Abstract
Zika is a mosquito-borne disease associated with neurological disorders that causes an on-going pandemic. The first outbreak was recorded in Micronesia in 2007, then in French Polynesia in 2014 from which it spread to South America in 2015 and ignited a widespread epidemic. Interestingly, Zika outbreaks in Asia remained of moderate intensity although the virus is circulating. To understand these epidemiological variations, we investigated the entomological determinants of ZIKV transmission in Asia. We used oral infection of mosquitoes collected in Singapore to identify the vector species, to quantify the blood infection threshold and to compare transmissibility between an Asian ZIKV strain (H/PF13) and an American strain collected in Brazil (BE H 815744). We have confirmed the vector status of Aedes aegypti and determined that 103 pfu/ml of blood is sufficient to infect mosquitoes. We showed that only the American strain was present in the saliva 3 days post-infection, and that this strain had a 30–40% higher rate of saliva infection in Ae. aegypti from 3 to 14 days post-infection than the Asian strain. Our data suggests that American strains are more efficiently transmitted than Asian strains, which raises concerns about the introduction of American strains in Asia.
Similar content being viewed by others
Introduction
ZIKA virus (ZIKV) is a newly emerging mosquito-borne virus that belongs to the Flavivirus genus of the Flaviviridae family. The virus can cause life-debilitating brain defects in adults and newborns from infected mothers1,2,3. ZIKV is responsible for an on-going pandemic that started in the Yap Island in 2007, then French Polynesia in 2014 and spread to South and Central America, where it resulted in a widespread epidemic4,5,6. Introduced on the American continent in early 2015, ZIKV has, in just two years, infected tens of thousands individuals and resulted in thousands confirmed cases of microcephaly in Brazil7,8,9. In response, the World Health Organization declared the cluster of microcephaly and other neurological disorder as a ‘Public Health Emergency of International Concern’ on the 1st of February 201610.
ZIKV is also present on the African and Asian continents but has not resulted in similarly large outbreaks, despite the presence of permissive mosquito vectors and favorable ecological conditions for transmission. Although the first human cases of ZIKA were detected in the 1950s in Uganda and Tanzania11, there have been no detectable outbreaks in Africa in the intervening years. For the past four decades, the virus has also been reported in many Southeast Asian countries12, 13. Since the emergence of the current epidemic in 2014, Singapore has reported its first autochthonous transmission in August 201614. By the end of 2016, however, the ZIKV introduction had only resulted in a minor epidemic (457 reported cases) and no new cases were reported in the last three weeks of the year (www.nea.gov.sg). In Cambodia, a recent study reported that the virus was circulating with a low prevalence between 2007 and 201615. While closely related genetically, Southeast Asian and American strains form different lineages16 and the Southeast Asian ZIKV might not be the direct source of the South American outbreaks17. A better characterization of the entomological determinants of ZIKV transmission in Asia could help understand the epidemiological variation between Asia and America.
ZIKV can be transmitted by Aedes aegypti and Aedes albopictus mosquito species18,19,20. ZIKV has also been reported to infect Culex quinquefasciatus 21, although this is highly controversial22,23,24 and may depend upon the origin of the mosquito colony and the virus strain. For instance, mosquito transmission efficiency for dengue virus, another epidemic flavivirus, depends upon mosquito origin25, 26 and mosquito-virus genetic interaction27.
Currently, there is no clear evidence to explain the relatively low prevalence of ZIKV infection in Southeast Asia, nor why there have been no large ZIKV outbreaks in Asia similar to those seen in South and Central America. Here, we endeavored to determine the entomological factors that might play a role in the epidemiology of ZIKV in Southeast Asia. Working with mosquito colonies collected in Singapore, we confirmed that Ae. aegypti mosquitoes are more susceptible to ZIKV infection than Ae. albopictus and ruled out Cx. quinquefasciatus as an efficient vector for ZIKV transmission. We also showed that Ae. aegypti is extremely susceptible to ZIKV infection by determining the blood infection threshold. Importantly, we found that a ZIKV strain from Brazil (BE H 815744) had higher transmissibility than a strain originating from the Asian lineage (H/PF13) in Ae. aegypti.
