Abstract
Many neurotransmitters directly inhibit neurons by activating G protein-gated inwardly rectifying K+ (GIRK) channels, thereby moderating the influence of excitatory input on neuronal excitability. While most neuronal GIRK channels are formed by GIRK1 and GIRK2 subunits, distinct GIRK2 isoforms generated by alternative splicing have been identified. Here, we compared the trafficking and function of two isoforms (GIRK2a and GIRK2c) expressed individually in hippocampal pyramidal neurons lacking GIRK2. GIRK2a and GIRK2c supported comparable somato-dendritic GIRK currents in Girk2 −/− pyramidal neurons, although GIRK2c achieved a more uniform subcellular distribution in pyramidal neurons and supported inhibitory postsynaptic currents in distal dendrites better than GIRK2a. While over-expression of either isoform in dorsal CA1 pyramidal neurons restored contextual fear learning in a conditional Girk2 −/− mouse line, GIRK2a also enhanced cue fear learning. Collectively, these data indicate that GIRK2 isoform balance within a neuron can impact the processing of afferent inhibitory input and associated behavior.
Similar content being viewed by others
Introduction
G protein-gated inwardly rectifying K+ (GIRK/Kir3) channels mediate the G protein-dependent, postsynaptic inhibitory effects of many neurotransmitters in the central nervous system, including GABA, dopamine, serotonin, acetylcholine, and glutamate1. GIRK channels are found almost exclusively in the somato-dendritic compartment and are enriched near excitatory synapses in spines2. Thus, these channels are well-positioned to temper the influence of excitatory input on neuronal activity3. Genetic and pharmacological investigations in mice have revealed important roles for GIRK channels in diverse settings, including seizures, learning/memory, nociception/analgesia, anxiety, and addiction4,5,6.
GIRK channels are homo- and heterotetrameric complexes formed by 4 subunits (GIRK1/Kir3.1, GIRK2/Kir3.2, GIRK3/Kir3.3, and GIRK4/Kir3.4)2. While GIRK1, GIRK2, and GIRK3 are widely expressed in the brain7, GIRK4 is found in only a few brain regions8, 9. Most, if not all, neuronal GIRK channels contain GIRK24. Indeed, Girk2 −/− mice exhibit small or absent synaptic and somato-dendritic GIRK currents in all neuron types evaluated to date10,11,12,13,14,15,16,17,18. GIRK1 cannot form functional homomeric channels, as it lacks an endoplasmic reticulum export signal19,20,21,22, and is thought to assemble with GIRK2 in most brain regions23, 24. Thus, the prototypical GIRK channel in the central nervous system is thought to contain both GIRK1 and GIRK2.
The diversity of neuronal GIRK channels is enhanced by alternative splicing which, in the case of GIRK2, yields at least 3 distinct proteins22, 25,26,27,28,29,30. Two GIRK2 variants (termed GIRK2a and GIRK2c) are particularly intriguing as they differ only by 11 amino acids in their intracellular C-terminal domains. GIRK2c is the longer of the 2 variants, and the 11 amino acid extension contains a conspicuous Class I PDZ interaction domain22, 31.
The impact of alternative splicing of the Girk2 (Kcnj6) gene on GIRK channel trafficking and function is unclear. GIRK2c (but not GIRK2a) was reported to bind PSD-95 in one study31, but not in others32, 33. GIRK2c (but not GIRK2a) was also reported to bind to SAP-97, and co-expression of SAP-97 with GIRK2c conferred both basal activity and G protein-dependent gating to GIRK2c homomeric channels expressed in Xenopus oocytes34. To date, however, the attributes of GIRK2a and GIRK2c have not been systematically compared in mammalian cell systems. In this study, we expressed GIRK2a and GIRK2c independently in neurons lacking GIRK2, and compared their subcellular distributions and respective abilities to rescue GIRK channel function and GIRK-dependent behavior.
Results
Comparison of GIRK2a and GIRK2c-containing channels in transfected mammalian cells
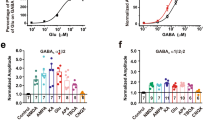
We began by comparing the functional properties of GIRK2a and GIRK2c expressed in mammalian (HEK) cells. As most neuronal GIRK channels are thought to be GIRK1/GIRK2 heteromers23, 24, HEK cells were transfected with GIRK1 and either GIRK2a or GIRK2c, along with the GABAB receptor (GABABR) subunits, GABABR1 and GABABR2. Whole-cell currents evoked by rapid bath application of the GABABR agonist baclofen (100 μM) were measured (Fig. 1A). While no response to baclofen was seen with expression of GABABR alone, cells expressing GABABR and either GIRK1/GIRK2a or GIRK1/GIRK2c exhibited baclofen-induced currents of comparable size (Fig. 1A,B), with similar activation and deactivation kinetics (Fig. 1C,D). Furthermore, no difference in the EC50 for baclofen activation of GIRK1/GIRK2a or GIRK1/GIRK2c channels was observed (Fig. 1E–G), indicating that alternative splicing of GIRK2 does not impact the sensitivity of the prototypical neuronal GIRK channel to GABABR activation in a typical mammalian expression system.
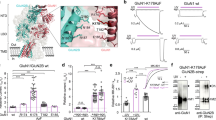
Functional comparison of GIRK2a and GIRK2c in HEK cells. (A) Whole-cell currents (Vhold = −70 mV) evoked by baclofen (100 μM) in HEK cells expressing GABABR, GIRK1, and either GIRK2a (red) or GIRK2c (blue). No current was evoked by baclofen in cells expressing only GABABR (control, black). Scale: 500 pA/10 s. (B) Summary of baclofen-induced, steady-state current densities (Ibaclofen, pA/pF) in HEK cells expressing GIRK1/GIRK2a or GIRK1/GIRK2c (t 19 = 1.1, P = 0.29; n = 10–11/group). Individual data points are represented as small squares overlapping the relevant bar in the plot. (C,D) Activation (t 19 = 1.5, P = 0.16) and deactivation (t 16 = 0.4, P = 0.72) kinetics for baclofen-induced currents in HEK cells expressing GIRK1/GIRK2a or GIRK1/GIRK2c (n = 9–11/group). (E) Representative concentration-response experiment for a HEK cell expressing GABABR and GIRK1/GIRK2c. Scale: 500 pA/10 s. (F,G) Summary of concentration-response experiments for baclofen-induced currents in HEK cells expressing GIRK1/GIRK2a or GIRK1/GIRK2c (t 17 = 1.0, P = 0.32; n = 9–10/group). Currents were normalized to the response evoked by 100 μM baclofen in each experiment.
GIRK2a and GIRK2c expression in the mouse hippocampus
HEK cells may lack factors that support GIRK2 isoform-specific contributions to channel trafficking and function, and thus, we next sought to compare the GIRK2 variants in a cell type in which they are normally expressed. Cell biological, biochemical, and electrophysiological studies involving wild-type and Girk2 −/− mice have confirmed the presence of GIRK2 in mouse hippocampal neurons10, 15. Using RNA-Seq and laser-captured tissue samples from the adult mouse, we probed for Girk2a and Girk2c mRNAs in the CA1 region of the hippocampus. Because the 3′-UTR regions of Girk2a and Girk2c mRNAs are distinct and may influence their subcellular trafficking29, we compared transcript levels in cell body (stratum pyramidale) and neuropil (stratum radiatum) samples. Both transcripts were detected in neuropil, albeit at significantly lower levels than seen in CA1 cell body samples (q = 0.0012 for Girk2a and 0.0125 for Girk2c), suggesting that an active RNA trafficking mechanism exists for both isoforms (Fig. 2A). Higher levels of Girk2c mRNA relative to Girk2a mRNA were detected, with the difference reaching statistical significance in neuropil (P = 0.04) but not cell bodies (P = 0.07). Interestingly, we detected considerable read coverage past the annotated terminal exon for both transcripts, suggesting transcription occurs past the annotated 3′ ends in hippocampal pyramidal neurons.
GIRK2 expression in the mouse hippocampus. (A) Girk2a and Girk2c mRNA levels as assessed by RNA-Seq in CA1 cell body and neuropil samples, taken from 3 adult mice. Main effects of isoform (F1,8 = 14.3, P < 0.01) and compartment (F1,8 = 27.1, P < 0.001) were observed, but there was no interaction between isoform and compartment (F1,8 = 0.07, P = 0.80). Symbols: *P < 0.05 vs. GIRK2a (within compartment); +,++ P < 0.05 and 0.01, respectively, vs. cell body (within isoform). (B) Representative GIRK2 immunoblots (from 3 independent experiments) of adult hippocampus and neonatal hippocampal culture (10–12 DIV) from wild-type (WT) and Girk2 −/− mice, probed with pan-GIRK2 or GIRK2c antibodies (upper blots), as well as a β-actin antibody (lower blots). (C,D) Representative images showing GIRK2 immunolabeling (as revealed with the pan-GIRK2 antibody) in cultured hippocampal neurons from wild-type (C) and Girk2 −/− (D) mice. Neurons were co-stained with a MAP2 antibody to highlight neuronal morphology and dendritic processes. Scale bar: 50 microns.
We next assessed whether GIRK2a and GIRK2c proteins are present in mouse hippocampus and cultured hippocampal neurons. A commercially-available GIRK2 antibody (pan-GIRK2) targeting a domain in both isoforms revealed a doublet in intact hippocampus and cultured hippocampal neurons from wild-type but not Girk2 −/− mice (Fig. 2B). We also used a custom GIRK2c-specific antibody, raised against the unique 11-amino acid domain of GIRK2c, to probe for the GIRK2c isoform in hippocampal samples. The GIRK2c antibody recognized GIRK2c, but not other GIRK subunits, when expressed in HEK cells (Supplementary Fig. S1). In hippocampal samples, this antibody revealed a single band that aligned with the upper band in the doublet seen with the pan-GIRK2 antibody (Fig. 2B). Thus, two GIRK2 splice isoform proteins are present in the mouse hippocampus, one definitively identified as GIRK2c and the other exhibiting an electrophoretic mobility consistent with that of GIRK2a.
