Abstract
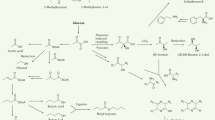
The ability of bacteria and fungi to communicate with each other is a remarkable aspect of the microbial world. It is recognized that volatile organic compounds (VOCs) act as communication signals, however the molecular responses by bacteria to fungal VOCs remain unknown. Here we perform transcriptomics and proteomics analyses of Serratia plymuthica PRI-2C exposed to VOCs emitted by the fungal pathogen Fusarium culmorum. We find that the bacterium responds to fungal VOCs with changes in gene and protein expression related to motility, signal transduction, energy metabolism, cell envelope biogenesis, and secondary metabolite production. Metabolomic analysis of the bacterium exposed to the fungal VOCs, gene cluster comparison, and heterologous co-expression of a terpene synthase and a methyltransferase revealed the production of the unusual terpene sodorifen in response to fungal VOCs. These results strongly suggest that VOCs are not only a metabolic waste but important compounds in the long-distance communication between fungi and bacteria.
Similar content being viewed by others
Introduction
Interactions and communication among organisms are central to understanding any ecosystem. The essential role of volatile organic compounds (VOCs) in the communication with other organisms, also known as infochemicals, has been acknowledged for more than 30 years1. However, their ecological functions have been mainly studied for aboveground plant-plant and plant-insect interactions2, 3. In recent years, VOCs are becoming increasingly important in the field of microbial ecology. Due to their unique nature (low molecular mass, high vapor pressure, low boiling point and a lipophilic moiety) VOCs play important roles in the long-distance interactions and communication within the microbial world4. In soil and in the rhizosphere, VOCs readily diffuse under atmospheric pressure and travel throughout the abundant air- and liquid-filled pockets of the soil4, 5.
Several soil-associated bacteria were shown to have positive effects on plant growth and resistance6,7,8, as well as to have the ability to control plant diseases via production of VOCs9. Various studies have documented that VOCs can have diverse roles in the interaction between physically separated microorganisms, ranging from infochemical molecules affecting the behavior, population dynamics and gene expression in responding microorganisms to interference competition tools suppressing or eliminating potential enemies4, 10,11,12,13. Interestingly, most studies have only examined the role of bacterial VOCs and their effect on plants and fungi. However, the role and function of fungal VOCs on bacteria remains largely unknown. Only few studies demonstrated that the growth of some bacterial species was suppressed by fungal VOCs14. For example, VOCs produced by the oyster mushroom Pleurotus ostreatus showed inhibitory effects on Bacillus cereus and Bacillus subtilis 15. Another study by Lutz et al.16 demonstrated that VOCs emitted by T. atroviride increased the expression of a biocontrol gene (phlA) of P. fluorescens.
Previously, we investigated the effect of fungal VOCs on the behavior of phylogenetically different soil bacteria17. In these experiments we showed that VOCs emitted by several fungi lead to phenotypical responses in bacteria, for example, by inducing a change in bacterial motility17. We observed that the plant pathogenic fungus Fusarium culmorum produced a unique cluster of VOCs, consisting primarily of terpenes. When exposed to the VOCs emitted by this fungus, the rhizobacterium Serratia plymuthica PRI-2C responded with an induction of motility.
It is plausible that in soil, microorganisms sense changes in their environments via shifts in VOCs blends and adapt their behavior accordingly11. Although several studies indicated that VOCs can be used as signaling molecules in microbial inter-species interactions, the following questions remain unanswered: how are VOCs perceived as signals by the interacting microorganism and which regulatory genes and pathways are involved in the response?
To answer these questions, the rhizosphere isolate S. plymuthica PRI-2C was grown exposed or unexposed to VOCs emitted by F. culmorum and the bacterial transcriptome and proteome were analyzed under each situation to identify the molecular basis of the bacterial response to fungal VOCs. Further, metabolomics analysis and heterologous expression studies were performed to confirm the production of an unusual terpene compound involved in this bacterial-fungal conversation.
Methods
Strains and growth conditions
Based on previous work17 S. plymuthica PRI-2C and F. culmorum PV were selected for this study. The bacterial strain S. plymuthica PRI-2C was isolated from maize rhizosphere in the Netherlands18, pre-cultured from frozen glycerol stocks on 0.1 Tryptic Soy Broth plates (TSB)19 and grown for 3 days at 20 °C before use. The fungal strain F. culmorum PV was isolated from a sandy dune soil in the Netherlands20, pre-cultured on 0.5 Potato Dextrose Agar plates (PDA)21 and incubated for 6 days at 20 °C before use.
S. plymuthica PRI-2C genome sequencing
Genomic DNA was extracted from S. plymuthica PRI-2C cells using the Qiagen MagAttract HMW DNA kit (Qiagen) and subjected to PacBio RS II sequencing at the Institute for Genome Sciences (IGS), Baltimore, Maryland, USA. PacBio RS II sequences were obtained from 1 SMRT cell. Sequences were filtered using SMRT Analysis server v2.3.0 with default settings and assembled using the RS_HGAP Assembly.3 (HGAP3) protocol, followed by a final Quiver correction using the RS_Resequencing protocol. The Illumina reads obtained from RNA-sequencing were filtered using Fastq MCF22 with default settings and aligned against the contigs using BWA V0.7.1223. The aligned reads were re-aligned with GATK V3.5.024 to ensure correct indel calls. Contigs were corrected using the re-aligned reads with Pilon25. Singular contigs were checked with a custom script and overlapping ends were trimmed. The final circular contig was rearranged to start at the dnaA gene in the forward direction. The sequences were annotated using Prokka 1.1126. The accession number of the genome after submission to NCBI is CP015613.
Interaction experiment between S. plymuthica PRI-2C and F. culmorum PV
To investigate the response of S. plymuthica PRI-2C to volatiles emitted by F. culmorum PV, a plate-within-a-plate system was used (Fig. 1a). F. culmorum PV plugs (6 mm in diameter) were inoculated on a 0.5 PDA Petri dish (3.5 cm) that was placed into the partitioned Petri dish (9 cm) and incubated for 3 days at 20 °C. On day 3 S. plymuthica PRI-2C inoculum, consisting of 50 μl washed cell suspension in a 10 mM phosphate buffer (pH 6.5) and containing 107 cells/ml, was spread on 1.5% water agar supplied with artificial root exudates (WA + ARE)17 of the partitioned Petri dish. As control S. plymuthica PRI-2C was exposed to 0.5 PDA medium only. Plates were incubated at 20 °C. Total RNA and cytosolic proteins were extracted after 48 h (time point 1) and 72 h (time point 2). All experiments were performed in triplicates.
Overview of experimental setup and fungal VOC effect on bacterial growth. Schematic overview of the plate-within-a-plate system (a). Growth of S. plymuthica PRI-2C exposed to fungal VOCs after 48 h (t1) and 72 h (t2) (b). Controls S. plymuthica PRI-2C without exposure to VOCs after 48 h (C t1) and 72 h (C t2). Error bars represent standard deviations between replicates (n = 3). Asterisk indicates statistical significance (p < 0.05) of each time point compared to the respective control.
To test the effect of fungal VOCs on the growth of S. plymuthica PRI-2C, the CFU/ml were determined after 48 h and 72 h (Fig. 1b). After scratching off the cells from the surface dilution series of each treatment were prepared in 10 mM phosphate buffer (pH 6.5). A volume of 100 µl of each serial dilution was plated on 1/10th TSB plates in duplicates. The plates were incubated for three days at 20 °C. Bacterial enumeration was carried out on an aCOlyte Colony Counter (Meintrup DWS Laborgeräte GmbH). A two-tailed tailed Student’s t-test (p < 0.05) was used to evaluate the differences between two groups using SPSS (SPSS, Chicago, IL, USA). Results are shown as mean ± S.D. Sample number (n) represents biological replicates.
