Abstract
The epithelial-mesenchymal transition (EMT) contributes to various processes in cancer progression, such as metastasis and drug resistance. Since we have already established a cell-based reporter system for identifying EMT-suppressive microRNAs (miRNAs) in the pancreatic cancer cell line Panc1, we performed a function-based screening assay by combining this reporter system and a miRNA library composed of 1,090 miRNAs. As a result, we identified miR-509-5p and miR-1243 as EMT-suppressive miRNAs, although the mechanisms for EMT-suppression induced by these miRNAs have yet to be clarified. Herein, we demonstrated that overexpression of miR-509-5p and miR-1243 increased the expression of E-cadherin through the suppression of EMT-related gene expression and that drug sensitivity increased with a combination of each of these miRNAs and gemcitabine. Moreover, miR-509-5p was associated with worse overall survival in patients with pancreatic cancer and was identified as an independently selected predictor of mortality. Our findings suggest that miR-509-5p and miR-1243 might be novel chemotherapeutic targets and serve as biomarkers in pancreatic cancer.
Similar content being viewed by others
Introduction
Pancreatic cancer is the most lethal and common cancer in the world1. According to the National Cancer Research Center Japan, the mortality rate from pancreatic cancer is the fourth highest among all cancers in Japan in 2014 (http://ganjoho.jp/reg_stat/statistics/stat/summary.html). Pancreatic ductal adenocarcinoma (PDAC) accounts for 90% of pancreatic cancers, and its prognosis remains very poor, with a 5-year survival of 7–8% to date. Most PDACs are diagnosed at an already advanced stage because there is currently no reliable diagnostic method for the diagnosis of early stage PDAC. Advanced-stage PDAC is characterized by invasion of the lymph nodes and vasculature or distant metastasis2. The current therapeutic strategies of chemotherapy and/or radiation therapy provide limited effects for such advanced PDAC3. In the last decade, microRNAs (miRNAs) have become firmly established as critical molecules in normal and neoplastic cells4. Thus, a detailed understanding of the miRNA-based molecular mechanisms by which pancreatic cancer is so malignant might provide useful insights into the identification of biomarkers and development of novel therapeutic strategies for this virulent tumor.
Epithelial-mesenchymal transition (EMT) plays important roles in cell differentiation, wound healing, and fibrosis in normal tissue and during embryonic development5. EMT also contributes to cancer progression, including invasion, metastasis and chemo-resistance6. When an epithelial cell is stimulated by EMT inducers, it can transform into a mesenchymal cell with the ability to migrate and invade into other tissues and organs7. This phenotypic transition is triggered by several extracellular signals, such as TGF-β and WNT, resulting in an activation of EMT-promoting transcription factors, such as the ZEB family, the Snail family, and Twist8. In addition, cancer cells can acquire stem cell-like characteristics and chemoresistant properties through EMT9. However, because EMT has phenotypic plasticity, a mesenchymal cell can transform into an epithelial cell through a mesenchymal-to-epithelial transition (MET)10. Taken together, inhibiting EMT is a potential therapeutic strategy for cancers.
miRNAs are endogenous, small non-coding, single-stranded RNAs of 20–25 nucleotides that regulate the expression of target genes at the post-transcriptional level by binding to complementary sequences within the 3′ untranslated region (3′ UTR) of their target gene mRNAs4. An individual miRNA usually has multiple target genes with partially complementary mRNA sequences, whereas a single gene can be targeted by several miRNAs11. miRNAs play crucial roles in the modulation of various biological processes, including EMT12, 13. In addition, miRNAs act as tumor suppressors and/or oncogenes in a cell type-dependent manner in various cancers14. A number of miRNAs function as crucial modulators of cell proliferation, migration and EMT15,16,17. For example, the miR-200 family inhibits EMT through the direct suppression of ZEB1/2 and increases the sensitivity of cancer cells to chemotherapeutic agents18, 19. Using a cell-based reporter assay system with the CDH1 promoter, we identified miR-655 as both an EMT-suppressive miRNA and a predictor for poor prognosis in esophageal squamous cell carcinoma20. EMT-related miRNAs might be important biomarkers for diagnosis of several cancer including pancreatic cancer and contribute to overcoming chemoresistance via combination therapy using anti-cancer drugs11, 21.
In the present study, we identified two novel EMT-suppressive miRNAs, miR-509-5p and miR-1243, through a combination of our cell-based reporter system and a miRNA library containing 1,090 synthetic miRNAs. We also clarified their functions, including the direct targets of each miRNA. Our screening system has already proved useful for the identification of miRNAs related to the phenotypic transformation of EMT20, 22. We demonstrated that miR-509-5p induced an MET phenotype by directly regulating VIM and HMGA2. By contrast, miR-1243 directly regulated SMAD2 and SMAD4, which regulate the TGF-β signaling pathway, resulting in an induction of the MET phenotype. In addition, we found that those miRNAs could increase the sensitivity of the pancreatic cancer cell line Panc1 to gemcitabine. Interestingly, the expression of miR-509-5p was significantly associated with a worse overall survival in patients with pancreatic cancer and was indicated as an independently selected predictor for overall survival. Taken together, our findings suggest that a novel therapeutic strategy for pancreatic cancer might involve a combination of gemcitabine and miR-509-5p or miR-1243, which regulate EMT, and that miR-509-5p might be useful as a prognostic biomarker in pancreatic cancer.
Results
EMT-suppressive miRNAs were extracted using a cell-based reporter system and miRNA library in Panc1 cells
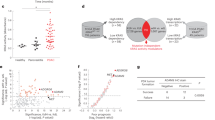
We previously established a cell-based reporter system with the CDH1/E-cadherin promoter in Panc1 cells (Supplementary Fig. S1)20. In the present study, we established two clones (PEcadZsG-Panc1 #1 and #2) that expressed ZsGreen1 protein by increased CDH1 promoter activity in Panc1 cells. To identify EMT-suppressive miRNAs, we performed a function-based screening by combining our cell-based reporter assay system and a miRNA library containing 1,090 miRNAs. A different number of cells were cultured for each clone because each clone had a different cell growth ability. After transfection of each miRNA in this library, we measured the cell growth and fluorescence intensity and calculated the relative fluorescence intensity (RFI) (Fig. 1a). Figure 1b shows the results of this screening in PEcadZsG-Panc1 #1 (upper) and #2 (bottom) 72 hours after transfection with each miRNA. We established the criteria for extracting EMT-suppressive miRNAs (RFI >1.4, cell survival rate >0.65) and then extracted 40 miRNAs from PEcadZsG-Panc1 #1 and 44 miRNAs from PEcadZsG-Panc1 #2 (Fig. 1c and Supplementary Table S1). Six miRNAs, miR-200c, miR-367*, miR-452*, miR-509-5p, miR-660 and miR-1243, which were common in both screening assays, were identified as candidate EMT-suppressive miRNAs (Table 1). miR-367* and miR-452* were excluded from these candidate miRNAs because these miRNAs are expressed at low levels in the body23. We next validated whether the remaining four candidate miRNAs increased the expression of E-cadherin in PEcadZsG-Panc1 #1 and Panc1 cells. Overexpression of miR-200c, miR-509-5p and miR-1243 induced the expression of endogenous E-cadherin in both cell lines (Fig. 1d). Because little has been reported about miR-509-5p and miR-1243 with respect to the EMT phenotype, we focused on the novel function of these miRNAs as EMT-suppressive miRNAs.