Subcellular distribution of GIRK2a and GIRK2c in Girk2 −/− pyramidal neurons
While GIRK2a and GIRK2c exhibited comparable labeling patterns when expressed in cultured rat hippocampal neurons22, endogenous GIRK2 might mask isoform-dependent differences in the trafficking and/or function of exogenously expressed subunits. Thus, we sought to examine the subcellular distribution of each isoform when expressed individually in cultured hippocampal neurons from Girk2 −/− mice. AAV vectors harboring coding sequence for GIRK2a or GIRK2c, inserted downstream of the CaMKIIα promoter, were used to drive expression of untagged GIRK2a or GIRK2c (and EGFP, bi-cistronically) in Girk2 −/− neurons. Strong overlap between recombinant GIRK2 and endogenous CaMKIIα immunolabeling was observed (Supplementary Fig. S2), confirming that the expression of GIRK2 was restricted primarily to pyramidal neurons.
Consistent with a previous report35, the pan-GIRK2 antibody yielded a punctate labeling pattern in wild-type cultures, with extensive labeling seen in dendritic processes (Fig. 2C). No such labeling was seen in hippocampal cultures from Girk2 −/− mice (Fig. 2D), confirming antibody specificity. The labeling patterns for recombinant GIRK2a and GIRK2c expressed in Girk2 −/− pyramidal neurons were overlapping but distinct (Fig. 3). While both subunits were detected in the soma and dendrites of infected neurons, GIRK2a labeling was more restricted to the cell body and proximal dendrites (Fig. 3A). In contrast, GIRK2c labeling was uniformly distributed throughout the neuron, including distal dendrites (Fig. 3B). Quantification of GIRK2 isoform labeling confirmed that GIRK2c exhibited a higher density of labeling than GIRK2a in secondary and tertiary dendrites of infected hippocampal pyramidal neurons (Fig. 3C).
Subcellular distribution of GIRK2a and GIRK2c in Girk2 −/− pyramidal neurons. (A,B) Representative images showing GIRK2 (red) and MAP2 (green) immunolabeling, and their overlay, in Girk2 −/− hippocampal pyramidal neurons expressing either GIRK2a (A) or GIRK2c (B). Scale bars: 50 microns. The insets highlight different densities of GIRK2a and GIRK2c puncta along proximal/primary (Ai,Bi) and distal/secondary (Aii,Bii) dendritic segments. Scale bars: 5 microns. (C) Quantification of GIRK2a and GIRK2c labeling in dendrites from infected Girk2 −/− pyramidal neurons. GIRK2 fluorescence intensity was measured in 2–3 primary (t 27 = 1.4, P = 0.17), secondary (t 29 = 2.1, *P < 0.05), and tertiary (t 27 = 5.2, ***P < 0.001) dendritic segments from 5 different neurons expressing each subunit. Fluorescence intensity is expressed as arbitrary units (AU) normalized to segment length. Only GIRK2 labeling that overlapped directly with the dendritic segment (identified by MAP2 labeling) was quantified in this analysis.
GIRK3 has been shown to mediate interactions between GIRK channels and proteins that can regulate GIRK channel trafficking (sorting nexin 27, or SNX-27)36 and receptor coupling efficiency (regulator of G protein signaling 2, or RGS2)37. As GIRK3 is also expressed in hippocampal neurons7, 15, 38, we examined the subcellular distribution of GIRK2a and GIRK2c expressed in cultured hippocampal pyramidal neurons from mice lacking both GIRK2 and GIRK3 (Girk2 −/− /Girk3 −/− mice). The labeling patterns for GIRK2a and GIRK2c in neurons from Girk2 −/− /Girk3 −/− mice were comparable to those seen in Girk2 −/− neurons (Supplementary Fig. S3), indicating that the GIRK2 isoform-dependent difference in subcellular distribution is not dependent on GIRK3.
Overlap between PSD-95 and GIRK2 splice isoforms in Girk2 −/− pyramidal neurons
GIRK2-containing channels are enriched near excitatory synapses15, 39, and some overlap between GIRK2 and PSD-95 was reported in cultured hippocampal neurons35. As the PDZ interaction motif found in GIRK2c may promote the association of GIRK channels with excitatory synapses, we next compared the overlap between endogenous PSD-95 and either GIRK2a or GIRK2c in Girk2 −/− pyramidal neurons (Fig. 4A,B). No significant difference in the extent of overlap between PSD-95 and either GIRK2a or GIRK2c was detected at baseline (Fig. 4C, control). We also explored the possibility that incubation with morphine, a treatment previously shown to increase the overlap between GIRK2 and PSD-95 in cultured hippocampal neurons35, might reveal an isoform-dependent difference. Consistent with published observations, we found that morphine treatment increased total GIRK2 protein level by approximately 2-fold relative to vehicle-treated controls (Supplementary Fig. S4A,B), while having no apparent effect on PSD-95 labeling intensity (Supplementary Fig. S4C). Morphine treatment increased the overlap between PSD-95 and both GIRK2 isoforms relative to control conditions, with GIRK2c showing a small but significantly higher degree of overlap with PSD-95 than did GIRK2a following treatment (Fig. 4C, morphine; Supplementary Fig. S4D,E).
Overlap of GIRK2a and GIRK2c with PSD-95 in pyramidal neurons. (A,B) Representative images showing GIRK2 (red) and PSD-95 (green) immunolabeling, and their overlay, in a Girk2 −/− hippocampal pyramidal neuron expressing GIRK2a (A) or GIRK2c (B). Scale bars: 50 microns. The inset (Bi) highlights the limited overlap between PSD-95 and GIRK2, as demonstrated with GIRK2c. Scale bars: 20 microns. (C) Quantification of overlap between PSD-95 and GIRK2a or GIRK2c, under control conditions and following morphine treatment (n = 8 per isoform and treatment condition). Main effects of isoform (F1,28 = 9.2, P < 0.01) and treatment (F1,28 = 22.1, P < 0.001) were observed, but there was no interaction between isoform and treatment (F1,28 = 0.3, P = 0.60). Symbols: *P < 0.05 vs. GIRK2a (within treatment); +,++ P < 0.05 and 0.01, respectively, vs. control (within isoform).
GIRK currents measured in Girk2 −/− pyramidal neurons expressing GIRK2a or GIRK2c
We next asked whether GIRK2a and GIRK2c conferred unique functionality to GIRK channels in cultured hippocampal pyramidal neurons. Whole-cell currents evoked by GABABR stimulation were measured in wild-type and Girk2 −/− pyramidal neurons expressing EGFP, and in Girk2 −/− pyramidal neurons expressing GIRK2a or GIRK2c. Baclofen evoked a prominent inward current in wild-type pyramidal neurons that was not seen in Girk2 −/− control neurons expressing EGFP alone (Fig. 5A,B). Viral expression of either GIRK2 isoform in Girk2 −/− pyramidal neurons yielded baclofen-induced currents that were larger than currents in wild-type controls. Activation (but not deactivation) kinetics of the baclofen-induced currents in Girk2 −/− pyramidal neurons expressing GIRK2a or GIRK2c also differed from those in wild-type control neurons, though no isoform-dependent differences were found (Fig. 5C,D). Similarly, while there was no isoform-dependent difference in channel sensitivity (EC50) to GABABR activation, GABABR-GIRK coupling efficiency in both cases was diminished relative to wild-type controls (Fig. 5E).
GPCR-GIRK currents in Girk2 −/− pyramidal neurons expressing GIRK2a or GIRK2c. (A) Whole-cell currents (Vhold = −70 mV) evoked by baclofen (100 μM) in a wild-type pyramidal neuron expressing EGFP (WT, black), as well as Girk2 −/− pyramidal neurons expressing EGFP (Girk2 −/−, gray), GIRK2a (red), or GIRK2c (blue). Scale: 1 nA/5 s. (B) Summary of baclofen-induced, steady-state current densities (pA/pF) in wild-type control and Girk2 −/− pyramidal neurons expressing EGFP, GIRK2a, or GIRK2c (F3,53 = 44.8, P < 0.001; n = 11–16/group). Symbols: ***P < 0.001 vs. WT. (C,D) Summary of activation (C; F2,35 = 19.2, P < 0.001) and deactivation (D) F2,33 = 2.76, P = 0.08) kinetics for baclofen-induced currents in infected wild-type and Girk2 −/− pyramidal neurons. Symbols: ***P < 0.001 vs. WT (n = 11–15 per group). (E) EC50 values for baclofen-induced currents in infected wild-type and Girk2 −/− pyramidal neurons expressing GIRK2a or GIRK2c (F2,17 = 8.84, P < 0.01; n = 6–8/group). Symbols: *,**P < 0.05 and 0.01, respectively, vs. WT. (F) Whole-cell currents evoked by adenosine (10 μM) in a wild-type pyramidal neuron expressing EGFP, as well as Girk2 −/− pyramidal neurons expressing EGFP, GIRK2a, or GIRK2c (Vhold = −70 mV). Scale: 500 pA/5 s. (G) Summary of adenosine-induced, steady-state current densities (Iadenosine, pA/pF) in wild-type control and Girk2 −/− pyramidal neurons expressing EGFP, GIRK2a, or GIRK2c (F3,36 = 32.1, P < 0.001; n = 9–13/group). Symbols: ***P < 0.001 vs. WT. (H) Whole-cell currents evoked by 8-OH-DPAT (10 μM) in a wild-type pyramidal neuron expressing EGFP, as well as Girk2 −/− pyramidal neurons expressing EGFP, GIRK2a, or GIRK2c (Vhold = −70 mV). As currents evoked by 8-OH-DPAT persisted following bath washout, the 5HT1AR-selective antagonist WAY100635 (WAY, 2 μM) was used to demonstrate reversibility of the 8-OH-DAT response. Scale: 500 pA/5 s. (I) Summary of 8-OH-DPAT-induced, steady-state current densities (I8-OH-DPAT, pA/pF) in wild-type control and Girk2 −/− pyramidal neurons expressing EGFP, GIRK2a, or GIRK2c (F3,30 = 7.92, P < 0.001; n = 5–11/group). Symbols: *P < 0.05 vs. WT. (J) Summary of current densities (pA/pF) evoked by 10 μM ML297 (IML297) in wild-type control and Girk2 −/− pyramidal neurons expressing EGFP, GIRK2a, or GIRK2c (F2,12 = 2.0, P = 0.16; n = 5–8/group).
