Abstract
Autophagy is an evolutionarily conserved mechanism in eukaryotes with roles in development and the virulence of plant fungal pathogens. However, few reports on autophagy in oomycete species have been published. Here, we identified 26 autophagy-related genes (ATGs) belonging to 20 different groups in Phytophthora sojae using a genome-wide survey, and core ATGs in oomycetes were used to construct a preliminary autophagy pathway model. Expression profile analysis revealed that these ATGs are broadly expressed and that the majority of them significantly increase during infection stages, suggesting a central role for autophagy in virulence. Autophagy in P. sojae was detected using a GFP-PsAtg8 fusion protein and the fluorescent dye MDC during rapamycin and starvation treatment. In addition, autophagy was significantly induced during sporangium formation and cyst germination. Silencing PsAtg6a in P. sojae significantly reduced sporulation and pathogenicity. Furthermore, a PsAtg6a-silenced strain showed haustorial formation defects. These results suggested that autophagy might play essential roles in both the development and infection mechanism of P. sojae.
Similar content being viewed by others
Introduction
Autophagy is a conserved cellular process in which cytoplasmic contents are degraded within a lysosome or vacuole, and the resulting macromolecular constituents are recycled1, 2. Cytoplasmic materials are degraded to produce amino acids and fatty acids during periods of autophagy3. Therefore, autophagy is essential for cell survival under various stress conditions, such as starvation4. Recent studies have revealed a wide variety of physiological roles for autophagy and its relevance to diseases. In many well-characterized plant pathogenic fungi, such as Magnaporthe grisea, Fusarium graminearum and Ustilago maydis, autophagy has been associated with development and virulence5,6,7,8,9.
During autophagy, cup-shaped, single-membrane-bound structures called isolation membranes appear and expand, resulting in cytosol and organelle sequestration10. Eventually, spherical, double-membrane-bound structures called autophagosomes are formed, which are delivered to lysosomes for fusion or vacuoles for degradation of contents11. This process requires the concerted actions of a distinctive set of proteins named ATG (autophagy-related). The core autophagy machinery can be divided into different subgroups: the Atg1 protein kinase complex (Atg1, Atg13 and Atg17–Atg31–Atg29 subcomplex) is essential for initiating induction12; the Atg14-containing phosphatidylinositol-3 kinase complex (beclin 1/Atg6, Vps34 and Vps15 with Atg14), which is essential for the recruitment of ATG proteins to the phagophore assembly site for vesicle nucleation; the Atg2-Atg18 complex and the Atg12-Atg5 protein conjugation system, which are involved in vesicle expansion and completion13, 14; and the Atg9-containing membrane protein recycling system15. ATG genes have been studied in many eukaryotic organisms, such as Saccharomyces cerevisiae, Arabidopsis thaliana, Caenorhabditis elegans and Homo sapiens but have received little attention in oomycete species16,17,18,19.
Oomycetes, classified in the kingdom Stramenopiles, contain many economically important eukaryotic plant pathogens, including Phytophthora sojae 20. Asexual sporangia (or zoosporangia) or zoospores play essential roles in both the initial infection and spread of P. sojae from host to host21. Zoospores are released from sporangia and guided by chemotaxis and electrotaxis towards host roots. Then, the zoospores germinate to form structures known as appressoria that breach the host epidermis. When the health of a colonized plant declines, sporangia and/or zoospores develop and move to another new host22, 23. In addition, Phytophthora species are hemibiotrophic pathogens, with a lifestyle that includes an abiotrophic phase, followed by a switch to necrotrophy24, 25. Although the physiological and genetic mechanisms of sporulation, spore germination, and hemibiotrophic disease cycles have been well-studied, little research has been performed on the role of autophagy in such processes.
In this study, 26 ATG homolog genes from the core autophagy machinery were identified, and a preliminary autophagy pathway model was posited based on core ATGs in oomycetes. P. sojae autophagy was induced with rapamycin and was also activated in P. sojae sporangia and germinating cysts. In addition, a gene orthologous to yeast Atg6, named PsAtg6a, was experimentally silenced in P. sojae, and the PsAtg6a-silenced transformants showed defects in sporangia production and virulence. These results will elucidate the effects of autophagy on the development and virulence of P. sojae.
Results
Identification of ATGs in P. sojae
Several putative ATGs with conserved sequences and function have been identified in yeast, filamentous fungi, plant, and mammalian genomes16, 17, 19, 26. To identify ATGs in P. sojae, a tBlastn search was performed using different ATG sequences from M. grisea, S. cerevisiae, A. thaliana, C. elegans and H. sapiens 27. To further confirm putative ATG homologues in P. sojae, the deduced ATG protein sequences were analyzed in the Pfam database (http://pfam.sanger.ac.uk/) based on S. cerevisiae and H. sapiens ATGs. After removing redundant sequences, a total of 26 putative ATGs were identified in P. sojae (Fig. 1 and Supplementary Table S1). The information returned for each ATG is listed in Supplementary Table S1. As in other species, autophagy in P. sojae can be divided into 4 steps: initiation of autophagy, vesicle nucleation at the preautophagosomal structure (PAS), vesicle expansion, and recycling (Fig. 2). In addition, the ATG prediction pipeline, including building and updating species-specific hidden Markov models (HMMs), was used to predict ATGs in three other oomycetes (24 candidates in Phytophthora capsici, 25 candidates in Pythium ultimum, and 26 candidates in Hyaloperonospora parasitica) and one diatom (12 candidates in Thalassiosira pseudonana) (Fig. 1 and Supplementary Table S1). ATG proteins in these species are categorized based on their function in autophagic processes based on yeast and mammal studies28, 29. The putative ATG distribution was mostly restricted to oomycete species. Furthermore, oomycetes, unlike fungi (M. grisea and S. cerevisiae), lack Atg19, Atg20, Atg21, Atg22, Atg23, Atg24, Atg29 and Atg31 proteins, but Atg1, Atg6, Atg15, Atg18 and Vps34 are expanded, suggesting that the autophagy pathway of oomycete species is more closely related to animals and plants than to fungi.