Results
Aedes aegypti is the main vector of Zika virus in Southeast Asia
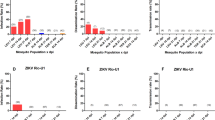
To identify the main vector of ZIKV in Southeast Asia, we compared the susceptibility of Ae. aegypti, Ae. albopictus and Cx. quinquefasciatus mosquitoes from Singapore to the French Polynesian (H/PF13) ZIKV strain that is representative of the Asian lineage13. The mosquitoes were orally fed with a mix of blood containing 106 or 105 pfu/ml in order to determine the influence of the inoculum concentration. At 7 days post-infection (p.i.), we detected and quantified the virus genome copies in whole mosquitoes. Infection rate and genome copies in Cx. quinquefasciatus were significantly lower than in Ae. aegypti for the two virus titers tested (Fig. 1A). Infection rates in Ae. albopictus mosquitoes were not different from Ae. aegypti, however the number of genome copies was significantly lower in Ae. albopictus.
Comparison of susceptibility to Zika virus infection between Ae. aegypti, Ae. albopictus and Cx. quinquefasciatus. Mosquitoes were orally infected with PF13 at a titer of 106 or 105 pfu/ml. (A) At 7 days p.i., ZIKV genome copies were quantified in whole mosquitoes. (B) At 14 days p.i., ZIKV genome copies were quantified in salivary glands. Infection rate and gRNA copies per infected mosquitoes are shown. Each point represents one mosquito sample. Line shows average of gRNA copies. Twenty mosquitoes were tested per condition. Asterisks indicate significant difference with Ae. aegypti in the same condition following a Z-test and T-test for infection rate and gRNA copies, respectively.
Transmission of arboviruses is dependent upon infection of salivary glands28. To differentiate the vector potential of Ae. aegypti and Ae. albopictus, we orally infected the two species with 106 or 105 pfu/ml and quantified the virus genome copies at 14 days p.i. in dissected salivary glands. While genome copies were not different between the two species, infection rate was significantly higher in salivary glands of Ae. aegypti (Fig. 1B). These results clearly demonstrate that Ae. aegypti is the main vector of ZIKV in Singapore and probably in Southeast Asia.
A very low blood titer threshold is required to infect Ae. aegypti mosquitoes
The capacity of the virus to infect mosquitoes greatly influences its transmission efficiency28. To determine the titer threshold for mosquito infection, we orally infected Ae. aegypti with H/PF13 at 105, 104, 103 or 102 pfu/ml of blood and quantified the virus genome copies in whole mosquitoes at 7 days p.i. All mosquitoes were infected up to 104 pfu/ml, 35% were infected with a titer of 103 pfu/ml and no mosquitoes were infected after feeding on 102 pfu/ml of blood (Fig. 2). ZIKV genome copies per infected mosquito gradually decreased in a dose-dependent manner. Average virus genome copies varied from 2 × 107, 4 × 106 to 8 × 105 after infection with 105, 104 and 103 pfu/ml, respectively. Our data determined the threshold of infection in Ae. aegypti.
Zika virus can infect mosquitoes at a titer of 103 pfu/ml in the blood. Aedes aegypti mosquitoes were orally infected with 105, 104, 103 or 102 pfu of PF13 per ml of blood. At 7 days p.i., ZIKV genome copies were quantified in whole mosquitoes. Each point represents one mosquito sample. Twenty mosquitoes were tested per condition. Line shows average of gRNA copies.
A strain from Brazil is more efficiently transmitted by Ae. aegypti than a strain from French Polynesia
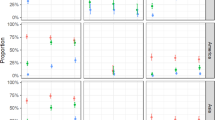
The extrinsic incubation period corresponds to the time between oral infection and presence of virus in the saliva of vectors. This period determines transmission efficiency of arboviruses28. To compare the efficiency of transmission for viruses from the Asian and the American lineages, we orally infected Ae. aegypti mosquitoes with H/PF13 (Asian lineage) or BE H 815744 (American lineage) with 105 pfu/ml of blood. At 3, 7, 10 and 14 days p.i., we quantified virus genome copies in whole mosquitoes and in saliva. In whole mosquitoes, BE H 815744 showed a higher infection rate early during our study period (day 3 p.i.), and had higher genome copies at each time point (Fig. 3A). To confirm this result, we titered the virus in whole mosquitoes at 3 days p.i. with H/PF13 or BE H 815744. Similarly to our results obtained with virus genome quantification, virus titer was significantly higher in mosquitoes infected with BE H 815744, although titers for both viruses were several orders lower than genome copies (Fig. S1). Strikingly, in the saliva, the Brazilian strain was present earlier (day 3 p.i.) and had higher rate of infection than H/PF13 at the other time points (Fig. 3B). Although only significant at 10 days p.i., the American strain had consistently higher average number of genome copies per infected saliva. These results indicate that the American strain BE H 815744 could be more efficiently transmitted than the Asia lineage strain H/PF13.