GIRK channels in hippocampal pyramidal neurons are also activated by adenosine and serotonin10, 40, 41. To test whether GIRK2 isoforms promote preferential associations between GIRK channels and specific GPCRs, we measured GIRK currents evoked via these signaling pathways in Girk2 −/− pyramidal neurons expressing GIRK2a or GIRK2c. Both adenosine (Fig. 5F) and the 5HT1A receptor-selective agonist 8-OH-DPAT (Fig. 5H) evoked reliable currents in wild-type neurons, but not Girk2 −/− pyramidal neurons expressing EGFP. Girk2 −/− pyramidal neurons expressing either GIRK2a or GIRK2c exhibited adenosine- and 8-OH-DPAT-induced currents, and these currents were comparable in magnitude to those observed in wild-type control neurons (Fig. 5G,I). To gauge the influence of GIRK3 on the coupling between the GIRK2 isoforms and specific GPCRs, baclofen-, adenosine-, and 8-OH-DPAT-induced currents were also measured in infected Girk2 −/− /Girk3 −/− pyramidal neurons. While no differences were found in Girk2 −/− /Girk3 −/− pyramidal neurons expressing GIRK2a or GIRK2c, the currents measured in these neurons were larger than those recorded in Girk2 −/− pyramidal neurons expressing these isoforms (Supplementary Fig. S5).
We also probed for differences in the interaction between the GIRK2 isoforms and endogenous GIRK1 in cultured Girk2 −/− hippocampal neurons. Whole-cell currents were evoked by a selective activator of GIRK1-containing channels (ML297)42, 43. ML297-induced currents of comparable size were observed in Girk2 −/− pyramidal neurons expressing GIRK2a or GIRK2c, and in wild-type control neurons (Fig. 5J), indicating that viral expression of either GIRK2 isoform restores a normal level of GIRK1-containing heteromeric channel activity.
Synaptic GIRK currents in Girk2 −/− pyramidal neurons expressing GIRK2a or GIRK2c
The lack of natural connectivity and normal afferent inputs to cultured neurons complicates efforts to study the full range of physiological contributions made by GIRK channels to pyramidal neuron excitability. Thus, we next examined the functional properties of GIRK2a and GIRK2c in organotypic hippocampal slices. Organotypic hippocampal slices were cultured from young wild-type or Girk2 −/− mice (P8–9), and the CA1 region was targeted with control or GIRK2 isoform-specific expression viruses (Fig. 6A). Following a 1-wk incubation period, we measured somatodendritic and synaptic GIRK-dependent responses in infected dorsal CA1 pyramidal neurons. Consistent with our observations in infected Girk2 −/− pyramidal neurons in culture, we observed larger baclofen-induced somatodendritic currents in dorsal Girk2 −/− CA1 pyramidal neurons expressing either GIRK2a or GIRK2c, as compared to wild-type controls (Fig. 6B ).
Synaptic GIRK currents in Girk2 −/− neurons expressing GIRK2a and GIRK2c. (A) Image showing EGFP expression in the CA1 region of an organotypic hippocampal slice, taken 7 d after infection with AAV8-hSyn-GIRK2a-IRES-EGFP. The dotted white line highlights key features of slice morphology. Scale bar: 500 microns (inset: 50 microns). (B) Summary of peak baclofen-induced somatodendritic current amplitudes (Ibaclofen) in dorsal CA1 pyramidal neurons, measured 7 d after CA1 infusion of control virus (con) to wild-type (WT) organotypic slices, or GIRK2 expression viruses (2a, 2c) to Girk2 −/− organotypic slices (F2,17 = 1.2, P < 0.01; n = 6–7/group). Symbols *,**P < 0.05 and 0.01, respectively, vs. WT. (C) Slow IPSCs evoked via stimulation of proximal (Schaffer collateral/SC) or distal (perforant path, PP) dendritic fields, in wild-type and Girk2 −/− CA1 pyramidal neurons expressing EGFP, as well as Girk2 −/− CA1 pyramidal neurons expressing GIRK2a or GIRK2c. The stimulus artifact and residual GABAAR-dependent responses were removed for clarity, and the triangles below the traces denote stimulation times. Scale: 100 pA/1 s. (D) Summary plot comparing the ratios of slow IPSC maximal amplitudes evoked by SC and PP stimulation in wild-type CA1 pyramidal neurons expressing EGFP, as well as Girk2 −/− CA1 pyramidal neurons expressing GIRK2a or GIRK2c (F2,37 = 51.9; P < 0.001; n = 6–8/group). Maximal responses evoked by stimulation of SC and PP fields were measured in each neuron, in counterbalanced fashion. Symbols: ***P < 0.001 vs. WT (control) and Girk2 −/− (GIRK2c).
Slow inhibitory postsynaptic currents (slow IPSCs) in dorsal CA1 pyramidal neurons were evoked by electrical stimulation within proximal (Schaeffer collateral/SC) or distal (perforant path/PP) dendritic fields. SC and PP slow IPSCs were reliably observed in wild-type CA1 pyramidal neurons (Fig. 6C), and were blocked completely by the GABABR receptor antagonist CGP54626 (2 μM, not shown). Slow IPSCs were not observed in Girk2 −/− CA1 neurons expressing EGFP. While viral expression of either GIRK2a or GIRK2c in Girk2 −/− slices restored both SC and PP slow IPSCs in CA1 pyramidal neurons, the relative amplitudes of slow IPSCs measured in SC and PP differed between the groups (Fig. 6C). Indeed, the ratio of slow IPSC amplitudes, measured in PP and SC for each neuron, was significantly smaller in Girk2 −/− CA1 pyramidal neurons expressing GIRK2a, as compared to both wild-type control neurons, and Girk2 −/− pyramidal neurons expressing GIRK2c (Fig. 6D).
Fear conditioning in CaMKIICre(+):Girk2 fl/fl mice following GIRK2a or GIRK2c reconstitution
Recently, we reported that mice lacking GIRK2 in forebrain pyramidal neurons (CaMKIICre(+):Girk2 fl/fl mice) were deficient in contextual fear learning, a hippocampal-dependent behavior6. As CaMKIICre(+):Girk2 fl/fl mice exhibited a loss of GIRK-dependent signaling in dorsal but not ventral CA1 pyramidal neurons6, we asked whether viral restoration of GIRK2a or GIRK2c in dorsal CA1 pyramidal neurons of CaMKIICre(+):Girk2 fl/fl mice could rescue normal fear learning. Adult CaMKIICre(+):Girk2 fl/fl mice received bilateral infusions of Cre-dependent variants of the GIRK2a and GIRK2c expression viruses into the dorsal CA1 (Fig. 7A). After a suitable post-surgical recovery period (2 wk), we tested the efficacy of our viral treatment by measuring baclofen-induced somato-dendritic currents in CA1 pyramidal neurons in acutely isolated slices. Currents in dorsal CA1 pyramidal neurons from CaMKIICre(+):Girk2 fl/fl mice treated with control virus were small (Fig. 7B,C), and comparable in size to those reported previously for untreated CaMKIICre(+):Girk2 fl/fl mice6. Dorsal CA1 pyramidal neurons from CaMKIICre(+):Girk2 fl/fl mice expressing GIRK2a or GIRK2c exhibited significantly larger baclofen-induced currents (Fig. 7B,C). Indeed, currents measured in dorsal CA1 pyramidal neurons expressing GIRK2a or GIRK2c were notably larger than those reported previously in wild-type or CaMKIICre(−):Girk2 fl/fl controls6, 15, suggesting that the GIRK2 isoforms were over-expressed in these neurons.
Fear learning in CaMKIICre(+):Girk2 fl/fl mice expressing GIRK2a or GIRK2c. (A) EGFP expression in the dorsal CA1 of a CaMKIICre(+):Girk2 fl/fl mouse, 2 wk after infusion of the AAV8-CaMKIIα-DIO-GIRK2a-IRES-EGFP virus. Scale bar: 500 microns. (B) Representative somato-dendritic currents (Vhold = −60 mV) evoked by baclofen (200 μM) and reversed by the GABABR antagonist CGP54626 (2 μM) in dorsal CA1 pyramidal neurons from CaMKIICre(+):Girk2 fl/fl mice, 2 wk after infusion of Cre-dependent control (con, mCherry), GIRK2a, or GIRK2c virus. Scale: 100 pA/100 s. (C) Summary of baclofen-induced currents in dorsal CA1 pyramidal neurons from CaMKIICre(+):Girk2 fl/fl mice, 2 wk after infusion of Cre-dependent control (con, mCherry), GIRK2a, or GIRK2c virus (F2,11 = 24.3, P < 0.001; n = 4–5/group). Symbols: ***P < 0.001 vs. control. (D) Contextual fear learning in untreated CaMKIICre(−):Girk2 fl/fl mice and CaMKIICre(+):Girk2 fl/fl mice, and in CaMKIICre(+):Girk2 fl/fl mice following infusion of Cre-dependent control (con, mCherry), GIRK2a, or GIRK2c virus in the dorsal CA1 (F4,55 = 14.7, P < 0.001; n = 6–23/group). Symbols: *,***P < 0.05 and 0.001, respectively, vs. Cre(−). (E) Cue fear learning in untreated CaMKIICre(−):Girk2 fl/fl mice and CaMKIICre(+):Girk2 fl/fl mice, and in CaMKIICre(+):Girk2 fl/fl mice following infusion of Cre-dependent control (con, mCherry), GIRK2a, or GIRK2c virus in the dorsal CA1 (F4,55 = 4.0, P < 0.01; n = 6–23/group). Symbols: **P < 0.01 vs. Cre(−).
