Abstract
The potato tuberworm, Phthorimaea operculella Zeller, is an oligophagous pest feeding on crops mainly belonging to the family Solanaceae. It is one of the most destructive pests of potato worldwide and attacks foliage and tubers in the field and in storage. However, the lack of a high-quality reference genome has hindered the association of phenotypic traits with their genetic basis. Here, we report on the genome assembly of P. operculella at the chromosomal level. Using Illumina, Nanopore and Hi-C sequencing, a 648.2 Mb genome was generated from 665 contigs, with an N50 length of 3.2 Mb, and 92.0% (596/648.2 Mb) of the assembly was anchored to 29 chromosomes. In total, 16619 genes were annotated, and 92.4% of BUSCO genes were fully represented. The chromosome-level genome of P. operculella will provide a significant resource for understanding the genetic basis for the biological study of this insect, and for promoting the integrative management of this pest in future.
Measurement(s) | whole genome sequencing |
Technology Type(s) | Illumina NovaSeq. 6000 • Oxford Nanopore Sequencing |
Sample Characteristic - Organism | Phthorimaea operculella |
Sample Characteristic - Location | Yunnan Province, Southwest China |
Similar content being viewed by others
Background & Summary
The potato tuberworm, Phthorimaea operculella Zeller (Lepidoptera: Gelechiidae), is one of the main pests affecting potatoes, Solanum tuberosum, worldwide (Fig. 1). As an oligophagous pest of plants in the family Solanaceae, it uses potato, tomato (S. lycopersicum), and tobacco (Nicotiana tabacum) as principal hosts. It was first described in California in 1856. Since then, its presence has been reported in over 90 countries1. Phthorimaea operculella larvae feed on potato leaves, stems and petioles in the field, and tubers in storage. Severe infestations can destroy the foliage and results in substantial yield loss; however, main damage is the one that affects tubers. For instance, in some developing countries, the larvae can cause a 50–90% economic loss in storage; within weeks, tubers can become unmarketable if left untreated2. Pesticide application is most widely used management strategy to control P. operculella. Unfortunately, it can cause the development of insecticide resistance and negatively impact agro-ecosystem3,4. Thus, to promote more innovative management strategies for this destructive pest, a deeper understanding of its genetics is required but remains to be accomplished.
P. operculella belongs to the family Gelechiidae which is one of the most diverse families of microlepidoptera. Gelechiidae includes over 4700 described species in more than 500 genera in the world5. Many species of this family are considered important agricultural pests and feed voraciously on Solanaceous crops. P. operculella, the Guatemalan potato tuber moth Tecia solanivora Povolny, the tomato leaf miner Tuta absoluta Meyrick, the tomato pinworm Keiferia lycopersicella Walsingham, etc are among the major pests of this family. The genomic information for this family, however, remains scarce. Tabuloc et al. recently constructed a draft genome assembly for T. absoluta; the group also sequenced a preliminary genome of K. lycopersicella and P. operculella, with which a panel of 21-SNP markers6. Accumulating genomic information in the Gelechiidae family could promote a better understanding of the supra-specific classification within species7. To promote future studies on the genetics, biology, and ecology of Gelechiidae, it is of importance to build a chromosomal-level high quality genome assembly for important species such as P. operculella.
In the current study, we present a high-quality P. operculella chromosome-level genome assembly and life cycle transcriptomes. Using Illumina short reads, Nanopore, and High-throughput chromosome conformation capture (Hi-C) data, a 648.2 Mb genome was generated from 665 contigs, with an N50 length of 3.2 Mb, and 92.0% (596/648.2 Mb) of the assembly was anchored to 29 chromosomes. The female-specific W chromosome of P. operculella was not dertermined in this genome, since the identification of W chromosome is challenging due to high degeneracy, being gene-poor and repeat-rich8. In total, 16441 genes were annotated. Our genomic features of P. operculella will lay a foundation for further research on this insect pest.
