Abstract
In the analysis of large-scale gene expression data, it is important to identify groups of genes with common expression patterns under certain conditions. Many biclustering algorithms have been developed to address this problem. However, comprehensive discovery of functionally coherent biclusters from large datasets remains a challenging problem. Here we propose a GPU-accelerated biclustering algorithm, based on searching for the largest Condition-dependent Correlation Subgroups (CCS) for each gene in the gene expression dataset. We compared CCS with thirteen widely used biclustering algorithms. CCS consistently outperformed all the thirteen biclustering algorithms on both synthetic and real gene expression datasets. As a correlation-based biclustering method, CCS can also be used to find condition-dependent coexpression network modules. We implemented the CCS algorithm using C and implemented the parallelized CCS algorithm using CUDA C for GPU computing. The source code of CCS is available from https://github.com/abhatta3/Condition-dependent-Correlation-Subgroups-CCS.
Similar content being viewed by others
Introduction
Clustering algorithms have been widely used to group genes based on their similarities in expression1,2,3,4. The gene expression coherence is often related to functional coherence. A recent comparative assessment of 21 existing clustering algorithms showed that the clustering algorithms report more functionally meaningful clusters by exploiting the relationships between all pairs of genes1. Clustering algorithms are also known to perform better grouping of co-functional genes if they search for similarities between expression patterns rather than similarities between expression values1, 5.
Functional relations between genes may vary over conditions6, 7. For example, a group of genes may act coherently under one set of conditions but may become inactive or perform different functions separately under another set of conditions. Clustering algorithms that obtain grouping based on similarities over all the samples in a dataset are not effective for detecting condition-dependent coexpression patterns. Biclustering algorithms have been developed to address this problem1, 8,9,10,11,12. A bicluster consists of a group of genes and a set of conditions under which the genes are co-expressed. Searching for bicluster is a challenging problem because the number of potential biclusters is exponential to the number of genes and samples. An important difference between various biclustering algorithms is how they apply heuristic rules to detect biclusters. Most biclustering algorithms use local search heuristics that may miss many biclusters. Conventional way of finding biclusters depends on the selection of random seeds of genes and/or samples followed by their augmentation based on a scoring function. However, random selection of initial seeds is often unable to efficiently cover the search space and to obtain a consistent set of biclusters from multiple runs on the same input data. The selection of scoring function for the heuristic search is also important for finding the biologically meaningful biclusters. Most common biclustering approaches adopt arithmetic mean of the gene expression or an up/down-regulation patterns on corresponding discretized data matrix. Both are inefficient for finding the interesting co-regulatory modules where genes are expressed with similar or opposite patterns of expression but the expression values are very different. Such modules are important as they may represent relations between genes in the same biological functions1, 13.
A Pearson Correlation Coefficient based scoring function was introduced by Correlated Pattern Biclusters (CPB)14. However, the performance CPB is still depends on the random selection of samples. Benefits of Pearson Correlation Coefficient based similarity measures over the conventional mean square residue based bicluster scores again was demonstrated by Bi-Correlation Clustering algorithm (BCCA) that looks for positively correlated biclusters and reports biclusters for each pair of genes present in a dataset13. Initially, for each pair of genes all the samples are selected and then a greedy search is made to eliminate samples one at a time until the gene pair shows positive correlation higher than a predefined high positive correlation threshold. The gene pair is then augmented with other genes that are correlated to form a bicluster. However the backward elimination of samples in a greedy search is subject to finding a local optimal subset of samples, hence in reality it imposes a restriction on the search space for finding the optimal set of biclusters. BIclustering by Correlated and Large number of Individual Clustered seeds (BICLIC) introduces another alternative correlation based biclustering. BICLIC forms biclusters by starting from a large number of clustered seeds followed by augmentation and deletion of rows and columns based on the correlations with seeds15. The main drawback of BICLIC which is also true for BCCA is finding a large number of overlapping biclusters which are very much identical. Moreover, BICLIC, BCCA and other correlation based biclustering methods, only look for finding positively correlated gene groups as bicluster. However, in reality negatively correlated genes are equally relevant and may represent important regulatory mechanisms in biological functions.