EMT-suppressive miRNAs are extracted using the function-based screening by combining the cell-based reporter system and an miRNA library in Panc1 clones. (a) Flow chart of the study. (b) Result of the function-based screening using Pre-miRTM miRNA Precursor Library-Human V15, which contains 1,090 miRNAs mimicking human mature miRNAs. The fluorescence intensity of ZsGreen1 was measured in duplicate with a fluorescence microplate reader. The relative fluorescence intensity in each transfectant was calculated using the fluorescence intensity in negative control cells and was normalized based on the cell survival rate, as measured by the crystal violet stain (Supplementary Table S1). The lower closed arrow indicates the 1090 miRNAs examined. (c) Venn diagram showing the overlap of six miRNAs between PEcadZsG-Panc1 #1 and #2. (d) The expression of E-cadherin in PEcadZsG-Panc1 #1 and parental Panc1 cells transfected with each candidate miRNA that was selected in the screening.
miR-509-5p and miR-1243 induced the MET phenotype through the suppression of ZEB1 and Snail
We first evaluated the expression of miR-509-5p and miR-1243 in 24 pancreatic cancer cell lines. The expression levels of these miRNAs tended to be lower in these cells compared with that in normal pancreatic tissue (Supplementary Fig. S2), suggesting that these miRNAs have tumor suppressive functions. To investigate the mechanism by which these miRNAs induced an MET phenotype, we evaluated the EMT marker genes E-cadherin, Vimentin, ZO-1, ZEB1 and Snail in several cancer cell lines, Panc1, KP4-4, SU.86.86, BxPC3 and MDA-MB-231 (Figs 2 and S3). Overexpression of these miRNAs increased the expression of E-cadherin at both the mRNA and protein levels in Panc1 cells, whereas the expression of Vimentin was reduced by these miRNAs (Fig. 2a–c). In addition, MDA-MB-231 cells, a breast cancer cell line, also exhibited an MET phenotype when transfected with miR-1243, resulting in the upregulation of E-cadherin and the downregulation of Vimentin. However, overexpression of miR-509-5p yielded no increase in the expression of E-cadherin (Fig. 2c). Interestingly, in KP4-4 cells, a SMAD4-depleted pancreatic cancer cell line, the MET phenotype was induced by miR-509-5p but not by miR-1243, suggesting that the mechanism by which the MET phenotype is induced by miR-1243 might depend on the TGF-β signaling pathway (Fig. 2c). Therefore, to probe whether miR-1243 depends on the TGF-β signaling pathway to induce the MET phenotype, we analyzed E-cadherin expression in Panc1 cells using TGF-β recombinant protein, which induces an EMT phenotype through the activation of the TGF-β signaling pathway24. As a result, we observed that the expression of E-cadherin in both miR-NC and miR-509-5p transfectants was suppressed by TGF-β, whereas the effect of TGF-β was weak in miR-1243 transfectants compared with miR-Negative Control (miR- NC) transfectants (Fig. 2d). These results indicate that miR-1243 induced the MET phenotype through the regulation of the TGF-β signaling pathway. We also checked the expression of ZO-1, ZEB1 and Snail, the representative EMT marker genes, in Panc1 cells transfected with each of the miRNAs and observed that the protein expression level of ZO-1 was increased by these miRNAs, whereas that of ZEB1 and Snail was reduced (Fig. 2e).
miR-509-5p and miR-1243 induce an MET phenotype in several cancer cell lines. (a) The expression of miR-509-5p, miR-1243 and EMT marker genes was confirmed by qRT-PCR in Panc1 cells 72 hours after transfection of miR-509-5p and miR-1243; the data are the mean ± SD (bars) for duplicate experiments. (b) Immunofluorescent staining of E-cadherin (green) and DAPI (blue) in Panc1 cells after transfection of 10 nmol/L of miR-NC, miR-509-5p or miR-1243. (c) Western blot analysis of E-cadherin and Vimentin protein levels in Panc1, KP4-4 and MDA-MB-231 cells 72 hours after transfection of 10 nmol/L of miR-NC, miR-509-5p or miR-1243. (d) The expression of E-cadherin in Panc1 cells 72 hours after treatment with or without TGF-β (5 ng/ml) and simultaneous transfection with miR-NC, miR-509-5p and miR-1243. The number under the figure indicates the relative ratio based on the miR-NC expression level. (e) Western blot analysis of the EMT-related proteins ZO-1, ZEB1 and Snail in Panc1 cells 72 hours after transient transfection with 10 nmol/L of miR-NC, miR-509-5p and miR-1243. Student’s t-test was used for statistical analysis, and asterisks represent P < 0.05 versus miR-NC transfectants.
miR-509-5p and miR-1243 suppressed cell migration and invasion and targeted EMT-related genes
To evaluate the functions of miR-509-5p and miR-1243, we utilized several biological approaches, including cell growth, migration and invasion assays. Overexpression of miR-509-5p did not affect cell growth in Panc1 cells, but miR-1243 suppressed cell growth (Fig. 3a). These results were consistent with the first screening (Table 1). The cell migration and invasion abilities in miR-509-5p or miR-1243 transfectants were reduced compared with miR-NC transfectants (Fig. 3b,c). To explore the mechanism through which these miRNAs could suppress migration and invasion abilities, we sought to determine their direct target genes using the TargetScan database. We extracted SMAD2, SMAD4, VIM, ZEB1 and HMGA2 as candidate target genes for each miRNA. SMAD2 and SMAD4 contain target sequences of miR-1243 and are well-known regulators of the TGF-β signaling pathway25, 26. VIM, ZEB1 and HMGA2 contain target sequences of miR-509-5p. HMGA2 is an EMT-promoting factor in several cancer types27, 28. We first checked the protein levels of these genes in miR-509-5p or miR-1243 transfectants using western blot analysis and observed that their protein expression levels were decreased in miR-509-5p or miR-1243 transfectants compared with miR-NC transfectants (Figs 2c,e and 3d). To determine whether these candidate target genes are directly regulated by miR-509-5p or miR-1243, we constructed reporter plasmids for each target gene (Supplementary Fig. S4a–c). We performed a luciferase reporter assay using plasmids containing the wild type or mutant 3′ UTR of SMAD2, SMAD4, VIM, ZEB1 and HMGA2. We detected significant reductions in luciferase activity in wild-type constructs but not in mutant constructs for SMAD2, SMAD4, VIM and HMGA2 (Fig. 3e). miR-509-5p could not bind to ZEB1 (Supplementary Fig. S4b), indicating that SMAD2 and SMAD4 were direct targets of miR-1243 and VIM and that HMGA2 was a direct target of miR-509-5p. To evaluate the biological effects of knockdown of these miRNAs, we performed several biological experiments with KMP3 and CFPAC1 cells, which show higher expression of miR-509-5p and miR-1243 than Panc1 cells. However, knockdown of each miRNA did not affect the EMT phenotype, cell growth, migration and invasion abilities (Supplementary Fig. S5).