Aedes aegypti mosquitoes infected with a strain from Brazil have a higher infection rate of saliva. Aedes aegypti mosquitoes were orally infected with 105 pfu/ml of PF13 or the Brazil virus. At 3, 7, 10 and 14 days p.i., Zika virus genome copies was quantified in (A) whole mosquitoes and (B) saliva. Percent of infection rate and average ± s.e.m. for genome copies per infected mosquitoes or saliva are presented. Thirty mosquitoes were analyzed per condition. Asterisks show differences between the two viruses within the same day following a Z-Test and T-test for infection rate and gRNA copies, respectively.
Discussion
The on-going ZIKA pandemic continues to spread throughout the tropics and sub-tropics but has not caused outbreaks of similar magnitude in all regions. As with other pathogenic Flaviviruses, ZIKV is transmitted through the bite of a mosquito that inoculates the virus along with its saliva when biting. Additionally, ZIKV can be transmitted through sexual intercourse9, 29, 30 and between the mother and her foetus31. While the rapid expansion of the virus across the tropical world is indicative of efficient transmission, variation in transmissibility could account for the epidemiological variation9. ZIKV is currently detected in Southeast Asia but has not been associated with outbreaks of the same scale as those observed in South and Central America. Our data reveal that, despite the presence of a very efficient vector in Southeast Asia, the Asian ZIKV strains may not be as efficiently transmitted as the American strains.
The vector status of mosquito species depends on their competence to transmit the virus28. Vector competence is determined by the ability of the virus to replicate to high titers throughout the mosquito body and particularly in the salivary glands, thus facilitating transmission during subsequent blood feeding32. Comparing Ae. aegypti, Ae. albopictus and Cx. quinquefasciatus mosquitoes collected in Singapore, we found that salivary glands of Ae. aegypti had the highest infection rate across different ZIKV blood titers. Furthermore, Ae. aegypti mosquitoes were susceptible to infection by ZIKV down to a titer of 103 pfu/ml of blood at 7 days p.i. Previous estimates have quantified that Ae. aegypti blood meal size is on average 2.8 µl33, suggesting that as little as 2.8 infectious particles are enough to infect a mosquito. For comparison, the infection threshold for twelve strains of dengue virus was determined in Ae. aegypti collected in Thailand34; infection rate was not higher than 15% 7 days p.i. after feeding on a blood titer of 103.7 pfu/ml. Our data confirmed the vector status of Ae. aegypti and provide new insight about the ZIKV infectivity to mosquitoes.
Though significantly lower than for Ae. Aegypti, our results have also shown a high rate of salivary gland infection for Ae. albopictus originating from Southeast Asia. Aedes albopictus mosquitoes have been spreading through Brazil for decades35 and its potential contribution to the outbreaks should also be taken into account. ZIKV has been detected in field-caught Ae. albopictus from Gabon36. Additionally, a number of other Aedes species mosquitoes are susceptible to ZIKV; viral isolations were reported from many species of Aedes in Africa37,38,39. For Cx. quinquefasciatus, our data clearly indicate that it is not a competent vector in Singapore.