Here we propose a more effective correlation-based biclustering algorithm named Condition-dependent Correlation Subgroup (CCS). It integrates several important features for developing an effective algorithm for comprehensive discovery of functionally coherent biclusters1. A significant challenge in genomic data analysis is to utilize the fast growing high performance computing capacity to process and analyze large complex datasets efficiently16,17,18. CCS is particularly suitable for parallel computing. We used the GPGPU computing for a parallel implementation of the algorithm in CUDA C which shows a substantial speedup compared to the sequential C program. The performance of CCS was compared with CPB, BCCA, BICLIC and ten other widely-used biclustering algorithms on 5 synthetic and 5 real gene expression datasets. CCS outperforms the other biclustering algorithms in all the comparisons. We also showed that there is equivalence between the CCS biclusters and condition-dependent coexpression network modules.
Methods
A bicluster is a group of genes with a related pattern of expression over a group of samples. We defined the related pattern of gene expression in terms of positive and negative Pearson correlation coefficients. Let us consider a gene expression data set D = G × E where G = {g1, g2, …, gn} is a set of “n” genes and E = {e1, e2, …, es, …, em} is a set of “m” samples. For each gene gi there is an m-dimensional vector xi. In vector xi, xis is the value of es for gene gi. In our algorithm we defined a bicluster as a group of genes “I” over a group of samples “J” where each gene gi in “I” is correlated to all the other genes gj in “I” with an absolute correlation value greater than a threshold θ. Mathematically a bicluster “C” is represented as C = (I; J) where “I” is a subset of “G” and “J” is a subset of “E”. The Pearson correlation coefficient between gi and gj over a subset of samples “J” is represented as r(gi, gj)J and defined as
where b is a bit vector of size “m”. If sample es is in “J” then we set the sth bit of b s = 1, otherwise we set b s = 0. The terms xis and xjs represent sth sample values for gene gi and gj respectively. Average expression values of gi and gj for samples in “J” are represented as \(\overline{{x}_{i}\times b}\) and \(\overline{{x}_{j}\times b}\) respectively. We considered a pair of genes gi and gj similar for a subset of samples “J” if \(|r{(gi,gj)}_{J}| > \theta \), where θ is the correlation threshold and \(|r{(gi,gj)}_{J}|\,\,\)is the absolute value of correlation between gi and gj for the subset of samples “J”. For each gene gi in a dataset “D”, CCS considers gi as the base gene to start forming a bicluster for each gi.
Search space sorting and base-gene selection
The gene with the lower variability over the samples (commonly known as housekeeping genes) are often considered as less significant for the corresponding biological conditions, hence the less important candidates for forming a bicluster. Prior to biclustering, the search space (input data matrix) was sorted by the standard deviations of the gene expression values. In an ordered data matrix the genes with the higher variability were placed before the other with the lower variability.
CCS algorithm selects ‘base_number’ of genes as base-gene from the sorted data matrix in the decreasing order of variability. Ordering of the search space ensured that a gene with higher variability considered for forming a bicluster and also for augmentation before the other with lower variability. The ‘base_number’ can be set between “1” to “n” (total number of genes). Here we set the base_number value equals 1,000 for the biological dataset to restrict the search time.
Similarity pattern classes and sample selections
CCS algorithm considers three classes of gene expression pattern similarities: (i) up-regulated positive correlation: gi and gj are positively correlated and the expression values of gi and gj for the selected samples are higher than the arithmetic mean expression values over all smaples; (ii) down-regulated positive correlation: gi and gj are positively correlated and the expression values of gi and gj for the selected samples are lower than the arithmetic mean expression values over all smaples; (iii) negative correlation: gi and gj are negatively correlated over the selected samples and their expression values are at the opposite sides to their respective arithmetic mean expressions.