Distribution of autophagy-related (ATG) proteins in representative species. ATG proteins identified from P. sojae (Ps), P. capsici (Pc), H. parasitica (Hp), Py. ultimum (Pu), T. pseudonana (Tp), A. thaliana (At), C. elegans (Ce), H. sapiens (Hs), S. cerevisiae (Sc), and M. grisea (Mg) were compared. The distribution of putative ATGs appeared to be mostly restricted to oomycete species. Evolutionarily, the autophagy pathway of oomycetes appears more closely related to that of plants and animals than to that of fungi.
In P. sojae, most ATGs seem to be highly conserved compared with different species (Supplementary Table S1). For example, putative Atg8 orthologs encode a similar amino acid sequence and domain structure to Atg8 proteins from other species30 (Fig. 3A). However, some ATGs, such as Atg1, Atg11 and Vps34, are specific to oomycetes (Supplementary Table S1 and Fig. 3). Atg1, the only kinase of the core autophagy machinery, is required for autophagosome formation and the Cvt pathway31. The oomycete protein Atg1b belongs to an ancient family that is conserved in almost all eukaryotic organisms, and the predicted protein shares a C-terminal Pkinase domain. However, an obvious difference from the above is that the DUF3543 domain in conserved PI3K is replaced by an Atg11 domain in Atg1a (Fig. 3B). In S. cerevisiae, Atg11 is the adaptor of the Cvt-specific receptor protein Atg19/Atg3432. However, an extra Atg17 domain aside from the Y-X-X-X-L/V/I-X-E-V/I-X-R-R-L sequences in oomycete Atg11 has been identified (Fig. 3C). In addition, a unique PI3K family that exists only in oomycetes has been reported in previous studies33. All of these data suggest that the core autophagy machinery is conserved in oomycetes with some differences in individual regulation.
Phylogenetic relationship and domain structures of selected ATGs. (A) Sequence features of Atg8 proteins. Atg8 proteins from H. sapiens (GABARAP and MAP1LC3B), A. thaliana (AtAtg8) and P. sojae (PsAtg8) are shown. Atg8 proteins of different species are highly conserved across their entire length, and very similar to each other in most positions. (B) and (C) Phylogenetic relationship and domain structures of Atg1 and Atg11. The phylogenetic trees of Atg1 (B) and Atg11 (C) proteins identified from P. sojae (Ps), P. capsici (Pc), H. parasitica (Hp), Py. ultimum (Pu), T. pseudonana (Tp), A. thaliana (At), C. elegans (Ce), H. sapiens (Hs), S. cerevisiae (Sc), and M. grisea (Mg). Pkinase: kinase domain. Atg11: Atg11 domain. DUF3543: DUF3543 domain. Atg17: Atg17 domain.
Expression profiles of P. sojae ATGs
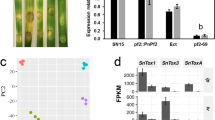
Increasing evidence suggests that autophagy is essential for the development and pathogenesis of many organisms5. To explore the potential role of ATGs in the development and virulence of P. sojae, the expression patterns of 24 ATGs (PsAtg8 and PsAtg12 were excluded) at 10 different stages were obtained from the published DGE transcriptional database34. Heat map analysis was performed based on the relative expression level of each ATG gene (Fig. 4). The majority of ATGs, including PsAtg1a, PsAtg1b, PsAtg2, PsAtg6a, PsAtg6b, PsAtg7, PsAtg10, PsAtg11, PsAtg13, PsAtg15a, PsAtg15b, PsAtg16, PsAtg18b, PsVps15, and PsPI3K1, show a relatively higher expression level (>2-fold expression change) during infection stages (2–14-fold; P ≤ 0.01) (Fig. 4), suggesting that these genes might play important virulence roles. In particular, PsAtg1b (9.0–12.7-fold) and PsAtg6a (7.0–14.0-fold) have a relatively high expression level of the assessed ATGs. Furthermore, the relative expression levels of 3 ATG genes (PsAtg6a, PsAtg8 and PsAtg9) were validated by quantitative RT-PCR (Supplementary Fig. S1). The analysis showed that PsAtg6a was dramatically up-regulated during infection stages, whereas PsAtg9 maintained a steady expression level; these results are comparable to transcriptome data. PsAtg8, which is required for autophagosome formation and is a reliable marker of autophagy induction and progression, visibly changes during infection (Supplementary Fig. S1). These data suggest that autophagy might function in P. sojae virulence.
