Abstract
The Indian Ocean has long been a hub of interacting human populations. Following land- and sea-based routes, trade drove cultural contacts between far-distant ethnic groups in Asia, India, the Middle East and Africa, creating one of the world’s first proto-globalized environments. However, the extent to which population mixing was mediated by trade is poorly understood. Reconstructing admixture times from genomic data in 3,006 individuals from 187 regional populations reveals a close association between bouts of human migration and trade volumes during the last 2,000 years across the Indian Ocean trading system. Temporal oscillations in trading activity match phases of contraction and expansion in migration, with high water marks following the expansion of the Silk Roads in the 5th century AD, the rise of maritime routes in the 11th century and a drastic restructuring of the trade network following the arrival of Europeans in the 16th century. The economic fluxes of the Indian Ocean trade network therefore directly shaped exchanges of genes, in addition to goods and concepts.
Similar content being viewed by others
Introduction
For more than 2,000 years, the Indian Ocean rim has been an area of intense interaction between African, Middle Eastern and Asian populations, driven by a strong tradition of wealthy maritime and land-based trading routes1, 2. The political unification of large territories in the third century BCE (Before Current Era) opened up new trading routes, most famously the Silk Roads and the maritime network along the coasts of the Indian Ocean3, 4. These in turn triggered sustained interactions between major geopolitical poles, including states in China, India, Indonesia, Arabia and East Africa1, 5, 6. Trade was both diverse and intense, benefiting from specialized local production, such as cotton and beads from India, gold from East Africa, spices from the Malacca city-states, incense from Yemen and silk from China1. With population growth and technical advances, notably in agriculture, the first century CE saw a major intensification in the movements of goods and people3. The development of new sailing techniques, particularly during the 11th century CE, enabled movements over very long distance. Indonesian traders reached as far as East Africa and the Swahili city-states7; Arab sailors installed trading posts on Madagascar and the west coast of India1; and Chinese traded across Island Southeast Asia and East India. Far from competing, the various maritime and terrestrial routes created an intertwined and dense network that rapidly diffused goods, but also knowledge, beliefs and values, proving a unifying force across a diverse set of partners1, 3, 6. New urban spaces acted as hubs to the flow of trade and culture, often growing into cosmopolitan cities with large immigrant populations, such as in Baghdad and Zanzibar8, 9. The intensity, stability and speed of these trading connections formed a large world-system, a precursor to the heavily globalized societies of today10. Yet whether Indian Ocean trade directly shaped the genetics of modern populations is less well understood.
Results
Genome-wide genetic variation in 3,006 individuals from 187 regional populations was used to build a picture of gene flow around the Indian Ocean rim over the past 2,000 years (Supplementary Table 1). The genetic landscape of the Indian Ocean rim today, as characterized by ADMIXTURE11 and EEMS12, is a structured space with distinct regional genetic ancestries allowing the fine-scale reconstruction of historical human migrations mediating gene flow (Supplementary Figures 1–6). Long corridors of genetic similarity can be seen along the coasts of East Africa, South Asia and the rim of the China Sea, but also strong genetic barriers such as one observed between South Asia and East Africa (Supplementary Figure 1). Despite these genetic barriers remarkable instance of gene flow can be identified such as the Asian gene flow to Madagascar (Supplementary Figures 4 and 5)13, 14.
To determine whether trade drove significant bouts of population mixing, the temporal pulses of genetic dispersal around the Indian Ocean were estimated with GLOBETROTTER15 and MALDER16 (Supplementary Tables 2 and 3). Consistent with the idea that trading activities stimulate biological contacts, both analyses indicate that migration is highly correlated with historical trading volumes9 (r 2 = 0.89, P = 0.00001; Fig. 1; Supplementary Tables 4, 5 and Supplementary Figure 7). This model is a significantly better fit to the data than a simple increase of the number of admixture events over time (P = 0.009; Supplementary Table 5), which might be expected due to the statistical bias of the software towards estimating more recent admixture events. While the overall intensity of trade and population interactions increased steadily over time, these activities instead directly track the periodicity of economic developments and recession9. Four major phases of trade have previously been described9, with intervening recessions not breaking the network but instead restructuring connections leading to a new dynamic9. These periods of increased trade are associated with bursts of human migration (F = 10.39, P = 0.0002; Fig. 1; Supplementary Table 6).
