2B). A more substantial difference found between the tol and pal (and ybgF) mutants was the frequent presence of big blebs at the cell surface of the former, implicating a defect in PG7. In line with the failure in construction of tolA knockout, these results conclude that Tol proteins are more critical to S. oneidensis cell morphology. Again, the SoybgC mutant was indistinguishable from the wild-type in these aspects. In addition, we found that FlhF-GFP fusions were mainly located to the cell pole in both the wild-type and ∆SoybgC strains (Fig. 2B). Despite morphological differences, the fusions in all other mutants were found mainly at the cell pole and/or division sites of the cell chain, suggesting that neither YbgC nor the Tol-Pal system affects FlhF localization. Similar results were obtained from these mutants without carrying the vector expressing FlhF-GFP fusions (Fig. S2).
In the case of motility, all mutants with growth defect displayed heavily impaired ability to move on soft agar plates (Fig. 2C). By comparing to the flagellin-free mutant (ΔflaAB), however, none of mutations completely abolishes motility. Owing to the growth and division defects, whether the reduced motilities associate with the flagellar system is difficult to assess. In contrast, the ΔSoybgC strain clearly had enhanced motility (Fig. 2C). To confirm this, we placed the gene under the control of IPTG-inducible promoter Ptac and assessed effects of its expression at varying levels on motility35. Because Ptac is slightly leaky31, 51 (Fig. S2), in the absence of IPTG the ΔSoybgC strain carrying the cloned gene displayed a significant reduction in motility (Fig. 2C). Expression of the gene at 0.05 mM IPTG restored motility to the wild-type level. Consistently, we found that the SoybgC promoter activity was comparable to that of Ptac at 0.05 mM IPTG by using a lacZ reporter (Fig. S3A). Motility decreased with IPTG concentrations up to 0.2 mM inversely, not necessarily in a proportional manner. But this phenomenon disappeared in the presence of IPTG at further increased levels, possibly due to that the effect of the mutation is saturated. In the case of growth, YbgC in excess did not elicit significant difference compared to the wild-type (Fig. S3B), excluding the possibility that the reduction in motility is a result of retarded growth. Based on all of these data, we conclude that SoYbgC is involved in motility.
Loss of SoYbgC enhances motile capacity of individual cells
Discovery of an unexpected motility phenotype raised an interesting question about the role played by SoYbgC in S. oneidensis. We have previously illustrated that the flagellar assembly in S. oneidensis is governed by an atypical four-tier regulatory system, which differs from that of V. cholerea in that the S. oneidensis FlrBC regulatory system is not essential to motility31. To determine if SoYbgC interferes with the flagellar assembly, we removed the SoybgC gene from strains lacking one of the flagellar regulators, including ΔflrA, ΔrpoN, ΔflrC, and ΔfliA. In the case of regulators that are essential to motility, the additional removal had no effect on motility of these mutants (Fig. 3A). In contrast, in the absence of FlrC, the SoYbgC depletion resulted in enhanced motilities, that are comparable to those of the ∆SoybgC strain (Fig. 3A). Moreover, none of flagellar regulators was found to mediate expression of SoybgC (Fig. S3A). These data, collectively, imply that SoYbgC may not play a role in the flagellar assembly.
S. oneidensis ybgC mutant assembles a normal flagellum. (A) The ybgC mutation did not interfere with regulation of flagellar assembly. Four regulators were under test. Please note that FlrC is not essential to flagellar assembly in S. oneidensis. (B) Estimation of the flagellar length with SDS-PAGE analysis of isolated flagellins, whose levels are proportional to flagellar filament length. Flagellins from the same volume of mid-log phase cultures (adjusted to the same optical density) were extracted and separated on SDS-PAGE. The band intensity was estimated by using ImageJ. Relative flagellin levels were calculated by normalizing to the average level of WT, which was set to 1 for presentation. (C) Estimation of ratio of flagellin FlaA to FlaB with LC-MS/MS analysis of flagellins obtained in B. Aliquots of flagellins were trypsin-digested and analyzed by LC-MS/MS to determine relative abundance of flagellins FlaA and FlaB. All experiments were performed at least three times with either representative results shown or with SEM presented as error bars.