For each pair of genes gi and gj the sample sets “J1: for up-regulated positive correlation”, “J2: for down-regulated positive correlation” and “J3: for negative correlation” are selected from sample selection rules 1, 2 and 3 respectively. For each sample es, s = 1:m, we defined the following three rules to determine J1, J2 and J3: Rule 1. IF \(({x}_{is}-\overline{{x}_{i}}) > 0\,AND\,({x}_{js}-\overline{{x}_{j}}) > 0\) THEN include es to J1 [Expression values of gi and gj for the samples in J1 are higher than the arithmetic mean expression values], Rule 2. IF \(({x}_{is}-\overline{{x}_{i}}) < 0\,AND\,({x}_{js}-\overline{{x}_{j}}) < 0\) THEN include es to J2 [Expression values of gi and gj for the samples in J2 are lower than the arithmetic mean expression values], Rule 3. IF \(({x}_{is}-\overline{{x}_{i}})\times ({x}_{js}-\overline{{x}_{j}}) < 0\) THEN include es to J3 [Expression values of gi and gj for the samples in J3 are opposite to their arithmetic mean expression values]. Here \(\overline{{x}_{i}}\) and \(\overline{{x}_{j}}\) are the mean expression value of gene gi and gj over all samples. The correlations between gi and gj for sample sets Jk=1:3 are determined first by computing Pearson Correlation Coefficient ‘r’ and then by comparing against a threshold ‘θ’ which is denoted by |r(gi, gj)Jk| > θ in the algorithm.
Defining biclusters as condition dependent correlation modules
CCS defines biclusters as condition dependent correlation modules where the genes in a bicluster are expected to show correlations only for the samples in that bicluster. CCS introduces a scoring function named BScore as defined in Equation (2). The BScore is designed to compare two sets of correlated gene pairs N and M. The correlations in set ‘N’ are computed over the samples that are included in a bicluster while the correlations in set ‘M’ are computed over the rest of the samples that are not included in the same bicluster. Thus, the BScore measures the degree to which the gene coexpression in a bicluster is condition-specific. In this work, we used a small BScore threshold (<0.01) to ensure the discovered biclusters are based on condition-specific coexpression.
Algorithm
The algorithm at each iteration of step 2 starts with a new base-gene gi. In each iteration of step 5, gi is paired with a gene gj (i < j ≤ n). In step 6, the sample sets are selected for gene pair gi, gj. In step 8, the algorithm computes the absolute value of the correlation |r(gi, gj)Jk| for each sample set Jk=1:3 (Equation (1)). If |r(gi, gj)Jk|\( > \theta \) then the algorithm starts a bicluster with an initial gene set Ii = {gi, gj}. In step 10, augmentation of the gene set is performed by including a new gene gp in Ii. In step 16, \({\rm{BScore}}\)(Ii,Ji) is compared against the previous best \({\rm{BScore}}\)(I,J) to update the sets I and J. The most condition dependent bicluster for each base gene gi is selected in step 22. In step 26, all the overlapping biclusters are merged while keeping \({\rm{BScore}} < 0.01\)
Input: (i) A gene expression data set D = G × H, G = {g1, g2,…, gn} is a set of n genes, H = {e1, e2,…, em} is a set of m samples for gene set G. (ii) A correlation threshold θ. |
Output: A set of biclusters S = {bicluster(g1), bicluster(g2),…, bicluster(gbase_number)}, base_number ≤ n. |
1. S ← NULL |
2. for all gi \(\in \,\)G, i ≤ base_number do |
3. I ← NULL |
4. J ← NULL |
5. for all gj \(\in \,\)G, i < j ≤ n do |
6. apply Rules(1,2,3) on {gi, gj} for sample sets Jk(k=1,2,3) |
7. for all Jk, k = 1:3 do |
8. if |r(gi, gj)Jk| > θ then |
9. Ii ← {gi, gj} |
10. for all gp not in Ii do |
11. if |r(gp, gq)Jk| > θ, for all gq \(\in \,\,\)Ii then |
12. Ii ← Ii ∪ gp |
13. end if |
14. end for |
15. end if |
16. If (BScore(Ii,Ji) < BScore(I,J) < 0.01) or |
(BScore(Ii,Ji) = BScore(I,J) < 0.01 and |Ii| > |I|) then |
17. I ← Ii |
18. J ← Ji |
19. end if |
20. end for |
21. end for |
22. if I ≠ NULL and J ≠ NULL then |
23. bicluster(gi) ← {I, J} |
24. end if |
25. end for |
26. for each bicluster({I, J}) do |
27. for each bicluster({K, L})\(\ne \,\)NULL do |
28. if BScore(I ∪ K, J ∪ L) < 0.01 and I ∩ K ≠ NULL then |
29. bicluster({I, J}) ← {I ∪ K, J ∪ L} |
30. bicluster({K, L}) ← NULL |
31. end if |
32. end for |
33. end for |
for the final set of biclusters ‘S’.