Association between human migration and trade volumes through time. The green line shows the number of migration events estimated by GLOBETROTTER15 per century. The red line shows the average volume of trade per century adapted from data detailed by Beaujard9. The blue line shows the average distance of migration per century, with light blue shading showing the variance. Dashed vertical lines mark the four trade phases9. Horizontal bars represent t-tests between successive phases with significance values shown for the variance of migration distances (light blue), the average migration distance (dark blue) and the number of migration events (green): **P < 0.01; *P < 0.05; -: not significant.
Despite technological advancements and the expansion of Indian Ocean trade, the average migration distance did not increase through time (P > 0.05; Fig. 1; Supplementary Table 5). However, migration distances did fluctuate, with periods of extreme migration followed by contractions in population movements, corresponding to the phases of trade (Fig. 1; F average = 5.34, P = 0.01; F variance = 4.33, P = 0.03; Supplementary Table 6).
The first phase (1st–5th century) reflects the rise of the Silk Roads1, 2, dominated by terrestrial and coastal migrations from China to Arabia (Fig. 2A; Supplementary Table 2). Major gene flows are detected in the northern part of India corresponding to the influential zone of the Gupta Empire (330–550 CE), as previously identified17. This gene flow highlights long-distance interactions with Southeast Asia and China, which were the main actors for trade in items sold to West Eurasian and Middle East markets. At this period, Arab merchants were already dominating the Near East trading routes, whose influence can be seen by the numerous gene flows originating from the Arabian peninsula (Fig. 2A; Supplementary Table 2)18, 19.
The consolidation of the second Pax Sinica by the Chinese Tang Empire stimulated the Indian Ocean world-system to expand further3, 4 (Phase II; 6th–10th century). This occurred in parallel with the spread of Islam by Arab merchants, marked by an intensification of gene flows from the Middle East to East Africa and Central Asia (Fig. 2B; Supplementary Table 2), as also reported in previous studies19, 20. Although causes of recessions are always multi-factorial, trade conditions likely declined at the end of Phase II due to the demographic collapse of urban centers in the Middle East, such as Baghdad following the fall of the Abbasid Caliphate in Arabia, and perhaps exacerbated by arid climatic conditions affecting agricultural production in Central Asia3. The end of Phase II is characterized by both a significant reduction in human migration and smaller migration distances (P < 0.05; Figs 1 and 2B; Supplementary Table 6).
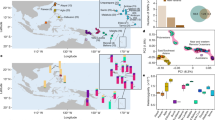
Maps showing gene flow during the four phases of Indian Ocean trade. Admixture events during two century periods are plotted for each phase: (A) Phase I, (B) Phase II, (C) Phase III and (D) the beginning of Phase IV. Arrow widths are proportional to the percentage of ancestry inherited from each source population. Colors are specific to each cluster, as defined by the fineSTRUCTURE31 results: green palette: sub-Saharan African Pygmies, Bantu, East African and Malagasy clusters; yellow palette: Middle East, Nilo-Saharan and Caucasus clusters; purple palette: Pakistan, India and Bengal clusters; blue palette: East and North Asian clusters; red palette: Indonesian clusters; brown and orange palettes: Negrito, Andaman and Papuan clusters. Blue dots illustrate the locations of sampled populations. Maps were generated using Global Mapper v.15. (http://www.bluemarblegeo.com/products/global-mapper.php).
Major technical improvements in sailing, such as the widespread adoption of the compass, likely triggered Phase III (11th–14th century) with the appearance of new maritime routes2 (Fig. 2C; Supplementary Table 2). Hindu Malay Empires, such as Srivijaya and Mojopahit in Island Southeast Asia3, dominated this reinvigorated maritime trade, notably with Chinese and Indian Empires, as can be seen by numerous gene flows occurring between these areas (Fig. 2C; Supplementary Table 2). This state of domination was followed by Austronesian settlements in the Comoros and in Madagascar21, which we had previously established13, 14. We note that no other Austronesian gene flow to the western rim of the Indian Ocean was inferred by our analyses, suggesting a direct route of migration to Madagascar. Along with the development of the Swahili Corridor22, which can be inferred from the high density of gene flows in East Africa at that time, Arab merchants developed trading posts on the East African coast, to increase their access to gold and slaves. These slaves were deported to Arabia and South Asia, as shown by the South African Bantu gene flow into Yemen and South Pakistan15, 23. Finally, we also detected gene flow between Mongols and populations from Central Asia and Anatolia24, as well as Turkish gene flows to Middle Eastern groups, converging with the Mongol migration which started in 1206 with the reign of Genghis Khan and reached Arabia in 1258. These vast migrations participated in the unification of Indian Ocean trade partners at an unprecedented geographical scale.