Flaviviruses have an error-prone replication system that can result in rapid evolution of their genome and generate new strains with variable epidemiological fitness. For instance, variation in transmission efficiency between dengue viruses can result in different epidemiological fitness34, 40. Here, we showed that a strain from Brazil (BE H 815744) had a higher transmissibility than a strain from the Asian lineage (H/PF13); the Brazilian strain had a shorter extrinsic incubation period and higher infection rate of saliva. To identify nucleotide variations that could account for the differences in vector competence, we compared H/PF13 and BE H 815744 genome sequences (Table 1). We first sequenced BE H 815744 from our virus stock to control for mutations acquired during expansion and although we did detect thirty mutations relative to the reference sequence (KU365780), all but two of these were conservative and did not affect protein coding (Table S1). Additionally, we also examined the low-frequency variants present in the viral inoculum and our results indicated that no further positions in the genome had greater than 6% divergence from the published sequence (Table S2)41. Overall, there were 5 nucleotides that differed between H/PF13 and BE H 815744 and changed the amino acid sequence, and 2 of these were fixed in Asian and American strains for which the whole genome was available (Table 1; Table S3). These two segregating substitutions were located in NS4B and 3′UTR. Phylogenetic analysis previously identified an on-going selection pressure on the NS4B gene42, which inhibits the innate immune response43. The 3′UTR of all flaviviruses produces a subgenomic flaviviral RNA (sfRNA) upon partial degradation of their genomic RNA44, 45. SfRNA can alter the innate immune response in both mammals and mosquitoes44, 46 and single mutations in 3′UTR sequence of dengue virus have been linked to increased epidemiological fitness by altering the interferon innate immune response in human47. While additional American and Asian strains need to be tested, we suggest that epidemiological differences between the Americas and Asia are due to variation in mosquito transmission efficiency and we highlighted fixed nucleotide differences between lineages. The ZIKV strains that caused the recent moderate outbreaks in Singapore48, Thailand49 and Cambodia15 all belonged to the Asian lineage. Our study then raises concerns about the introduction of American ZIKV strains in Asia where there are competent mosquito vectors and climatic conditions suitable for transmission.
Material and Methods
Mosquito colonies
The Ae. aegypti, and Ae. albopictus colonies were established in 2010 from eggs collected in Singapore. Culex quinquefasciatus colony were derived from larvae collected by enforcement officers of the National Environment Agency (NEA), Singapore from residential premises during routine house to house inspections in 2010. Eggs were hatched in tap water, larvae were fed a mix of fish food (TetraMin fish flakes) and liver powder (MP Biomedicals), and adults were held in rearing cages (Bioquip) supplemented with 10% sucrose and fed pig’s blood (Primary Industries Pte Ltd) twice weekly. The insectary was held at 28 °C with 50% humidity on a 12:12 h dark:light cycle.
Virus isolates
H/PF13 (H/PF/2013) was collected from human serum in French Polynesia in 2013 and obtained from the European Virus Archive (EVA). The Brazilian strain (BE H 815744) was collected in the Paraiba state (northeast region), Brazil, in 2015 from a febrile non-pregnant woman with a rash. Viruses were propagated in C6/36 cells and used after 6 and 3 passages for H/PF/13 and BE H 815744, respectively. Viruses were titrated trice by plaque assay in BHK-21 cells as previously described50.
Oral infection
Three to five day-old mosquito females were sugar-deprived for 24 h and subsequently offered a blood meal containing a 40% volume of washed erythrocytes from SPF pig’s blood (PWG Genetics), 5% of 100 mM ATP (Thermo Scientific), 5% human serum (Sigma) and 50% volume of virus in RPMI (Gibco). The blood viral titers for both ZIKV strains were confirmed by plaque assays50. Mosquitoes were exposed to the artificial blood meal for one hour using a Hemotek membrane feeder system (Discovery Workshops) with a porcine intestine membrane. Fully engorged females were selected and maintained with free access ad libitum to a 10% sugar solution in an incubation chamber with conditions similar for insect rearing until analysis.
Saliva collection
Orally infected mosquitoes were cold-anesthetized and severed from their wings and legs. Their proboscis was inserted into a 10 µl filter tip containing 10 µl of SPF blood. Mosquitoes were allowed to expectorate for 30 min.