Heat map of expression profiles for P. sojae ATGs. Color bar represents the log2 expression values, ranging from green (0) to red (8). MY, mycelia; SP, zoosporangia; ZO, zoospores; CY, cysts; GC, germinated cysts; IF1.5 to IF24, indicates samples from 1.5, 3, 6, 12 and 24 h after infection of soybean leaves.
Autophagy induction by rapamycin in P. sojae
Rapamycin, a lipophilic macrolide antibiotic, is a well-established autophagy inducer35. Here, we used three different assays to test rapamycin-induced P. sojae autophagy. The first assay measured the altered location of a GFP-Atg8 fusion protein after processing. During autophagy, Atg8 is normally incorporated into the autophagosome, delivered into the vacuolar lumen, and degraded by vacuolar hydrolases. When the fusion protein undergoes the same process, the stable GFP protein is released36. Thus, Atg8 is the most common marker used to monitor autophagy. In P. sojae, one highly conserved homolog to Atg8 (PsAtg8) was identified (Figs 1 and 3A), and a plasmid with a GFP-PsAtg8 fusion gene driven by the constitutive Ham34 promoter was used for transformation. GFP-PsAtg8-overexpressing transgenic lines were generated using the polyethylene glycol (PEG)-mediated protoplast stable transformation method in P. sojae 37. Using fluorescence microscopy, we found that GFP-PsAtg8 is targeted to the PAS in control (treated with DMSO) mycelia and appeared as elliptical spots. When mycelia were treated with 100 nM rapamycin for 4 h, a decrease in the GFP-PsAtg8 signal was clearly observed (Fig. 5A). Some GFP-PsAtg8 puncta persisted within the vacuole lumen, whereas the majority of PsAtg8 molecules were degradated, and GFP molecules were released into the cytoplasm (Fig. 5A).
Induction of autophagy in response to rapamycin treatment in P. sojae. (A) Autophagy indicated by GFP-PsAtg8. GFP-PsAtg8-expressing P. sojae hyphae were incubated in 10% V8 juice. After treated with 100 nM rapamycin or DMSO (control) for 4 h, hyphae were analyzed by confocal microscopy. Bars = 10 μm. (B) Autophagy indicated by MDC dye. P. sojae wild-type hyphae were incubated in 10% V8 juice. After treated with 100 nM rapamycin or DMSO (control) for 4 h, hyphae samples concurrently stained with MDC were analyzed by confocal microscopy. Bars = 10 μm.
The second assay examined autophagosomes stained with monodansylcadaverine (MDC), a dye that has been widely used as an indicator of autophagic activity38. As shown in Fig. 5B, control P. sojae mycelia exhibit hardly any MDC fluorescence. After mycelia were incubated in 100 nM rapamycin for 4 h, the punctate fluorescent dots from MDC-stained autophagosomes substantially increased in number. It was also found by electron microscopy that more autophagosomes accumulated in the cytoplasm in rapamycin treatment hyphal cells (Supplementary Fig. S2A and S2B). These observations indicated that autophagy was induced in P. sojae by rapamycin as in other eukaryotes.
Autophagic activation in P. sojae sporangia and germinating cysts
Sporangia, part of the asexual life cycle of Phytophthora species, play key roles in diffusion and infection39. In P. sojae, sporangia are induced under nitrogen starvation conditions, which a method used to induce cell autophagy in fungi such as S. cerevisiae and various filamentous fungi4. The dependence on starvation indicates the possible involvement of autophagy in sporulation. To test whether autophagy is activated during sporulation in P. sojae, sporangia were collected and stained with MDC. As shown in Fig. 6A, MDC fluorescence was hardly detected in mycelia cultured in 10% V8 juice. When fresh hyphae were washed with sterile distilled water 3 times and incubated in sterile distilled water for 4 h for sporulation, punctate fluorescent MDC-stained autophagosomal dots increased robustly in some hypha apexes. Furthermore, strong MDC fluorescence was observed in immature sporangia, but very little staining was observed in mature sporangia (Fig. 6A). In addition, when the PsAtg6a gene was silenced in P. sojae, substantially reduced sporulation was detected (Fig. 6B,C). These results suggest that autophagy might play an important role in sporulation and that it is specifically activated during the proliferation of sporangia from the expansion of hypha apex to the subdivision of cytoplasm in sporangia but during not the release stage.
P. sojae sporangial production is regulated by autophagy. (A) Visualization of autophagic activation with MDC in P. soaje mycelia (MY) and zoosporangia (SP). Wild-type hyphae were incubated in 10% V8 juice for 30 h. After 3 washes and incubation with sterile distilled water for 4 h, hyphae were concurrently stained for MDC and analyzed by confocal microscopy. Bars = 10 μm. (B) Numbers of sporangia produced by PsAtg6a transgenic lines. Sporangia of the indicated samples were counted 12 h after the induction of sporangial production. Representative data are shown from three separate experiments. The bars indicate standard errors, and stars above bars indicate that the difference from the wild-type value was significant. **P < 0.01 (t-test). (C) Sporangial phenotypes forPsAtg6a transgenic lines. Micrographs were taken 12 h after sporangial induction. Black arrows indicate sporangia. Bars = 500 μm.