The input parameter for our biclustering algorithm is the correlation threshold θ, which is the minimum absolute value of correlation between genes in a bicluster. Allocco et al. investigated the relation between co-regulation and coexpression and came up with a conclusion that for a correlation greater than 0.84 there is at least 50% chance of co-regulation19. Here we set θ = 0.8. If the number of biclusters is zero for θ = 0.8 then we decrease θ by 0.05 until we get a nonempty set of biclustering result or θ drops to 0.
GPU computing for biclustering analysis of large datasets
GPU was initially introduced for rendering video graphics on display devices. The GPU designing company NVIDIA introduced their CUDA architecture in 2007 for general purpose computing on graphics processing unit (GPGPU computing). The single instruction set and multiple dataset architecture (SIMD) of GPU is particularly useful for executing a single instruction over thousands of core processors in a GPU card. CUDA is currently the most popular programming model for general purpose GPU computing. CUDA C is an extension of regular C that supports GPGPU computing. The parallel part is implemented as a CUDA C kernel function. Thousands of instances of a CUDA C kernel function run parallel on a group of GPU core processors. Each group of GPU cores is also known as a thread block. Each thread block is a virtual processor formed from one or more GPU core processors, registers, local memories, shared memory and global memory. CUDA C kernel functions are sent to the GPU for parallel execution, and sequential parts of the code run on the CPU.
We used GPGPU computing to reduce the execution time. In this work, we executed our CUDA C code for CCS on NVIDIA Tesla K20 (2,496 CUDA cores) and K80 GPU (4,992 CUDA cores) accelerator. For compilation of CUDA C code we installed CUDA 7.5 toolkit in our Linux workstation. Transferring big datasets between CPUs and GPUs is time consuming. To reduce the data transfer overhead we moved the entire process of finding new biclusters to GPU.
Step 2 of CCS executes “n” times when “base_number” is set at “n” for a dataset with “n” rows. For each execution the inner loop at step 5 executes a maximum “n-1” times. To achieve increased speed from parallel GPGPU computing, we removed the step 2 loop from CCS algorithm and implement steps 5 to 25 of CCS as a CUDA C kernel function.
Synthetic datasets
Five synthetic datasets were generated using the BiBench-0.2 python library8. The synthetic datasets contains two types of biclustering: Constant biclusters and Shift-Scale biclusters. Constant biclusters are sub-matrices with a constant value and Shift-Scale biclusters are biclusters that are formed from both shifting and scaling a base row by random numbers. Table 1 describes the synthetic datasets for their types, dimensions and number of true biclusters.
Gene expression datasets
Five publicly available gene expression datasets GDS531, GDS589, GDS3603, GDS3966 and GDS4794 were downloaded from Gene Expression Omnibus (GEO). Genes with standard deviation less than 2.0 were removed. For genes appeared multiple times, only the row (probe set) with the highest standard deviation was kept. The dataset information is summarized in Table 2.