We utilized a trace fear conditioning protocol to assess the impact of viral treatment on fear learning behavior, as both contextual and cue fear learning in this model are supported by the dorsal hippocampus44, 45. Untreated CaMKIICre(+):Girk2 fl/fl mice, as well as CaMKIICre(+):Girk2 fl/fl mice treated with control virus, exhibited a deficit in contextual fear learning in this model (Fig. 7D), consistent with the contribution of GIRK-dependent signaling in dorsal CA1 to this behavior. Over-expression of either GIRK2a or GIRK2c in dorsal CA1 pyramidal neurons restored contextual fear learning in CaMKIICre(+):Girk2 fl/fl mice (Fig. 7D). Interestingly, while untreated CaMKIICre(+):Girk2 fl/fl mice, and CaMKIICre(+):Girk2 fl/fl mice treated with control virus, exhibited normal cue fear learning, over-expression of GIRK2a but not GIRK2c in dorsal CA1 pyramidal neurons from CaMKIICre(+):Girk2 fl/fl mice significantly enhanced cue fear learning (Fig. 7E). Thus, GIRK2a and GIRK2c exert overlapping but distinct influences on dissociable forms of hippocampal-dependent fear learning.
Discussion
In this study, we probed for differences between GIRK2a and GIRK2c in an array of expression systems. Our findings suggest that these two isoforms differ primarily in terms of subcellular trafficking, with GIRK2c achieving a more uniform distribution throughout neurons, including distal dendrites. This difference in subcellular trafficking likely underlies the differing abilities of GIRK2a and GIRK2c to support slow IPSCs evoked via the electrical stimulation of proximal and distal dendritic fields. Indeed, our findings suggest that GIRK2c is critical for ensuring robust synaptic inhibitory GIRK-dependent responses in the distal dendritic fields of CA1 pyramidal neurons.
We observed some interesting differences between GIRK-dependent signaling in transfected HEK cells and infected Girk2 −/− pyramidal neurons. Notably, GIRK1/GIRK2 channels expressed in HEK cells were more sensitive to GABABR activation (i.e., they exhibited a lower EC50) than the GIRK channels found in wild-type pyramidal neurons or Girk2 −/− neurons expressing GIRK2a or GIRK2c. This difference could be explained by a higher density of receptor and channel expressed in HEK cells, and/or by different receptor-channel ratios achieved in the different cell systems. The difference in coupling efficiency could also reflect the presence of negative regulatory elements in neurons that are not found in HEK cells. Indeed, we reported previously that a complex consisting of RGS7 and Gβ5 decreases the sensitivity of GIRK channels to GABABR activation in hippocampal neurons46,47,48, while also prominently accelerating GABABR-GIRK current deactivation kinetics. In support of this contention, the deactivation kinetics of GABABR-GIRK currents were significantly faster in neurons than in HEK cells.
Another difference between HEK cell and neuron models is the presence of GIRK3, which has been reported to decrease the activity and surface trafficking of GIRK1/GIRK2 heteromers and GIRK2 homomers in expression systems22, 49. We show here that while the relative dendritic labeling patterns for GIRK2a and GIRK2c were comparable in Girk2 −/− and Girk2 −/− /Girk3 −/− neurons, GPCR-GIRK currents measured in Girk2 −/− /Girk3 −/− neurons expressing GIRK2a or GIRK2c were larger than those measured in Girk2 −/− neurons. These results are consistent with the possible role of GIRK3 as a negative regulator of GIRK channel surface trafficking. In midbrain DA neurons, GIRK3 mediates an interaction between the GIRK channel, a GIRK2c/GIRK3 heteromer13, and SNX-2736, 50, 51, a protein implicated in the surface trafficking and internalization of several channels and receptors52,53,54,55,56. The interaction between SNX-27 and GIRK3 is conferred by a Class I PDZ interaction domain at the GIRK3 C-terminus, which is identical to that found in GIRK2c33. At present, the relevance of SNX-27 and GIRK3 to GIRK channel trafficking in hippocampal neurons is unclear.
Previous immunoelectron microscopy studies have shown that GIRK1 and GIRK2 labeling is concentrated at the peri- and extra-synaptic positions relative to the postsynaptic density (PSD) in hippocampal CA1. In addition, GIRK2 (but not GIRK1) labeling was detected within the PSD of CA1 pyramidal neurons15. The presence of a PDZ domain on the C-terminus of GIRK2c suggested that this isoform might promote the distribution of GIRK channels within or around the PSD, by virtue of its interaction with a scaffolding protein such as PSD-95. As noted above, GIRK2c was reported to bind PSD-9531, as well as SAP-9734. Moreover, SAP97 conferred both basal activity and G protein-dependent gating to GIRK2c-containing channels expressed in Xenopus oocytes34. Our data here, however, show that GIRK2a and GIRK2c assume comparable subcellular distributions relative to PSD-95 in mouse hippocampal pyramidal neurons. Thus, alternative splicing of the coding sequence of the Girk2 gene does not seem to influence the position of GIRK channels relative to the PSD. We did, however, detect significantly more Girk2c than Girk2a mRNA in the neuropil of CA1 in vivo, suggesting that RNA localization may play a role in setting the amount of GIRK2 isoforms in CA1 dendrites. Moreover, our RNA-Seq data suggest that the 3′ terminal exons of both isoforms may be extended in vivo, which could further regulate the targeting, translation, and turnover of these transcripts. Follow up studies looking at the significance of this extended transcription are needed.
The level of GIRK1 protein is significantly reduced in the hippocampus of constitutive Girk2 −/− mice15, 24. Our experiments with the GIRK1-selective agonist ML297, however, suggest that viral expression of GIRK2 isoforms in Girk2 −/− pyramidal neurons restored a normal level of functional GIRK1/GIRK2 heteromeric channels. Baclofen-induced currents in Girk2 −/− pyramidal neurons expressing either GIRK2a or GIRK2c were significantly larger than those measured in wild-type controls, however, indicating that some of the GABABR-GIRK current was carried by GIRK2 homomeric or GIRK2/GIRK3 heteromeric channels. Interestingly, expression of GIRK1 in VTA dopamine neurons, which normally express a GIRK2c/GIRK3 heteromeric channel, increased GIRK channel sensitivity to GABABR activation37. Moreover, GIRK2/GIRK3 heteromers are less sensitive to G protein activation than GIRK2 homomers37. Thus, an altered proportion of GIRK1-containing and GIRK1-lacking channels in Girk2 −/− pyramidal neurons expressing GIRK2a or GIRK2c could explain why some features of the baclofen-induced current, including the EC50, differed from those in wild-type neurons. Notably, adenosine and 5HT1AR-dependent currents in Girk2 −/− neurons expressing GIRK2a or GIRK2c did not differ in terms of amplitude from wild-type controls, suggesting that adenosine and 5HT1A receptors may couple preferentially to GIRK1-containining channels, while GABABR may be able to couple effectively to GIRK channels irrespective of channel subtype.
Multiple forms of plasticity involving GIRK-dependent signaling in hippocampal neurons have been reported. For example, Jan and colleagues showed that stimuli capable of inducing long-term potentiation (LTP) also enhanced GABABR-GIRK signaling in CA1 pyramidal neurons57. Subsequently, this group showed that NMDA receptor activation triggered a rapid increase in surface trafficking of GIRK channels in hippocampal cultures58, 59. This adaptation involved the dephosphorylation of an N-terminal GIRK2 residue (Ser-9) found in both GIRK2a and GIRK2c, and yielded enhanced basal GIRK channel activity and GIRK channel responses to adenosine receptor activation, but had no effect on GABABR-GIRK signaling. Slesinger and colleagues showed that prolonged morphine treatment in hippocampal cultures shifted the subcellular distribution of GIRK2 from dendritic shafts to spines, leading to enhanced basal and 5HT-induced GIRK currents, but no change in GABABR-GIRK signaling35. Our data suggest that this morphine-induced adaptation does not require elements within the Girk2 promoter or untranslated domains in the Girk2 mRNAs, as only coding sequence was included in the viral reconstitution vectors. The unique GIRK2c C-terminus is also not required, as both GIRK2a and GIRK2c were similarly impacted by morphine treatment. Interestingly, the strengthening of GIRK-dependent signaling in VTA dopamine neurons evoked by burst firing was blocked by inclusion of a peptide mimic of the GIRK2c (and GIRK3) C-terminal domain60. These data indicate that the GIRK2c C-terminal domain may be a critical determinant of plasticity involving specific GIRK channel subtypes, including the GIRK1-lacking channels expressed in midbrain dopamine neurons13, 31.