Real-Time quantitative PCR quantification of Zika virus genome copies
Single mosquitoes or total volume of blood used to collect saliva were homogenized in 350 µl of TRK lysis buffer (E.Z.N.A. Total RNA kit I (OMEGA Bio-Tek)) using a bead Mill homogenizer (FastPrep-24, MP Biomedicals) for mosquitoes and by pipetting for blood. Total RNA was extracted using E.Z.N.A. Total RNA kit I (OMEGA Bio-Tek) and eluted in 30 µl of DEPC-treated water. Genomic RNA (gRNA) copies was quantified with a one-step RT-qPCR with Sensifast SYBR No-ROX one-step kit (BioLine). Primers, targeting conserved region in the envelope, were: 5′- AGGACAGGCCTTGACTTTTC-3′ and 5′-TGTTCCAGTGTGGAGTTC-3′. The 10 µl reaction mix contained 400 nM of forward and reverse primers, and 3 µl of RNA extract. Quantification was conducted on a CFX96 Touch Real-Time PCR Detection System (Bio-Rad). Thermal profile was 45 °C for 10 min, 95 °C for 1 min and 40 cycles of 95 °C for 5 sec and 60 °C for 20 sec.
An absolute standard curve was generated by amplifying fragments containing the qPCR targets using the qPCR forward primers tagged with a T7 promoter and the qPCR reverse primer. The fragment was reverse transcribed using MegaScript T7 transcription kit (Ambion) and purified using E.Z.N.A. Total RNA kit I. The total amount of RNA was quantified using a Nanodrop (ThermoScientific) to estimate copy number. Ten times serial dilutions were made and used to generate an absolute standard equation. In each subsequent RT-qPCR plate with samples, we quantified four standard aliquot dilutions to adjust for threshold variation between plates.
Virus Titration
Individual mosquitoes were homogenized in 500 µl of RPMI, filtered through 0.22 µm filter (Sartorius) and titered using plaque assay with BHK-21(ATCC® CCL-10) cells as previously described47.
Virus Sequencing
Total RNA from 500 µl of the virus stocks were extracted using RNAzol RT (MRC). Library preparation was done using NEBNext Ultra Directional RNA Library Prep Kit (NEB) and sequenced on an Illumina MiSeq instrument at the Duke-NUS Genome Biology Facility. Quality control was performed on the paired-end raw reads using FastQC v0.10.151 and trimmed using Trim Galore v0.4.0. These reads were then competitively mapped against all publicly available complete ZIKV genomes from NCBI using BWA v0.7.12-r103952 to identify the best initial reference with the highest number of mapped reads, using SAMtools v0.1.1953, 54 “idxstats” command. The consensus sequence was then generated using “bam2cons.py v0.1” script from the Viral Pipeline Runner (ViPR), with parameter setting “MIN_COV = 0” and the best initial reference.
Virus genome from NCBI
Zika Virus Resource from NCBI was searched for complete genome sequences for Zika virus from Asia and America on 22nd December 2016 (Table S1). We excluded sequences from virus collected in Japan and China as they were potentially imported cases. H/PF13 (H/PF/2013) and BE H 815744 codes were KX369547 and KU365780, respectively.
References
Cao-Lormeau, V. M. et al. Guillain-Barre Syndrome outbreak associated with Zika virus infection in French Polynesia: a case-control study. Lancet 387, 1531–1539, doi:10.1016/S0140-6736(16)00562-6 (2016).
Li, C. et al. Zika Virus Disrupts Neural Progenitor Development and Leads to Microcephaly in Mice. Cell stem cell 19, 120–126, doi:10.1016/j.stem.2016.04.017 (2016).
Garcez, P. P. et al. Zika virus impairs growth in human neurospheres and brain organoids. Science 352, 816–818, doi:10.1126/science.aaf6116 (2016).
Lessler, J. et al. Assessing the global threat from Zika virus. Science 14 (2016).
Hennessey, M., Fischer, M. & Staples, J. E. Zika Virus Spreads to New Areas - Region of the Americas, May 2015-January 2016. MMWR Morb Mortal Wkly Rep 65, 55–58, doi:10.15585/mmwr.mm6503e1 (2016).
Hayes, E. B. Zika virus outside Africa. Emerg. Infect. Dis. 15, 1347–1350, doi:10.3201/eid1509.090442 (2009).
Campos, G. S., Bandeira, A. C. & Sardi, S. I. Zika Virus Outbreak, Bahia, Brazil. Emerging Infectious Disease journal 21, 1885–1886, doi:10.3201/eid2110.150847 (2015).