Evaluating biclustering results
We evaluated the results of the biclustering algorithms on synthetic datasets using Recovery and Relevance metrics8, 20. They compare the set of found biclusters ‘F’ against the expected biclusters ‘E’ (the true bicluster). Both for Recovery and Relevance the scores are ranging between ‘0’ and ‘1’. The highest score ‘1’ for \({\rm{Recovery}}\) denotes that all the expected biclusters are found. Similarly, the highest ‘1’ for \({\rm{Relevance}}\) denotes that all the found biclusters are expected.
To evaluate the results on the real gene expression datasets, we performed functional enrichment tests for each bicluster1, 8, 13, 20. We used gProfileR21 for selecting the enriched gene annotation terms. gProfilerR retrieves a comprehensive set of gene annotations from Gene Ontology (GO) terms, biological pathways, regulatory motifs in DNA, microRNA targets and protein complexes. From gProfiler search, all the annotation terms with a Benjamini-Hochberg FDR less than 0.01 were considered as enriched. From the biclustering results on all five gene expression datasets we computed the average number of enriched annotation terms. Higher average number of enriched terms indicates better functional grouping. We also evaluated the performance of biclustering algorithms from percentage of total biclusters that have one or more enriched annotation terms.
Results and Discussions
We obtained the biclustering results from CCS and 13 other biclustering algorithms namely Cheng and Church Biclustering algorithm (CC)22, Iterative Signature Algorithm (ISA)23, Plaid24, Spectral25, Order Preserving Sub Matrix (OPSM)26, xMotifs27, SAMBA28, Bimax20, BicSPAM29, UniBic30, CPB14, BICLIC15 and BCCA13. We used BicAT20 software package for ISA, CC, OPSMs, xMotifs and Bimax. We also used ‘biclust’ R package (https://cran.r-project.org/web/packages/biclust/) for Plaid and Spectral. For UniBic, BicSPAM, BCCA, CPB and BICLIC the published versions of the software packages were used. SAMBA biclustering was performed using the Expander package31.
Performance on synthetic data
Figure 1 shows the Recovery and Relevance scores on five synthetic datasets (Table 1) for CCS and the best among the other 13 algorithms for comparison. For all of the five synthetic datasets CCS significantly outperformed other algorithms for finding the expected biclusters (recovery) and the most relevant set of biclusters (relevance). For constant bicluster on a datasets of 100 rows and 100 columns, CCS obtained 0.88 score for both recovery and relevance. For larger size datasets and shift and scale biclusters, still the recovery and relevance scores are higher than the other algorithms which clearly demonstrate the accuracy of CCS for recovering the relevant biclusters irrespective of the number and pattern of the biclusters and size of the datasets.
Performance on gene expression data
CCS biclustering algorithm obtained total 25 biclusters from five gene expression datasets (Table 2). Figure 2 shows the average number of enriched terms from gProfiler enrichment analysis. Figure 2 shows there are more enriched terms per bicluster from CCS than the others. Figure 3 compares the biclustering algorithms for the percentage of biclusters with at least one enriched term. Again, CCS obtained the highest percentage of enriched biclusters (Figure 3).
GPU accelerated biclustering
The CPU version of the CCS algorithm is computationally expensive. If the ‘base_number’ is ‘n’ then step 5 executes ‘n’ times. Again for ‘n’ iterations of step 5 the augmentation task at step 10 executes ‘n’ times. Hence, the worst case time complexity of CCS is O(n3). The CUDA C implementation of CCS eliminates the outer loop and runs from step 5 to step 25 as CUDA C kernel function. This reduces the time complexity to O(n2). We compared the execution time of the CPU and GPU implementations of CCS. Figure 4 shows the speedup gained from CCS running on the NVIDIA Tesla K20 and K80 GPUs. For larger datasets the GPU-based CCS runs more than 20 times faster than the CPU-based CCS.