However, the most drastic change occurred in the middle of the 14th century (Fig. 2D; Supplementary Table 6). Outbreaks of plague in Asia, Africa and Europe combined with climatic changes led to a major demographic crisis9. This is reflected in major geopolitical restructuring, such as the fall of the Chinese Yuan Empire, which controlled the terrestrial Silk Roads, and the prohibition of trade between China and Southeast Asia, dictated by the Ming Empire in 1433 AD3. These events are mostly noticeable in our analyses by reduced gene flow in Island Southeast Asia and increased migration within China. An additional shock included the arrival of Europeans in the 16th century, a major disruptive factor. This recession was followed by Phase IV (15th–16th centuries to the present), with the Industrial Revolution driving another long period of strong international trade.
All of these trade spikes were paired with physical population movements, showing that the flow of goods and ideas was linked to the movements of the people who brought them. At least in the Indian Ocean, at a global scale, trade and migration were coupled forces, and the Indian Ocean trade network therefore provides an early example of globalization, showing connections between human trade and mobility that are still apparent around the world today.
Methods
Dataset
Our dataset is based on previously published studies of populations around the Indian Ocean rim (Supplementary Table 1). This dataset was built to trade-off large population diversity with the high number of overlapping SNPs, necessary for Identity-by-Descent (IBD) based methods, given the wide spectrum of genotyping platforms used by the scientific community. To avoid any statistical bias that could be introduced by a size effect of over-represented populations, we randomly selected a maximum of 25 individuals in each group, such that each population has sample size between 3 and 25. Quality controls were applied using Plink v1.925 to filter for i) close relatives, using an IBD estimation with upper threshold of 0.25 (second degree relatives); ii) SNPs that failed the Hardy-Weinberg exact (HWE) test (P < 10−6) were excluded; iii) samples with a call rate <0.99 and displayed missing rates >0.05 across all samples in each population were excluded. After filtering, our dataset included a total of 3,006 individuals, genotyped for 215,335 SNPs, from 187 different populations located in Southeast Asia, South Asia, East Asia, Middle East, East Africa, South Africa, and Europe (Supplementary Table 1). All genotypes were phased together with SHAPEIT v2.r79026 using the 1000Genomes phased data27 as reference panel and the HapMap phase 2 genetic map28. The same dataset was in parallel pruned for Linkage Disequilibrium (LD; r2 < 0.2) with Plink v1.925 resulting in an alternative dataset of 100,830 SNPs for specific analyses.
Statistical Analyses
The genetic diversity of our dataset pruned for LD was first analyzed by EEMS v112 to define genetic barriers and corridors. Using geographic coordinates (with the noticeable exception of HGDP CEU samples placed in Germany for graphical convenience) and a genetic dissimilarity matrix between populations, we set a map of the Indian Ocean rim defining a grid of 1,000 demes. Depending on their location, several populations may be included in one deme. 3 × 106 MCMC iterations were run before checking for convergence of the MCMC chain. Plots were generated in R following the EEMS v112 manual (Supplementary Figures 1 and 2). ADMIXTURE v1.2311 was used with default settings to decompose genetic ancestries of the pruned dataset. Ten iterations with randomized seeds were run and compiled with CLUMPAK v129. We use the minimum average cross-validation value to define the most descriptive K components, and the major modes defined by CLUMPAK v129 are reported. Plots were obtained using Genesis v.0.2.530. The lowest cross-validation value was obtained for K = 29 (Supplementary Figures 4–6). Both of these analyses were used together to define the genetic diversity of our dataset.