miR-509-5p and miR-1243 suppress cell motility and invasion through an MET phenotype alteration and directly target VIM, HMGA2, SMAD2 and SMAD4. (a) The number of viable cells 24–96 hours after transfection with 5 nmol/L of miR-NC, miR-509-5p and miR-1243 in Panc1 cells was assessed by the WST-8 assay. Each data point represents the mean of triplicate experiments (bars, SD). (b,c) Transwell migration and invasion assays were performed in 24-well modified Boyden chambers without and with Matrigel, respectively. Panc1 cells (4 × 104 cells per well [migration and invasion assay]) that were transfected with each miRNA were transferred into the upper chamber, and the migrated or invaded cells on the lower surface of the filters were fixed, stained and counted after 24 hours of incubation. Experiments were performed in triplicate, and each data point represents the mean (bars, SD). (d) Western blot analysis of SMAD2, SMAD4 and HMGA2 in Panc1 cells 72 hours after transfection with 10 nmol/L of miR-NC, miR-509-5p and miR-1243. (e) Results of the luciferase reporter assays in Panc1 cells after co-transfection of the pmirGLO Dual-Luciferase vectors containing wild-type (WT) of SMAD2, SMAD4, VIM and HMGA2 or mutant variants of these genes and miR-NC, miR-509-5p and miR-1243. Student’s t-test was used for statistical analysis, and asterisks represent P < 0.05 versus miR-NC transfectants.
Suppression of HMGA2 inhibited cell migration and invasion abilities via an MET phenotype change
To investigate whether these direct target genes of each miRNA induce an MET phenotype, we performed knockdown experiments using specific siRNAs against these direct target genes. We first confirmed that the expression of E-cadherin in si-SMAD2, si-SMAD4 and si-HMGA2 transfectants was increased compared with the si-NC transfectant in Panc1 cells, resulting in the induction of an MET phenotype (Fig. 4a). The cell growth rate was not affected by knockdown of these genes (Fig. 4b). We next investigated the cell migration and invasion abilities in each of the siRNA transfectants. Knockdown of SMADs did not reduce the migration and invasion abilities, suggesting that the inhibition of cell migration and invasion by miR-1243 might be unrelated to the expression of SMADs (Fig. 4c,d). However, the altered MET phenotype could be induced by suppressing SMADs. In addition, we analyzed the effect of TGF-β in each of the siRNA-transfected cells and observed that suppressing SMADs reduced the effect of TGF-β (Supplementary Fig. S6). Although knockdown of SMADs did not affect cell proliferation under the addition of recombinant TGF-β, knockdown of SMAD4 or SMAD2/SMAD4 reduced the migration and invasion. Therefore, these results were consistent with the miR-1243 overexpressed phenotype, indicating that miR-1243 increased the expression of E-cadherin by regulating SMADs (Fig. 4e). On the other hand, miR-509-5p induced an MET phenotype by regulating HMGA2 (Fig. 4e).
Suppression of HMGA2 inhibits cell motility and invasion through an MET phenotype alteration, whereas these abilities are not affected by suppression of SMADs. (a) Knockdown of SMAD2, SMAD4, SMAD2 plus SMAD4 (left) and HMGA2 (right) via specific siRNAs was confirmed by western blotting in Panc1 cells. The protein expression of these endogenous genes was downregulated by each specific siRNA compared with control siRNA. (b) The number of viable cells 24–72 hours after transfection of each siRNA was assessed by the WST-8 assay and is presented as the mean ± SD (bars) for triplicate experiments. (c,d) Transwell migration and invasion assays were performed in 24-well modified Boyden chambers without and with Matrigel, respectively. siRNA-transfected Panc1 cells (4 × 104 cells per well [migration and invasion assay]) were transferred into the upper chamber, and the migrated or invaded cells on the lower surface of the filters were fixed, stained and counted after 24 hours of incubation. Experiments were performed in triplicate, and each data point represents the mean (bars, SD). Student’s t-test was used for statistical analysis, and asterisks represent P < 0.05 versus si-NC transfectants. (e) Model for the miR-509-5p- and miR-1243-mediated pathway in EMT.
miR-509-5p and miR-1243 enhanced the effect of gemcitabine
Because EMT can contribute to the resistance of anti-cancer drugs29, we evaluated whether overexpression of these miRNAs increased their sensitivity to gemcitabine, an anti-cancer drug that is classified as an anti-metabolite. After treatment with several concentrations of gemcitabine (0.1, 1, 10, 50, 100 and 200 nmol/L) and PBS as a control in Panc1 cells, we measured real-time cell growth for 120 hours using the xCELLigence RTCA DP system (ACEA Biosciences, San Diego, CA, USA) (Fig. 5). In 0.1 nmol/L gemcitabine-treated cells, cell growth did not change in each miRNA transfectant at 120 hours post-treatment (Fig. 5a), whereas treating the miR-509-5p and miR-1243 transfectants with 100 nmol/L gemcitabine showed an enhanced effect of gemcitabine compared with miR-NC transfectants (Fig. 5b). Moreover, we measured the IC50 of gemcitabine in each miRNA transfectant (miR-NC: 119 nmol/L, miR-509-5p: 65.3 nmol/L, miR-1243: 27.8 nmol/L) (Fig. 5c). Together, our data show that inducing an MET phenotype by miR-509-5p and miR-1243 enhanced the cytotoxic effect of gemcitabine. Thus, a combination of each of these miRNAs and gemcitabine might be a novel therapy for pancreatic cancer.
Overexpression of miR-509-5p and miR-1243 enhances the effect of gemcitabine. (a,b) Effect of a combination of miRNA and gemcitabine on cell index curves (RTCA) in Panc1 cells. A total of 5 nmo/L of miR-NC, miR-509-5p or miR-1243 was transfected, and cells were treated with gemcitabine (0.1, 1, 10, 50, 100 and 200 nmol/L) after 6 hours. Figure 5a shows 0.1 nmol/L gemcitabine-treated cells, and Fig. 5b shows 100 nmol/L gemcitabine-treated cells. Cell indexes were normalized with the last time point before treatment with gemcitabine. (c) Representative curves of the growth-suppressive effects at 120 hours following treatment with gemcitabine in cells transfected with miR-NC, miR-509-5p and miR-1243, and the IC50 in each miRNA transfectant.