CCS biclusters are equivalent to condition-dependent coexpression network modules
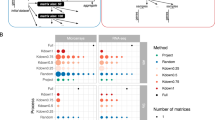
A network module is usually defined as a highly connected sub-network. A module in coexpression network consists of a group of genes whose expression levels are highly correlated32. The correlative relations in a coexpression network often depends on conditions such as genotype, environment, treatment, cell type, disease state or developmental stage33,34,35,36,37,38. Therefore, coexpression network modules may also change with those conditions. There is an interesting relation between CCS biclusters and coexpression network modules. A CCS bicluster consists of genes whose expression levels are highly correlated under a set of conditions. Therefore, a CCS bicluster is equivalent to a condition-dependent module in a coexpression network. This has been illustrated here from two CCS-biclusters. We picked two CCS-biclusters from GDS589 dataset. The first bicluster (bicluster 1) includes 166 genes and 70 samples. The second bicluster (bicluster 2) includes 81 genes and 60 samples. We also identified all the neighboring genes of the biclusters. Neighboring genes are the genes that are not from a bicluster but they are correlated with at least one gene in that bicluster. Figure 5(A) shows the coexpression network of the genes in two biclusters and their neighboring genes. Because this network shows coexpression over all the samples, none of the biclusters are forming a network module. In contrast, when the coexpression network is constructed using correlations over the samples of bicluster 1, the genes of this bicluster form a network module, but the genes of bicluster 2 do not form a module (Figure 5(B)). Similarly, genes of bicluster 2 form a module when the coexpression network is based on correlations over samples of bicluster 2, but genes from bicluster 1 do not form a module in this case (Figure 5(C)).
The coexpression network related to two CCS biclusters from GDS589. The blue, green and white nodes represent the genes from bicluster 1, bicluster 2 and the neighboring genes respectively. Red nodes are common genes between bicluster 1 and 2. Edges represent correlation >=0.8 or <=−0.8. (A) Coexpression network based on correlations over all the samples in GDS589. (B) Coexpression network based on correlations over the samples of bicluster 1 (C) Coexpression network based on correlations over the samples of bicluster 2.
Conclusion
In this work we designed a novel algorithm, CCS, to discover the functionally coherent biclusters from large-scale gene expression datasets. The performance of CCS was compared with thirteen widely used biclustering algorithms. CCS consistently outperforms all the other algorithms in the comparison for obtaining true biclusters from synthetic datasets and discovering functionally enriched biclusters from the real gene expression datasets. Moreover, the CCS biclusters are equivalent to condition-dependent modules in coexpression networks. This important feature makes the CCS algorithm also useful for the study of the condition-dependent structural characteristics of the coexpression networks. The biclusters and network modules of co-regulated and co-functional genes discovered by the CCS algorithm may provide an important entry point for many other analyses such as gene set enrichment analysis, regulatory network inference and disease genes identification.
The recent development of fast and accurate data acquisitions and quantifications in genomics, transcriptomics and proteomics greatly increased the data volume that needs to be processed by clustering algorithms. To enable rapid discovery of high quality biclusters, we implemented CCS algorithm using CUDA C for parallel GPGPU computing. For large datasets, the GPU implementation of CCS achieved about 20 fold speedup compared to the sequential version of CCS. We expect that the GPU- accelerated biclustering will have important applications in the time-sensitive analysis of genomic medicine data.
References
Bhattacharya, A., Chowdhury, N. & De, R. K. Comparative Analysis of Clustering and Biclustering Algorithms for Grouping of Genes: Co-Function and Co-Regulation. Current Bioinformatics 7, 63–76 (2012).
Andreopoulos, B., An, A., Wang, X. & Schroeder, M. A roadmap of clustering algorithms: finding a match for a biomedical application. Brief Bioinform 10, 297–314 (2009).
Rui, X. & Wunsch, D. C. Clustering Algorithms in Biomedical Research: A Review. Biomedical Engineering, IEEE Reviews in 3, 120–154 (2010).
Thalamuthu, A., Mukhopadhyay, I., Zheng, X. & Tseng, G. C. Evaluation and comparison of gene clustering methods in microarray analysis. Bioinformatics 22, 2405–2412 (2006).
Bhattacharya, A. & De, R. K. Divisive Correlation Clustering Algorithm (DCCA) for grouping of genes: detecting varying patterns in expression profiles. Bioinformatics 24, 1359–66 (2008).