Before estimating admixture scenarios, we defined clusters of populations. We first performed a fineSTRUCTURE v2.0731 analysis using the phased dataset to define genetic clusters31 (Supplementary Figure 3). This method detects shared IBD fragments between each pair of individuals, without self-copying, calculated with CHROMOPAINTER v2.031 (default settings) to perform a model-based Bayesian clustering of genotypes. Mutational rates (Mu) and effective population size (Ne) were estimated with an Estimation-Maximization (EM) algorithm running in CHROMOPAINTER v2.031, and was performed on all 22 autosomes for the entire dataset (10 iterations). The weighted average of these parameters, according to the SNP coverage of each chromosome and the number of individuals, was then used to compute the chromosome painting. Using fineSTRUCTURE v2.0731 with 2 × 106 Markov-Chain-Monte-Carlo (MCMC) iterations, discarding the first 1 × 106 iterations as “burn-in”, sampling from the posterior distribution every 10,000 iterations following the burn-in, a coancestry heat map and a dendrogram were inferred to visualize the number of clusters defined statistically that best describe the data.
This analysis defined clusters of populations that share a similar genetic history, so that populations in one cluster cannot be used as a parental group for another population in the same cluster. This criterion is critical to avoid statistical bias for the following analyses, notably GLOBETROTTER v2.015. Population clusters were defined in two steps. From the fineSTRUCTURE v2.0731 results, each cluster was defined by: (i) high posterior probabilities given for the nodes of the population dendogram (>0.8); (ii) at least 100 individuals per cluster. This last criteria, although arbitrary, allowed us to define uniform clusters from statistically robust branches higher up in the tree, in a similar approach to that reported previously31. Subsequently an FST matrix was calculated with Eigensoft v5.0.232 between populations within each cluster to define outliers with FST values greater than one standard deviation from the mean (Supplementary Table 7). Eight populations called as outliers could only be defined as ‘surrogates’ within their respective cluster (Supplementary Table 7). Those outliers were not analysed as a ‘target’ as they all show positive f3-statistics33 results and are known to have no recent history of admixture33,34,35,36,37 (Supplementary Table 7). After all criteria were applied, 22 clusters were defined (Supplementary Table 1 and Supplementary Figure 4).
To test different scenarios of admixture we performed GLOBETROTTER v2.015 analyses for each population (excluding outliers) defining surrogates populations from all clusters but its own. Note that the numbers of populations within a given cluster is not correlated with the estimated dates of admixture (P = 0.95). The painted chromosomes obtained by CHROMOPAINTER v2.031 for each population were used in GLOBETROTTER v2.015 to estimate the ratios and dates of the potential admixture events that characterize them. Coancestry curves were estimated with and without standardization using a ‘NULL’ individual, and consistency for each estimated parameter was checked. 100 bootstrap resamplings were performed to estimate the p-value of the admixture events (considering the ‘NULL’ individual) and the 95% confidence interval for the obtained dates. The ‘best-guess’ scenario given by GLOBETROTTER v2.015 was considered for each target population. Admixture events whose estimated 95% confidence intervals of dates between both ‘NULL0’ and’NULL1’ models do not overlap were subsequently reclassified as ‘uncertain’, as described by Hellenthal et al.15. To obtain a second estimate of potential admixture scenarios, we ran MALDER v116, a modified version of the ALDER v1.316 software to observe any multiple admixture events, using the parental populations defined by GLOBETROTTER v2.015. Both analyses give highly correlated admixture dates (r2 = 0.65; P < 0.00001; Supplementary Figure 7). The estimated dates likely reflect the midpoint or end of noticeable admixture events rather than the exact date of migration (which could occur prior to any admixture). The number of migrations per century is equivalent to the cumulative number of parental populations, given by the ‘best-matching’ scenario in GLOBETROTTER v2.015, and involved in each admixture event not defined as ‘uncertain’ (Supplementary Tables 3 and 4). Dates of admixture, given in generations, were converted to chronological time using a generation interval of 25 years.