The expression level of miR-509-5p was associated with overall survival in patients with pancreatic cancer
To assess whether the expression of miR-509-5p and miR-1243 is associated with disease outcome in patients with pancreatic cancer, we performed miRNA in situ hybridization (ISH) assay in 50 primary pancreatic tumors. We first prepared positive and negative controls by transfecting Panc1 cells with each miRNA and staining with a miR-509-5p or miR-1243 miRNA probe (Figs 6a and S7a). After we confirmed that these miRNA probes were available, 50 primary pancreatic tumors were subjected to RNA ISH using each probe (Figs 6b and S7b). Kaplan-Meier survival estimates showed that low expression of miR-509-5p was significantly associated with worse overall survival in all cases (P = 0.0175, log-rank test) (Fig. 6c), whereas the expression of miR-1243 was not related to the prognosis (P = 0.4842, log-rank test) (Supplementary Fig. S7c). Moreover, we examined the clinicopathological significance of the expression of miR-509-5p and miR-1243 in 50 primary pancreatic tumors based on the RNA ISH staining pattern of each miRNA. We found that the expression of miR-1243 was associated with the venous invasion category (P = 0.03) (Supplementary Table S2). In the Cox proportional hazard regression model (Supplementary Table S3), univariate analyses demonstrated that the expression of miR-509-5p and the lymphatic invasion category in the tumor-lymph node-metastases (TNM) classification were significantly associated with overall survival. In the multivariate analysis using a stepwise Cox regression procedure, the expression of miR-509-5p based on the TNM classification was identified as an independently selected predictive factor for overall survival in both forward and backward procedures (P = 0.0063). We also evaluated correlation between clinical features and each miRNA using the TCGA database, in which expression data of 1881 miRNAs on 178 pancreatic cancer samples have been stored. We examined the correlation of prognosis and expression of miR-509-5p and miR-1243 using 141 samples excluding Stage IV, macroscopic residual tumors (R2) or unevaluable presence of tumors (RX), and tumor-types other than PDAC. However, Kaplan-Meier survival estimates showed the expression of miR-509-5p and miR-1243 not to be associated with the prognosis (P = 0.6975, P = 0.3416, log-rank test, respectively), respectively (Supplementary Fig. S8).
The expression of miR-509-5p is associated with overall survival in human PDAC. (a,b) Representative results of the in situ hybridization for miR-509-5p. (a) FFPE of Panc1 cells 24 hours after transfection with miR-NC (upper) and miR-509-5p (bottom). (b) Primary PDAC with negative staining (upper) and positive staining (bottom). (c) Kaplan-Meier curves for the overall survival rates of patients with primary PDAC. A lack of miR-509-5p expression in tumor cells was significantly associated with a worse overall survival (P = 0.0175, log-rank test).
Discussion
In our previous study, we identified an EMT-suppressive miRNA, miR-655, using a function-based screening assay, which combined a cell-based reporter system with a miRNA library containing 470 miRNAs20. In the present study, we reliably observed that miR-655 increased the expression of E-cadherin and induced an MET phenotype (Supplementary Table S1). The function-based screening assay performed in the current study also identified miR-509-5p and miR-1243 as MET inducers. These two miRNAs were not explored in our previous study because these miRNAs were absent in the miRNA library V3 that was used. Several studies have demonstrated that miR-509-5p suppresses cell proliferation, invasion and migration of cervical cancer, hepatocellular carcinoma, renal carcinoma and breast cancer30,31,32. In addition, a recent study reported that miR-509-5p acts as an EMT-suppressive miRNA and inhibits cell migration and invasion by targeting FOXM1 in non-small cell lung cancer33. However, only a few studies have focused on the correlation between EMT and miR-509-5p. Furthermore, the function of miR-1243 has not yet been clarified.
We revealed that overexpression of miR-509-5p and miR-1243 induced an MET phenotype by downregulating ZEB1 and Snail. However, our experiment using TGF-β showed different functional mechanisms between miR-509-5p and miR-1243: miR-1243 depended on the SMAD signaling pathway, whereas miR-509-5p did not. Thus, these findings suggest that each miRNA has different target genes for the induction of the MET phenotype. We additionally evaluated the EMT marker genes, E-cadherin and vimentin, with two pancreatic cancer cell lines, SMAD4-intact SU.86.86 and SMAD4-homozygously-deleted BxPC3. Although overexpression of miR-509-5p reduced expression of vimentin, overexpression of miR-509-5p or miR-1243 did not show up-regulation of E-cadherin in these cell lines (Supplementary Fig. S3). In general, pancreatic cancer cell lines, which have the EMT plasticity, are very few. In fact, since these cell lines have the epithelial phenotype, they were not induced the MET phenotype by each miRNA. Transfections of each miRNA inhibitor (anti-miRNA) did not affect EMT, cell growth, migration and invasion abilities in CFPAC1 and KMP3 cells (Supplementary Fig. S5). Furthermore, through TCGA database, almost PDAC shows low expression of miR-509-5p and miR-1243. Taken together, we concluded that inhibition of these miRNAs could not induce the EMT phenotype.
We next explored representative EMT-related genes from the Target Scan Human 7.1 database (http://www.targetscan.org/vert_71/) for the direct target-genes of each miRNA, and found that miR-509-5p regulates the expression of VIM and HMGA2 by directly binding to their 3′ UTR regions. We next focused on the function of HMGA2 because it is an important regulator of cell growth, differentiation and EMT27, 28. HMGA2 is highly expressed in embryonic tissues and many malignant tumors, and overexpression of HMGA2 is associated with EMT, metastasis and poor prognosis in several cancers34,35,36. Because recent studies have revealed that some miRNAs, such as miR-101, miR-221, miR-485-5p and let-7a, inhibit EMT by targeting HMGA2, HMGA2 is considered a key regulator of EMT37,38,39,40. In addition, the Lin28b-let-7-HMGA2 axis contributes to the maintenance of the stemness phenotype in hematopoietic cells41. Thus, our findings strongly suggest that miR-509-5p induces an MET phenotype by regulating HMGA2. In addition, low expression levels of miR-509-5p were significantly associated with worse overall survival in pancreatic cancer in our miRNA-ISH assay (Fig. 6). On the other hand, an additional evaluation of the correlation between miR-509-5p expression and the prognosis in patients with PDAC (n = 141 samples) in TCGA showed no significant value (P = 0.6975, log-rank test) in the Kaplan-Meier survival curve analysis (Supplementary Fig. S8). The statistical difference between TCGA and ours may be due to TCGA database not including Japanese PDAC tumors or a difference in the guidelines for the management of patients with pancreatic cancer between Japan and other countries such as the US and Europe.