To further investigate the possibility that the hypermotility resulting from the SoYbgC loss is due to changes in the flagellum per se, we examined other flagellar factors that impact motility in S. oneidensis. In the wild-type population grown at the edge of bacterial swarms on semi-solid agar LB plates, approximately 60% were flagellated (Table 2). This percentage is in excellent agreement with the results of previous studies28, 37. In the ΔSoybgC population, a similar portion of cells possessed a flagellum, supporting that the flagellar assembly is not affected by the mutation as suggested above. We then compared the swimming speed of individual cells between the wild-type and ΔSoybgC strains (Table 2). While cells of the wild-type were monitored to swim at ∼53 µm per second, ΔSoybgC cells revealed an increase in swimming rate to ∼68 µm per second, approximately 130% relative to the wild-type, suggesting that the mutation enhances locomotive capacity of individual cells.
In bacteria with a single polar flagellum, the length, composition, and rotation rate of the filament dictate motility. Length of filaments was estimated by SDS-PAGE analysis of flagellin quantities from cells of the similar numbers. Evidently, comparable amounts of flagellins were produced by both the wild-type and ΔSoybgC strains (Fig. 3B), indicating filament length unlikely a critical factor in explaining the motility difference. Furthermore, we examined the ratio of two flagellins, FlaA and FlaB, a factor which also plays an important role in motility31. This is because, with respect to motility, FlaB is predominant but effect of FlaA is negligible27, 29. The consequence is that motility increases with the ratio of FlaB to FlaA31. By using LC/MS/MS, we found that the ratio of two flagellins, based on averaged intensities of unique signature peptides for each flagellin, was not significantly affected by the SoybgC mutation (Fig. 3C). These observations thus manifest that neither the filament length nor the composition is critically altered in the ΔSoybgC strain. Thus, we propose that the rotation rate of the filament is likely accountable for the hypermotility of the SoybgC mutant although we were unable to accurately measure it.
Thioesterase activity is required for the role of SoYbgC in motility
S. oneidensis YbgC is annotated to be a thioesterase and its counterparts in bacteria, such as E. coli, H. influenzae, and H. pylori, show thioesterase activity although their substrates differ17, 19, 52. A sequence analysis revealed a comfortable sequence similarity (against EcYbgC, E-value, 5e-40) between S. oneidensis YbgC and those whose thioesterase activity had been established (Fig. 4A). However, it shares the consensus sequence [DTD-X(2)-GVV-X-H-X(2)-Y] that defines the active site core53, suggesting that the protein likely has thioesterase activity. To test this, we overproduced the recombinant SoYbgC with the 6x His-tag at the N-terminus in E. coli and purified it to homogeneity (Fig. S4). Various acyl-CoAs were used as the substrates in thioesterase activity assay with SoYbgC and DTNB. As shown in Table 3, SoYbgC apparently had a preferred activity for short-chain acyl-CoA substrates such as acetyl-CoA and propionyl-CoA. Thioesterase activity of the enzyme reduced with length of acyl-CoA substrates; both butyryl-CoA and octanoyl-CoA were consumable substrates. In contrast, no activity was detected for lauroyl-CoA or palmitoyl-CoA, indicating that SoYbgC does not work with acyl-CoA substrates that have 12 carbons or more. These data confirm that SoYbgC is an enzyme with thioesterase activity.
Thioesterase activity is required for the role of SoYbgC in motility. (A) Sequence alignment of SoYbgC with YbgC proteins from E. coli, H. influenzae, and H. pylori, whose thioesterase activity has been confirmed. The consensus sequence for the active site core of thioesterase is in bold. Three residues that are essential to thioesterase activity are in red, of which Asp15 is subjected to mutation to create mutants deficient in thioesterase activity. (B) Effect of SoYbgCD15N on motility. Expressed SoYbgCD15N in the ∆ybgC strain under Ptac in the presence of 0.05 mM IPTG could not function as the wild-type with respect to motility. Vec and YbgCWT represent the empty plasmid and the vector carrying the wild-type YbgC, respectively. The experiment was performed at least three times with SEM presented as error bars.
To determine whether thioesterase activity of SoYbgC is required for motility regulation, we made attempts to express mutant proteins whose thioesterase activity is abolished. According to reports on other YbgC proteins16, Tyr11 and the active site core residue Asp15 and His22 in SoYbgC are crucial residues for hydrolysis of acyl-CoA substrates (Fig. 4A). In particular, the essentiality of Asp15 to thioesterase activity has been firmly established on the findings that the replacement of the Asp15 counterpart by Asn inactivates such enzymes, such as HiYbgC and a Pseudomonas thioesterase17, 54. Hence, we performed site-directed mutagenesis for generating SoYbgCD15N. In the ΔybgC strain, production of the resulting mutant proteins was driven by the Ptac promoter as for the wild-type SoYbgC used above. In the presence of 0.05 mM IPTG, a concentration at which the wild-type motility was restored with the wild-type SoYbgC, the ΔybgC strain producing SoYbgCD15N remained similarly hypermotile (Fig. 4B). To confirm that the mutant protein loses thioesterase activity, we purified SoYbgCD15N in the same manner as SoYbgC and found that it was virtually unable to catalyze the hydrolysis of acetyl-CoA (Table 3). Based on these data, we conclude that thioesterase activity is essential to the regulatory role of SoYbgC in motility.