RNA Extraction
S. plymuthica PRI-2C cells were suspended in 2 ml of sterile phosphate buffer and transferred to a double volume of RNAprotect Bacteria Reagent (Qiagen). The mixture was centrifuged at 14,000 rpm for 1.5 min at 4 °C. The supernatant was discarded and the cell pellets were directly frozen in liquid N2 and stored at −80 °C until RNA extraction. RNA was extracted using the Aurum Total RNA Mini Kit (Bio-Rad), following the manufacturer’s recommendations. Contaminating DNA was removed using the TURBO DNA-free Kit (Ambion). RNA concentrations and quality were measured on a NanoDrop Spectrophotometer (Isogen Life Science) and on a Fragmentanalyzer RNA 6000 Pico Chip (Agilent).
RNA sequencing and differential expression analysis
RNA sequencing was performed at Erasmus Center for Biomics, The Netherlands. RNA was depleted of rRNA using the Ribo-Zero rRNA Removal Bacteria Kit (Illumina). A sequencing library was constructed using the KAPA Stranded RNA-Seq Kit (Kapa Biosystems), and sequenced according to the Illumina TruSeq V3 protocol on the HiSeq2000 with a single read 50 bp and 7 bp index.
The obtained Illumina reads from the RNA-sequencing were filtered using Fastq MCF and aligned against the cDNA sequences of S. plymythica PRI-2C using Bowtie 2 (2.2.5)27 with the following settings: –no-mixed –no-discordant –gbar 1000 –end-to-end. Transcript abundance was estimated using RSEM V1.1.2628 and differential expression between the treatments was analyzed using edgeR V3.229, 30. Differentially expressed genes (DEGs) were defined with a cutoff at log fold change of greater than 0.585 or less than −0.585, an FDR of less than 0.05. Data were filtered with a p-value of 0.001 (Table S2). Venn diagrams were produced using Venny31. Functional prediction and assignment of genes to TIGRFAMs32 was accomplished by the in-house developed analysis pipeline ‘Prophane 2.0′ (http://www.prophane.de; Schneider et al.33. Voronoi treemaps were generated using Paver (Decodon, Greifswald, Germany, http://www.decodon.com/).
Validation of RNA sequencing data
Validation of RNA sequencing data was performed by Quantitative Real-Time PCR (qRT-PCR) analysis of a subset of differentially expressed genes related to terpene production, motility and a gene encoding for isocitrate lyase. Two housekeeping genes, rpoB and gyrB, were used as internal standards (Table 1). All experiments were performed in triplicates.
First strand cDNA was synthesized from 2 μg previously extracted total RNA with random hexamer primers using the SuperScript VILO cDNA Synthesis Kit (Invitrogen) according to the manufacturer’s protocol. A volume of 4 μl cDNA of each treatment was subjected to qRT-PCR using QuantiNova SYBR Green I PCR master mix (Qiagen) a Qiagen Research Rotor- Gene Q thermal cycler (Qiagen) with the following conditions: initial cycle 95 °C for 2 min, followed by 40 cycles of 95 °C for 10 sec, and 60 °C for 30 sec. Standard curves were established for each cDNA sample to calculate the expression values (Ct-value). Log fold changes in gene expression were determined using the ΔΔCt method34, data were normalized against the two housekeeping genes (rpoB and gyrB), and compared to log fold changes obtained from RNA sequencing data (Figure S1).
Protein extraction and MS sample preparation
S. plymuthica PRI-2C cells were suspended in 2 ml of sterile phosphate buffer and centrifuged at 14,000 rpm for 1.5 min at 4 °C. Cytosolic proteins were extracted using a bead-beating protocol. Fractions were transferred into 2 ml Eppendorf tubes with 0.25 ml of 0.1 mm glass beads (BioSpec Products) and placed into FastPrep-24 lyser tubes (MP Biomedicals). Eight rounds of 30 s lysis at 6.5 m s−1 followed by 5 min incubation on ice were used to lyse the cells. Afterwards, cell debris was removed by centrifugation; determination of cytosolic protein concentration was done using Roti®-Nanoquant (Roth). Reduction of disulfide bonds with TCEP, alkylation of thiolgroups with iodoacetamide and subsequent in-solution digestion of proteins using trypsin was performed as described by Muntel et al.35. Desalting of peptides prior to mass spectrometry analysis using Stage tips was done according to the protocol by Rappsilber et al.36.
Setup for label-free protein quantification (LC-IMSE) and data analysis
The nanoACQUITYTM UPLCTM system (Waters) was used to separate and to introduce peptides into the Synapt G2 (Waters) mass spectrometer. Parameters for liquid chromatography and IMSE (MSE with ion mobility separation) were used as described previously37, 38.
LC-IMSE data were processed using PLGS v3.0.1 Processing parameters were set as follows: Chromatographic peak width and MS TOF resolution were set to automatic, lock mass charge 2 set to 785.8426 Da/e with a lock mass window of 0.25 Da, low energy threshold 200.0 counts, elevated energy threshold 20.0 counts, intensity threshold 750 counts. The data were searched against a randomized S. plymuthica PRI-2C database (version February 2016) with added laboratory contaminants and yeast ADH1 sequence (10,090 entries). For positive protein identification the following criteria had to be met: 1 fragment ion matched per peptide, 5 fragment ions matched per protein, 1 peptide matched per protein; 2 missed cleavages allowed, primary digest reagent: trypsin, fixed modification: carbamidomethylation C (+57.0215), variable modifications: deamidation N, Q (+0.9840), oxidation M (+15.9949), pyrrolidonecarboxylacid N-TERM (−27.9949). The protein false discovery rate (FDR) was set to 5%. For the final analysis only identifications based on at least two peptides were considered. In total 9 MS-runs per time point were conducted (3 biological replicates with 3 technical replicates each). For positive identification and reliable quantification, a replicate filter of 4/9 was applied. This reduced the false discovery rate per sample to less than 0.7%. Absolute protein quantification of identified proteins was performed using top3 peptide intensity with spiked-in yeast alcohol dehydrogenase (Waters; concentration of 50 fmol/µl sample) as a reference (Hi3 approach, Silva et al.39). Data were corrected for detector saturation effects by implementing a correction factor as recently described38. To allow comparison between samples absolute protein amounts calculated by PLGS (given in fmol) were normalized (fmol/ng total protein).
Statistical analysis was done using MeV v4.8.140 for proteins that were present in at least four out of nine replicates. Hierarchical clustering and Student’s t-test were performed with the following settings: unequal group variances were assumed (Welch approximation), p-values based on all permutation with p = 0.01, significance determined by adjusted Bonferroni correction. Only proteins showing at least 1.5 fold changes in addition to statistical significance were considered for further analysis. So-called ‘off/on’ proteins needed to be detected in at least four replicates of one treatment and absent from all replicates of another treatment.
Functional prediction and assignment of proteins to TIGRFAMs32, respectively, was accomplished by the in-house developed analysis pipeline ‘Prophane 2.0’ (http://www.prophane.de; Schneider et al.33. Voronoi treemaps were generated using Paver (Decodon, Greifswald, Germany, http://www.decodon.com/). The mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository41 with the dataset identifier PXD004819.
Volatile trapping
For the collection of volatiles produced during the interaction, partitioned glass Petri dishes with lids connected to a steel trap containing 150 mg Tenax TA and 150 mg Carbopack B (Markes International Ltd., Llantrisant, UK) were used42. F. culmorum PV was grown on 0.5 PDA at 20 °C for 3 days. On day 3 S. plymuthica PRI-2C inoculum, consisting of 50 μl washed cell suspension in a 10 mM phosphate buffer (pH 6.5) and containing 107 cells/ml, was spread on 1.5% WA + ARE. As control S. plymuthica PRI-2C was exposed to 0.5 PDA medium only. Plates were incubated for 48 h and 72 h at 20 °C. Volatiles were collected during 48 h and 72 h of incubation at 20 °C. All experiments were performed in triplicates. Traps were removed, capped and stored at 4 °C until analysis using GC-Q-TOF.