Next, more tissues, including cysts and germinating cysts, were stained with MDC. Zoospores are generally short-lived and quickly differentiate to form cysts, so fluorescence images of swimming zoospores were not detected. As shown in Fig. 7, little staining was detected in resting cysts. However, MDC fluorescence remarkably increased when cysts started to germinate, and there was a slight reduction in fluorescence as the germ tube grew. This pattern of autophagosome formation indicated that the autophagy process was activated during cyst germination, making it plausible that autophagy plays a role in infection.
Construction of PsAtg6a transgenic lines
To further investigate the role of autophagy in the sporulation and pathogenicity of P. sojae, PsAtg6a-silenced transgenic lines were generated using the polyethylene glycol (PEG)-mediated protoplast stable transformation method in P. sojae 37. Atg6, the first identified marker protein for autophagy, is part of a lipid kinase complex and plays a central role coordinating the cytoprotective function of autophagy. In many cases, Atg6 is examined to identify the role of autophagy40. In P. sojae, two Atg6 homologs, PsAtg6a (Ps128490) and PsAtg6b (Ps136998), were identified (Fig. 1). Predictions of the conserved domains showed three conserved β-sheet-α-helix repeats in the autophagy-specific BARA domain (Supplementary Fig. S3A). The phylogenetic tree analyses indicated that PsAtg6a forms a close cluster with AtAtg6, whereas PsAtg6b is close to the Atg6 proteins of H. sapiens (Supplementary Fig. S3B). Because the transcriptional and qRT-PCR analysis showed that PsAtg6a genes were highly up-regulated during infection, PsAtg6a was selected for further study (Fig. 4, Supplementary Fig. S1). The required transformants were preliminarily identified using genomic PCR analysis (Fig. 8A), and the expression levels of target genes were measured using qRT-PCR. As shown in Fig. 8B, PsAtg6a transcription levels were reduced to 50%, 60% and 14% of the wild-type value in lines ST6–17, ST6-21 and ST6-39, respectively. The PsAtg6a gene in T9 was similar to WT, which was used as a control strain. The PsAtg6b gene in all of the transgenic lines was similar to WT. The growth rates of PsAtg6a-silenced strains were similar to that of the wild-type and control T9 (Supplementary Fig. S4A and S4B). Compared to WT, PsAtg6a-silenced strains showed decreased accumulation of autophagic bodies in hyphal cells after culture in sterile distilled water for 4 h (Fig. 8C). These results suggest that the autophagic pathway in PsAtg6a-silenced strains was affected and PsAtg6a might be responsible for autophagy in P. sojae.
Construction of PsAtg6a-silenced lines. (A) Verification of incorporation into genomic DNA by PCR using oligonucleotides from the Ham34 promoter and terminator regions as primers. The transgene should yield an amplified fragment 1520 bp in length. +: plasmid DNA; WT: wild-type strain P6497; M: molecular markers. (B) qRT-PCR measurement of the relative transcript levels of PsAtg6a and PsAtg6b in control (P6497), control (T9) and silenced (ST17, ST21 and ST39) transformants. The relative expression levels were calculated using TEF1 as the reference gene. The bars indicate standard errors, and stars above the bars indicate that the difference from the wild-type value was significant. (C) Visualization of autophagic activation with MDC in wild type and ST39. P. sojae wild-type or ST39 hyphae were incubated in 10% V8 juice for 30 h. After 3 washes and incubation with sterile distilled water for 4 h, hyphae samples were concurrently stained with MDC and analyzed by confocal microscopy. Bars = 10 μm.
PsAtg6a silencing in P. sojae reduces virulence
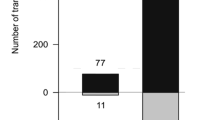
To determine whether PsAtg6a affected P. sojae virulence, we inoculated etiolated seedlings of the susceptible soybean cultivar Williams with zoospore suspensions of WT, T9, ST17, ST21, and ST39. Compared to WT and CK (T9), the silenced lines (ST17, ST21 and ST39) exhibited an obvious reduction in virulence against seedlings (Fig. 9A,B). As shown in Fig. 9B, lesions from silenced lines were smaller by approximately 50–80% of WT.
PsAtg6a silencing reduced P. sojae virulence. (A) Phenotypes of soybean etiolated seedlings inoculated with wild-type P. sojae and recombinant strains. Approximately 250 zoospores were used for each inoculation. Photographs were taken 24 and 36 hpi. The edges of the lesions are indicated by arrows. (B) Lesion lengths measured at 24 hpi and 36 hpi on five seedlings in each of four independent experiments. The bars indicate the standard errors. **P < 0.01, *P < 0.05 (t-test). (C) Trypan blue staining of hyphae infecting soybean epidermal cells. Photographs were taken at 30 hpi. Black arrows: haustoria.
To examine the extent that plant tissue colonization was compromised in PsAtg6a-silenced strains, we stained hyphae-infected soybean epidermal cells with trypan blue (Fig. 9C). Many infected hyphae and finger-like haustoria were observed upon infection with WT or CK (T9). In contrast, ST39 hyphae and rare haustoria were observed in the infected soybean epidermal cells. Thus, low expression levels of the PsAtg6a gene resulted in a significant reduction in virulence, and the reduced virulence of the PsAtg6a-silenced lines seemed attributable to the inability to produce haustoria.