Regulation of motility by YbgC proteins is not universal
To date, this is the first report that suggests a link between an YbgC thioesterase and motility. There are two other YbgC-family thioesterases encoded in the genome, SO_1256 (137 a.a.) and SO_4375 (144 a.a.), which are similar in length and share modest sequence similarities to SoYbgC (E-value of BLASTp, 9e-13 and 4e-11, respectively) (Fig. S5). To determine whether the hypermotility phenotype of the ∆SoybgC strain is attributable to any YbgC-family thioesterase, we constructed strains lacking either SO_1256 or SO_4375 and examined their motility. Clearly, difference in motilities of these two mutants and the wild-type was insignificant (Fig. 5), manifesting that SoYbgC is the only member of YbgC-family thioesterases that participates in motility regulation.
Unique features of SoYbgC may account for its regulatory role in motility. Effect of various thioesterases on motility. Various thioestereases, including YbgC proteins from E. coli, H. influenzae, and H. pylori, under Ptac were expressed in the ∆ybgC strain in the presence of 0.05 mM IPTG. The experiment was performed at least three times with SEM presented as error bars.
We next addressed whether YbgC proteins in other bacteria can function as SoYbgC with respect to motility regulation. To this end, we performed heterogeneous complementation with HiYbgC, HpYbgC, and EcYbgC as we did with SoYbgC, all of which are proven to have thioesterase activity16. Production of all these three proteins was confirmed by using GFP fusion and GFP fusions were likely to be functional because GFP-SoYbgC complemented the phenotype of ∆SoybgC as effective as SoYbgC (Fig. S6). In presence of 0.05 mM IPTG, none showed any effect on motility of the ΔybgC strain (Fig. 5), ruling out the possibility that these YbgC proteins are able to regulate motility. All together, these findings implicate that SoYbgC has some unique characteristics that influence motility.
The SoybgC mutation reduces intracellular c-di-GMP levels
Thioesterases are essential in biosynthesis of diffusible signal factor (DSF), which contributes to bacterial virulence, formation of biofilms, antibiotic tolerance, and various types of locomotion55, 56. To test whether DSF factors are associated with the pheontype caused by the ybgC mutation in S. oneidensis, we examined relevant processes57, 58; based on the results (Fig. S7), we propose that S. oneidensis is unlikely to produce DSF molecules.
Second messenger molecule c-di-GMP controls a variety of cellular processes, including motility, to mediate transition between motile and sedentary forms of bacterial life21. Recent studies have linked c-di-GMP to thioesterases via new sensor proteins for perception of DSF-family signals that modulates c-di-GMP turnover23, 59. Despite the lack of DSF in S. oneidensis, these findings motivated us to look for a possible relationship between SoYbgC and intracellular c-di-GMP levels in S. oneidensis. Intracellular levels of c-di-GMP were measured by LC/MS-MS. Depletion of SoYbgC caused a substantial decrease in the intracellular c-di-GMP concentration, only ∼30% relative to that of the wild-type (Fig. 6A). When the SoybgC gene was expressed in trans, c-di-GMP levels increased with IPTG concentrations, but this effect was not observed in the ΔybgC strain producing SoYbgCD15N (Fig. S8). These data indicate that SoYbgC mediates c-di-GMP homeostasis and thioesterase activity is essential to this role.
SoYbgC-mediated regulation of motility functions through c-di-GMP. (A) Relative intracellular levels of c-di-GMP measured by LC-MS/MS. P. fluorescens WspRR129C, a constitutive active c-di-GMP synthetase, under Ptac was expressed in the wild-type and ∆ybgC strains in the presence of IPTG at indicated levels. The levels of c-di-GMP in WT carrying the empty vector were set to 1. (B) Relative motility. Motilities of the strains cultivated under the same condition as in A were compared. In both A and B, a second-order polynomial best fit for each strain is given and con represents the experimental condition that both strains carry the empty vector.