Constitutive Girk2 −/− mice exhibit an array of neurological phenotypes, including impaired fear learning6, 61. The global nature of the genetic manipulation, however, complicates attempts to link the phenotype to a defined neuron population. Accordingly, we developed conditional Girk2 −/− (Girk2 fl/fl) mice and have crossed them with transgenic Cre driver lines to permit GIRK channel ablation in a neuron-specific manner6, 61. In CaMKIICre(+):Girk2 fl/fl mice, Girk2 ablation is driven by Cre recombinase, expressed under the control of the CaMKIIα promoter. While the CaMKIICre driver line is generally considered to promote gene ablation in postnatal forebrain pyramidal neurons, the scope of recombination achieved with this driver line depends on the “floxed” gene in question, with some studies showing highly restricted target loss in dorsal CA1 pyramidal neurons and others showing recombination in multiple forebrain pyramidal neuron populations62,63,64,65,66,67. We found that GIRK2 expression was preserved in most forebrain structures in CaMKIICre(+):Girk2 fl/fl mice, but that GIRK-dependent signaling was blunted in dorsal (but not ventral) CA1 pyramidal neurons6.
To further probe the contribution of GIRK-dependent signaling in the hippocampus to fear learning, we assessed the performance of CaMKIICre(+):Girk2 fl/fl mice in trace fear conditioning test. In this paradigm, the trace interval inserted between the presentation of the conditioned (cue/tone) and unconditioned (footshock) stimulus activates temporal memory mechanisms within the hippocampus, thus rendering both contextual and cue fear learning dependent on the hippocampus44, 68. Viral-mediated over-expression of either GIRK2a or GIRK2c in dorsal CA1 pyramidal neurons rescued contextual fear learning in CaMKIICre(+):Girk2 fl/fl mice, suggesting that the fear learning deficit in CaMKIICre(+):Girk2 fl/fl mice is linked to GIRK2 loss in these neurons. Although we did not see a difference in cue fear learning between untreated CaMKIICre(−):Girk2 fl/fl and CaMKIICre(+):Girk2 fl/fl mice, over-expression of GIRK2a, but not GIRK2c, in dorsal CA1 pyramidal neurons significantly enhanced cue learning. The enhanced cue learning seen in CaMKIICre(+):Girk2 fl/fl mice over-expressing GIRK2a in dorsal CA1 pyramidal neurons, together with our evidence that the GIRK2 isoforms contribute in distinct fashion to the processing of inhibitory input to proximal and distal dendritic fields, suggests that trace cue learning may be differentially modulated by inputs from the PP and SC.
In summary, we demonstrate in this study that alternative splicing of the mouse Girk2 gene generates at least two variants, and that these splice variants can give rise to channel subtypes that differ in terms of their subcellular trafficking and the neurophysiological processes in which they participate. As GIRK-dependent signaling plays an important role in moderating glutamatergic input to neurons, our findings raise the intriguing possibility that altering the balance of GIRK2 splice variants could represent a means by which neurons fine-tune the processing of excitatory synaptic input, in a subcellular compartment-dependent manner.
Methods
Animals
All animal studies were approved by the Institutional Animal Care and Use Committee at the University of Minnesota or NIEHS, and were conducted in accordance with National Institutes of Health guidelines for the care and use of animals in research. The generation of Girk2 −/−, Girk2 −/− /Girk3 −/−, and CaMKIICre(+):Girk2 fl/fl mice was described previously6, 12, 24. The Amigo2-EGFP line was acquired from GENSAT (founder line LW244), and bred for at least 10 generations onto C57BL6/n background. Mice were maintained on a 12 h light/dark cycle (lights on at 0700), with food and water available ad libitum.
Chemicals
Baclofen (R-(+)-β-(aminomethyl)-4-chlorobenzenepropanoic acid hydrochloride), adenosine, CGP54626, kynurenic acid, and picrotoxin were purchased from Sigma-Aldrich (St. Louis, MO). (R)-(+)-8-OH-DPAT hydrobromide and WAY100635 maleate were purchased from Tocris (Bristol, UK). ML297 was synthesized in the Vanderbilt Center for Neuroscience Drug Discovery.
DNA constructs and virus production
pcDNA3-based expression constructs containing coding sequence for rat GIRK1, mouse GIRK2a (NM_001025584.2, initially referred to as Girk2-1: NP_001020755, AAC34284), and mouse GIRK2c (NM_010606.2, initially referred to as Girk2-A or Girk2A-1: NP_034736, AAC34285, P48542), as well as mouse GABABR1 and GABABR2 (provided by Dr. Paul Slesinger), were used for HEK293 cell transfection and electrophysiology studies. AAV vectors used for expression of GIRK2a or GIRK2c were generated by sub-cloning GIRK2a/c-IRES-EGFP cassettes into the backbones of pAAV-hSyn-TRβ-IRES-EGFP (provided by Dr. Bernd Gloss), pAAV-CaMKIIα-hChR2(C128S/D156A)-mCherry (provided by Dr. Karl Deisseroth; Addgene plasmid #35502)69, and pAAV-hSyn-DIO-hM4D(Gi)-mCherry (provided by Dr. Bryan Roth; Addgene plasmid #44362)70 plasmids. The AAV Helper-Free System was used for viral packaging (Agilent Technologies; Santa Clara, CA). AAV particles were obtained by co-transfecting pHelper (provided by Dr. David Armstrong), pAAV-RC2/8 (UPenn Viral Vector Core), and pShuttle plasmids into AAV293 cells using the polyethylenimine (PEI) method (Polysciences, Inc.; Warrington, PA). For each 15-cm plate, a total of 100 μg of DNA (33.3 μg of each plasmid) was added to a 150 mM NaCl solution, in a final volume of 2 mL; 12.5 μL of PEI (16 mg/ml) was then added while mixing constantly to favor the formation of DNA/PEI complexes. After 10 min of incubation, the complexes were added drop wise to AAV293 cells maintained in DMEM supplemented with 10% FBS, 4 mM L-glutamine, 1 mM sodium pyruvate at 37 °C and 5% CO2. After 72 h, cells were collected for viral particle extraction and purification. Viral titering was performed by qPCR with specific primers for the WPRE region of the AAV vectors: 5′-CCGTTGTCAGGCAACGTG-3′ (forward) and 5′-AGCTGACAGGTGGTGGCAAT-3′ (reverse). Control viruses (AAV8-CaMKII-EGFP, AAV8-hSyn-EGFP, and AAV8-hSyn-DIO-mCherry) were obtained from the UNC Viral Vector Core (Chapel Hill, NC).
RNA-Seq and analysis
Total RNA was extracted from laser-captured tissue microdissected from hippocampal CA1 cell body and neuropil layers from 3 adult (6–8 wk) male Amigo2-EGFP transgenic mice using the micro RNeasy kit (QIAGEN; Hilden, Germany), including on-column DNase-treatment. Total RNA samples were analysed for RNA integrity (RIN > 8.5) and concentration using the 2100 Bioanalyzer instrument (Agilent; Santa Clara, CA) and RNA 6000 Pico assay (Agilent). Stranded RNA-Seq libraries were prepared using 1–5 ng of total RNA per sample and region (1 ng for neuropil samples and 5 ng for cell body samples) with the Ovation RNA-Seq Systems 1–16 for mouse (NuGEN; San Carlos, CA), according to manufacturer instructions. Libraries were analyzed for size and concentration using the 2100 Agilent Bioanalyzer and the High Sensitivity DNA assay. Libraries were multiplexed and run on a NextSeq500 instrument (Illumina; San Diego, CA), acquiring an average of 66 ± 17 million 100 bp reads per sample. Reads were trimmed using Sickle software (https://github.com/najoshi/sickle) and only paired reads with a quality score >20 and a minimum length of 20 bp were pseudo-aligned and quantified to merged GENCODE (M9) and RefSeq gene models using Kallisto71. Samples were normalized and pairwise comparisons (cell body to neuropil) were made using Sleuth software (https://github.com/pachterlab/sleuth)72, with a false discovery rate (FDR, q value) of 0.05 (reported in Results). Scatter plots for Kncj6 transcripts corresponding to accession numbers NM_001025584.2 (Girk2a) and NM_010606.2 (Girk2c) were made using log2 normalized counts.
Cell culture and transfection/infection
HEK293 cells (ATCC; Manassas, VA) were cultured at 37 °C/5%CO2 according to supplier recommendations. For electrophysiological studies, cells were plated on 8-mm glass coverslips (15,000 cells/well), and transfected using calcium phosphate with a mixture of pcDNA3-based expression plasmids harboring coding sequences for EGFP (10 ng/well), GABABR1 (25 ng/well), GABABR2 (25 ng/well), GIRK1 (25 ng/well), and either GIRK2a or GIRK2c (25 ng/well); experiments were conducted 16–24 h later. For immunoblotting experiments, cells were plated on 6-well plates (80,000 cells/well) and transfected using Lipofectamine 2000 reagent (Thermo Fisher Scientific; Waltham, MA); 2 μg of GIRK1, GIRK2a, GIRK2c or GIRK3 expression plasmids were used per well, and cells were harvested 48 h after transfection.
Primary cultures of hippocampal neurons were prepared as described46, with minor modifications. Briefly, hippocampi were extracted from neonatal (P1–4) pups and placed into an ice-cold modified Hank’s Balanced Salt Solution (HBSS, Sigma-Aldrich) containing 1 mM kynurenic acid, 12 mM MgCl2, 3 mM HEPES, and 5.5 mM D-glucose. The tissue was digested for 20 min with papain and DNAse I in the modified HBSS solution at 37 °C with occasional inversion. Tissue was then washed twice with modified HBSS, twice with modified HBSS plus 20% FBS, and twice with growth medium: Neural-basal A medium containing 2% B27 supplement, 0.5 mM Glutamax, and antibiotic/antimycotic (Thermo Fisher Scientific). Hippocampi were mechanically-dissociated in growth medium using trituration in 1 mL pipettes. Neurons were pelleted by centrifugation (150 × g, for 5 min at 4 °C), and plated onto 8-mm glass coverslips pre-coated with 0.0005% poly-L-lysine (Sigma-Aldrich) in 48-well plates. Neurons were maintained in culture at 37 °C/5%CO2, and half of the medium was replaced with fresh growth medium every 3–4 d. After 10 d in culture, neurons were infected with control or GIRK expression viruses; electrophysiological analysis or immunofluorescence microscopy was conducted 4–5 d after infection.