Discussion
Autophagy, a protein degradation system conserved in eukaryotic cells, is used to recycle macromolecules and aid cellular survival under nutritional starvation conditions1, 2. A set of genes involved in this process are called ATGs and have been extensively studied in yeast and mammals19. In this study, 26 ATGs belonging to 20 different groups in P. sojae were identified on the basis of sequence similarity to ATG proteins in yeast and mammals, suggesting that the core components of autophagy are conserved among different species. In P. sojae, the compositions of domains within each class were similar with three exceptions (Atg1, Atg11 and Vps34), suggesting possible functional redundancy among different members of the same group. However, the conserved and specific functions of ATGs in the same group still must be further elucidated. In addition, 26 ATGs can be divided into four major functional groups, namely, the Atg1 complex, the phosphoinositide-3-kinase (PI3K) complex, the ubiquitin-like conjugation system and the Atg9 recycling complex.
Over recent decades, autophagy has been shown to play roles in the development and infection process of different pathogenic microbes, including M. grisea, F. graminearum and U. maydis 5,6,7,8,9. However, the roles of autophagy in P. sojae development and pathogenicity are still unknown. Two autophagy markers36, 38, a GFP-PsAtg8 fusion protein and the fluorescent dye MDC, were used to label autophagosomes in P. sojae. As in other higher eukaryotes, rapamycin and starvation induced autophagy in P. sojae cells. Furthermore, autophagy was significantly induced during sporangium formation and cysts germination, and PsAtg6a silencing suppressed sporulation. These results are consistent with the research on Phytophthora infestans showing that autophagy was high in sporangium41. Thus, we suggest that autophagy might play an important role in the development of Phytophthora species.
According to transcriptome analysis, the relative expression levels of most ATGs were significantly increased during infection stages, implying a common role for autophagy in virulence. PsAtg6a, a component of the class III PI3K complex, was shown to be essential for pathogenicity. PsAtg6a-silenced strains exhibited decreased virulence, and few finger-like haustoria were observed in infected soybean cells. Unfortunately, we were unable to assess autophagy in haustoria using MDC staining because the strong fluorescence signal in the plant cells interfered with observation. In addition, the ST39 line showed no obvious loss of ability to penetrate host cells. It is possible that PsAtg6b might complement a function of PsAtg6a in penetration. Nonetheless, it is clear that autophagy has extensive roles during P. sojae development and virulence, and there is a pressing need for further research on the molecular mechanisms underlying autophagy. In addition to these experiments, we attempted to silence other ATG genes (PsAtg1, PsAtg6b, PsAtg8 and PsAtg9) in P. sojae, but no stable genetic transformants were obtained. CRISPR/Cas9 technology has potential as a tool for Phytophthora genetic manipulation, and comparing the roles of autophagy during different developmental and infectious stages should be assessed in future studies.
Methods
ATGs identification in oomycetes
To identify ATG proteins in oomycetes, we obtained known ATG proteins from GenBank. Databases for oomycetes were obtained from their original sources: P. sojae 42, H. parasitica 43, P. capsici 44, P. ultimun 45 and T. pseudonana 46. Reciprocal Blastp was used with an e-value cut-off of 1e-5. The retrieved ATG genes were used as the query to search oomycete databases with the tBlastN algorithm47. Significant hits (E-value < 1e-5) were assessed for the presence of an obvious ATG-related function domain using Pfam48. The MUSCLE algorithm was used for multiple protein sequence alignment. The phylogenetic tree of ATG proteins was constructed using MEGA 4.1 with the neighbor-joining method and 1000 replicates were performed for bootstrap analysis49.
RNA isolation and qRT-PCR
Total RNA was extracted from mycelia, sporangia and infected soybean leaves using the RNAsimple Total RNA Kit (Tiangen, China) following the recommended protocol. Quantitative RT-PCR was performed in 20 μL reactions including 20 ng cDNA, 0.2 μM gene-specific primer for the reference P. sojae TEF1 gene (EU079791), 10 μL SYBR Premix ExTaq (Takara), and 6.8 μL ddH2O. PCR reactions were performed on an ABI PRISM 7300 fast Real-Time PCR System (Applied Biosystems) under the following conditions: 95 °C for 30 s, 40 cycles at 95 °C for 5 s, and 60 °C for 31 s to calculate cycle threshold values, followed by a dissociation program of 95 °C for 15 s, 60 °C for 1 min, and 95 °C for 15 s to obtain melt curves. The 7300 System Sequence Detection Software (version 1.4; SDS) was used to obtain relative expression levels for each sample. The transcript levels for test genes were determined according to the function ΔCT = CT (test gene) − CT (reference gene). To compare untreated and treated expression levels, the function ΔΔCT was determined using the equation ΔΔCT = ΔCT (treatment) − ΔCT (control), in which the control was mock-treated P. sojae P6497 mycelia. The induction ratio of treatment/control was calculated using the equation 2−ΔΔ CT.
Rapamycin treatment and monodansylcadaverine (MDC) staining
For rapamycin treatment, hyphae were incubated in V8 for up to 30 h, and 100 nmol/L rapamycin (dissolved in DMSO) was added to 10% V8 medium for 4 h before MDC staining. The same concentration of DMSO was used as a control.