Methods
Sample collection and sequencing
In 2014, P. operculella adults (n = 500) were collected from a potato field in Yunnan Province, China. The insect colony was maintained in the climate chamber at 27 ± 2°C, 60% RH and photoperiod of 12 h L: 12 h D. As in 2022, the colony has 100 generations of P. operculella. The chromosomal sex determination of P. operculella takes the form of female heterogamety (females are WZ, males ZZ)9. The male genome of P. operculella was thus sequenced to avoid the complications expected from the W chromosome of Lepidoptera10. DNA for both Illumina and Oxford Nanopore sequencing was obtained from 16 male pupae to avoid the contamination of eggs, and for Hi-C sequencing it was obtained from 200 mg fresh eggs.
The high-quality genomic DNA of P. operculella was prepared by the CTAB method and purified with QIAGEN® Genomic kit (QIAGEN, USA) at Grandomic Biosciences Co., Ltd (Wuhan, China), which was used for preparing the Illumina and Oxford Nanopore (ONT) sequencing libraries. The Illumina NovaSeq 6000 platform generated ~61 Gb of data with 150 bp paired-end reads, with an average insert size of 300~500 bp (Table 1). The Nanopore PromethION 24 platform generated ~68 Gb of sequencing data, and the adapters were removed using Porechop (https://github.com/rrwick/Porechop). The Hi-C library was constructed at Annoroad Gene Technology Co., Ltd (Beijing) following the standard library preparation protocol, and ~101 Gb of data with 50 bp paired-end sequencing raw reads were generated.
With the Illumina sequencing data, we estimated the P. operculella genome size of ~636 Mb directly from kmer coverage from jellyfish v 2.0.0 analysis11; meanwhile, we used the Genomescope v1.0.0 method12 and estimated the genome size of ~560 Mb. The result suggested that the size of P. operculella may range from 560 to 636 Mb.
RNA sequencing and analysis
Newly laid eggs, 1st, 2nd, 3rd and 4th instar larvae, mature larvae, pupae, and newly emerged adult moths were collected for transcriptome sequencing and gene expression analysis. Total RNA was isolated from eggs, larvae, and adults samples collected above, using Trizol reagent (Invitrogen, USA) following the manufacturer’s protocol. Illumina sequencing and complementary DNA (cDNA) library construction were performed at Grandomic Biosciences Co., Ltd (Wuhan, China). Clean data were obtained by removing adapters, low-quality reads, and high-content unknown sequences. Clean reads from each sample were mapped to the genome assembly to measure gene transcript levels using the reported analysis pipeline13.
De novo genome assembly
Nanopore sequenced reads with a length of at least 8 kb were used for genome assembly by the Canu v1.814 with parameters of “maxThreads = 60 genomeSize = 636 m -nanopore-raw”. For the primary assembly, the purge_dups15 was used to remove haplotypic duplication sequences, and Pilon16 and Racon17 were used to polish the assembly. Bacterial sequences that were identified by aligning against the NCBI nt database were also removed. After removing the mitogenome and bacterial sequences, we obtained the 665 contigs with size of 648.2 Mb, which was similar to the predicted size of ~560–636 Mb. The contig N50 size was 3.2 Mb. The analysis of Hi-C data helped to anchor 337 (50.7%) contigs of 596.3 (92.0%) Mb sequence to 29 chromosomes18 (Table 1 and Fig. 2). The 328 (49.3%) un-anchored scaffolds contained 51.9 (8.0%) Mb sequence. The mitochondrial genome of 15,267 bp was also obtained (Table 2 and Fig. 3).
Repeat annotation
Transposable elements (TE), low complexity sequences and simple repeats were identified by RepeatMasker open-4.0.5 (http://www.repeatmasker.org) and RepeatScout19. Firstly, we used RepeatMasker to analyze low complexity sequences and simple repeats, as well as reference based TEs based on Repbase sequences v19.0620. Then we used the de novo method to discover TE families by running RepeatScout analysis, and these TE families were used for repeat annotation by running RepeatMasker analysis. In the 648.2 Mb genome assembly, 55.0% was repeat sequences21, including 54.2% of transposable elements (TEs), and 0.8% of simple repeats and low complexity sequences22.