Zanluca, C. et al. First report of autochthonous transmission of Zika virus in Brazil. Mem. Inst. Oswaldo Cruz 110, 569–572, doi:10.1590/0074-02760150192 (2015).
Faria, N. R. et al. Zika virus in the Americas: Early epidemiological and genetic findings. Science 352, 345–349, doi:10.1126/science.aaf5036 (2016).
Heymann, D. L. et al. Zika virus and microcephaly: why is this situation a PHEIC? Lancet 387, 719–721, doi:10.1016/S0140-6736(16)00320-2 (2016).
Dick, G. W. A. Zika virus (II). Pathogenicity and physical properties. Trans. R. Soc. Trop. Med. Hyg. 46, 521–534, doi:10.1016/0035-9203(52)90043-6 (1952).
Wiwanitkit, V. The current status of Zika virus in Southeast Asia. Epidemiol Health 16 (2016).
Hamel, R. et al. Zika virus: epidemiology, clinical features and host-virus interaction. Microbes Infect. 18, 441–449, doi:http://dx.doi.org/10.1016/j.micinf.2016.03.009 (2016).
Leo, Y. S. & Chow, A. Zika virus has arrived in Singapore. Lancet Infectious Diseases 16, 1317–1319, doi:10.1016/S1473-3099(16)30448-0 (2016).
Duong, V. et al. Low circulation of Zika virus, Cambodia, 2007–2016. Emerg. Infect. Dis. 23, 161432–299, doi:10.3201/eid2302.161432 (2017).
Ye, Q. et al. Genomic characterization and phylogenetic analysis of Zika virus circulating in the Americas. Infect. Genet. Evol. 43, 43–49, doi:10.1016/j.meegid.2016.05.004 (2016).
Kochakarn, T. et al. Comparative genome analysis between Southeast Asian and South American Zika viruses. Asian Pac J Trop Med 9, 1048–1054, doi:10.1016/j.apjtm.2016.10.002 (2016).
Wong, P.-S. J., Li, M.-zI., Chong, C.-S., Ng, L.-C. & Tan, C.-H. Aedes (Stegomyia) albopictus (Skuse): A potential vector of Zika virus in Singapore. PLoS Negl. Trop. Dis. 7, e2348, doi:10.1371/journal.pntd.0002348 (2013).
Chouin-Carneiro, T. et al. Differential susceptibilities of Aedes aegypti and Aedes albopictus from the Americas to Zika virus. PLoS Negl Trop Dis 10, e0004543, doi:10.1371/journal.pntd.0004543 (2016).
Li, M. I., Wong, P. S. J., Ng, L. C. & Tan, C. H. Oral susceptibility of Singapore Aedes (Stegomyia) aegypti (Linnaeus) to Zika virus. PLoS Negl Trop Dis 6, e1792, doi:10.1371/journal.pntd.0001792 (2012).
Guo, X. X. et al. Culex pipiens quinquefasciatus: a potential vector to transmit Zika virus. Emerg Microbes Infect 5, 102, doi:10.1038/emi.2016.102 (2016).
Fernandes, R. S. et al. Culex quinquefasciatus from Rio de Janeiro is not competent to transmit the local Zika virus. PLoS Negl Trop Dis 10 (2016).
Huang, Y. S. et al. Culex Species Mosquitoes and Zika Virus. Vector Borne Zoonotic Diseases 24, 24 (2016).
Haddow, A. D. et al. Genetic characterization of Spondweni and Zika viruses and susceptibility of geographically distinct strains of Aedes aegypti, Aedes albopictus and Culex quinquefasciatus (Diptera: Culicidae) to Spondweni virus. PLoS Negl Trop Dis 10, e0005083, doi:10.1371/journal.pntd.0005083 (2016).
Goncalves, C. M. et al. Distinct variation in vector competence among nine field populations of Aedes aegypti from a Brazilian dengue-endemic risk city. Parasit Vectors 7, 1756–3305, doi:10.1186/1756-3305-7-320 (2014).
Gubler, D. J., Nalim, S., Tan, R., Saipan, H. & Suliantisaroso, J. Variation in susceptibility to oral infection with dengue viruses among geographic strains of Aedes aegypti. Am. J. Trop. Med. Hyg. 28, 1045–1052 (1979).