MDC staining was performed as described by Contento et al.50. Briefly, tissues were stained with 50 μmol/L MDC in PBS for 10 min in the dark, and then washed 3 times with PBS before microscopic observation.
Plasmid construction and P. sojae manipulation
GFP-PsAtg8 is fused with a AceGFP gene and the PsAtg8 gene. To generate pHamGFP-PsAtg8, the NptII gene of pHAMT35N was replaced with the AcGFP1 gene (AB255038.1) that is amplified using the primers GFP-F and GFP-R. Then, PsAtg8 was amplified using the primers PsAtg8GFP-F and PsAtg8GFP -R and inserted the 3′ terminal of AceGFP gene. To generate pHamPsAtg6a for P. sojae gene silencing manipulation, the PsAtg6a gene was amplified by PCR using primers PsAtg6a-F and PsAtg6a-R. PsAtg6a gene was substituted for the NptII gene of pHAMT35N51.
P. sojae strain P6497 was routinely grown and maintained on V8 agar. P. sojae transformation was performed as previously described37. P. sojae transformants were selected on V8 medium with 50 μg/mL G418 and mycelia were harvested for extraction of DNA or RNA. Mycelial genomic DNA was used to screen for transgenes via PCR using the primers HamF and HamR. Transcription levels were measured using qRT-PCR assay of RNA extracts.
P. sojae development and infection assays
P. sojae development was analyzed using previously described methods52. Assays for P. sojae developmental markers such as vegetative growth, sporangia, zoospores, cysts and cysts germination were performed as previously described37. Virulence levels were determined by infecting etiolated soybean (the susceptible cultivar Williams) seedlings with 10 µL zoospore suspension (25 zoospores/µL). The lesion lengths of etiolated seedlings were measured at 24 and 36 hours post infection. Trypan blue staining was performed as described previously37. All of the experiments were repeated at least three times, and the data were analyzed using a t-test.
Microscopy
MDC-stained tissues and fluorescent mycelia expressing GFP and GFP-PsAtg8 were examined with the aid of a ZEISS LSM700 Confocal Laser Scanning Microscope (CLSM). ZEN 2010 software was used for fluorescence intensity analysis. An Olympus IX71 instrument was used to count sporangia and to monitor trypan blue staining during infection.
For rapamycin treatment, hyphae were incubated in V8 for up to 30 h, and 100 nmol/L rapamycin (dissolved in DMSO) was added to 10% V8 medium for 4 h. The same concentration of DMSO was used as a control. The mycelial growth was collected, thoroughly washed in distilled water, fixed overnight at 4 °C in modified Karmovsky’s fixative containing 2% paraformaldehyde and 2.5% (vol/vol) glutaraldehyde in 0.1 M phosphate buffer (pH 7.2). The fixed samples were washed three times, for 10 min each time, with 0.1 M phosphate buffer (pH 7.2). The samples were postfixed in 1% OsO4 for 2 h at 25 °C, washed three times with phosphate buffer as before, dehydrated in a graded ethanol series, embedded in Spurr resin, and stained with 2% uranyl acetate and Reynold’s lead solution before sectioning. The ultrathin sections were examined under a JEM-1230 electron microscope (JEOL, Tokyo, Japan) operating at 70 kV. Autophagosomes were counted using Image J software from electron microscopy images of P. sojae cells in the presence/absence of chemical inhibitors. Autophagosomes were counted using Image J software from electron microscopy images of P. sojae cells.
References
Feng, Y., He, D., Yao, Z. & Klionsky, D. J. The machinery of macroautophagy. Cell research 24, 24–41, doi:10.1038/cr.2013.168 (2014).
Klionsky, D. J. et al. A comprehensive glossary of autophagy-related molecules and processes (2nd edition). Autophagy 7, 1273–1294, doi:10.4161/auto.7.11.17661 (2011).
Chen, Y. & Klionsky, D. J. The regulation of autophagy - unanswered questions. J Cell Sci 124, 161–170, doi:10.1242/jcs.064576 (2011).
He, C. C. & Klionsky, D. J. Regulation Mechanisms and Signaling Pathways of Autophagy. Annu Rev Genet 43, 67–93, doi:10.1146/annurev-genet-102808-114910 (2009).
Liu, X. H. et al. Autophagy in plant pathogenic fungi. Semin Cell Dev Biol 57, 128–137, doi:10.1016/j.semcdb.2016.03.022 (2016).
Khan, I. A., Lu, J. P., Liu, X. H., Rehman, A. & Lin, F. C. Multifunction of autophagy-related genes in filamentous fungi. Microbiological research 167, 339–345, doi:10.1016/j.micres.2012.01.004 (2012).
Long, N. N. et al. Autophagy-related lipase FgATG15 of Fusarium graminearum is important for lipid turnover and plant infection. Fungal Genetics and Biology 48, 217–224, doi:10.1016/j.fgb.2010.11.004 (2011).
Nadal, M. & Gold, S. E. The autophagy genes atg8 and atg1 affect morphogenesis and pathogenicity in Ustilago maydis. Molecular plant pathology 11, 463–478, doi:10.1111/J.1364-3703.2010.00620.X (2010).
Veneault-Fourrey, C., Barooah, M., Egan, M., Wakley, G. & Talbot, N. J. Autophagic fungal cell death is necessary for infection by the rice blast fungus. Science (New York, N.Y.) 312, 580–583, doi:10.1126/science.1124550 (2006).