The trapped VOCs were desorbed from the traps using an automated thermo desorption unit (Unity TD-100, Markes International Ltd., Llantrisant, UK) at 210 °C for 12 min (He flow 50 ml/min) and trapped on a cold trap at −10 °C. The trapped volatiles were introduced into the GC-Q-TOF (model Agilent 7890B GC and the Agilent 7200 A QTOF, Santa Clara, USA) by heating the cold trap for 3 min to 280 °C with split ratio set to 1:20. The column used was a 30 mm × 0.25 mm ID RXI-5MS, film thickness 0.25 μm (Restek 13424-6850, Bellefonte, PA, USA). VOCs were detected by the MS operating at 70 eV in EI mode. Mass spectra were acquired in full scan mode (30–400 amu, 4 scans/s). MassHunter Qualitative Analysis Software V B.06.00 Build 6.0.633.0 (Agilent Technologies, Santa Clara, CA, USA) was used to control the instrument and for data acquisition and analysis. Mass-spectra were extracted with MassHunter Qualitative Analysis Software V B.06.00 Build 6.0.633.0 (Agilent Technologies, Santa Clara, USA) and compared with the data of the NIST 2014 V2.20 library and the spectrum of sodorifen published in ref. 43.
Production of sodorifen in Escherichia coli
PCR-generated DNA encompassing the complete coding sequence of the sodorifen synthase (SpSS, Q5A_011535) and the methyltransferase (SpMT, Q5A_011540) of S. plymuthica PRI-2C were inserted into MCS1 (BamHI, NotI) and MCS2 (NdeI, KpnI) cloning sites of the expression vector pACYCDuet-1 (Novagen, CmR), respectively. The insertion was confirmed by colony PCR and sequencing. The constructed plasmids, pACYCDuet SpSS, pACYCDuet SpMT, pACYCDuet SpSS/SpMT and pACYCDuet empty, were transformed into chemically competent E. coli BL21 DE3 carrying the plasmid pMEV, which is a kanamycin-resistant version of the mevalonate pathway-expressing plasmid i-pBbA5c-MevT(CO)-MBIS(CO)-NptII44 (Addgene, Peralta-Yahya et al.45), which was kindly supplied by Thamara Hendricks. Transformants were plated on LB plates supplemented with kanamycin (50 µg/ml), chloramphenicol (50 µg/ml) and 1% glucose. A starter culture of all 4 clones was grown overnight in 5 ml LB medium supplemented with kanamycin (50 µg/ml), chloramphenicol (50 µg/ml) and 1% glucose at 37 °C, 250 rpm. 500 µl of the starter culture was inoculated into 50 ml LB supplemented with kanamycin (50 µg/ml) and chloramphenicol (50 µg/ml) in 250 ml Erlenmeyer flasks. The culture was incubated at 37 °C, 250 rpm until the OD600 reached 0.4. At this point 100 µl of each culture were spread on a glass Petri dish with 20 ml LB agar supplemented with kanamycin (50 µg/ml), chloramphenicol (50 µg/ml) and 0.1 M IPTG. Steel trap containing 150 mg Tenax TA and 150 mg Carbopack B were added into the lids and the plates were incubated at 25 °C for 48 h. All experiments were performed in duplicates. Traps were removed, capped and stored at 4 °C until analysis by using GC-Q-TOF. The traps were measured as described in the previous section with a split ratio set to 1:80. Mass-spectra were extracted with MassHunter Qualitative Analysis Software V B.06.00 Build 6.0.633.0 (Agilent Technologies, Santa Clara, USA) and compared with the data of the NIST 2014 V2.20 library and the spectrum of sodorifen published in von Reuss et al.43 and Domik et al.46.
Results
Genomic features of S. plymuthica PRI-2C
The complete genome of S. plymuthica PRI-2C was sequenced to facilitate the interpretation of proteomics and transcriptomics data and to gain better insights into the bacterial perception and response to fungal VOCs. The complete genome of S. plymuthica PRI-2C is 5,464,425 bp in size. Annotation of the genome predicted 5,002 protein-coding genes, 88 tRNAs and 22 rRNAs (Accession number CP015613).
S. plymuthica PRI-2C harbors several gene clusters encoding for secondary metabolites. Based on in silico prediction using antiSMASH a total of 36 secondary metabolite gene clusters were identified. One of those gene clusters belonged to the class of terpenes, one to the class of arylpolyene-siderophore, one to the class of butyrolactones, one to nonribosomal peptide synthetase (nrps) -Type I polyketide synthase (t1pks), one to homoserine lactone (hserlactones), one to others, two to nonribosomal peptide synthetase (nrps). Out of the remaining 28 gene clusters, four were identified as putative fatty acids, four as putative saccharides and twenty as non-identified putative gene clusters.
Important for microbial interactions and communication are signal transduction systems that enable bacteria to detect and respond to changes in the environment (Stock et al.47). We investigated the genome for signal transduction systems using the MIST2 database48. The genome of S. plymuthica PRI-2C encodes 428 one-component systems (1CSs) and 69 two-component systems (TCSs). Two extracytoplasmic function (ECF) sigma factors were found, which comprise the largest group among the σ70 family49. Additionally, 8 genes involved in chemotaxis systems were found (Table S1).
Effect of fungal VOCs on the growth of S. plymuthica PRI-2C
After 48 h of exposure to fungal VOCs the cell density of S. plymuthica PRI-2C was 4.13 × 109 CFU/ml and after 72 h of exposure 4.63 × 109 CFU/ml. We observed a significant increase in the growth of S. plymuthica PRI-2C exposed to VOCs after both time points as compared to S. plymuthica PRI-2C without exposure to VOCs (Fig. 1b).
Global changes in the transcriptome and proteome of S. plymuthica PRI-2C in response to fungal VOCs
To study the response of S. plymuthica PRI-2C to fungal VOCs, we investigated changes at the transcriptome and proteome level after 48 h (t1) and 72 h (t2) of exposure to F. culmorum VOCs. After 48 h of exposure to fungal VOCs 204 differentially expressed genes (DEGs) were identified (199 up- and 5 down-regulated), while after 72 h of exposure only 6 DEGs were detected (5 up- and 1 down-regulated) (Fig. 2a, Table S2). Only 2 DEGs were unique for t2 and 4 DEGs were expressed at both times points (Fig. 2b). These 4 genes were Q5A_006040 (coding for the Pyrimidine-specific ribonucleoside hydrolase RihA), Q5A_018030 (coding for the putative HTH-type transcriptional regulator YahB), Q5A_003275 (coding for the UDP-N-acetylmuramoyl-L-alanyl-D-glutamate–2,6-diaminopimelate ligase) and Q5A_011375 (coding for a hypothetical protein). Most of the DEGs at t1 were assigned to functions involved in energy metabolism, transport and binding proteins, regulatory functions, cell envelope, cellular processes, protein synthesis and fate as well as unknown function. For t2 DEGs were assigned to functions involved in DNA metabolism, regulatory functions, cell envelope and unknown function (Fig. 2c, Table S2).