We also explored direct target genes of miR-1243 and identified SMAD2 and SMAD4 as direct target genes of miR-1243. Inhibition of SMAD2 and/or SMAD4 by miR-1243 or their respective siRNAs increased the expression of E-cadherin. Although cell migration and invasion abilities were suppressed by miR-1243, knockdown of SMAD2 and/or SMAD4 did not affect their function. In general, SMAD2 and SMAD4 are well-known tumor suppressor genes25, 26. SMAD4 is frequently inactivated by genomic alterations, such as deletion or mutation, contributing to cancer progression in pancreatic cancer, colorectal cancer and prostate cancer42,43,44,45. Furthermore, loss of SMAD4 causes head and neck cancer in mice and promotes metastasis in human pancreatic cancer46. In this study, SMAD2 and SMAD4 inhibition showed no significant effect on cell proliferation of Panc1 cells. This finding was indeed consistent with previous studies in which SMAD2 and SMAD4 suppression did not induce cell growth in Panc1 cells in vitro 47, 48. SMAD2 and SMAD4 have an important role of the regulation of EMT/MET in TGF-β pathway. We also evaluated the cell growth, migration and invasion abilities by knockdown of SMAD2 and/or SMAD4 with treatment of recombinant TGF-β. Knockdown of SMADs with TGF-β did not affect cell growth, whereas TGF-β-induced migration and invasion were suppressed by knockdown of SMAD4 or SMAD2/SMAD4 (Supplementary Fig. S6), suggesting that the induction of MET phenotype by miR-1243 might be relevant to the suppression of SMAD2 and SMAD4 which resulted in changing the cellular phenotype in part. However, in the present study, we could not identify direct target-genes of miR-1243, which regulate cell growth. Our findings suggest that the function of miR-1243 in pancreatic cancer is likely dependent on the status of SMAD2 and SMAD4 as well as cancer-related genes other than SMAD2 and SMAD4, further indicating the possible dependence on cellular-context manner14, 49. miR-1243 might have some crucial role in TGF-β pathway via regulating the expression of SMAD2/4 in PDAC with wild-type SMAD2/4.
On the other hand, the expression of miR-1243 in our miRNA-ISH assay was not associated with the prognosis in pancreatic cancer in contrast to miR-509-5p. Since there were only 50 PDAC samples in the present study, we may need to examine the relationship between the prognosis and expression of miR-1243 in large scale with more number of cases.
Although EMT contributes to cancer progression, metastasis and drug resistance6, the mechanism by which it contributes to drug resistance is poorly understood. The role of miRNAs in regulating drug resistance through EMT in cancer also remains unclear. Our findings showed that overexpression of miR-509-5p and miR-1243 increased the sensitivity of pancreatic cancer cells to gemcitabine. This result suggests that patients with high expression of these miRNAs in PDAC may have the sensitivity to gemcitabine compared to patients without the expression of these miRNAs. Thus, the expression status in these two miRNAs might predict gemcitabine efficacy in patients with pancreatic cancer. In addition, since the expression miR-509-5p was associated with the prognosis, miR-509-5p might be useful for a biomarker in the clinical setting of pancreatic cancer. Moreover, interestingly, recent studies have revealed that the inhibition of EMT in lung and pancreatic cancers increase drug sensitivity18, 50. Thus, combinatorial therapeutics with anti-cancer drugs and overexpression of miR-509-5p or miR-1243 might serve as a novel cancer therapy that can overcome chemoresistance. Indeed, overexpression of miR-509-5p and miR-1243 increased the sensitivity of pancreatic cancer cells to gemcitabine. We hypothesize that gemcitabine-induced cytotoxicity was increased via the induction of the MET phenotype through the downregulation of HMGA2, which in turn was caused by the overexpression of miR-509-5p. Because cell death could be induced by a single treatment of miR-1243, the combination of miR-1243 and gemcitabine may have produced a synergistic effect on cell growth, i.e., the MET phenotype and other effects caused by miR-1243 might increase the sensitivity to gemcitabine. Taken together, these results suggest that such combinatorial therapeutics might be useful for chemoresistant cancer.
In conclusion, we have established a cell-based reporter system to monitor the promoter activity of CDH1 and identified miR-509-5p and miR-1243 as EMT-suppressive miRNAs using this system. Overexpression of miR-509-5p and miR-1243 markedly induced the MET phenotype and inhibited cell motility and invasion in vitro through the regulation of the target genes of each miRNA. In addition, miR-509-5p and miR-1243 enhanced the effect of gemcitabine on cell growth. The expression level of miR-509-5p could predict prognosis in patients with pancreatic cancer. Our findings implicate the EMT-suppressive miR-509-5p and miR-1243 as potential therapeutic targets for pancreatic cancer and suggest that miR-509-5p might be a prognostic biomarker. Importantly, our cell-based reporter system, which was used to identify EMT-related molecules, can be utilized with other libraries of cDNA, siRNA, shRNA and therapeutic compounds.
Materials and Methods
Cell Lines and Primary Tumor Samples
The pancreatic cancer cell lines, Panc1 and KMP3 were maintained in Dulbecco’s Modified Eagle Medium containing 10% fetal bovine serum (FBS). The pancreatic cancer cell line KP4-4, BxPC3, CFPAC1 and SU.86.86 were maintained in Roswell Park Memorial Institute medium 1640 containing 10% FBS. The human mammary carcinoma cell line MDA-MB-231 was purchased from the American Type Culture Collection (Manassas, VA, USA) and was maintained in L-15 medium containing 10% FBS. Pancreatic cancer cell lines were authenticated in our previous studies of array-CGH analyses51.
A total of 50 primary PDAC samples were obtained from patients with PDAC who underwent pancreatectomy at Kyoto Prefectural University of Medicine between 2000 and 2011. These samples were embedded in paraffin after 24 hours of formalin fixation. None of these patients underwent preoperative chemotherapy or radiotherapy, and none had metachronous multiple cancers in other organs. All samples were obtained with the informed consent of each patient after approvals by the local ethics committees of the Medical Research Institute and Faculty of Medicine in Tokyo Medical and Dental University (approval number: 2015-001) and Kyoto Prefectural University of Medicine (approval number: ERB-C-67-2). The methods were carried out in accordance with the approved guidelines and regulations.
Screening system using the promoterless expression vector pZsGreen1
The two stable clones, PEcadZsG-Panc1 #1 and #2 cells, were established by the limiting dilution method after transfection of the reporter construct into Panc1 cells. First, PEcadZsG-Panc1 #1 (5 × 103 cells per well) and #2 (3 × 103 cells per well) were seeded on 96-well plates. After 24 hours, each clone was transfected in duplicate with one of 1090 dsRNAs from the Pre-miRTM miRNA Precursor Library-Human V15 (Thermo Fisher Scientific, CA, USA) or with a negative control RNA, using an RNA concentration of 10 nmol/L. After 72 hours, the fluorescence intensity of the ZsGreen1 protein was measured by ARVO mx (Perkin Elmer, MA, USA). At the same time, cell survival was assessed by the crystal violet staining assay. The relative fluorescence intensity was calculated by the formula: fluorescence intensity/cell survival rate22.