Protein-coding genes prediction and other annotation of the genome
Based on the genome sequence, we used Augustus23 and Genemark24 for ab initio gene prediction. And based on evidence from RNA-seq alignments and NCBI refseq invertebrate homology (https://ftp.ncbi.nlm.nih.gov/refseq/release/invertebrate/), we used Braker25 to infer gene models under three rounds of prediction (i.e., Braker + RNA-seq, Braker + refseq, Braker + RNA-seq + refseq). Then, we assigned priority to five gene sets (i.e., Braker + RNA-seq + refseq > Braker + RNA-seq > Braker + refseq > Genemark > Augustus), and selected genes supported by at least two methods or genes supported by only one method but containing functional domains. Moreover, all the selected genes may have similarity to reported invertebrate proteins or have RNA-seq evidence. Finally, we obtained 16,619 predicted protein-coding genes.
For five genomes of Samia ricini, Dendrolimus punctatus, Drepana arcuata, T. absoluta and Carposina sasakii without gene sets available at NCBI, we performed gene prediction analysis based on genome sequences using Augustus, Genemark and Braker + refseq methods, similar to those for the prediction of P. operculella genes. A total of 14,015, 14,483, 13,387, 17,607 and 15,873 genes were identified for S. ricini, D. punctatus, D. arcuata, T. absoluta and C. sasakii, respectively26.
For the amino acids of gene sets from 23 genomes within Lepidoptera and L. decemlineata genome within Coleoptera22, we used Benchmarking Universal Single-Copy Orthologs (BUSCO) v3.0.127 to evaluate their quality. The “eukaryota_odb9” dataset at the BUSCO website (https://busco-archive.ezlab.org/v3/) was downloaded for analysis. The BUSCO completeness of >90% and >80% were found for gene sets from 16 and 20 genomes, respectively (Fig. 4).
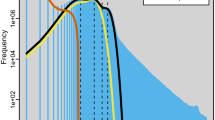
Comparison of characteristics between Phthorimaea operculella and 22 other Lepidopteran genomes. (a) GC contents of 23 lepidopteran genomes including species of 12 superfamily. (b) BUSCO scores for 23 assembled lepidopteran genomes. (c) Relationships between Contig N50 sizes and Contig numbers for 19 lepidopteran genomes with complete BUSCO scores above 80%. (d) Relationships between genome sizes and gene numbers for 19 lepidopteran genomes with complete BUSCO scores above 80%. The abbreviations for the name of each species were marked with blue.
To perform functional annotation, we aligned gene sequences against Pfam22,28, NCBI refseq invertebrate (https://ftp.ncbi.nlm.nih.gov/refseq/release/invertebrate/), UniProt29 and KOG30 databases using BLASTP with E-value cutoff of 1e-5 (Fig. 5). And pathway annotation was analyzed by KAAS31 online database server22. The P450 genes were annotated by aligning amino acids of genes against the collected data on Cytochrome P450 database (http://www.p450.unizulu.ac.za/)32. The genes from four sub-families (mito, CYP2, CYP3 and CYP4) were confirmed according to their annotation and orthogroup information (as described in the “Comparative genomic analysis”). The ABC transporters were identified based on UniProt annotation and orthogroup information of genes. All other metabolic enzyme genes were annotated based on domain annotations. The chemosensory genes containing the odorant receptor, olfactory receptor, the chemosensory receptor, PBP/GOBP family, and insect pheromone-binding family domains were annotated as OR, ORother, GR, OBP, and CSP genes, respectively. Genes of ligand-gated ion channels were annotated as IR genes. The sub-families (delta, epsilon, sigma, zeta, omega, theta and unclassified) of GST genes were annotated by comparing sequences against GSTs of Plutella xylostella33. These metabolic enzyme genes and chemosensory genes from 20 lepidopteran genomes with BUSCO completeness of larger than 80% were annotated22 (Fig. 6).