Immunoblotting
Intact adult hippocampi and primary hippocampal cultures from wild-type and Girk2 −/− mice were homogenized in RIPA lysis buffer containing Halt phosphatase and protease inhibitor cocktail (Thermo Fisher Scientific) and centrifuged at 4 °C for 20 min at 16,000 × g. The supernatant was taken and protein concentrations were determined by BCA assay. Equal amounts of protein from each sample were mixed with SDS sample buffer and heated at 85 °C for 10 min. Samples were then separated by 12% SDS-PAGE and transferred to nitrocellulose membranes. Membranes were blocked in 5% milk/PBS, incubated overnight at 4 °C with rabbit polyclonal antibodies directed against GIRK1 (1:200, Alomone Labs; Jerusalem, Israel), GIRK2 (1:200, Alomone Labs), GIRK3 (1 μg/mL, Frontier Institute Co.; Ishikari, Japan), or β-actin (0.27 μg/mL, Abcam; Cambridge, MA), and diluted in 5% milk/PBS/0.1% Tween 20. The custom GIRK2c antibody was generated and affinity-purified using a peptide corresponding to the unique 11-amino acid C-terminal domain (DVANLENESKV, Pacific Immunology Corp.; Ramona, CA). Membranes were washed with PBS/0.1% Tween 20 and incubated with donkey anti-rabbit (0.2 μg/mL, LI-COR Biotechnology; Lincoln, NE) or donkey anti-mouse (0.5 μg/mL; LI-COR) secondary antibodies. Blots were developed using the Odyssey infrared imaging system (LI-COR), and the integrated density of each band was measured using NIH ImageJ software (Bethesda, MD).
Hippocampal organotypic culture and infection
Slice cultures were prepared as described73, with small modifications. Briefly, neonatal mice (7–9 d) were anesthetized with halothane in an induction chamber and decapitated. Brains were quickly removed and hippocampi dissected into ice-cold and oxygenated (95% O2/5% CO2) cutting medium: Minimal Essential Media (MEM) supplemented with 25 mM HEPES, 10 mM Tris-base, 10 mM glucose, and 3 mM MgCl2 (pH 7.2). Transverse hippocampal slices (400 μm) were cut using a tissue slicer (Stoelting Co.; Wood Dale, IL). Slices were placed onto transwell membrane inserts (Corning, Inc.; Corning, NY) in 6-well plates; each well was pre-filled with 1.6 mL of a 2:1 mixture of Basal Medium Eagle (Sigma) and Earle’s Balanced Salt Solution (Sigma) supplemented with (in mM): 20 NaCl, 5 NaHCO3, 0.2 CaCl2, 1.7 MgSO4 48 glucose, 26.7 HEPES, penicillin (40,000 U/L) (Thermo Fisher Scientific), streptomycin (40 mg/L) (Thermo Fisher Scientific), insulin (1 mg/L) (Sigma-Aldrich), 0.5 ascorbic acid and 5% horse serum (Thermo Fisher Scientific) (pH 7.2). Slices were maintained in an incubator at 34 °C/5% CO2, and were fed 3 times per week by replacing half of the growth medium with fresh medium. Slices were infected with purified AAV viruses in a 1:50 dilution with 5% glycerol/10% sucrose PBS solution, using a Nanoject III injector (Drummond Scientific Company; Broomall, PA) one day after plating. Recordings were made 6–7 d post-infection.
Electrophysiology in cultured cells
Coverslips containing transfected HEK cells or infected hippocampal neurons were transferred to a chamber containing a low-K+ bath solution (in mM): 130 NaCl, 5.4 KCl, 1 CaCl2, 1 MgCl2, 5.5 D-glucose, 5 HEPES/NaOH (pH 7.4). Neurons were visualized using an Olympus IX-70 microscope. Fire-polished borosilicate patch pipettes (3–5 MΩ) were filled with K-gluconate pipette solution (in mM): 140 K-gluconate, 2 MgCl2, 1.1 EGTA, 5 HEPES, 2 Na2-ATP, 0.3 Na-GTP, and 5 phosphocreatine (pH 7.2). Upon achieving whole-cell access, cells were held in voltage-clamp mode at −70 mV (with no correction for liquid junction potential). GIRK currents were measured in a high-K+ bath solution containing (in mM): 120 NaCl, 25 KCl, 1 CaCl2, 1 MgCl2, 5.5 D-glucose, 5 HEPES/NaOH (pH 7.4); GPCR agonists were dissolved in the high-K+ bath solution, which was then applied directly to the cell using a ValveLink 8.2 rapid perfusion system (AutoMate Scientific, Inc.; Berkeley, CA). Whole-cell currents were measured at room temperature with hardware (Axopatch-200B amplifier, Digidata 1322A) and software (pCLAMP v. 8.2) from Molecular Devices, LLC (Sunnyvale, CA). All currents were low-pass filtered at 2 kHz, digitized at 10 kHz, and stored on computer hard disk for subsequent analysis. Only those experiments in which the access resistances were stable and low (<18 MΩ) were included in the analysis. Peak and steady-state currents were measured for each experiment. Current activation and deactivation rates were extracted from 1-term Boltzmann/exponential fits of the recording segments corresponding to the onset and offset, respectively, of drug effect.
Electrophysiology in organotypic slices
Organotypic hippocampal slices were cut out of the culture insert and positioned in a recording chamber superfused (2–2.5 mL/min) with an external solution containing (in mM): 125 NaCl, 2.5 KCl, 25 NaHCO3, 1.3 MgCl2, 2 CaCl2, 0.4 ascorbic acid and 10 glucose (pH 7.4) and bubbled with 95% O2/5% CO2. Kynurenic acid (2 mM) and picrotoxin (100 μM) were added to the bath solution to block ionotropic glutamate and GABAA receptors, respectively. Bath and chamber temperatures were maintained at 29–30 °C using a TC344C Dual Automatic Temperature Controller (Warner Instruments; Hamden, CT). Whole-cell recordings (Vhold = −60 mV) were made from EGFP-positive CA1 pyramidal neurons using borosilicate pipettes (3–5 MΩ) filled with a K-gluconate pipette solution. All measured and command potentials factored in a junction potential (−15 mV) predicted using JPCalc software (Molecular Devices, LLC). Recordings were amplified with an EPC10 HEKA amplifier, filtered at 2 kHz, digitized at 5–10 kHz using Patchmaster 2x73.2 software (HEKA Elektronik; Bellmore, NY) and stored on hard disk. Stimulation was delivered with a cluster-type stimulating electrode (FHC, Bowdoin, ME) and a DS3 stimulator (Digitimer North America, LLC; Ft. Lauderdale, FL). Stimulating electrodes were placed in the stratum radiatum or stratum lacunosum moleculare, the dendritic fields that receive Schaffer Collateral (SC) and Perforant Path (PP) inputs, respectively. Stimuli of increasing intensity (0.2–25 mA) were applied at a 0.05 Hz frequency to establish maximal slow IPSC amplitude. Slow IPSCs evoked by SC and PP were made in each neuron, and the order of stimulation was counterbalanced across experiments. Only experiments with stable (<20% variation) and low series resistances (<30 MΩ) were analyzed.
Immunofluorescence microscopy
Cultured hippocampal neurons were fixed with 4% paraformaldehyde and 0.4% sucrose in PBS for 10 min, permeabilized with 0.2% Triton X-100 in PBS for 10 min, washed with PBS, blocked with 10% normal goat serum in PBS for 1 h, and then incubated at 4 °C overnight with primary antibodies: pan-GIRK2 (1:500, Alomone Labs), PSD-95 (2 μg/mL, Thermo Fisher Scientific), MAP2 (1:1000, PhosphoSolutions; Aurora, CO) and CaMKIIα (1:1000, EMD Millipore; Billerica, MA). After washing 3–5 times in PBS, cells were incubated with secondary antibodies tagged with FITC (1:2000, Jackson ImmunoResearch Laboratories, Inc.; West Grove, PA) or Alexa-595/647 (2 μg/mL, Thermo Fisher Scientific) for 1 h at room temperature and protected from light. After 3 washes in PBS, coverslips were mounted onto glass slides for image acquisition. Fluorescent images were captured under a 40X or 60X (oil-immersion) objectives from a BX51W1 upright microscope with Disk Spinning Unit confocal system (Olympus; Center Valley, PA) and digital CCD camera (C10600-10B, Hamamatsu Photonic System Corp; Bridgewater, NJ). Images were processed and analyzed with Metamorph Advanced 7.7.7.0 (Molecular Devices) and NIH ImageJ software.
For quantification of GIRK2 fluorescence in dendritic processes, only infected neurons that were clearly separated from other infected cells were evaluated. MAP2 labeling was used to identify clearly the dendrites; 2–3 relatively straight segments (35–50 microns) of primary, secondary and tertiary dendrites exhibiting even MAP2 labeling from each neuron were selected for analysis. The primary dendrite fragments were located at least 20 microns from the soma. GIRK2 fluorescence intensity measurements within these processes were acquired following background subtraction, and data are presented as arbitrary units (AU). For GIRK2 isoform and PSD-95 co-localization analysis, the soma and surrounding area (20 microns from the edge of the soma) was removed from the image prior to analysis. Quantification was performed using the NIH ImageJ 1.29 plug-in, Puncta Analyzer. MAP2 labeling was used to delineate dendritic processes, and red (GIRK2) and green (PSD-95) channels were thresholded to highlight visible puncta without introduction of background noise. The percentage of co-localized GIRK puncta overlapping with total PSD-95 puncta was calculated for each analyzed neuron.