Fansiri, T. et al. Genetic mapping of specific interactions between Aedes aegypti mosquitoes and dengue viruses. Plos Genetics 9, e1003621, doi:10.1371/journal.pgen.1003621 (2013).
Conway, M. J., Colpitts, T. M. & Fikrig, E. Role of the vector in arbovirus transmission. Annual Review of Virology 1, 71–88, doi:10.1146/annurev-virology-031413-085513 (2014).
Tang, W. W. et al. A mouse model of Zika virus sxual transmission and vaginal viral replication. Cell Rep 17, 3091–3098, doi:10.1016/j.celrep.2016.11.070 (2016).
Foy, B. D. et al. Probable non-vector-borne transmission of Zika virus, Colorado, USA. Emerg. Infect. Dis. 17, 880–882, doi:10.3201/eid1705.101939 (2011).
Besnard, M., Lastere, S., Teissier, A., Cao-Lormeau, V. & Musso, D. Evidence of perinatal transmission of Zika virus, French Polynesia, December 2013 and February 2014. Euro Surveill 19, 20751, doi:10.2807/1560-7917.ES2014.19.13.20751 (2014).
Kramer, L. D. & Ebel, G. D. In Adv. Virus Res. Vol. 60 187–232 (Academic Press, 2003).
Knaus, R. M., Foil, L. D., Issel, C. J. & Leprince, D. J. Insect blood meal studies using radiosodium 24Na and 22Na. J. Am. Mosq. Control Assoc. 9, 264–268 (1993).
Pongsiri, A. et al. Differential susceptibility of two field Aedes aegypti populations to a low infectious dose of dengue virus. PLoS ONE 9, e92971, doi:10.1371/journal.pone.0092971 (2014).
Kraemer, M. U. et al. The global distribution of the arbovirus vectors Aedes aegypti and Ae. albopictus. eLife 4, 10.7554/eLife.08347 (2015).
Grard, G. et al. Zika virus in Gabon (Central Africa) – 2007: A new threat from Aedes albopictus? PLoS Negl Trop Dis 8, e2681, doi:10.1371/journal.pntd.0002681 (2014).
Diallo, D. et al. Zika virus emergence in mosquitoes in southeastern Senegal, 2011. PLoS ONE 9, e109442, doi:10.1371/journal.pone.0109442 (2014).
Dick, G. W., Kitchen, S. F. & Haddow, A. J. Zika virus. I. Isolations and serological specificity. Trans. R. Soc. Trop. Med. Hyg. 46, 509–520, doi:10.1016/0035-9203(52)90042-4 (1952).
Diagne, C. T. et al. Potential of selected Senegalese Aedes spp. mosquitoes (Diptera: Culicidae) to transmit Zika virus. BMC Infect Dis 15, 015–1231, doi:10.1186/s12879-015-1231-2 (2015).
Hanley, K., Nelson, J., Schirtzinger, E., Whitehead, S. & Hanson, C. Superior infectivity for mosquito vectors contributes to competitive displacement among strains of dengue virus. BMC Ecol. 8, 1, doi:10.1186/1472-6785-8-1 (2008).
Wilm, A. et al. LoFreq: a sequence-quality aware, ultra-sensitive variant caller for uncovering cell-population heterogeneity from high-throughput sequencing datasets. Nucleic Acids Res. 40, 11189–11201, doi:10.1093/nar/gks918 (2012).
Sironi, M., Forni, D., Clerici, M. & Cagliani, R. Nonstructural proteins are preferential positive selection targets in Zika virus and related flaviviruses. PLoS Negl. Trop. Dis. 10, e0004978, doi:10.1371/journal.pntd.0004978 (2016).
Muñoz-Jordán, J. L. et al. Inhibition of alpha/beta interferon signaling by the NS4B protein of faviviruses. J. Virol. 79, 8004–8013, doi:10.1128/jvi.79.13.8004-8013.2005 (2005).
Schnettler, E. et al. Noncoding flavivirus RNA displays RNA interference suppressor activity in insect and mammalian cells. J. Virol. 86, 13486–13500, doi:10.1128/jvi.01104-12 (2012).