In parallel with the admixture dates obtained from GLOBETROTTER15 and MALDER16, we computed geographical distances and historical trading volumes in order to perform correlation tests. Euclidian distances between a given target group, whose admixture scenario is not defined as ‘uncertain’ in GLOBETROTTER v2.015, and each of its parental sources, given by the ‘best-matching’ scenario, were calculated using the great circle formula. We used the estimated value of 6,371 km for the Earth’s radius. Average volumes of trade per century were calculated from historical data9. Five measures of trade volume were taken for each century to obtain an average per century. This allows us to compare these data to the number of admixture events per century. However we note that this also smoothes the exact evolution of trade, as more precisely described in Beaujard9, as for example the drastic decrease observed at the end of Phase III9 which appears more progressive in our representation (Fig. 1). Therefore this measure does not reflect the exact evolution of trade volume but rather its overall trend across the centuries. Descriptive statistics of the distance of gene flow per century, correlation tests, t-tests and analyses of variance were computed with SPSS v20.038. Curves were generated in SPSS v20.038 using the spline interpolation. We performed correlation tests between our variables as we did not put any assumption on the dependence of one to another (for example, trade on the number of migrations). We performed an F-test to compare the models with time and volume of trade using SPSS v20.038. When required, the Bonferroni multiple testing correction was applied. Networks were generated with Cytoscape v3.2.139 and maps were generated using Global Mapper v.15. (http://www.bluemarblegeo.com/products/global-mapper.php).
References
Chaudhuri, K. N. Trade and Civilization in the Indian Ocean: An Economic History from the Rise of Islam to 1750. (Cambridge University Press, 1985).
Lawler, A. Sailing Sinbad’s seas. Science 344, 1440–1445, doi:10.1126/science.344.6191.1440 (2014).
Beaujard, P. Les mondes de l’ocean indien. Vol. 2: L’océan Indien, au coeur des globalisations de l’Ancien Monde (7e-15e siècles). Vol. 2 (Armand Collin, 2012).
Beaujard, P. Les mondes de l’océan Indien. Vol. 1: De la formation de l’État au premier système-monde afro-eurasien (4e millénaire av. J.-C.-6e siècle apr. J.-C.). Vol. 1 (Armand Collin, 2012).
Edens, C. Comments on Frank, 1993. Current Anthropology 34, 408–409 (1993).
Frank, G. A. Bronze Age World System Cycles. Current Anthropology 34, 383–405 (1993).
Boivin, N., Crowther, A., Helm, R. & Fuller, D. Q. East Africa and Madagascar in the Indian Ocean world. J World Prehist 26, 213–281 (2013).
LaViolette, A. Swahili Cosmopolitanism in Africa and the Indian Ocean World, A.D. 600–1500. Archaeologies: Journal of the World Archaeological Congress 4, 24–49 (2008).
Beaujard, P. The Indian Ocean in Eurasian and African World-Systems before the Sixteenth Century. Journal of World History 16, 441–465 (2005).
Wallerstein, I. The Modern World System I: Capitalist Agriculture and the Origins of the European World-Economy in the Sixteenth Century. (Academic Press Inc., 1976).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664, doi:10.1101/gr.094052.109 (2009).
Petkova, D., Novembre, J. & Stephens, M. Visualizing spatial population structure with estimated effective migration surfaces. Nat Genet 48, 94–100, doi:10.1038/ng.3464 (2016).
Brucato, N. et al. Malagasy Genetic Ancestry Comes from an Historical Malay Trading Post in Southeast Borneo. Mol Biol Evol 33, 2396–2400, doi:10.1093/molbev/msw117 (2016).
Pierron, D. et al. Genome-wide evidence of Austronesian–Bantu admixture and cultural reversion in a hunter-gatherer group of Madagascar. PNAS 111, 936–941, doi:10.1073/pnas.1321860111 (2014).
Hellenthal, G. et al. A genetic atlas of human admixture history. Science 343, 747–751, doi:10.1126/science.1243518 (2014).
Loh, P. R. et al. Inferring admixture histories of human populations using linkage disequilibrium. Genetics 193, 1233–1254, doi:10.1534/genetics.112.147330 (2013).
Basu, A., Sarkar-Roy, N. & Majumder, P. P. Genomic reconstruction of the history of extant populations of India reveals five distinct ancestral components and a complex structure. Proc Natl Acad Sci USA 113, 1594–1599, doi:10.1073/pnas.1513197113 (2016).
Cerny, V., Cizkova, M., Poloni, E. S., Al-Meeri, A. & Mulligan, C. J. Comprehensive view of the population history of Arabia as inferred by mtDNA variation. Am J Phys Anthropol 159, 607–616, doi:10.1002/ajpa.22920 (2016).
Haber, M. et al. Genome-wide diversity in the levant reveals recent structuring by culture. PLoS Genet 9, e1003316, doi:10.1371/journal.pgen.1003316 (2013).