Transfection of miRNAs, siRNAs and miRNA inhibitors and treatment with TGF-β
The dsRNA mimicking mature human miRNA for miR-200c (MC11714), miR-509-5p (MC13068), miR-660 (MC11216), miR-1243 (MC13161) and negative control miRNA (negative control #1) were purchased from Thermo Fisher Scientific. The siRNA for Smad2 (L-003561-00), Smad4 (L-003902-00), HMGA2 (L-013495-00), and negative control siRNAs (D-001810-05) were purchased from GE Healthcare (Buckinghamshire, UK). The miRNA inhibitors for anti-miR-509-5p (MH13068), anti-miR-1243 (MH13161) and negative control anti-miRNA (negative control #1) were purchased from Thermo Fisher Scientific. miRNAs, siRNAs and miRNA inhibitors were transfected individually into cells at the indicated concentrations using Lipofectamine RNAiMAX (Thermo Fisher Scientific) according to the manufacturer’s instructions. After each miRNA transfectant was treated with TGF-β (5 ng/ml), the expression of E-cadherin was evaluated by western blot analysis.
Quantitative RT–PCR (qRT-PCR)
Total RNA was extracted using TRIsure reagent (BIOLINE, London, UK) according to standard procedures. For miRNA, total RNA was reverse transcribed using Taqman Reverse Transcription Kit followed by qRT-PCR performed using Custom Taqman miRNA Assays kit (Applied Biosystems). The miRNA expression was normalized to endogenous control RNU6B. Single-stranded cDNA generated from the total RNA was amplified with a gene-specific primer set. Gene expression was normalized to the housekeeping gene glyceraldehyde-3-phosphate dehydrogenase (GAPDH). The qRT-PCR was performed using an ABI PRISM 7500 sequence detection System (Applied Biosystems, Foster City, CA, USA) according to the manufacturer’s instructions. The following primers were used for the Taqman assay (Thermo Fisher Scientific): human miR-509-5p (002235), miR-1243 (002854), RNU6B (001093), CDH1 (Hs01023894_m1), VIM (Hs00958111_m1) and GAPDH (Hs02758991_g1).
Western blotting and immunofluorescent staining
The following primary antibodies were used for western blotting or immunofluorescent staining: anti-E-Cadherin (#610181) (BD Biosciences, Franklin Lakes, NJ, USA), anti-HMGA2 (#8179), anti-Snail (#3879), anti-TCF8/ZEB1 (#3396), anti-ZO-1 (#5406) (Cell Signaling Technology, Danvers, MA, USA), anti-Vimentin Ab-2 (Clone V9) (Thermo Fisher Scientific), anti-Smad2 (ab33875) (Abcam, Cambridge, UK), anti-β-actin (Sigma, St. Louis, MO, USA) and anti-Smad4 (sc-7966) (Santa Cruz Biotechnology, Santa Cruz, CA, USA). Western blotting and immunofluorescent staining were performed as described elsewhere52.
Cell survival, migration and invasion assays
The number of living cells at various time points after transfection was assessed by a colorimetric water-soluble tetrazolium salt (WST) assay or crystal violet staining assay, as described elsewhere53. Transwell migration and invasion assays were performed in 24-well modified chambers precoated without (migration) or with (invasion) Matrigel (BD BioCoat, BD Biosciences), as described elsewhere54. Panc1, CFPAC1 and KMP3 cells (4 × 104 per well) in serum-free medium were transferred into the upper chamber. After 24 hours of incubation, the cells that migrate to the lower chamber, which contained 10% FBS as a chemoattractant, were fixed, stained with the Diff-Quik stain (Sysmex, Kobe, Japan) and counted in five random fields.
Luciferase activity assay
Luciferase reporter plasmids were made by inserting the 3′-UTR of Smad2, Smad4, VIM, HMGA2 and ZEB1 downstream of the luciferase gene within a pmirGLO Dual-Luciferase miRNA Target Expression Vector (Promega, Madison, WI, USA). All site-specific mutations used the GeneTailor site-directed mutagenesis system (Thermo Fisher Scientific). Luciferase reporter plasmid or control plasmid (pmirGLO) was transfected into Panc1 cells using Lipofectamine 2000 (Thermo Fisher Scientific), and 10 nmol/L of miRNA (miR-NC, miR-509-5p or miR-1243) was also transfected 6 hours later. After two days, Firefly and Renilla luciferase activities were measured using the Dual-Luciferase Reporter Assay System (Promega), and relative luciferase activity was calculated by normalizing the Firefly luciferase reading with its corresponding internal Renilla luciferase control.
Cytotoxicity study
Panc1 cells (5 × 103) were seeded in wells of the E-Plate 16 (ACEA Biosciences, San Diego, CA, USA). Approximately 24 hours later, 5 nmol/L of each miRNA (miR-NC, miR-509-5p or miR-1243) was transfected into Panc1 cells. Six hours later, these transfectants were treated with gemcitabine (0.1, 1, 10, 50, 100 and 200 nmol/L) and PBS. Cell-electrode impedance was monitored using the xCELLigence RTCA DP system (ACEA Biosciences) to produce time-dependent cell response dynamic curves. Data were collected every 10 min after treatment with gemcitabine for the first four hours, every 15 min for the next 20 hours, and then every 1 hour for an additional 4 days.
miRNA-ISH assay
The ISH assay was performed on formalin-fixed and paraffin-embedded (FFPE) tissue sections according to the manufacturer’s instructions (miRCURY LNA microRNA ISH Optimization Kit; Exiqon Inc., Vedbaek, Denmark). Briefly, the sections were deparaffinized in xylene, rehydrated with graded ethanol and incubated with proteinase-K for 10 min at 37 °C. Then, the sections were hybridized with the miR-509-5p and miR-1243 double-digoxigenin (DIG)-labeled LNA probes for 1 hour at 55 °C and were washed stringently prior to incubation with blocking for 15 min and probing with specific anti-DIG antibody directly conjugated with alkaline phosphatase. Finally, the sections were counterstained with nuclear red. We classified samples stained even a little as each miRNA-positive groups and samples with no stain as each miRNA-negative groups.
Public data sets
To explore the generality of the miRNA expression and clinical features among pancreatic cancer, we examined the publicity dataset from TCGA (http://cancergenome.nih.gov) retrieved on 20th February 2017. We took the primary pancreatic cancer data (TCGA-PAAD) from the TCGA data set, which included mRNA data on 178 samples and 1881 miRNAs, and examined correlation of prognosis and expression of miR-509-5p and miR-1243 using 141 samples excluding Stage IV, macroscopic residual tumors (R2) or unevaluable presence of tumors (RX), and tumor-types other than PDAC. Expression of miR509-5p was taken as the sum of expression of miR-509-1 and miR-509-2.