Ohsumi, Y. Historical landmarks of autophagy research. Cell research 24, 9–23, doi:10.1038/cr.2013.169 (2014).
Turco, E. & Martens, S. Insights into autophagosome biogenesis from in vitro reconstitutions. Journal of structural biology, doi:10.1016/j.jsb.2016.04.005 (2016).
Kofinger, J., Ragusa, M. J., Lee, I. H., Hummer, G. & Hurley, J. H. Solution structure of the Atg1 complex: implications for the architecture of the phagophore assembly site. Structure 23, 809–818, doi:10.1016/j.str.2015.02.012 (2015).
Obara, K. & Ohsumi, Y. PtdIns 3-Kinase Orchestrates Autophagosome Formation in Yeast. Journal of lipids 2011, 498768–9, doi:10.1155/2011/498768 (2011).
Nakatogawa, H. Two ubiquitin-like conjugation systems that mediate membrane formation during autophagy. Essays Biochem 55, 39–50, doi:10.1042/Bse0550039 (2013).
Yorimitsu, T. & Klionsky, D. J. Autophagy: molecular machinery for self-eating. Cell Death Differ 12(Suppl 2), 1542–1552, doi:10.1038/sj.cdd.4401765 (2005).
Suzuki, K., Akioka, M., Kondo-Kakuta, C., Yamamoto, H. & Ohsumi, Y. Fine mapping of autophagy-related proteins during autophagosome formation in Saccharomyces cerevisiae. J Cell Sci 126, 2534–2544, doi:10.1242/jcs.122960 (2013).
Kim, S. H., Kwon, C., Lee, J. H. & Chung, T. Genes for plant Autophagy: Functions and interactions. Mol Cells 34, 413–423, doi:10.1007/s10059-012-0098-y (2012).
Tian, Y. et al. C. elegans screen identifies autophagy genes specific to multicellular organisms. Cell 141, 1042–1055, doi:10.1016/j.cell.2010.04.034 (2010).
Meijer, W. H., van der Klei, I. J., Veenhuis, M. & Kiel, J. A. ATG genes involved in non-selective autophagy are conserved from yeast to man, but the selective Cvt and pexophagy pathways also require organism-specific genes. Autophagy 3, 106–116, doi:10.4161/auto.3595 (2007).
Kamoun, S. Molecular genetics of pathogenic oomycetes. Eukaryotic cell 2, 191–199, doi:10.1128/EC.2.2.191-199.2003 (2003).
Walker, C. A. & van West, P. Zoospore development in the oomycetes. Fungal Biology Reviews 21, 10–18, doi:10.1016/j.fbr.2007.02.001 (2007).
Judelson, H. S. & Blanco, F. A. The spores of Phytophthora: weapons of the plant destroyer. Nature reviews. Microbiology 3, 47–58, doi:10.1038/nrmicro1064 (2005).
Tyler, B. M. Molecular basis of recognition between phytophthora pathogens and their hosts. Annual review of phytopathology 40, 137–167, doi:10.1146/annurev.phyto.40.120601.125310 (2002).
Koeck, M., Hardham, A. R. & Dodds, P. N. The role of effectors of biotrophic and hemibiotrophic fungi in infection. Cellular microbiology 13, 1849–1857, doi:10.1111/j.1462-5822.2011.01665.x (2011).
Horbach, R., Navarro-Quesada, A. R., Knogge, W. & Deising, H. B. When and how to kill a plant cell: Infection strategies of plant pathogenic fungi. J Plant Physiol 168, 51–62, doi:10.1016/j.jplph.2010.06.014 (2011).
Kershaw, M. J. & Talbot, N. J. Genome-wide functional analysis reveals that infection-associated fungal autophagy is necessary for rice blast disease. Proceedings of the National Academy of Sciences of the United States of America 106, 15967–15972, doi:10.1073/pnas.0901477106 (2009).
Xie, Z. & Klionsky, D. J. Autophagosome formation: core machinery and adaptations. Nature cell biology 9, 1102–1109, doi:10.1038/ncb1007-1102 (2007).
Feng, Y. H. et al. Global analysis of protein structural changes in complex proteomes. Nat Biotechnol 32, 1036–1044, doi:10.1038/nbt.2999 (2014).
Tanida, I. Autophagy basics. Microbiol Immunol 55, 1–11, doi:10.1111/j.1348-0421.2010.00271.x (2011).
Shpilka, T., Weidberg, H., Pietrokovski, S. & Elazar, Z. Atg8: an autophagy-related ubiquitin-like protein family. Genome Biol 12, doi:Artn 22610.1186/Gb-2011-12-7-226 (2011).
Mizushima, N. The role of the Atg1/ULK1 complex in autophagy regulation. Current opinion in cell biology 22, 132–139, doi:10.1016/j.ceb.2009.12.004 (2010).
Yorimitsu, T. & Klionsky, D. J. Atg11 links cargo to the vesicle-forming machinery in the cytoplasm to vacuole targeting pathway. Mol Biol Cell 16, 1593–1605, doi:10.1091/mbc.E04-11-1035 (2005).