To confirm that c-di-GMP is responsible for the hypermotility phenotype of the SoybgC mutant, we manipulated intracellular c-di-GMP levels by producing Pseudomonas fluorescens WspRR129C to varying levels39. WspRR129C, a mutant of WspR which is a diguanylate cyclase, is constitutively active. When produced, WspRR129C increased intracellular c-di-GMP concentrations in both the wild-type and ∆SoybgC strains with IPTG (Fig. 6A). However, R2 (coefficient of determination) values for the wild-type and ∆SoybgC strains were 0.98 and 0.85, respectively. This difference suggests that factors other than YbgC are also involved in c-di-GMP homeostasis under experimental conditions. In line with this, the responding patterns of c-di-GMP concentrations to IPTG levels, based on polynomial regression, between these two strains were also apparently different. In the wild-type, c-di-GMP concentrations increased with IPTG (up to 0.2 mM) nearly in a linear manner. On the contrary, effects of WspRR129C on c-di-GMP concentrations in the ∆ybgC strain were much more drastic with IPTG at relatively high levels (0.1 and 0.2 mM) than those with low levels (no more than 0.05 mM), implying that depletion of YbgC counteracts the activity of WspRR129C. In the case of motility, expression of WspRR129C reduced motilities of the wild-type and ∆SoybgC strains with IPTG levels (Fig. 6B), a scenario consistent with the notion that c-di-GMP inhibits motility60. Evidently, the SoybgC mutant was more resistant to WspRR129C overproduction, especially when IPTG levels were low. These data, collectively, conclude that the SoybgC mutation enhances c-di-GMP degradation in S. oneidensis.
Discussion
Thioesterases catalyze hydrolysis of the thioester bond between a carbonyl group and a sulfur atom16. As thioesters are widely found in a variety of metabolites from numerous biological processes, thioesterases play a critical role in metabolism, membrane biosynthesis, signal transduction, and gene regulation16. Based on enzyme function and substrate specificities, thioesterases are grouped into 23 families almost unrelated to one another by primary structure, of which YbgC and YbgC-like proteins constitute number nine, namely TE916. The majority of thioesterases, including YbgC, have a hot-dog fold, whose signature is a five-stranded antiparallel β–sheet around an elongated α–helix61.
A recent report about the E. coli Tol-Pal system has revealed that YbgF coordinates envelope machines facilitating septal PG synthesis and OM constriction (Tol system), leaving YbgC the only protein encoded in the tol-pal cluster without a clearly defined role12. Compared to established Tol-Pal members, YbgC bears two significant differences. First, YbgC is located in the cytoplasm, contrasting IM-associated TolA, TolQ, and TolR, OM-associated Pal, and soluble TolB and YbgF in the periplasm10, 12; hence it may not directly work with these Tol-Pal members at the IM constriction site of the periplasmic side. Second, in certain bacteria the ybgC gene is missing in the tol-pal cluster12. Proteins in the hotdog fold superfamily are characterized by the low sequence homology but the same structural fold, leading to a weak correlation between the sequence similarity and protein function53. Take E. coli as an example. Although its YbgC is the closest homologue to a cyanobacterial hotdog fold thioesterase involved in phylloquinone biosynthesis, it does not play an equivalent role. Rather, YdiI, one of eight other hotdog fold thioesterases, performs the function52. In addition, EntH and YdiI, two hotdog fold thioesterases highly homologous to each other (E-value, 5e-48; identity, 60%), carry out completely different physiological roles52. As a consequence, whether a genuine ybgC gene is present in the bacteria with an ybgC-free tol-pal cluster remains enigmatic.
YbgC proteins, found only in bacteria, have acyl-CoA hydrolase activity16. S. oneidensis YbgC, as revealed in this study, hydrolyze primarily short-chain acyl-CoA thioesters. While YbgCs with a similar preference have been reported, others favor long-chain acyl-CoA thioesters17, 19. However, there is a caveat for the statement: all thioesterases under examination are genuine YbgC. SoYbgC and HiYbgC, representatives for the former, are no doubt YbgC proteins because their coding genes are clustered with the tol-pal genes17. The identity of HpYbgC, a representative for the latter, remains uncertain as it is not associated with the tol-pal genes19.
Although YbgC proteins are firmly established to be thioesterase, their cellular role of YbgC proteins has remained elusive. We showed that S. oneidensis YbgC is linked to motility, a phenomenon relying on its thioesterase activity. None of HiYbgC, EcYbgC, and HpYbgC is able to complement the SoYbgC loss. The failure with HpYbgC is reasonable because their substrates are distinct, but the same result with HiYbgC is unexpected. However, this provides further evidence to support the characteristics of hotdog fold family proteins, that is, a weak correlation between the sequence similarity and protein function53. It is worth mentioning that SoYbgC, as well as the Tol-Pal proteins, are not required for flagellar positioning.