Statistical analysis
The association between clinicopathological characteristics and the status of miR-509-5p or miR-1243 expression in patients with PDAC was evaluated with the χ2 or Fisher’s exact test. In Kaplan-Meier curves, differences between subgroups were tested with the log-rank test. Univariate and multivariate survival analyses were performed using the likelihood ratio test of the stratified Cox proportional hazards model. Differences between subgroups were tested with the Student’s t-test and considered significant at P < 0.05.
References
Hariharan, D., Saied, A. & Kocher, H. M. Analysis of mortality rates for pancreatic cancer across the world. HPB (Oxford) 10, 58–62, doi:10.1080/13651820701883148 (2008).
Kamisawa, T., Wood, L. D., Itoi, T. & Takaori, K. Pancreatic cancer. The Lancet 388, 73–85, doi:10.1016/s0140-6736(16)00141-0 (2016).
Garrido-Laguna, I. & Hidalgo, M. Pancreatic cancer: from state-of-the-art treatments to promising novel therapies. Nat Rev Clin Oncol 12, 319–334, doi:10.1038/nrclinonc.2015.53 (2015).
Ha, M. & Kim, V. N. Regulation of microRNA biogenesis. Nature reviews. Molecular cell biology 15, 509–524, doi:10.1038/nrm3838 (2014).
Kalluri, R. & Weinberg, R. A. The basics of epithelial-mesenchymal transition. J Clin Invest 119, 1420–1428, doi:10.1172/JCI39104 (2009).
Tsai, J. H. & Yang, J. Epithelial-mesenchymal plasticity in carcinoma metastasis. Genes Dev 27, 2192–2206, doi:10.1101/gad.225334.113 (2013).
De Craene, B. & Berx, G. Regulatory networks defining EMT during cancer initiation and progression. Nature reviews. Cancer 13, 97–110, doi:10.1038/nrc3447 (2013).
Thiery, J. P., Acloque, H., Huang, R. Y. & Nieto, M. A. Epithelial-mesenchymal transitions in development and disease. Cell 139, 871–890, doi:10.1016/j.cell.2009.11.007 (2009).
Mani, S. A. et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133, 704–715, doi:10.1016/j.cell.2008.03.027 (2008).
Tam, W. L. & Weinberg, R. A. The epigenetics of epithelial-mesenchymal plasticity in cancer. Nat Med 19, 1438–1449, doi:10.1038/nm.3336 (2013).
Iorio, M. V. & Croce, C. M. MicroRNA dysregulation in cancer: diagnostics, monitoring and therapeutics. A comprehensive review. EMBO molecular medicine 4, 143–159, doi:10.1002/emmm.201100209 (2012).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Zhang, J. & Ma, L. MicroRNA control of epithelial-mesenchymal transition and metastasis. Cancer metastasis reviews 31, 653–662, doi:10.1007/s10555-012-9368-6 (2012).
Esquela-Kerscher, A. & Slack, F. J. Oncomirs - microRNAs with a role in cancer. Nature reviews. Cancer 6, 259–269, doi:10.1038/nrc1840 (2006).
Qian, P. et al. Pivotal role of reduced let-7g expression in breast cancer invasion and metastasis. Cancer research 71, 6463–6474, doi:10.1158/0008-5472.CAN-11-1322 (2011).
Sun, L. et al. MiR-200b and miR-15b regulate chemotherapy-induced epithelial-mesenchymal transition in human tongue cancer cells by targeting BMI1. Oncogene 31, 432–445, doi:10.1038/onc.2011.263 (2012).
Yu, F. et al. let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 131, 1109–1123, doi:10.1016/j.cell.2007.10.054 (2007).
Fischer, K. R. et al. Epithelial-to-mesenchymal transition is not required for lung metastasis but contributes to chemoresistance. Nature 527, 472–476, doi:10.1038/nature15748 (2015).
Gregory, P. A. et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nature cell biology 10, 593–601, doi:10.1038/ncb1722 (2008).
Harazono, Y. et al. miR-655 Is an EMT-suppressive microRNA targeting ZEB1 and TGFBR2. PloS one 8, e62757, doi:10.1371/journal.pone.0062757 (2013).
Hayes, J., Peruzzi, P. P. & Lawler, S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends in molecular medicine 20, 460–469, doi:10.1016/j.molmed.2014.06.005 (2014).
Yanaka, Y., Muramatsu, T., Uetake, H., Kozaki, K. & Inazawa, J. miR-544a induces epithelial-mesenchymal transition through the activation of WNT signaling pathway in gastric cancer. Carcinogenesis 36, 1363–1371, doi:10.1093/carcin/bgv106 (2015).
Lau, N. C., Lim, L. P., Weinstein, E. G. & Bartel, D. P. An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294, 858–862, doi:10.1126/science.1065062 (2001).
Xu, J., Lamouille, S. & Derynck, R. TGF-beta-induced epithelial to mesenchymal transition. Cell Res 19, 156–172, doi:10.1038/cr.2009.5 (2009).
Eppert, K. et al. MADR2 maps to 18q21 and encodes a TGFbeta-regulated MAD-related protein that is functionally mutated in colorectal carcinoma. Cell 86, 543–552 (1996).
Hahn, S. A. et al. DPC4, a candidate tumor suppressor gene at human chromosome 18q21.1. Science 271, 350–353 (1996).
Watanabe, S. et al. HMGA2 maintains oncogenic RAS-induced epithelial-mesenchymal transition in human pancreatic cancer cells. The American journal of pathology 174, 854–868, doi:10.2353/ajpath.2009.080523 (2009).
Morishita, A. et al. HMGA2 is a driver of tumor metastasis. Cancer research 73, 4289–4299, doi:10.1158/0008-5472.CAN-12-3848 (2013).
Singh, A. & Settleman, J. EMT, cancer stem cells and drug resistance: an emerging axis of evil in the war on cancer. Oncogene 29, 4741–4751, doi:10.1038/onc.2010.215 (2010).
Zhang, W. B., Pan, Z. Q., Yang, Q. S. & Zheng, X. M. Tumor suppressive miR-509-5p contributes to cell migration, proliferation and antiapoptosis in renal cell carcinoma. Irish journal of medical science 182, 621–627, doi:10.1007/s11845-013-0941-y (2013).
Ren, Z. J. et al. Mir-509-5p joins the Mdm2/p53 feedback loop and regulates cancer cell growth. Cell death & disease 5, e1387, doi:10.1038/cddis.2014.327 (2014).
Xing, F. et al. miR-509 suppresses brain metastasis of breast cancer cells by modulating RhoC and TNF-alpha. Oncogene 34, 4890–4900, doi:10.1038/onc.2014.412 (2015).
Ma, N. et al. The Tumor Suppressive Role of MiRNA-509-5p by Targeting FOXM1 in Non-Small Cell Lung Cancer. Cellular physiology and biochemistry: international journal of experimental cellular physiology, biochemistry, and pharmacology 38, 1435–1446, doi:10.1159/000443086 (2016).