Changes in transcriptome of S. plymuthica PRI-2C in response to fungal VOCs. Heatmap of 206 DEGs with at least +/−1.5-fold changes and a p-value less than 0.001 as a cut-off (a). Hierarchical clustering was performed using the Pearson Correlation metric. Venn diagram of uniquely and commonly expressed DEGs at t1 and t2 (b). Voronoi treemaps visualizing changes in the transcriptome of S. plymuthica PRI-2C exposed to fungal VOCs at t1 (C left) and at t2 (C right) (c). Functional classification of genes was carried out by Prophane 2.0 and is based on TIGRFAMS. Red color indicates higher expression of the respective gene and blue color indicates lower expression as compared to the control. Each cell represents a single gene; functional classes are separated by thicker black lines. Numbers indicate the following classes: 1 - Fatty acid and phospholipid metabolism, 2 - Biosynthesis of cofactors, prosthetic groups, and carriers, 3 - Purines, pyrimidines, nucleosides, and nucleotides, 4 - Amino acid biosynthesis, 5 - Central intermediary metabolism, 6 - Mobile and extrachromosomal element functions, 7 - Transcription, 8 - DNA metabolism, 9 - Signal transduction.
On the proteome level absolute amounts for a total of 1,574 proteins were obtained by applying the LC-MSE approach39 (Table S3). A large number of these proteins were assigned to functions associated with metabolic processes (584 proteins; e.g. biosynthesis of amino acids and cofactors; fatty acid, energy and central intermediary metabolism) and genetic information processing (311 proteins; e.g. protein synthesis and protein fate, DNA metabolism and transcription). Moreover, a high number of so far uncharacterized or unclassified proteins (448 proteins) were detected (Fig. 3, Table S3).
Voronoi treemaps visualizing changes in the proteome composition of S. plymuthica PRI-2C exposed to fungal VOCs. Quantitative expression values of S. plymuthica PRI-2C proteins at t1 of exposure (a) and at t2 (b). Functional classification of proteins was carried out by Prophane 2.0 and is based on TIGRFAMS. Red color indicates higher expression of the respective protein and blue color indicates lower expression as compared to the control. Each cell represents a single protein. Functional classes are separated by thicker black lines. Numbers indicate the following classes: 1 - Fatty acid and phospholipid metabolism, 2 - Biosynthesis of cofactors, prosthetic groups, and carriers, 3 - Purines, pyrimidines, nucleosides, and nucleotides, 4 - Central intermediary metabolism, 5 - Transcription, 6 - Mobile and extrachromosomal element functions, 7 - Regulatory functions, 8 - Signal transduction.
Longer exposure (72 h) to fungal VOCs led to higher number of proteins with a significantly changed amount as compared to the control. Overall 169 proteins were differentially expressed (significance determined by t-test and expression change of more than 1.5 fold) when exposed to fungal VOCs as compared to the non-exposed control (Table S3). At t1 51 differentially expressed proteins (DEPs) were identified (28 up- and 23 down-regulated) while at t2 124 DEPs were identified (78 up- and 46 down-regulated). For both time points mainly proteins involved in metabolic processes (amino acid biosynthesis, energy and central intermediary metabolism, biosynthesis of cofactors), signal transduction, transport and binding proteins and several uncharacterized proteins were found to be differentially expressed (Fig. 4, Table S3).
Impact of fungal volatiles on S. plymuthica PRI-2C protein content. The percentage of proteins with at least 1.5 fold change in relation to the number of identified proteins is depicted. Black bars indicate proteins with changed amount in both, early (t1, 48 h) and late (t2, 72 h) sampling points, grey bars indicate changed amounts on late sampling point, white bars indicate changed amount in early sampling point. Functional annotation is based on TIGRFAMS.
Functional gene and protein categories affected by fungal VOCs
Differentially expressed genes and proteins related to chemotaxis and motility
The expression of one gene, Q5A_015275 (coding for the flagellar protein FliS) relating to chemotaxis and motility was up-regulated at t1 when S. plymuthica PRI-2C was exposed to fungal VOCs (Table S2). One protein, Q5A_015505 (Methyl-accepting chemotaxis protein IV) was identified at t1 (Table S3) and is known to act as a sensor detecting attractants and promoting bacterial movement towards suitable sites for colonization50.
Differentially expressed genes and proteins related to signal transduction
29 genes related to signal transduction were up-regulated in our transcriptomics dataset (Table S2). Due to the high number we focus here on genes coding for proteins related to small molecule interactions, protein interactions and two-component systems. Cyclic di-GMP phosphodiesterase YhjH (Q5A_007585) is known to be involved in regulating the levels of c-di-GMP that control cell motility (the flagella) and adhesion (adhesive curli fimbriae)51. Phytochrome-like protein Cph2 (Q5A_015970) acts as a photoreceptor that perceives light signals52. The putative diguanylate cyclase AdrA (Q5A_005955) is an integral component of the membrane that induces cellulose synthesis, cell adherence to abiotic surfaces and swimming and swarming motility53. Serine/threonine-protein kinase HipA (Q5A_018950) can inhibit cell growth and induces multidrug tolerance54. Sensor histidine kinase GlrK (Q5A_019275) belongs to a two-component regulatory system that up-regulates transcription of sRNA55. The two-component system Nitrogen assimilation regulatory protein (Q5A_025300) was up-regulated in both, transcriptomics and proteomics data at t1. The expression of the corresponding gene is usually activated in response to nitrogen limitation56. Nitrate/nitrite response regulator protein NarL (Q5A_015065 and Q5A_018475) is involved in the modulate transcription of genes needed for anaerobic respiration with nitrate or nitrite as electron acceptors57. Transcriptional regulatory protein CitB (Q5A_019710) is involved in the expression of citrate-specific fermentation genes58.
In the proteomics dataset, the glucose-specific phosphotransferase enzyme IIA component (Q5A_018245) and nitrogen regulatory protein (Q5A_022765) were induced at t1 and t2 respectively (Table S3). Glucose-specific phosphotransferase enzyme IIA component is part of the sugar phosphotransferase system that is involved in glucose transport across the cell membrane59. The nitrogen regulatory protein is involved in the regulation of nitrogen utilization60. The sensor protein QseC (Q5A_021135) was only identified at t2, which is a member of the two-component regulatory system QseBC that activates the flagella regulon by activating transcription of FlhDC in E. coli 61. The PTS system fructose-specific EIIBC component (Q5A_017065), which is involved in the translocation of fructose across the membrane (Prior and Kornberg62), was repressed at t2.
Differentially expressed genes and proteins related to the cell envelope
Five genes coding for proteins related to the cell envelope were up-regulated at t1 of exposure to fungal VOCs (Table S2). The outer membrane protein W (Q5A_013980) is assumed to be involved in the protection of bacteria against various forms of environmental stress63. UDP-3-O-(3-hydroxymyristoyl)glucosamine N-acyltransferase (Q5A_019950) is involved in the biosynthesis of lipid A of the outer membrane64. Fimbria A protein (Q5A_015665) is a major structural component of mannose-resistant fimbriae65 and the putative fimbrial chaperone YfcS (Q5A_021485) is involved in pilus organization. At both t1 and t2, the gene Q5A_003275 (UDP-N-acetylmuramoyl-L-alanyl-D-glutamate–2,6-diaminopimelate ligase) was up-regulated, which is involved in the biosynthesis of bacterial cell-wall peptidoglycan66.
At t2 on the proteomics level, two proteins, UDP-N-acetylmuramate–L-alanyl-gamma-D-glutamyl-meso-2,6-diaminoheptandioate ligase (Q5A_001880) and UDP-N-acetylglucosamine 1-carboxyvinyltransferase (Q5A_022690) were present in higher levels. These proteins are involved in the peptidoglycan recycling pathway, part of cell wall biogenesis67, 68.