Proteins possessing thioesterease activity that have been previously implicated a role in motility are those responsible for DSF generation, including RpfF of some bacteria such as Xanthomonas and DfsA of B. cepacia that are active with acyl-ACP rather than acyl-CoA25, 62. Based on the lack of homologue to RpfF or DfsA, the negligible role of the spent medium from a SoYbgC overproducing strain on motility, and the irrelevance of a SofabA mutation in the phenotype of the SoybgC mutant, we propose that S. oneidensis is unlikely to produce DSF molecules.
Enhanced motility of the SoybgC mutant is attributed to reduced c-di-GMP levels. In addition, the mutation counteracts the effects of expressed c-di-GMP synthetase, especially at low amounts. It is therefore conceivable that the SoybgC mutant has a stronger c-di-GMP degradation capability. Cyclic di-GMP is synthesized by diguanylate cyclases, characterized by a canonical GGDEF motif and hydrolyzed by phosphodiesterases, characterized by conserved EAL or HD-GYP motifs, respectively21. S. oneidensis is renowned for its large repertoire of proteins involved in c-di-GMP turnover and signaling, including 51 diguanylate cylases, 27 phophodiesterases, and 20 hybrid diguanylate cylase or phophodiesterase proteins24. Many of these proteins contain additional domains, such as Che, Per-Arnt-Sim (PAS), and NIT domains, which are important signaling modules shown to respond to various environmental and cellular cues60, 63. For example, the PAS domain of Burkholderia cenocepacia RpfR, a hybrid protein with domains organization of PAS-GGDEF-EAL, accounts for sensing a DSF family signal and subsequently mediates c-di-GMP turnover23. We expect one or some of these proteins may link the YbgC function with c-di-GMP. Efforts to test this notion are under way.
References
Silhavy, T. J., Kahne, D. & Walker, S. The bacterial cell envelope. Cold Spring Harb Perspect Biol 2, a000414, doi:10.1101/cshperspect.a000414 (2010).
Braun, V. Energy‐coupled transport and signal transduction through the Gram‐negative outer membrane via TonB‐ExbB‐ExbD‐dependent receptor proteins. FEMS Microbiol Rev 16, 295–307 (1995).
Lloubès, R. et al. The Tol-Pal proteins of the Escherichia coli cell envelope: an energized system required for outer membrane integrity? Res Microbiol 152, 523–529 (2001).
Santos, T. M., Lin, T. Y., Rajendran, M., Anderson, S. M. & Weibel, D. B. Polar localization of Escherichia coli chemoreceptors requires an intact Tol-Pal complex. Mol Microbiol 92, 985–1004, doi:10.1111/mmi.12609 (2014).
Bernadac, A., Gavioli, M., Lazzaroni, J. C., Raina, S. & Lloubès, R. Escherichia coli tol-pal mutants form outer membrane vesicles. J Bacteriol 180, 4872–4878 (1998).
Heilpern, A. J. & Waldor, M. K. CTXphi infection of Vibrio cholerae requires the tolQRA gene products. J Bacteriol 182, 1739–1747 (2000).
Llamas, M. A., Ramos, J. L. & Rodríguez-Herva, J. J. Mutations in each of the tol genes of Pseudomonas putida reveal that they are critical for maintenance of outer membrane stability. J Bacteriol 182, 4764–4772 (2000).
Dubuisson, J. F., Vianney, A., Hugouvieux-Cotte-Pattat, N. & Lazzaroni, J. C. Tol-Pal proteins are critical cell envelope components of Erwinia chrysanthemi affecting cell morphology and virulence. Microbiology 151, 3337–3347, doi:10.1099/mic.0.28237-0 (2005).
Yeh, Y.-C., Comolli, L. R., Downing, K. H., Shapiro, L. & McAdams, H. H. The Caulobacter Tol-Pal complex is essential for outer membrane integrity and the positioning of a polar localization factor. J Bacteriol 192, 4847–4858, doi:10.1128/jb.00607-10 (2010).
Derouiche, R., Benedetti, H., Lazzaroni, J., Lazdunski, C. & Lloubes, R. Protein complex within Escherichia coli inner membrane. TolA N-terminal domain interacts with TolQ and TolR proteins. J Biol Chem 270, 11078–11084, doi:10.1074/jbc.270.19.11078 (1995).