Motoyama, K. et al. Clinical significance of high mobility group A2 in human gastric cancer and its relationship to let-7 microRNA family. Clinical cancer research: an official journal of the American Association for. Cancer Research 14, 2334–2340, doi:10.1158/1078-0432.CCR-07-4667 (2008).
Hristov, A. C. et al. HMGA2 protein expression correlates with lymph node metastasis and increased tumor grade in pancreatic ductal adenocarcinoma. Modern pathology: an official journal of the United States and Canadian Academy of Pathology, Inc 22, 43–49, doi:10.1038/modpathol.2008.140 (2009).
Wang, X. et al. Overexpression of HMGA2 promotes metastasis and impacts survival of colorectal cancers. Clinical cancer research: an official journal of the American Association for. Cancer Research 17, 2570–2580, doi:10.1158/1078-0432.CCR-10-2542 (2011).
Jiang, W. et al. miRNA-101 Suppresses Epithelial-to-Mesenchymal Transition by Targeting HMGA2 in Pancreatic Cancer Cells. Anti-cancer agents in medicinal chemistry 16, 432–439 (2016).
Wang, Y. C. et al. miR221 targets HMGA2 to inhibit bleomycininduced pulmonary fibrosis by regulating TGFbeta1/Smad3-induced EMT. International journal of molecular medicine, doi:10.3892/ijmm.2016.2705 (2016).
Chen, Z., Li, Q., Wang, S. & Zhang, J. miR-485-5p inhibits bladder cancer metastasis by targeting HMGA2. International journal of molecular medicine 36, 1136–1142, doi:10.3892/ijmm.2015.2302 (2015).
Mayr, C., Hemann, M. T. & Bartel, D. P. Disrupting the pairing between let-7 and Hmga2 enhances oncogenic transformation. Science 315, 1576–1579, doi:10.1126/science.1137999 (2007).
Copley, M. R. et al. The Lin28b-let-7-Hmga2 axis determines the higher self-renewal potential of fetal haematopoietic stem cells. Nature cell biology 15, 916–925, doi:10.1038/ncb2783 (2013).
Ding, Z. et al. SMAD4-dependent barrier constrains prostate cancer growth and metastatic progression. Nature 470, 269–273, doi:10.1038/nature09677 (2011).
Zhang, B. et al. Antimetastatic role of Smad4 signaling in colorectal cancer. Gastroenterology 138, 969–980 e961–963, doi:10.1053/j.gastro.2009.11.004 (2010).
Bardeesy, N. et al. Smad4 is dispensable for normal pancreas development yet critical in progression and tumor biology of pancreas cancer. Genes Dev 20, 3130–3146, doi:10.1101/gad.1478706 (2006).
Wong, S. H. et al. Genome-wide association and sequencing studies on colorectal cancer. Semin Cancer Biol 23, 502–511, doi:10.1016/j.semcancer.2013.09.005 (2013).
Malkoski, S. P. & Wang, X. J. Two sides of the story? Smad4 loss in pancreatic cancer versus head-and-neck cancer. FEBS Lett 586, 1984–1992, doi:10.1016/j.febslet.2012.01.054 (2012).
Chen, Y. W. et al. SMAD4 loss triggers the phenotypic changes of pancreatic ductal adenocarcinoma cells. BMC cancer 14, 181, doi:10.1186/1471-2407-14-181 (2014).
Duda, D. G. et al. Restoration of SMAD4 by gene therapy reverses the invasive phenotype in pancreatic adenocarcinoma cells. Oncogene 22, 6857–6864, doi:10.1038/sj.onc.1206751 (2003).
Kozaki, K. & Inazawa, J. Tumor-suppressive microRNA silenced by tumor-specific DNA hypermethylation in cancer cells. Cancer science 103, 837–845, doi:10.1111/j.1349-7006.2012.02236.x (2012).
Zheng, X. et al. Epithelial-to-mesenchymal transition is dispensable for metastasis but induces chemoresistance in pancreatic cancer. Nature 527, 525–530, doi:10.1038/nature16064 (2015).
Suzuki, A. et al. Identification of SMURF1 as a possible target for 7q21.3–22.1 amplification detected in a pancreatic cancer cell line by in-house array-based comparative genomic hybridization. Cancer science 99, 986–994, doi:10.1111/j.1349-7006.2008.00779.x (2008).
Muramatsu, T. et al. YAP is a candidate oncogene for esophageal squamous cell carcinoma. Carcinogenesis 32, 389–398, doi:10.1093/carcin/bgq254 (2011).
Muramatsu, T. et al. The hypusine cascade promotes cancer progression and metastasis through the regulation of RhoA in squamous cell carcinoma. Oncogene. doi:10.1038/onc.2016.71 (2016).
Ono, H. et al. SIX1 promotes epithelial-mesenchymal transition in colorectal cancer through ZEB1 activation. Oncogene 31, 4923–4934, doi:10.1038/onc.2011.646 (2012).
Acknowledgements
We thank R. Mifuji and E. Tonouchi for technical assistance. We thank Dr. H. Ikoma for providing primary PDAC samples. This study was supported by the Joint Usage/Research Program of Medical Research Institute, Tokyo Medical and Dental University (TMDU). This work was supported by KAKENHI (15H05908, 25250019, 15K18401, 26890012) from the Ministry of Education, Culture, Sports, Science, and Technology (MEXT), and partially supported by the Project for Development of Innovative Research on Cancer Therapeutics (P-DIRECT) and the Project for Cancer Research And Therapeutic Evolution (P-CREATE) from Japan Agency for Medical Research and development, AMED. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
H.H. was involved in research design, performed the experiments, analyzed data and wrote the manuscript. D.I., S.Y. and E.O. contributed materials. K.T. contributed to analysis of TCGA data. T.M. and J.I. was involved in research design and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hiramoto, H., Muramatsu, T., Ichikawa, D. et al. miR-509-5p and miR-1243 increase the sensitivity to gemcitabine by inhibiting epithelial-mesenchymal transition in pancreatic cancer. Sci Rep 7, 4002 (2017). https://doi.org/10.1038/s41598-017-04191-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-04191-w
This article is cited by
-
MiRNA-mediated EMT and CSCs in cancer chemoresistance
Experimental Hematology & Oncology (2021)
-
Association of sonic hedgehog signaling pathway genes IHH, BOC, RAB23a and MIR195-5p, MIR509-3-5p, MIR6738-3p with gastric cancer stage
Scientific Reports (2021)
-
Cancer-associated miRNAs and their therapeutic potential
Journal of Human Genetics (2021)
-
Cancer drug resistance induced by EMT: novel therapeutic strategies
Archives of Toxicology (2021)
-
Circular RNAs: Novel target of diabetic retinopathy
Reviews in Endocrine and Metabolic Disorders (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.