The expression of three proteins was reduced at t1 and t2. The methionine-binding lipoprotein MetQ (Q5A_007015) was reduced at t1 and is involved in the transport of methionine69. D-alanyl-D-alanine carboxypeptidase DacA (Q5A_005980) was reduced at t2 and plays a role in the pathway peptidoglycan biosynthesis. Mannose-1-phosphate guanylyltransferase 1 (Q5A_007950) was repressed at t2 and is involved in the synthesis of GDP-alpha-D-mannose.
Differentially expressed genes and proteins related to energy metabolism
Three genes, Q5A_001620 (fumarate reductase subunit D), Q5A_011355 (Glutathione S-transferase GST-6.0) and Q5A_024300 (Putative 3-oxopropanoate dehydrogenase) were upregulated at t1 (Table S2). The latter one is involved in the degradation of beta-alanine70.
On the transcriptome level, alpha-acetolactate decarboxylase (Q5A_018170) and isocitrate lyase (Q5A_023425) were downregulated at t1, while the latter one was found in higher amounts on the proteome level at t1. Isocitrate lyase catalyzes the bypass of decarboxylation steps of the TCA cycle (glyoxylate shunt). The Glyoxylate shunt is upregulated under different stress conditions, e.g. oxidative stress and antibiotic stress.
Differentially expressed genes and proteins related to biosynthesis of natural products
We observed an upregulation of gene expression of Q5A_011535, which encodes a putative terpene synthase, at t1 of exposure to VOCs (Table S2). The corresponding protein showed the highest induction rate of all proteins in the proteomics dataset at t2 (Table S3). This gene is part of a 3-gene cluster comprising genes from the DXP pathway for isoprenoid biosynthesis (DXP synthase Q5A_011545), isopentenyl diphosphate isomerase (Idi, Q5A_011550) and a methyl transferase (Q5A_011540).
Volatile analysis of S. plymuthica PRI-2C and F. culmorum PV interaction and heterologous expression of sodorifen
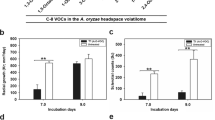
As in both, transcriptomics and proteomics datasets, the putative terpene synthase (Q5A_011535) was upregulated in response to F. culmorum VOCs we further investigated the production of the terpene compound encoded by this gene. By comparing the VOC emission of S. plymuthica PRI-2C exposed to F. culmorum VOCs to S. plymuthica PRI-2C unexposed to fungal VOCs, we observed one particular compound produced in higher amounts in response to fungal VOCs. This compound was identified as sodorifen (C16H26; mass = 218.2033; RT = 28.62) (Fig. 5).
Sodorifen is a recently discovered terpene, which is likely derived from farnesyl pyrophosphate (FPP) but would need an additional methylation to reach its final number of carbons (C16). To confirm that the terpene synthase and methyltransferase can mediate the production of sodorifen, the synthase and methyltransferase genes Q5A_011535 and Q5A_011540 were expressed in an E. coli strain overproducing FPP. Upon co-expression of both genes, formation of sodorifen was detected in the headspace of E. coli cultures, confirming involvement of these genes in the synthesis of sodorifen (Fig. 6a). No products were detected upon expression of either Q5A_011535 (now called SpSS) or Q5A_011540 (now called SpMT) alone. In addition to sodorifen we observed a cluster of other unidentified terpene compounds following the peak of sodorifen (Fig. 6b). This suggests that the terpene synthase is a multi-product terpene synthase that produces a set of sodorifen like terpenes simultaneously. Indeed a number of these peaks were also found in the headspace of Serratia (Fig. 6b).
Comparison of the sodorifen cluster in other bacterial species
We compared the presence of the sodorifen cluster with other bacterial species using antiSMASH71. The results show that the sodorifen cluster of S. plymuthica PRI-2C is present in several closely related isolates, including the originally described sodorifen producer S. plymuthica 4Rx13 (Fig. 7).
Comparison of the sodorifen cluster within other bacteria. The sodorifen synthase (Q5A_011535) forms a cluster with three additional genes, methyltransferase (Q5A_011540), 1-deoxy-D-xylulose-5-phosphate synthase (Q5A_011545) and isopentenyl-diphosphate delta-isomerase (Q5A_011550), which are positioned in the sense direction within the genome of S. plymuthica PRI-2C. The cluster is bordered by genes that are in antisense orientation. Red indicates biosynthesis genes, blue indicates regulatory genes and grey indicates other genes.
Discussion
The ability of microorganisms to communicate is well recognized, however, deciphering the language and the mechanism of communication remains a challenging task. In the recent years several independent studies, including our own, indicated VOCs as ideal info-chemicals for mediating both, short- and long-distance interactions4, 10, 11. Recently, we have shown that VOCs produced by Fusarium culmorum affect the motility of the rhizosphere isolate Serratia plymuthica without affecting bacterial cell numbers17. In the current study we applied both transcriptomic and proteomic approaches to understand the mechanisms underlying bacterial perception and responses to fungal VOCs. Integrated data of our complementary Omics-approach are summarized in Fig. 8.
Schematic overview of the most important changes in S. plymuthica PRI-2C transcriptome and proteome in response to fungal VOCs. The overview includes genes and proteins involved in major pathways that have been detected in this analysis and genes and proteins whose amounts were either decreased (red) or increased (green) after exposure to fungal VOCs.
Access to the complete genomic sequence of an organism is crucial for a reliable interpretation of functional omics data as it enables to correctly identify differentially expressed genes and proteins. Hence, prior to the transcriptomic and proteomic analysis the complete genome of S. plymuthica PRI-2C was sequenced and submitted to NCBI. Both transcriptomics and proteomics approaches identified several genes and proteins related to motility, which is in line with the induced S. plymuthica motility in response to fungal VOCs. Among these, FliS was described as a flagellin-specific chaperone that prevents premature polymerization of flagellins by facilitating its export72. Fimbria A protein was identified in S. marcescens as a major structural component of mannose-resistant fimbriae involved in cell-adhesion65. On the proteomics level the methyl-accepting chemotaxis protein IV was identified, which acts as chemotactic-signal sensor that detects attractants and promotes bacterial movement towards suitable sites for colonization50. Bacterial motility is an important strategy to move toward a favorable stimulus or away from an unfavorable one73. We speculate that the increased motility in S. plymuthica in response to F. culmorum VOCs indicates that fungal VOCs act either as chemo attractants or repellents in these fungal-bacterial interactions.
Next to motility our results revealed several differentially expressed genes and proteins involved in signal transduction. These genes and proteins were involved in the regulation of motility, cell growth, and multidrug tolerance or were part of the two-component system, involved in the regulation of nutrient uptake across the membrane and in the regulation of nutrient limitation. Based on the fact that VOCs modulate antimicrobial activity, Kim et al.12 hypothesized that VOCs could modulate multidrug resistance. Indeed, the study by Kovač et al.74 showed that the monoterpene (−)-α-pinene modulates antibiotic resistance in Campylobacter jejuni and decreased membrane permeability, which was related to the increased expression of the outer membrane protein product of the omp50 gene. The authors speculated that the up-regulation of omp50 is a strategy for bacterial adaptation contributing to decreased membrane permeability and decreased antimicrobial influx. Overexpression of the observed HipA protein has been described to lead to multidrug tolerance in E. coli (Coerreia et al. 2006). Thus, we suggest that fungal VOCs may as well modulate multidrug tolerance in bacteria.
Some VOCs might serve as energy source under nutrient limited conditions and stimulate the growth of other organisms11, 75. Recently Tyc et al.75 revealed that bacterial response to VOCs can vary from growth stimulation to growth inhibition. In this study, we observed a significant increase in S. plymuthica PRI-2C cell numbers when exposed to fungal VOCs. It seems plausible that bacteria take up carbon- or nitrogen-containing volatiles produced by other microorganisms in their surroundings.