Lee, K. et al. Proteome-wide remodeling of protein location and function by stress. Proceedings of the National Academy of Sciences 111, E3157–E3166 (2014).
Segal, E. et al. Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data. Nat Genet 34, 166–176 (2003).
Eren, K., Deveci, M., Kucuktunc, O. & Catalyurek, U. V. A comparative analysis of biclustering algorithms for gene expression data. Brief Bioinform 14, 279–92 (2013).
Oghabian, A., Kilpinen, S., Hautaniemi, S. & Czeizler, E. Biclustering Methods: Biological Relevance and Application in Gene Expression Analysis. PLoS ONE 9, e90801 (2014).
Verma, N. K. et al. A comparison of biclustering algorithms. International Conference on Systems in Medicine and Biology (ICSMB), 90–97 (2010).
Pontes, B., Giráldez, R. & Aguilar-Ruiz, J. S. Biclustering on expression data: A review. Journal of Biomedical Informatics 57, 163–180 (2015).
Padilha, V. A. & Campello, R. J. G. B. A systematic comparative evaluation of biclustering techniques. BMC Bioinformatics 18, 55 (2017).
Bhattacharya, A. & De, R. K. Bi-correlation clustering algorithm for determining a set of co-regulated genes. Bioinformatics 25, 2795–801 (2009).
Bozdağ, D., Parvin, J. D. & Catalyurek, U. V. A biclustering method to discover co-regulated genes using diverse gene expression datasets. In Bioinformatics and Computational Biology 151–163 (Springer, 2009).
Yun, T. & Yi, G. S. Biclustering for the comprehensive search of correlated gene expression patterns using clustered seed expansion. BMC Genomics 14, 144 (2013).
Zou, Q. et al. Survey of MapReduce frame operation in bioinformatics. Briefings in Bioinformatics 15, 637–647 (2014).
Zou, Q., Hu, Q., Guo, M. & Wang, G. HAlign: Fast multiple similar DNA/RNA sequence alignment based on the centre star strategy. Bioinformatics 31, 2475–2481 (2015).
Ocaña, K. & De Oliveira, D. Parallel computing in genomic research: Advances and applications. Advances and Applications in Bioinformatics and Chemistry 8, 23–35 (2015).
Allocco, D. J., Kohane, I. S. & Butte, A. J. Quantifying the relationship between co-expression, co-regulation and gene function. BMC Bioinformatics 5, 18 (2004).
Prelic, A. et al. A systematic comparison and evaluation of biclustering methods for gene expression data. Bioinformatics 22, 1122–9 (2006).
Reimand, J., Arak, T. & Vilo, J. g: Profiler–a web server for functional interpretation of gene lists (2011 update). Nucleic Acids Res 39, W307–15 (2011).
Cheng, Y. & Church, G. M. Biclustering of expression data. Proc Int Conf Intell Syst Mol Biol 8, 93–103 (2000).
Bergmann, S., Ihmels, J. & Barkai, N. Iterative signature algorithm for the analysis of large-scale gene expression data. Phys Rev E Stat Nonlin Soft Matter Phys 67, 031902 (2003).
Lazzeroni, L. & Plaid, O. A. models for gene expression data. Stat Sin 12, 61–86 (2000).
Kluger, Y., Basri, R., Chang, J. T. & Gerstein, M. Spectral biclustering of microarray data: coclustering genes and conditions. Genome Res 13, 703–16 (2003).
Ben-Dor, A., Chor, B., Karp, R. & Yakhini, Z. Discovering local structure in gene expression data: the order-preserving submatrix problem. J Comput Biol 10, 373–84 (2003).
Murali, T. M. & Kasif, S. Extracting conserved gene expression motifs from gene expression data. Pac Symp Biocomput, 77–88 (2003).
Tanay, A., Sharan, R., Kupiec, M. & Shamir, R. Revealing modularity and organization in the yeast molecular network by integrated analysis of highly heterogeneous genomewide data. Proc Natl Acad Sci USA 101, 2981–6 (2004).