Lu, S. et al. Intracellular and Extracellular Phosphatidylinositol 3-Phosphate Produced by Phytophthora Species Is Important for Infection. Mol Plant 6, 1592–1604, doi:10.1093/mp/sst047 (2013).
Ye, W. et al. Digital gene expression profiling of the Phytophthora sojae transcriptome. Molecular plant-microbe interactions: MPMI 24, 1530–1539, doi:10.1094/MPMI-05-11-0106 (2011).
Carames, B. et al. Autophagy activation by rapamycin reduces severity of experimental osteoarthritis. Annals of the rheumatic diseases 71, 575–581, doi:10.1136/annrheumdis-2011-200557 (2012).
Nair, U., Thumm, M., Klionsky, D. J. & Krick, R. GFP-Atg8 protease protection as a tool to monitor autophagosome biogenesis. Autophagy 7, 1546–1550, doi:10.4161/auto.7.12.18424 (2011).
Chen, L. L. et al. Phytophthora sojae TatD Nuclease Positively Regulates Sporulation and Negatively Regulates Pathogenesis. Mol Plant Microbe In 27, 1070–1080, doi:10.1094/Mpmi-05-14-0153-R (2014).
Biederbick, A., Kern, H. F. & Elsasser, H. P. Monodansylcadaverine (MDC) is a specific in vivo marker for autophagic vacuoles. European journal of cell biology 66, 3–14 (1995).
Tyler, B. M. Phytophthora sojae: root rot pathogen of soybean and model oomycete. Molecular plant pathology 8, 1–8, doi:10.1111/j.1364-3703.2006.00373.x (2007).
Cao, Y. & Klionsky, D. J. Physiological functions of Atg6/Beclin 1: a unique autophagy-related protein. Cell research 17, 839–849, doi:10.1038/cr.2007.78 (2007).
Luo, Q., Wang, F. X., Zhong, N. Q., Wang, H. Y. & Xia, G. X. The role of autophagy during development of the oomycete pathogen Phytophthora infestans. Journal of genetics and genomics=Yi chuan xue bao 41, 225–228, doi:10.1016/j.jgg.2014.03.004 (2014).
Tyler, B. M. et al. Phytophthora genome sequences uncover evolutionary origins and mechanisms of pathogenesis. Science (New York, N.Y.) 313, 1261–1266, doi:10.1126/science.1128796 (2006).
Baxter, L. et al. Signatures of adaptation to obligate biotrophy in the Hyaloperonospora arabidopsidis genome. Science (New York, N.Y.) 330, 1549–1551, doi:10.1126/science.1195203 (2010).
Lamour, K. H. et al. Genome Sequencing and Mapping Reveal Loss of Heterozygosity as a Mechanism for Rapid Adaptation in the Vegetable Pathogen Phytophthora capsici. Mol Plant Microbe In 25, 1350–1360, doi:10.1094/Mpmi-02-12-0028-R (2012).
Levesque, C. A. et al. Genome sequence of the necrotrophic plant pathogen Pythium ultimum reveals original pathogenicity mechanisms and effector repertoire. Genome Biol 11, doi:Artn R7310.1186/Gb-2010-11-7-R73 (2010).
Armbrust EV1, B. J. et al. The genome of the diatom Thalassiosira pseudonana: ecology, evolution, and metabolism. Science (New York, N.Y.) 306(5693), 79–86, doi:10.1126/science.1101156 (2004).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic Local Alignment Search Tool. J Mol Biol 215, 403–410, doi:10.1016/S0022-2836(05)80360-2 (1990).
Finn, R. D. et al. The Pfam protein families database. Nucleic Acids Res 36, D281–D288, doi:10.1093/Nar/Gkm960 (2008).
Tamura, K., Dudley, J., Nei, M. & Kumar, S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24, 1596–1599, doi:10.1093/molbev/msm092 (2007).
Contento, A. L., Xiong, Y. & Bassham, D. C. Visualization of autophagy in Arabidopsis using the fluorescent dye monodansylcadaverine and a GFP-AtATG8e fusion protein. Plant J 42, 598–608, doi:10.1111/j.1365-313X.2005.02396.x (2005).
Judelson, H. S., Tyler, B. M. & Michelmore, R. W. Transformation of the Oomycete Pathogen, Phytophthora infestans. Mol Plant Microbe In 4, 602–607, doi:10.1094/MPMI-4-602 (1991).
LaVallie, E. R. et al. A thioredoxin gene fusion expression system that circumvents inclusion body formation in the E. coli cytoplasm. Bio/technology (Nature Publishing Company) 11, 187–193, doi:10.1038/nbt0293-187 (1993).
Acknowledgements
This project is supported by grants from the National Natural Science Foundation of China (31501594) and the Special Fund for Agro-Scientific Research in the Public Interest (No.201503112).
Author information
Authors and Affiliations
Contributions
H.L., D.D. and L.C. conceived of the study, participated in its design and coordination and drafted the manuscript. X.Z. performed the bioinformatics analysis, L.C., W.W. and X.G. performed the experiments. All of the authors participated in the data analysis and interpretation. All of the authors read and approved of the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, L., Zhang, X., Wang, W. et al. Network and role analysis of autophagy in Phytophthora sojae . Sci Rep 7, 1879 (2017). https://doi.org/10.1038/s41598-017-01988-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-01988-7
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.