In addition we observed a number of upregulated genes and proteins relating to cell envelope biogenesis. Induction of these genes and proteins could imply that the bacterial cells activate membrane biosynthetic pathways when exposed to certain fungal VOCs that causing damages in the bacterial cell wall and thus require repair. Similar findings have been observed by Yung et al.76 when E. coli was exposed to N-methyl-2-pyrrolidone. However, at the same time several genes and proteins relating to cell wall/envelope biogenesis were repressed.
In a previous study we observed that F. culmorum emits a cluster of terpenes17. The interaction between terpenes and the bacterial cell of pathogenic bacteria wall has been mainly described as increased permeability of the cell membrane leading to a loss of microbial viability77, 78. Kovač et al.74 described a concentration dependent effect of (−)-α-pinene on antimicrobial resistance in C. jejuni, leading to either inhibition of antimicrobial efflux in lower concentrations while in higher concentrations leading to increased permeability of the cell membrane thereby promoting the influx of antimicrobials. Our results indicate that terpenes emitted by F. culmorum make the cell wall more permeable for VOCs to enter and to be taken up by the cell.
Volatiles may play a role as elicitors that activate gene clusters involved in secondary metabolites production. Likewise, antibiotic production triggered by volatiles mediated interactions was observed in Pseudomonas aeruginosa during co-culture with Enterobacter aerogenes and in P. fluorescens Pf0-1 in response to volatiles produced by Collimonas pratensis 42, 79.
Moreover, in Chromobacterium violaceum and P. aeruginosa, several monoterpenes increased violacein and pyocyanin production, respectively80. Here we observed an up-regulation of a terpene synthase gene encoding the synthesis of a terpene volatile compound. This compound was identified as sodorifen, an unusual volatile with extraordinary structure, where every carbon atom is substituted with either methyl or methylene group81. However, little is known about the biosynthesis of sodorifen other than the study by Domik et al.81 that reports that a terpene synthase is indispensable for the synthesis of sodorifen and that it descends from the terpene metabolism. Similarly, knockout mutants of the methyl transferase found in the same operon showed no sodorifen production46. We confirmed in our work that a terpene synthase and methyltransferase are essential for the biosynthesis of sodorifen by co-expressing SpSS and SpMT in E. coli. Furthermore, we demonstrated that methylation of FPP is necessary before the cyclisation by the terpene synthase. This was similarly demonstrated by Wang and Cane82 for the biosynthesis of methylisoborneol, a methylated monoterpene with an earthy odor, from Streptomyces coelicolor.
The sodorifen synthase from Serratia is a multiproduct terpene synthase and sodorifen was produced together with a cluster of other terpene compounds. This is in agreement with the analysis of knockout mutants of either the terpene cyclase or the methyl transferase of the sodorifen operon which lacked the production of sodorifen including a number of other terpenoids81. Sodorifen has been described to be produced by several Serratia species83. Even though S. plymuthica PRI-2C was previously reported as a non-producer of sodorifen, this work reveals that the compound is produced under certain conditions and in even higher amounts in response to fungal VOCs. Moreover, in a recent study that described the sodorifen cluster in S. plymuthica 4Rx1381, the cluster was not detected in S. plymuthica PRI-2C when using antiSMASH. This is most likely due to the fact that at the time of the study the genome annotation of this strain was still in draft state and the described cluster could not be found. Comparison of the cluster with the complete genome of S. plymuthica PRI-2C shows a very high sequence similarity at the amino acid level of all four genes of the cluster to other Serratia strains including S. plymuthica 4Rx13.
This work indicates that terpenes may be important molecules in the communication between different microorganisms and hence, may serve as a lingua franca in fungal- bacteria interactions.
References
Dicke, M. & Sabelis, M. W. Infochemical Terminology: Based on Cost-Benefit Analysis Rather than Origin of Compounds? Functional Ecology 2, 131–139, doi:10.2307/2389687 (1988).
Baldwin, I. T., Halitschke, R., Paschold, A., von Dahl, C. C. & Preston, C. A. Volatile signaling in plant-plant interactions: “talking trees” in the genomics era. Science 311, 812–815, doi:10.1126/science.1118446 (2006).
Babikova, Z. et al. Underground signals carried through common mycelial networks warn neighbouring plants of aphid attack. Ecology letters 16, 835–843, doi:10.1111/ele.12115 (2013).
Schmidt, R., Cordovez, V., de Boer, W., Raaijmakers, J. & Garbeva, P. Volatile affairs in microbial interactions. The ISME journal, doi:10.1038/ismej.2015.42 (2015).
Morath, S. U., Hung, R. & Bennett, J. W. Fungal volatile organic compounds: A review with emphasis on their biotechnological potential. Fungal Biology Reviews 26, 73–83, doi:10.1016/j.fbr.2012.07.001 (2012).
Blom, D. et al. Production of plant growth modulating volatiles is widespread among rhizosphere bacteria and strongly depends on culture conditions. Environmental microbiology 13, 3047–3058, doi:10.1111/j.1462-2920.2011.02582.x (2011).
Ryu, C. M. et al. Bacterial volatiles induce systemic resistance in Arabidopsis. Plant physiology 134, 1017–1026, doi:10.1104/pp.103.026583 (2004).
Ryu, C. M. et al. Bacterial volatiles promote growth in Arabidopsis. Proc Natl Acad Sci USA 100, 4927–4932 (2003).
Kai, M., Effmert, U., Berg, G. & Piechulla, B. Volatiles of bacterial antagonists inhibit mycelial growth of the plant pathogen Rhizoctonia solani. Archives of microbiology 187, 351–360, doi:10.1007/s00203-006-0199-0 (2007).
Effmert, U., Kalderas, J., Warnke, R. & Piechulla, B. Volatile mediated interactions between bacteria and fungi in the soil. Journal of chemical ecology 38, 665–703, doi:10.1007/s10886-012-0135-5 (2012).
Garbeva, P., Hordijk, C., Gerards, S. & de Boer, W. Volatile-mediated interactions between phylogenetically different soil bacteria. Frontiers in microbiology 5, 289, doi:10.3389/fmicb.2014.00289 (2014).
Kim, K. S., Lee, S. & Ryu, C. M. Interspecific bacterial sensing through airborne signals modulates locomotion and drug resistance. Nature communications 4, 1809, doi:10.1038/ncomms2789 (2013).
Hol, W. H. et al. Non-random species loss in bacterial communities reduces antifungal volatile production. Ecology 96, 2042–2048 (2015).
Werner, S., Polle, A. & Brinkmann, N. Belowground communication: impacts of volatile organic compounds (VOCs) from soil fungi on other soil-inhabiting organisms. Applied Microbiology and Biotechnology 100, 8651–8665, doi:10.1007/s00253-016-7792-1 (2016).
Pauliuc, B. Antibacterial activity of Pleurotus ostreatus gemmotherapic extract. J Hortic Forest Biotech 17, 242–245 (2013).
Lutz, M. P., Wenger, S., Maurhofer, M., Défago, G. & Duffy, B. Signaling between bacterial and fungal biocontrol agents in a strain mixture. FEMS microbiology ecology 48, 447–455, doi:10.1016/j.femsec.2004.03.002 (2004).
Schmidt, R. et al. Microbial Small Talk: Volatiles in Fungal-Bacterial Interactions. Frontiers in microbiology 6, 1495, doi:10.3389/fmicb.2015.01495 (2015).
Garbeva, P., van Veen, J. A. & van Elsas, J. D. Microbial diversity in soil: selection microbial populations by plant and soil type and implications for disease suppressiveness. Annual review of phytopathology 42, 243–270, doi:10.1146/annurev.phyto.42.012604.135455 (2004).
Garbeva, P. & de Boer, W. Inter-specific interactions between carbon-limited soil bacteria affect behavior and gene expression. Microbial ecology 58, 36–46, doi:10.1007/s00248-009-9502-3 (2009).