Bouveret, E., Bénédetti, H., Rigal, A., Loret, E. & Lazdunski, C. In vitro characterization of peptidoglycan-associated lipoprotein (PAL)–peptidoglycan and PAL–TolB interactions. J Bacteriol 181, 6306–6311 (1999).
Gray, A. N. et al. Coordination of peptidoglycan synthesis and outer membrane constriction during Escherichia coli cell division. eLife 4, e07118, doi:10.7554/eLife.07118 (2015).
Sturgis, J. N. Organisation and evolution of the tol-pal gene cluster. J Mol Microbiol Biotech 3, 113–122 (2001).
Vianney, A. et al. Characterization of the tol-pal region of Escherichia coli K-12: translational control of tolR expression by TolQ and identification of a new open reading frame downstream of pal encoding a periplasmic protein. J Bacteriol 178, 4031–4038 (1996).
Llamas, M. A., Ramos, J. L. & Rodríguez-Herva, J. J. Transcriptional organization of the Pseudomonas putida tol-oprL genes. J Bacteriol 185, 184–195, doi:10.1128/jb.185.1.184-195.2003 (2003).
Cantu, D. C., Chen, Y. & Reilly, P. J. Thioesterases: A new perspective based on their primary and tertiary structures. Protein Science 19, 1281–1295, doi:10.1002/pro.417 (2010).
Zhuang, Z., Feng, S., Martin, B. M. & Dunaway-Mariano, D. Dunaway-Mariano D. The YbgC protein encoded by the ybgC gene of the tol-pal gene cluster of Haemophilus influenzae catalyzes acyl-coenzyme A thioester hydrolysis. FEBS lett 516, 161–163 (2002).
Gully, D. & Bouveret, E. A protein network for phospholipid synthesis uncovered by a variant of the tandem affinity purification method in Escherichia coli. PROTEOMICS 6, 282–293, doi:10.1002/pmic.200500115 (2006).
Angelini, A., Cendron, L., Goncalves, S., Zanotti, G. & Terradot, L. Structural and enzymatic characterization of HP0496, a YbgC thioesterase from Helicobacter pylori. Proteins 72, 1212–1221, doi:10.1002/prot.22014 (2008).
McDougald, D., Rice, S. A., Barraud, N., Steinberg, P. D. & Kjelleberg, S. Should we stay or should we go: mechanisms and ecological consequences for biofilm dispersal. Nat Rev Micro 10, 39–50 (2012).
Römling, U., Galperin, M. Y. & Gomelsky, M. Cyclic di-GMP: the first 25 years of a universal bacterial second messenger. Microbiol Mol Biol Rev 77, 1–52, doi:10.1128/mmbr.00043-12 (2013).
Ryan, R. P. et al. Cell–cell signaling in Xanthomonas campestris involves an HD-GYP domain protein that functions in cyclic di-GMP turnover. Proc Natl Acad Sci USA 103, 6712–6717, doi:10.1073/pnas.0600345103 (2006).
Deng, Y. et al. Cis-2-dodecenoic acid receptor RpfR links quorum-sensing signal perception with regulation of virulence through cyclic dimeric guanosine monophosphate turnover. Proc Natl Acad Sci USA 109, 15479–15484, doi:10.1073/pnas.1205037109 (2012).
Fredrickson, J. K. et al. Towards environmental systems biology of Shewanella. Nat Rev Micro 6, 592–603 (2008).
Fu, H., Yuan, J. & Gao, H. Microbial oxidative stress response: Novel insights from environmental facultative anaerobic bacteria. Arch Biochem Biophys 584, 28–35, doi:10.1016/j.abb.2015.08.012 (2015).
Paulick, A. et al. Two different stator systems drive a single polar flagellum in Shewanella oneidensis MR-1. Mol Microbiol 71, 836–850 (2009).
Wu, L., Wang, J., Tang, P., Chen, H. & Gao, H. Genetic and molecular characterization of flagellar assembly in Shewanella oneidensis. PLoS One 6, e21479, doi:10.1371/journal.pone.0021479 (2011).
Sun, L. et al. Posttranslational modification of flagellin FlaB in Shewanella oneidensis. J Bacteriol 195, 2550–2561, doi:10.1128/jb.00015-13 (2013).
Sun, L. et al. Two residues predominantly dictate functional difference in motility between Shewanella oneidensis flagellins FlaA and FlaB. J Biol Chem 289, 14547–14559, doi:10.1074/jbc.M114.552000 (2014).