Eaaswarkhanth, M. et al. Traces of sub-Saharan and Middle Eastern lineages in Indian Muslim populations. Eur J Hum Genet 18, 354–363, doi:10.1038/ejhg.2009.168 (2010).
Crowther, A. et al. Ancient crops provide first archaeological signature of the westward Austronesian expansion. Proc Natl Acad Sci USA 113, 6635–6640, doi:10.1073/pnas.1522714113 (2016).
Horton, M. The Swahili Corridor. Scientific American 255, 86–93 (1986).
Blench, R. In Afriques on East Africa and the Indian Ocean (eds T. Vernet & P. Beaujard) 2–18 (2008).
Mezzavilla, M. et al. Genetic landscape of populations along the Silk Road: admixture and migration patterns. BMC Genet 15, 131, doi:10.1186/s12863-014-0131-6 (2014).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4, 7, doi:10.1186/s13742-015-0047-8 (2015).
Delaneau, O., Marchini, J. & Zagury, J. F. A linear complexity phasing method for thousands of genomes. Nature methods 9, 179–181, doi:10.1038/nmeth.1785 (2012).
Delaneau, O. & Marchini, J., Genomes Project, C. & Genomes Project, C. Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel. Nat Commun 5, 3934, doi:10.1038/ncomms4934 (2014).
International HapMap, C. A haplotype map of the human genome. Nature 437, 1299–1320, doi:10.1038/nature04226 (2005).
Kopelman, N. M., Mayzel, J., Jakobsson, M., Rosenberg, N. A. & Mayrose, I. Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Resour 15, 1179–1191, doi:10.1111/1755-0998.12387 (2015).
Genesis v.0.2.5, URL http://www.bioinf.wits.ac.za/software/genesis (2014).
Lawson, D. J., Hellenthal, G., Myers, S. & Falush, D. Inference of population structure using dense haplotype data. PLoS Genet 8, e1002453, doi:10.1371/journal.pgen.1002453 (2012).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet 2, e190, doi:10.1371/journal.pgen.0020190 (2006).
Patterson, N. J. et al. Ancient admixture in human history. Genetics 192, 1065–1093, doi:10.1534/genetics.112.145037 (2012).
Morseburg, A. et al. Multi-layered population structure in Island Southeast Asians. Eur J Hum Genet. doi:10.1038/ejhg.2016.60 (2016).
Kusuma, P. et al. Contrasting Linguistic and Genetic Influences during the Austronesian Settlement of Madagascar. Scientific Reports 6, 26066, doi:10.1038/srep26066 (2016).
Metspalu, M. et al. Shared and unique components of human population structure and genome-wide signals of positive selection in South Asia. Am J Hum Genet 89, 731–744, doi:10.1016/j.ajhg.2011.11.010 (2011).
Behar, D. M. et al. The genome-wide structure of the Jewish people. Nature 466, 238–242, doi:10.1038/nature09103 (2010).
SPSS Statistics v.20, URL https://www.ibm.com/analytics/us/en/technology/spss/ (2011).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504, doi:10.1101/gr.1239303 (2003).
Acknowledgements
We wish to acknowledge support from the laboratoire d’Anthopologie Moléculaire et Imagerie de Synthèse (UMR5288, France) and the GenoToul bioinformatics facility of Génopôle Toulouse Midi Pyrénées, France. This research was supported by French ANR grant number ANR-14-CE31-0013-01 (grant OceoAdapto to F.-X.R.), the French Ministry of Foreign and European Affairs (French Archaeological Mission in Borneo (MAFBO) to F.-X.R.), and the French Embassy in Indonesia through its Cultural and Cooperation Services (Institut Français en Indonésie).
Author information
Authors and Affiliations
Contributions
All authors (N.B., P.K., M.P.C., F.-X.R., H.S., P.B.) contributed to the design of the study. N.B. performed the computational analyses. N.B., P.K., M.P.C., F.-X.R. wrote the manuscript based on input from all other authors (P.B. and H.S.).
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Brucato, N., Kusuma, P., Beaujard, P. et al. Genomic admixture tracks pulses of economic activity over 2,000 years in the Indian Ocean trading network. Sci Rep 7, 2919 (2017). https://doi.org/10.1038/s41598-017-03204-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-03204-y
This article is cited by
-
Genes flow by the channels of culture: the genetic imprint of matrilocality in Ngazidja, Comoros Islands
European Journal of Human Genetics (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.