De Boer, W. et al. Anti-fungal properties of chitinolytic dune soil bacteria. Soil Biology and Biochemistry 30, 193–203, doi:10.1016/S0038-0717(97)00100-4 (1998).
Fiddaman, P. J. & Rossall, S. The production of antifungal volatiles by Bacillus subtilis. J Appl Bacteriol 74, 119–126 (1993).
Aronesty, E. Command-line tools for processing biological sequencing data; http://code.google.com/p/ea-utils (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760, doi:10.1093/bioinformatics/btp324 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Research 20, 1297–1303, doi:10.1101/gr.107524.110 (2010).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One 9, e112963, doi:10.1371/journal.pone.0112963 (2014).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069, doi:10.1093/bioinformatics/btu153 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Meth 9, 357-359, 10.1038/nmeth.1923 http://www.nature.com/nmeth/journal/v9/n4/abs/nmeth.1923.html - supplementary-information (2012).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. Bmc Bioinformatics 12, 1–16, doi:10.1186/1471-2105-12-323 (2011).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, doi:10.1093/bioinformatics/btp616 (2010).
Zhou, X. B., Lindsay, H. & Robinson, M. D. Robustly detecting differential expression in RNA sequencing data using observation weights. Nucleic Acids Res 42, doi:ARTN e9110.1093/nar/gku310 (2014).
Oliveros, J. Venny. An interactive tool for comparing lists with Venn diagrams. BioinfoGP, CNB-CSIC, doi:citeulike-article-id:6994833.
Haft, D. H. et al. TIGRFAMs and Genome Properties in 2013. Nucleic Acids Res 41, D387–395, doi:10.1093/nar/gks1234 (2013).
Schneider, T. et al. Structure and function of the symbiosis partners of the lung lichen (Lobaria pulmonaria L. Hoffm.) analyzed by metaproteomics. Proteomics 11, 2752–2756, doi:10.1002/pmic.201000679 (2011).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408, doi:10.1006/meth.2001.1262 (2001).
Muntel, J., Hecker, M. & Becher, D. An exclusion list based label-free proteome quantification approach using an LTQ Orbitrap. Rapid Commun Mass Spectrom 26, 701–709, doi:10.1002/rcm.6147 (2012).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat Protoc 2, 1896–1906, doi:10.1038/nprot.2007.261 (2007).
Muntel, J. et al. Comprehensive absolute quantification of the cytosolic proteome of Bacillus subtilis by data independent, parallel fragmentation in liquid chromatography/mass spectrometry (LC/MS(E). Mol Cell Proteomics 13, 1008–1019, doi:10.1074/mcp.M113.032631 (2014).
Zuhlke, D. et al. Costs of life - Dynamics of the protein inventory of Staphylococcus aureus during anaerobiosis. Sci Rep 6, 28172, doi:10.1038/srep28172 (2016).
Silva, J. C., Gorenstein, M. V., Li, G. Z., Vissers, J. P. & Geromanos, S. J. Absolute quantification of proteins by LCMSE: a virtue of parallel MS acquisition. Mol Cell Proteomics 5, 144–156, doi:10.1074/mcp.M500230-MCP200 (2006).
Saeed, A. I. et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34, 374–378 (2003).
Vizcaino, J. A. et al. ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat Biotechnol 32, 223–226, doi:10.1038/nbt.2839 (2014).
Garbeva, P., Hordijk, C., Gerards, S. & de Boer, W. Volatiles produced by the mycophagous soil bacterium Collimonas. FEMS microbiology ecology 87, 639–649, doi:10.1111/1574-6941.12252 (2014).
von Reuss, S. H., Kai, M., Piechulla, B. & Francke, W. Octamethylbicyclo[3.2.1]octadienes from the rhizobacterium Serratia odorifera. Angew Chem Int Ed Engl 49, 2009–2010, doi:10.1002/anie.200905680 (2010).
Martin, V. J., Pitera, D. J., Withers, S. T., Newman, J. D. & Keasling, J. D. Engineering a mevalonate pathway in Escherichia coli for production of terpenoids. Nat Biotechnol 21, 796–802 (2003).
Peralta-Yahya, P. P. et al. Identification and microbial production of a terpene-based advanced biofuel. Nature communications 2, 483, doi:10.1038/ncomms1494 http://www.nature.com/articles/ncomms1494 - supplementary-information (2011).
Domik, D., Magnus, N. & Piechulla, B. Analysis of a new cluster of genes involved in the synthesis of the unique volatile organic compound sodorifen of Serratia plymuthica 4Rx13. FEMS Microbiol Lett 363, doi:10.1093/femsle/fnw139 (2016).
Stock, A. M. et al. Two-component signal transduction. Annu Rev Biochem 69, 183–215 (2000).
Ulrich, L. E. & Zhulin, I. B. The MiST2 database: a comprehensive genomics resource on microbial signal transduction. Nucleic acids research 38, D401–407, doi:10.1093/nar/gkp940 (2010).
Ulrich, L. E., Koonin, E. V. & Zhulin, I. B. One-component systems dominate signal transduction in prokaryotes. Trends in microbiology 13, 52–56, doi:10.1016/j.tim.2004.12.006 (2005).
Li, Z. et al. Methyl-accepting chemotaxis proteins 3 and 4 are responsible for Campylobacter jejuni chemotaxis and jejuna colonization in mice in response to sodium deoxycholate. J Med Microbiol 63, 343–354, doi:10.1099/jmm.0.068023-0 (2014).
Pesavento, C. et al. Inverse regulatory coordination of motility and curli-mediated adhesion in Escherichia coli. Genes Dev 22, 2434–2446, doi:10.1101/gad.475808 (2008).
Park, C. M. et al. A second photochromic bacteriophytochrome from Synechocystis sp. PCC 6803: spectral analysis and down-regulation by light. Biochemistry 39, 10840–10847 (2000).
Da, Re,S. & Ghigo, J. M. A CsgD-independent pathway for cellulose production and biofilm formation in Escherichia coli. J Bacteriol 188, 3073–3087, doi:10.1128/jb.188.8.3073-3087.2006 (2006).
Germain, E., Castro-Roa, D., Zenkin, N. & Gerdes, K. Molecular mechanism of bacterial persistence by HipA. Mol Cell 52, 248–254, doi:10.1016/j.molcel.2013.08.045 (2013).
Yamamoto, K. et al. Functional characterization in vitro of all two-component signal transduction systems from Escherichia coli. J Biol Chem 280, 1448–1456, doi:10.1074/jbc.M410104200 (2005).
Feng, B. & Marzluf, G. A. Interaction between major nitrogen regulatory protein NIT2 and pathway-specific regulatory factor NIT4 is required for their synergistic activation of gene expression in Neurospora crassa. Mol Cell Biol 18, 3983–3990 (1998).
Zhang, J. H., Xiao, G., Gunsalus, R. P. & Hubbell, W. L. Phosphorylation triggers domain separation in the DNA binding response regulator NarL. Biochemistry 42, 2552–2559, doi:10.1021/bi0272205 (2003).
Yamamoto, K. et al. Anaerobic regulation of citrate fermentation by CitAB in Escherichia coli. Biosci Biotechnol Biochem 72, 3011–3014, doi:10.1271/bbb.80301 (2008).
Saffen, D. W., Presper, K. A., Doering, T. L. & Roseman, S. Sugar transport by the bacterial phosphotransferase system. Molecular cloning and structural analysis of the Escherichia coli ptsH, ptsI, and crr genes. J Biol Chem 262, 16241–16253 (1987).
Huergo, L. F., Chandra, G. & Merrick, M. P. II Signal transduction proteins: nitrogen regulation and beyond. FEMS Microbiol Rev 37, 251–283 (2013).