Chapman, E. G. et al. The structural basis of pathogenic subgenomic flavivirus RNA (sfRNA) production. Science 344, 307–310, doi:10.1126/science.1250897 (2014).
Akiyama, B. M. et al. Zika virus produces noncoding RNAs using a multi-pseudoknot structure that confounds a cellular exonuclease. Science 354, 1148–1152, doi:10.1126/science.aah3963 (2016).
Manokaran, G. et al. Dengue subgenomic RNA binds TRIM25 to inhibit interferon expression for epidemiological fitness. Science 350, 217–221, doi:10.1126/science.aab3369 (2015).
Maurer-Stroh, S. et al. South-east Asian Zika virus strain linked to cluster of cases in Singapore, August 2016. Euro Surveill 21, 1560–7917, doi:10.2807/1560-7917.ES.2016.21.38.30347 (2016).
Buathong, R. et al. Detection of Zika virus infection in Thailand, 2012–2014. Am. J. Trop. Med. Hyg. 93, 380–383, doi:10.4269/ajtmh.15-0022 (2015).
Manokaran, G. et al. Dengue subgenomic RNA binds TRIM25 to inhibit interferon expression for epidemiological fitness. Science (New York, N.Y.) 350, 217–221, doi:10.1126/science.aab3369 (2015).
Andrews, S. FQC: A quality control tool for high throughput sequence data. Reference Source (2010).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595, doi:10.1093/bioinformatics/btp698 (2010).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079, doi:10.1093/bioinformatics/btp352 (2009).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993, doi:10.1093/bioinformatics/btr509 (2011).
Acknowledgements
We are grateful to Duane J. Gubler for helping to gain access to the Brazilian strain of Zika virus. We are also in depth to Mariano Garcia-Blanco for constant support. We thank EVA for sharing its stock of H/PF13. Support for this research came from a grant from the National Research Foundation Singapore under its NIG scheme (NMRC/BNIG/2032/2015) awarded to J.P. and from the Duke-NUS Signature Research Programme funded by the Agency for Science, Technology and Research (A*STAR), Singapore, and the Ministry of Health, Singapore. RMV was partially supported by IRD through JEAI DENCHICTHAI. PFCV was partially supported by the Brazilian Ministry of Health, IEC and grants received from CAPES (Zika fast track grant; Coordenação de Apoio aos Profissionais de Nível Superior do Ministério da Educação – Coordination of Professionals of High Level Degree of Ministry of Education); CNPq (grants: 301641/2010-2,303999/2016-0, 439971/2016-0 and 440405/2016-5; Conselho Nacional para o Desenvolvimento Científico e Tecnológico - the Brazilian Agency for Scientific and Technological Deveelopment), and FINEP (Zika call; Financiadora de Estudos e Projetos do Ministério da Ciência, Tecnologia e Inovação– Agency for Financial support of Studies and Projects of the Ministry of Science, Technology and Innovation).
Author information
Authors and Affiliations
Contributions
J.P., R.M.V. and D.M. designed the experiments. R.M.V., M.M. and T.V. performed the experiments. J.P. analyzed the results. O.M.S. and J.H.T. performed the sequencing. C.H.T. and L.C.N. provided the Culex quinquefasciatus colony. P.C.V. collected the Brazilian ZIKV strain. J.P. and D.M. wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pompon, J., Morales-Vargas, R., Manuel, M. et al. A Zika virus from America is more efficiently transmitted than an Asian virus by Aedes aegypti mosquitoes from Asia. Sci Rep 7, 1215 (2017). https://doi.org/10.1038/s41598-017-01282-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-01282-6
This article is cited by
-
Residence in infected neighborhoods and fertility decline during the Zika epidemic in Singapore
Population and Environment (2022)
-
Evaluation on the effects of different plant matter in water on Aedes aegypti development (Diptera: Culicidae)
International Journal of Tropical Insect Science (2022)
-
Recent African strains of Zika virus display higher transmissibility and fetal pathogenicity than Asian strains
Nature Communications (2021)
-
Assessing the role of two populations of Aedes japonicus japonicus for Zika virus transmission under a constant and a fluctuating temperature regime
Parasites & Vectors (2020)
-
Experimental adaptation of dengue virus 1 to Aedes albopictus mosquitoes by in vivo selection
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.