Henriques, R. & Madeira, S. C. BicSPAM: flexible biclustering using sequential patterns. Bmc Bioinformatics 15 (2014).
Wang, Z. J., Li, G. J., Robinson, R. W. & Huang, X. Z. UniBic: Sequential row-based biclustering algorithm for analysis of gene expression data. Scientific Reports 6 (2016).
Shamir, R. et al. EXPANDER–an integrative program suite for microarray data analysis. BMC Bioinformatics 6, 232 (2005).
Fuller, T., Langfelder, P., Presson, A. & Horvath, S. Review of Weighted Gene Coexpression Network Analysis. in Handbook of Statistical Bioinformatics (eds. Lu, H.H.-S., Schölkopf, B. & Zhao, H.) 369–388 (Springer Berlin Heidelberg, 2011).
de la Fuente, A. F. ‘differential expression’ to ‘differential networking’ – identification of dysfunctional regulatory networks in diseases. Trends in Genetics 26, 326–333 (2010).
Li, W. et al. Pattern Mining Across Many Massive Biological Networks. In Functional Coherence of Molecular Networks in Bioinformatics (eds. Koyutürk, M., Subramaniam, S. & Grama, A.) 137–170 (Springer New York, 2012).
Li, H. et al. Integrative Genetic Analysis of Transcription Modules: Towards Filling the Gap between Genetic Loci and Inherited Traits. Hum. Mol. Genet. 15, 481–492 (2006).
Bao, L. et al. An integrative genomics strategy for systematic characterization of genetic loci modulating phenotypes. Hum. Mol. Genet. 16, 1381–1390 (2007).
Bao, L. et al. Combining gene expression QTL mapping and phenotypic spectrum analysis to uncover gene regulatory relations. Mammalian Genome 17, 575–583 (2006).
Miyairi, I. et al. Host Genetics and Chlamydia Disease: Prediction and Validation of Disease Severity Mechanisms. PLoS ONE 7, e33781 (2012).
Tian, E. et al. The role of the Wnt-signaling antagonist DKK1 in the development of osteolytic lesions in multiple myeloma. N Engl J Med 349, 2483–94 (2003).
Walker, J. R. et al. Applications of a rat multiple tissue gene expression data set. Genome Res 14, 742–9 (2004).
Boni, J. P. et al. Population pharmacokinetics of CCI-779: correlations to safety and pharmacogenomic responses in patients with advanced renal cancer. Clin Pharmacol Ther 77, 76–89 (2005).
Xu, L. et al. Gene expression changes in an animal melanoma model correlate with aggressiveness of human melanoma metastases. Mol Cancer Res 6, 760–9 (2008).
Sato, T. et al. PRC2 overexpression and PRC2-target gene repression relating to poorer prognosis in small cell lung cancer. Sci Rep 3, 1911 (2013).
Acknowledgements
The authors gratefully acknowledge the editorial assistance of Dr. Martha M. Howe provided through the Writing Assistance Program of the College of Graduate Health Sciences. Funding: This work was partly supported by The University of Tennessee Center for Integrative and Translational Genomics. A.B. was supported in part by P-41-RR24851.
Author information
Authors and Affiliations
Contributions
A.B. and Y.C. conceived the idea of the work, A.B. implemented and tested the algorithms, A.B. and Y.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bhattacharya, A., Cui, Y. A GPU-accelerated algorithm for biclustering analysis and detection of condition-dependent coexpression network modules. Sci Rep 7, 4162 (2017). https://doi.org/10.1038/s41598-017-04070-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-04070-4
This article is cited by
-
Two-mode clustering through profiles of regions and sectors
Empirical Economics (2022)
-
Rank-preserving biclustering algorithm: a case study on miRNA breast cancer
Medical & Biological Engineering & Computing (2021)
-
BicBioEC: biclustering in biomarker identification for ESCC
Network Modeling Analysis in Health Informatics and Bioinformatics (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.