Sperandio, V., Torres, A. G. & Kaper, J. B. Quorum sensing Escherichia coli regulators B and C (QseBC): a novel two-component regulatory system involved in the regulation of flagella and motility by quorum sensing in E. coli. Mol Microbiol 43, 809–821 (2002).
Prior, T. I. & Kornberg, H. L. Nucleotide sequence of fruA, the gene specifying enzyme IIfru of the phosphoenolpyruvate-dependent sugar phosphotransferase system in Escherichia coli K12. J Gen Microbiol 134(10), 2757–2768 (1988).
Li, W. et al. Contribution of the outer membrane protein OmpW in Escherichia coli to complement resistance from binding to factor H. Microb Pathog 98, 57–62, doi:10.1016/j.micpath.2016.06.024 (2016).
Bartling, C. M. & Raetz, C. R. Crystal structure and acyl chain selectivity of Escherichia coli LpxD, the N-acyltransferase of lipid A biosynthesis. Biochemistry 48, 8672–8683, doi:10.1021/bi901025v (2009).
Mizunoe, Y. et al. Cloning and sequence of the gene encoding the major structural component of mannose-resistant fimbriae of Serratia marcescens. J Bacteriol 170, 3567–3574 (1988).
McGroty, S. E. et al. Biochemical characterization of UDP-N-acetylmuramoyl-L-alanyl-D-glutamate: meso-2,6-diaminopimelate ligase (MurE) from Verrucomicrobium spinosum DSM 4136(T.). PLoS One 8, e66458, doi:10.1371/journal.pone.0066458 (2013).
Mengin-Lecreulx, D., van Heijenoort, J. & Park, J. T. Identification of the mpl gene encoding UDP-N-acetylmuramate: L-alanyl-gamma-D-glutamyl-meso-diaminopimelate ligase in Escherichia coli and its role in recycling of cell wall peptidoglycan. J Bacteriol 178, 5347–5352 (1996).
Skarzynski, T. et al. Structure of UDP-N-acetylglucosamine enolpyruvyl transferase, an enzyme essential for the synthesis of bacterial peptidoglycan, complexed with substrate UDP-N-acetylglucosamine and the drug fosfomycin. Structure 4, 1465–1474 (1996).
Gal, J., Szvetnik, A., Schnell, R. & Kalman, M. The metD D-methionine transporter locus of Escherichia coli is an ABC transporter gene cluster. J Bacteriol 184, 4930–4932 (2002).
Yao, X., He, W. & Lu, C. D. Functional characterization of seven gamma-Glutamylpolyamine synthetase genes and the bauRABCD locus for polyamine and beta-Alanine utilization in Pseudomonas aeruginosa PAO1. J Bacteriol 193, 3923–3930, doi:10.1128/jb.05105-11 (2011).
Medema, M. H. et al. antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic acids research 39, W339–346, doi:10.1093/nar/gkr466 (2011).
Galeva, A. et al. Bacterial flagellin-specific chaperone FliS interacts with anti-sigma factor FlgM. J Bacteriol 196, 1215–1221, doi:10.1128/jb.01278-13 (2014).
Alexandre, G. Chemotaxis Control of Transient Cell Aggregation. J Bacteriol 197, 3230–3237 (2015).
Kovač, J. et al. Antibiotic Resistance Modulation and Modes of Action of (−)-α-Pinene in Campylobacter jejuni. PLoS One 10, e0122871, doi:10.1371/journal.pone.0122871 (2015).
Tyc, O., Zweers, H., de Boer, W. & Garbeva, P. Volatiles in Inter-Specific Bacterial Interactions. Frontiers in microbiology 6, 1412, doi:10.3389/fmicb.2015.01412 (2015).
Yung, P. Y. et al. Global transcriptomic responses of Escherichia coli K-12 to volatile organic compounds. Scientific Reports 6, 19899, doi:10.1038/srep19899 http://www.nature.com/articles/srep19899 - supplementary-information (2016).
Nazzaro, F., Fratianni, F., De Martino, L., Coppola, R. & De Feo, V. Effect of Essential Oils on Pathogenic Bacteria. Pharmaceuticals 6, 1451–1474, doi:10.3390/ph6121451 (2013).
Trombetta, D. et al. Mechanisms of Antibacterial Action of Three Monoterpenes. Antimicrobial Agents and Chemotherapy 49, 2474–2478, doi:10.1128/aac.49.6.2474-2478.2005 (2005).
Venkataraman, A., Rosenbaum, M. A., Werner, J. J., Winans, S. C. & Angenent, L. T. Metabolite transfer with the fermentation product 2,3-butanediol enhances virulence by Pseudomonas aeruginosa. ISME J 8, 1210–1220, doi:10.1038/ismej.2013.232 (2014).
Ahmad, A., Viljoen, A. M. & Chenia, H. Y. The impact of plant volatiles on bacterial quorum sensing. Lett Appl Microbiol 60, 8–19 (2015).
Domik, D. et al. A Terpene Synthase Is Involved in the Synthesis of the Volatile Organic Compound Sodorifen of Serratia plymuthica 4Rx13. Frontiers in microbiology 7 (2016).
Wang, C. M. & Cane, D. E. Biochemistry and molecular genetics of the biosynthesis of the earthy odorant methylisoborneol in Streptomyces coelicolor. Journal of the American Chemical Society 130, 8908–8909, doi:10.1021/ja803639g (2008).
Weise, T. et al. VOC emission of various Serratia species and isolates and genome analysis of Serratia plymuthica 4Rx13. (FEMS Microbiol Lett. 2014 Mar; 352(1):45–53, doi:10.1111/1574-6968.12359. Epub 2014 Jan 7).
Acknowledgements
This work is supported by The Netherlands Organization for Scientific Research (NWO) VIDI personal grant to Paolina Garbeva (864.11.015). This is publication 6238 of the NIOO-KNAW.
Author information
Authors and Affiliations
Contributions
R.S. designed and conducted transcriptomics, proteomics and metabolomics experiments, analyzed the data and wrote the manuscript. V.D.J. contributed to the analysis of the transcriptomics data and provided genome assembly and annotation; D.Z. assisted in the proteomics design, conduction of the experiment, data analysis and in writing the manuscript; C.W. assisted in the proteomics experiment; J.B. assisted in the proteomics data analysis; K.C. performed heterologous expression experiments; J.B. contributed to the design of the heterologous expression experiments and edited the manuscript; W.V.I. and F.S. performed RNA-Sequencing. W.D.B. edited the manuscript; K.R. contributed to the design of the proteomics experiments and edited the manuscript; P.G. assisted in the design of all experiments and edited the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schmidt, R., Jager, V., Zühlke, D. et al. Fungal volatile compounds induce production of the secondary metabolite Sodorifen in Serratia plymuthica PRI-2C. Sci Rep 7, 862 (2017). https://doi.org/10.1038/s41598-017-00893-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-00893-3
This article is cited by
-
The microbial biosynthesis of noncanonical terpenoids
Applied Microbiology and Biotechnology (2024)
-
Investigation of rust mycoparasite Sphaerellopsis filum (TNAU SF1) volatilome during mono and di-partite interaction with Puccinia arachidis, causing Peanut rust
European Journal of Plant Pathology (2024)
-
Activity of volatiles induced by microbes and natural plants stifled the growth of Pythium aphanidermatum - the damping off in Tomato
BMC Plant Biology (2023)
-
Antagonistic fungal volatiles as potential biocontrol countermeasure for microbial postharvest fruit diseases
Egyptian Journal of Biological Pest Control (2023)
-
Repeated exposure of wheat to the fungal root pathogen Bipolaris sorokiniana modulates rhizosphere microbiome assembly and disease suppressiveness
Environmental Microbiome (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.