Viral expression of GIRK2a and GIRK2c in dorsal CA1
CaMKIICre(+):Girk2 fl/fl mice (7–8 wk) were placed in a stereotaxic device (David Kopf Instruments; Tujunga, CA) under isoflurane anesthesia. Microinjectors were created by affixing a 33-gauge stainless steel hypodermic tube within a shorter 26-gauge stainless steel hypodermic tube, and were attached to polyethylene-20 tubing affixed to 10 μL Hamilton syringes. Microinjectors were lowered bilaterally through burr holes in the skull to the dorsal CA1 (bregma: −2.20 mm AP, ±1.60 mm ML, −1.40 mm DV) and 400 nL of AAV8-hSyn-DIO-GIRK2a-IRES-EGFP, AAV8-hSyn-DIO-GIRK2c-IRES-EGFP, or AAV8-hSyn-DIO-mCherry control virus (~1.3 × 1013 viral particles/mL) was injected per side over 4 min using a Pump 11 Elite infusion pump (Harvard Apparatus; Holliston, MA). The syringe was left in place for 10 min following infusion to reduce solution backflow along the infusion track. Behavioral experiments were performed 2 wk after surgery to allow for full recovery and viral expression. The accuracy of viral targeting was assessed in all experiments using fluorescence confocal microscopy. Images were acquired with MetaMorph Advanced Acquisition v. 7.7.7.0 software. GIRK-dependent signaling in CA1 pyramidal neurons expressing GIRK2a or GIRK2c, indicated by EGFP expression, was assessed by measuring baclofen-induced somato-dendritic currents in acutely isolated slices, as described6.
Fear conditioning
To most effectively isolate the contribution of the hippocampus to context fear learning, we utilized a trace fear conditioning protocol45. In brief, mice were acclimated to and handled within the testing room for 2 d prior to testing. For the training session (Day 1), mice were placed in fear conditioning chambers (Med Associates, Inc.; St Albans, VT) with metal bar floors and white wall insert that were cleaned with 75% ethanol. A 30-s white noise (65 dB/3 kHz) was used as the conditioned stimulus (CS, cue), and a 2-s footshock (0.50 mA) was used as the unconditioned stimulus (US). Training consisted of 2 CS-US pairings in which the US was delivered 30 s after the end of the CS; CS presentations were separated by 90 s (7.5 min total conditioning time). On Day 2, the level of contextual fear learning was assessed by returning mice to the training chambers and measuring freezing behavior for 5 min. On Day 3, the chambers were reconfigured (i.e., floor bars were covered with a plastic insert and white wall inserts were replaced with black tent inserts to alter both the color and shape of the chamber) and cleaned with 0.1% acetic acid. Mice were then assessed for freezing behavior in response to 2 × 3-min CS presentations. Freezing was assessed by Video Freeze software (Med Associates, Inc.) and data are presented as percent time spent freezing.
Statistical analysis
Data are presented throughout as the mean ± SEM. Statistical analyses were performed using Prism 6 or 7 (GraphPad Software, Inc.; La Jolla, CA). Data were analyzed with Student’s two-tailed t-test, 1-way ANOVA, 2-way ANOVA, or 2-way ANOVA with repeated measures, as appropriate. Tukey, Holm-Sidak, Dunnett’s, and Bonferroni multiple comparison post hoc tests were used, if justified. Differences were considered significant if P < 0.05.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Slesinger, P. A. & Wickman, K. Structure to Function of G Protein-Gated Inwardly Rectifying (GIRK) Channels, In International Review of Neurobiology Vol. 123 (eds R. A. Harris & P. Jenner) 365 (Elsevier, 2015).
Lujan, R. & Aguado, C. Localization and targeting of GIRK channels in mammalian central neurons. Int. Rev. Neurobiol 123, 161–200 (2015).
Otmakhova, N. A. & Lisman, J. E. Contribution of Ih and GABAB to synaptically induced afterhyperpolarizations in CA1: a brake on the NMDA response. J. Neurophysiol. 92, 2027–2039 (2004).
Lujan, R., Marron Fernandez de Velasco, E., Aguado, C. & Wickman, K. New insights into the therapeutic potential of GIRK channels. Trends Neurosci. 37, 20–29 (2014).
Mayfield, J., Blednov, Y. A. & Harris, R. A. Behavioral and genetic evidence for GIRK channels in the CNS: role in physiology, pathophysiology, and drug addiction. Int. Rev. Neurobiol 123, 279–313 (2015).
Victoria, N. C. et al. G protein-gated K+ channel ablation in forebrain pyramidal neurons selectively impairs fear learning. Biol. Psychiatry 80, 796–806 (2016).
Karschin, C., Dissmann, E., Stuhmer, W. & Karschin, A. IRK(1-3) and GIRK(1-4) inwardly rectifying K+ channel mRNAs are differentially expressed in the adult rat brain. J. Neurosci. 16, 3559–3570 (1996).
Wickman, K., Karschin, C., Karschin, A., Picciotto, M. R. & Clapham, D. E. Brain localization and behavioral impact of the G-protein-gated K+ channel subunit GIRK4. J. Neurosci. 20, 5608–5615 (2000).
Perry, C. A. et al. Predisposition to late-onset obesity in GIRK4 knockout mice. Proc. Natl. Acad. Sci. USA 105, 8148–8153 (2008).
Luscher, C., Jan, L. Y., Stoffel, M., Malenka, R. C. & Nicoll, R. A. G protein-coupled inwardly rectifying K+ channels (GIRKs) mediate postsynaptic but not presynaptic transmitter actions in hippocampal neurons. Neuron 19, 687–695 (1997).
Slesinger, P. A., Stoffel, M., Jan, Y. N. & Jan, L. Y. Defective gamma-aminobutyric acid type B receptor-activated inwardly rectifying K+ currents in cerebellar granule cells isolated from weaver and Girk2 null mutant mice. Proc. Natl. Acad. Sci. USA 94, 12210–12217 (1997).
Torrecilla, M. et al. G-protein-gated potassium channels containing Kir3.2 and Kir3.3 subunits mediate the acute inhibitory effects of opioids on locus ceruleus neurons. J. Neurosci. 22, 4328–4334 (2002).
Cruz, H. G. et al. Bi-directional effects of GABAB receptor agonists on the mesolimbic dopamine system. Nat. Neurosci. 7, 153–159 (2004).
Beckstead, M. J., Grandy, D. K., Wickman, K. & Williams, J. T. Vesicular dopamine release elicits an inhibitory postsynaptic current in midbrain dopamine neurons. Neuron 42, 939–946 (2004).
Koyrakh, L. et al. Molecular and cellular diversity of neuronal G-protein-gated potassium channels. J. Neurosci. 25, 11468–11478 (2005).
Marker, C. L., Lujan, R., Colon, J. & Wickman, K. Distinct populations of spinal cord lamina II interneurons expressing G-protein-gated potassium channels. J. Neurosci. 26, 12251–12259 (2006).
Arora, D. et al. Altered neurotransmission in the mesolimbic reward system of Girk −/− mice. J. Neurochem 114, 1487–1497 (2010).
Hearing, M. et al. Repeated cocaine weakens GABA(B)-GIRK signaling in layer 5/6 pyramidal neurons in the prelimbic cortex. Neuron 80, 159–170 (2013).
Krapivinsky, G. et al. The G-protein-gated atrial K+ channel IKACh is a heteromultimer of two inwardly rectifying K+-channel proteins. Nature 374, 135–141 (1995).
Kennedy, M. E., Nemec, J. & Clapham, D. E. Localization and interaction of epitope-tagged GIRK1 and CIR inward rectifier K+ channel subunits. Neuropharmacology 35, 831–839 (1996).
Kennedy, M. E., Nemec, J., Corey, S., Wickman, K. & Clapham, D. E. GIRK4 confers appropriate processing and cell surface localization to G-protein-gated potassium channels. J. Biol. Chem. 274, 2571–2582 (1999).
Ma, D. et al. Diverse trafficking patterns due to multiple traffic motifs in G protein-activated inwardly rectifying potassium channels from brain and heart. Neuron 33, 715–729 (2002).
Liao, Y. J., Jan, Y. N. & Jan, L. Y. Heteromultimerization of G-protein-gated inwardly rectifying K+ channel proteins GIRK1 and GIRK2 and their altered expression in weaver brain. J. Neurosci. 16, 7137–7150 (1996).
Signorini, S., Liao, Y. J., Duncan, S. A., Jan, L. Y. & Stoffel, M. Normal cerebellar development but susceptibility to seizures in mice lacking G protein-coupled, inwardly rectifying K+ channel GIRK2. Proc. Natl. Acad. Sci. USA 94, 923–927 (1997).
Lesage, F. et al. Cloning provides evidence for a family of inward rectifier and G-protein coupled K+ channels in the brain. FEBS Lett 353, 37–42 (1994).
Lesage, F. et al. Molecular properties of neuronal G-protein-activated inwardly rectifying K+ channels. J. Biol. Chem. 270, 28660–28667 (1995).
Isomoto, S. et al. A novel ubiquitously distributed isoform of GIRK2 (GIRK2B) enhances GIRK1 expression of the G-protein-gated K+ current in Xenopus oocytes. Biochem. Biophys. Res. Commun. 218, 286–291 (1996).
Nelson, C. S., Marino, J. L. & Allen, C. N. Cloning and characterization of Kir3.1 (GIRK1) C-terminal alternative splice variants. Brain Res. Mol. Brain Res. 46, 185–196 (1997).
Wei, J. et al. Characterization of murine Girk2 transcript isoforms: structure and differential expression. Genomics 51, 379–390 (1998).