Shi, M. et al. Exoprotein production correlates with morphotype changes of nonmotile Shewanella oneidensis mutants. J Bacteriol 195, 1463–1474, doi:10.1128/jb.02187-12 (2013).
Shi, M., Gao, T., Ju, L., Yao, Y. & Gao, H. Effects of FlrBC on flagellar biosynthesis of Shewanella oneidensis. Mol Microbiol 93, 1269–1283, doi:10.1111/mmi.12731 (2014).
Gao, T., Shi, M., Ju, L. & Gao, H. Investigation into FlhFG reveals distinct features of FlhF in regulating flagellum polarity in Shewanella oneidensis. Mol Microbiol 98, 571–585, doi:10.1111/mmi.13141 (2015).
Jin, M. et al. Unique organizational and functional features of the cytochrome c maturation system in Shewanella oneidensis. PLoS One 8, e75610, doi:10.1371/journal.pone.0075610 (2013).
Gao, H. et al. Physiological roles of ArcA, Crp, and EtrA and their interactive control on aerobic and anaerobic respiration in Shewanella oneidensis. PLoS One 5, e15295, doi:10.1371/journal.pone.0015295 (2010).
Luo, Q., Dong, Y., Chen, H. & Gao, H. Mislocalization of rieske protein PetA predominantly accounts for the aerobic growth defect of tat mutants in Shewanella oneidensis. PLoS One 8, e62064, doi:10.1371/journal.pone.0062064 (2013).
Shi, M., Wan, F., Mao, Y. & Gao, H. Unraveling the mechanism for the viability deficiency of Shewanella oneidensis oxyR null mutant. J Bacteriol 197, 2179–2189, doi:10.1128/jb.00154-15 (2015).
Bubendorfer, S. et al. Specificity of motor components in the dual flagellar system of Shewanella putrefaciens CN-32. Mol Microbiol 83, 335–350 (2012).
Jin, M., Fu, H., Yin, J., Yuan, J. & Gao, H. Molecular underpinnings of nitrite effect on CymA-dependent respiration in Shewanella oneidensis. Front. Microbiol. 7, 1154, doi:10.3389/fmicb.2016.01154 (2016).
Spiers, A. J. & Rainey, P. B. The Pseudomonas fluorescens SBW25 wrinkly spreader biofilm requires attachment factor, cellulose fibre and LPS interactions to maintain strength and integrity. Microbiology 151, 2829–2839, doi:10.1099/mic.0.27984-0 (2005).
Fu, H., Jin, M., Ju, L., Mao, Y. & Gao, H. Evidence for function overlapping of CymA and the cytochrome bc 1 complex in the Shewanella oneidensis nitrate and nitrite respiration. Environ Microbiol 16, 3181–3195, doi:10.1111/1462-2920.12457 (2014).
Fu, H. et al. Crp-dependent cytochrome bd oxidase confers nitrite resistance to Shewanella oneidensis. Environ Microbiol 15, 2198–2212, doi:10.1111/1462-2920.12091 (2013).
Yuan, J., Wei, B., Shi, M. & Gao, H. Functional assessment of EnvZ/OmpR two-component system in Shewanella oneidensis. PLoS One 6, e23701, doi:10.1371/journal.pone.0023701 (2011).
Waters, C. M., Lu, W., Rabinowitz, J. D. & Bassler, B. L. Quorum sensing controls biofilm formation in Vibrio cholerae through modulation of cyclic di-GMP levels and repression of vpsT. J Bacteriol 190, 2527–2536, doi:10.1128/jb.01756-07 (2008).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7, 539–539, doi:10.1038/msb.2011.75 (2011).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9, 671–675, doi:10.1038/nmeth.2089 (2012).
Kazmierczak, B. I. & Hendrixson, D. R. Spatial and numerical regulation of flagellar biosynthesis in polarly flagellated bacteria. Mol Microbiol 88, 655–663 (2013).
Treuner-Lange, A. & Søgaard-Andersen, L. Regulation of cell polarity in bacteria. J Cell Biol 206, 7–17 (2014).
Yamaichi, Y. et al. A multidomain hub anchors the chromosome segregation and chemotactic machinery to the bacterial pole. Genes Dev 26, 2348–2360 (2012).
Rossmann, F. et al. The role of FlhF and HubP as polar landmark proteins in Shewanella putrefaciens CN-32. Mol Microbiol 98, 727–742, doi:10.1111/mmi.13152 (2015).
Cowles, K. N. et al. The putative Poc complex controls two distinct Pseudomonas aeruginosa polar motility mechanisms. Mol Microbiol 90, 923–938 (2013).
Fu, H., Jin, M., Wan, F. & Gao, H. Shewanella oneidensis cytochrome c maturation component CcmI is essential for heme attachment at the non-canonical motif of nitrite reductase NrfA. Mol Microbiol 95, 410–425, doi:10.1111/mmi.12865 (2015).
Chen, M. et al. Identification of a hotdog fold thioesterase involved in the biosynthesis of menaquinone in. Escherichia coli. J Bacteriol 195, 2768–2775, doi:10.1128/JB.00141-13 (2013).
Dillon, S. C. & Bateman, A. The Hotdog fold: wrapping up a superfamily of thioesterases and dehydratases. BMC Bioinform 5, 109–109, doi:10.1186/1471-2105-5-109 (2004).
Thoden, J. B., Holden, H. M., Zhuang, Z. & Dunaway-Mariano, D. X-ray crystallographic analyses of inhibitor and substrate complexes of wild-type and mutant 4-hydroxybenzoyl-CoA thioesterase. J Biol Chem 277, 27468–27476, doi:10.1074/jbc.M203904200 (2002).
Ryan, R. P., An, S.-q, Allan, J. H., McCarthy, Y. & Dow, J. M. The DSF family of cell–cell signals: an expanding class of bacterial virulence regulators. PLoS Pathogens 11, e1004986, doi:10.1371/journal.ppat.1004986 (2015).
Bi, H., Yu, Y., Dong, H., Wang, H. & Cronan, J. E. Xanthomonas campestris RpfB is a fatty acyl-CoA ligase required to counteract the thioesterase activity of the RpfF diffusible signal factor (DSF) synthase. Mol Microbiol 93, 262–275, doi:10.1111/mmi.12657 (2014).
Parsons, J. B. & Rock, C. O. Bacterial lipids: Metabolism and membrane homeostasis. Prog Lipid Res 52, 249–276, doi:10.1016/j.plipres.2013.02.002 (2013).
Luo, Q., Shi, M., Ren, Y. & Gao, H. Transcription factors FabR and FadR regulate both aerobic and anaerobic pathways for unsaturated fatty acid biosynthesis in Shewanella oneidensis. Front Microbiol 5, 736, doi:10.3389/fmicb.2014.00736 (2014).
McCarthy, Y. et al. A sensor kinase recognizing the cell–cell signal BDSF (cis-2-dodecenoic acid) regulates virulence in Burkholderia cenocepacia. Mol Microbiol 77, 1220–1236, doi:10.1111/j.1365-2958.2010.07285.x (2010).
Chao, L., Rakshe, S., Leff, M. & Spormann, A. M. PdeB, a cyclic di-GMP-specific phosphodiesterase that regulates Shewanella oneidensis MR-1 motility and biofilm formation. J Bacteriol 195, 3827–3833, doi:10.1128/jb.00498-13 (2013).
Leesong, M., Henderson, B. S., Gillig, J. R., Schwab, J. M. & Smith, J. L. Structure of a dehydratase–isomerase from the bacterial pathway for biosynthesis of unsaturated fatty acids: two catalytic activities in one active site. Structure 4, 253–264, doi:10.1016/S0969-2126(96)00030-5 (1996).
Cheng, Z. et al. Structural basis of the sensor-synthase interaction in autoinduction of the quorum sensing signal DSF biosynthesis. Structure 18, 1199–1209, doi:10.1016/j.str.2010.06.011 (2010).
Taylor, B. L. & Zhulin, I. B. PAS domains: internal sensors of oxygen, redox potential, and light. Microbiol Mol Biol Rev 63, 479–506 (1999).
Acknowledgements
This research was supported by National Natural Science Foundation of China (41476105), and the Fundamental Research Funds for the central Universities (2015FZA6001, 2016FZA6003).
Author information
Authors and Affiliations
Contributions
H.G. conceived the idea and designed the project. T.G. and Q.M. carried out the experiments. T.G. and H.G. analyzed data. T.G. and H.G. wrote the paper. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gao, T., Meng, Q. & Gao, H. Thioesterase YbgC affects motility by modulating c-di-GMP levels in Shewanella oneidensis . Sci Rep 7, 3932 (2017). https://doi.org/10.1038/s41598-017-04285-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-04285-5
This article is cited by
-
Functional roles of multiple Ton complex genes in a Sphingobium degrader of lignin-derived aromatic compounds
Scientific Reports (2021)
-
Flagellation of Shewanella oneidensis Impacts Bacterial Fitness in Different Environments
Current Microbiology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.