Inanobe, A. et al. Molecular cloning and characterization of a novel splicing variant of the Kir3.2 subunit predominantly expressed in mouse testis. J. Physiol 521(Pt 1), 19–30 (1999).
Inanobe, A. et al. Characterization of G-protein-gated K+ channels composed of Kir3.2 subunits in dopaminergic neurons of the substantia nigra. J. Neurosci. 19, 1006–1017 (1999).
Nehring, R. B. et al. Neuronal inwardly rectifying K+ channels differentially couple to PDZ proteins of the PSD-95/SAP90 family. J. Neurosci. 20, 156–162 (2000).
Balana, B. et al. Mechanism underlying selective regulation of G protein-gated inwardly rectifying potassium channels by the psychostimulant-sensitive sorting nexin 27. Proc. Natl. Acad. Sci. USA 108, 5831–5836 (2011).
Hibino, H. et al. Anchoring proteins confer G protein sensitivity to an inward-rectifier K+ channel through the GK domain. EMBO J 19, 78–83 (2000).
Nassirpour, R. et al. Morphine- and CaMKII-dependent enhancement of GIRK channel signaling in hippocampal neurons. J. Neurosci. 30, 13419–13430 (2010).
Lunn, M. L. et al. A unique sorting nexin regulates trafficking of potassium channels via a PDZ domain interaction. Nat. Neurosci. 10, 1249–1259 (2007).
Labouebe, G. et al. RGS2 modulates coupling between GABAB receptors and GIRK channels in dopamine neurons of the ventral tegmental area. Nat. Neurosci. 10, 1559–1568 (2007).
Fernandez-Alacid, L., Watanabe, M., Molnar, E., Wickman, K. & Lujan, R. Developmental regulation of G protein-gated inwardly-rectifying K+ (GIRK/Kir3) channel subunits in the brain. Eur. J. Neurosci 34, 1724–1736 (2011).
Kulik, A. et al. Compartment-dependent colocalization of Kir3.2-containing K+ channels and GABAB receptors in hippocampal pyramidal cells. J. Neurosci. 26, 4289–4297 (2006).
Sodickson, D. L. & Bean, B. P. Neurotransmitter activation of inwardly rectifying potassium current in dissociated hippocampal CA3 neurons: interactions among multiple receptors. J. Neurosci. 18, 8153–8162 (1998).
Leaney, J. L. Contribution of Kir3.1, Kir3.2A and Kir3.2C subunits to native G protein-gated inwardly rectifying potassium currents in cultured hippocampal neurons. Eur. J. Neurosci. 18, 2110–2118 (2003).
Kaufmann, K. et al. ML297 (VU0456810), the first potent and selective activator of the GIRK potassium channel, displays antiepileptic properties in mice. ACS Chem. Neurosci 4, 1278–1286 (2013).
Wydeven, N. et al. Mechanisms underlying the activation of G-protein-gated inwardly rectifying K+ (GIRK) channels by the novel anxiolytic drug, ML297. Proc. Natl. Acad. Sci. USA 111, 10755–10760 (2014).
Misane, I. et al. Time-dependent involvement of the dorsal hippocampus in trace fear conditioning in mice. Hippocampus 15, 418–426 (2005).
Raybuck, J. D. & Lattal, K. M. Double dissociation of amygdala and hippocampal contributions to trace and delay fear conditioning. PloS One 6, e15982 (2011).
Xie, K. et al. Gbeta5 recruits R7 RGS proteins to GIRK channels to regulate the timing of neuronal inhibitory signaling. Nat. Neurosci. 13, 661–663 (2010).
Fajardo-Serrano, A. et al. Association of RGS7/Gb5 complexes with GIRK channels and GABAB receptors in hippocampal CA1 pyramidal neurons. Hippocampus 23, 1231–1245 (2013).
Ostrovskaya, O. et al. RGS7/Gb5/R7BP complex regulates synaptic plasticity and memory by modulating hippocampal GABABR-GIRK signaling. eLife 3, e02053 (2014).
Kofuji, P., Davidson, N. & Lester, H. A. Evidence that neuronal G-protein-gated inwardly rectifying K+ channels are activated by G beta gamma subunits and function as heteromultimers. Proc. Natl. Acad. Sci. U.S.A. 92, 6542–6546 (1995).
Nassirpour, R. & Slesinger, P. A. Subunit-specific regulation of Kir3 channels by sorting nexin 27. Channels 1, 331–333 (2007).
Munoz, M. B. & Slesinger, P. A. Sorting nexin 27 regulation of G protein-gated inwardly rectifying K+ channels attenuates in vivo cocaine response. Neuron 82, 659–669 (2014).
Valdes, J. L. et al. Sorting nexin 27 protein regulates trafficking of a p21-activated kinase (PAK) interacting exchange factor (beta-Pix)-G protein-coupled receptor kinase interacting protein (GIT) complex via a PDZ domain interaction. J. Biol. Chem. 286, 39403–39416 (2011).
Cai, L., Loo, L. S., Atlashkin, V., Hanson, B. J. & Hong, W. Deficiency of sorting nexin 27 (SNX27) leads to growth retardation and elevated levels of N-methyl-D-aspartate receptor 2C (NR2C). Mol. Cell. Biol. 31, 1734–1747 (2011).
Wang, X. et al. Loss of sorting nexin 27 contributes to excitatory synaptic dysfunction by modulating glutamate receptor recycling in Down syndrome. Nat. Med 19, 473–480 (2013).
Hussain, N. K., Diering, G. H., Sole, J., Anggono, V. & Huganir, R. L. Sorting Nexin 27 regulates basal and activity-dependent trafficking of AMPARs. Proc. Natl. Acad. Sci. USA 111, 11840–11845 (2014).
Loo, L. S., Tang, N., Al-Haddawi, M., Dawe, G. S. & Hong, W. A role for sorting nexin 27 in AMPA receptor trafficking. Nat. Comm 5, 3176 (2014).
Huang, C. S. et al. Common molecular pathways mediate long-term potentiation of synaptic excitation and slow synaptic inhibition. Cell 123, 105–118 (2005).
Chung, H. J. et al. G protein-activated inwardly rectifying potassium channels mediate depotentiation of long-term potentiation. Proc. Natl. Acad. Sci. USA 106, 635–640 (2009).
Chung, H. J., Qian, X., Ehlers, M., Jan, Y. N. & Jan, L. Y. Neuronal activity regulates phosphorylation-dependent surface delivery of G protein-activated inwardly rectifying potassium channels. Proc. Natl. Acad. Sci. USA 106, 629–634 (2009).
Lalive, A. L. et al. Firing modes of dopamine neurons drive bidirectional GIRK channel plasticity. J Neurosci 34, 5107–5114 (2014).
Kotecki, L. et al. GIRK channels modulate opioid-induced motor activity in a cell type- and subunit-dependent manner. J. Neurosci. 35, 7131–7142 (2015).
Tsien, J. Z. et al. Subregion- and cell type-restricted gene knockout in mouse brain. Cell 87, 1317–1326 (1996).
Sonner, J. M. et al. Alpha 1 subunit-containing GABA type A receptors in forebrain contribute to the effect of inhaled anesthetics on conditioned fear. Mol. Pharmacol. 68, 61–68 (2005).
Pan, B. X., Vautier, F., Ito, W., Bolshakov, V. Y. & Morozov, A. Enhanced cortico-amygdala efficacy and suppressed fear in absence of Rap1. J. Neurosci. 28, 2089–2098 (2008).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Tolson, K. P. et al. Postnatal Sim1 deficiency causes hyperphagic obesity and reduced Mc4r and oxytocin expression. J. Neurosci. 30, 3803–3812 (2010).
Horn, K. E. et al. DCC expression by neurons regulates synaptic plasticity in the adult brain. Cell Rep 3, 173–185 (2013).
Raybuck, J. D. & Lattal, K. M. Bridging the interval: theory and neurobiology of trace conditioning. Behav. Processes 101, 103–111 (2014).
Yizhar, O., Fenno, L. E., Davidson, T. J., Mogri, M. & Deisseroth, K. Optogenetics in neural systems. Neuron 71, 9–34 (2011).
Krashes, M. J. et al. Rapid, reversible activation of AgRP neurons drives feeding behavior in mice. J. Clin. Invest. 121, 1424–1428 (2011).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Bio 34, 525–527 (2016).
Pimentel, H., Bray, N., Puente, S., Melsted, P. & Pachter, L. Differential analysis of RNA-Seq incorporating quantification uncertainty. bioRxiv in press (2017).
Gu, Z., Lamb, P. W. & Yakel, J. L. Cholinergic coordination of presynaptic and postsynaptic activity induces timing-dependent hippocampal synaptic plasticity. J. Neurosci. 32, 12337–12348 (2012).
Acknowledgements
The authors would like to thank Drs. Nicole Wydeven and José Colón Sáez for their early contributions to this project. This work was supported by NIH grants to K.W. (MH061933, DA034696, and MH107399), C.D.W. (MH107399), and S.M.D. (ES100221; the Intramural Research Program of NIEHS).
Author information
Authors and Affiliations
Contributions
K.W. and E.M.F.d.V. conceived the idea for the project and designed the overall scope of the study. K.W., E.M.F.d.V., L.Z., B.V., M.T., S.F., Z.X, A.A., N.C., C.D.W., and S.M.D. were responsible for experimental design, execution of experiments, and/or data analysis. K.W. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Marron Fernandez de Velasco, E., Zhang, L., N. Vo, B. et al. GIRK2 splice variants and neuronal G protein-gated K+ channels: implications for channel function and behavior. Sci Rep 7, 1639 (2017). https://doi.org/10.1038/s41598-017-01820-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-01820-2
This article is cited by
-
Alcohol reverses the effects of KCNJ6 (GIRK2) noncoding variants on excitability of human glutamatergic neurons
Molecular Psychiatry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.