Distribution of detoxification and chemosensory genes in Lepidoptera species. P450, cytochrome P450 monooxygenase; ABC, ATP-binding cassette transporter; GST, glutathione S-transferase; COE, carboxylesterase; APN, aminopeptidase N; UGT, uridine diphosphate-glycosyltransferase; OR, olfactory receptor; GR, gustatory receptor; IR, ionotropic receptor; OBP, odorant-binding protein; CSP, chemosensory proteins.
Comparative genomic analysis for lepidopteran species
Twenty-four genomes were used for performing a comparative genomic analysis22, including 23 Lepidoptera genomes and one Coleoptera genome (L. decemlineata). These Lepidoptera genomes were from 17 families, with T. absoluta and P. operculella from the Gelechiidae family. The OrthoFinder v2.3.1134 detected 69,067 orthogroups for genes from these 24 genomes22, including 85 single-copy gene groups. Each orthogroup was considered as one gene family in the following analysis.
For each single-copy gene, we used MUSCLE v3.8.3135 to perform sequence alignment of amino acids. All the aligned genes were assembled by an in-house perl script (global_alignment_single_copy_genes.pl; https://github.com/linrm2010/global_alignment_single_copy_genes/). Then Gblock v0.91b36 was used to remove ambiguously aligned regions. The ProtTest v3.437 identified the best model of JTT + I + F + G for constructing the phylogenetic trees. We used RAxML38 to construct the maximum likelihood phylogenetic tree for the 24 genomes (Fig. 7). After that, we analyzed the potential gene family emergence extinction according to the description in a previous study39, and applied CAFE v3.140 to examine the expansion and contraction of gene families across the phylogenetic tree of genomes (Figs. 8,9).
Gene family changes among lepidopteran insects. (a) Gene family changes associated with the origin and evolution of Lepidoptera. The topology of a phylogenetic tree constructed of 19 lepidopteran species. Leptinotarsa decemlineata (Coleoptera) was used as an outgroup. Gene family birth (+) and death (−) in 20 species are shown. (b) Expansion gene families in P. operculella, compared to other lepidopteran insects and L. decemlineata.
Data Records
The genome sequence and gene sequence had been deposited at the National Center for Biotechnology Information (NCBI), under the accession number of JANFCV000000000.1, and can be download from (ftp.ncbi.nlm.nih.gov/genomes/all/GCA/024/500/475/GCA_024500475.1_ASM2450047v1/)41. The NCBI BioProject accession number is PRJNA848272. The raw data of Nanopore, Illumina and Hi-C sequencing were submitted to NCBI SRA with the accession number of SRP40534042.Meanwhile, the genome sequence and gene sequence were also publicly available in National Genomic Data Center (NGDC), under the accession number of GWHBJUP00000000 (nuclear genome) and GWHBJUO01000000 (mitogenome). The gene expression data were publicly available in NGDC, under the accession number of OMIX001281. All data were related to the BioProject PRJCA010352.
Technical Validation
We assessed the quality of genome assembly in the following aspects: (i) We obtained the complete mitogenome sequence of P. operculella. (ii) We aligned the Illumina sequencing reads against the nuclear genome using BWA v 0.7.17-r118843, and found that 99.41% reads matched to genome sequences. (iii) The Core Eukaryotic Genes Mapping Approach (CEGMA) defined 458 core eukaryotic genes, and 248 of them were the most highly-conserved core genes, which could be used to assess the completeness of the genome or annotations44. We aligned the P. operculella genes against these 248 core genes, and identified that 243 (97.98%) core genes have homologous genes in the P. operculella gene sets. (iv) The BUSCO20 analysis showed that 96.4% of gene orthologs were identified in P. operculella, including complete and fragment scores of 92.4% and 4.0%, respectively. These results showed that we obtained the high-quality genome of P. operculella.
Code availability
All software and pipelines used in this study were executed according to the manual and protocols of the published bioinformatic tools. The versions of software have been described in Methods.
The parameters of software/programs are as follows:
Jellyfish: count -C -m 21.
Genomescope: Rscript genomescope-1.0.0/genomescope.R NGS_reads.histo 21 150 NGS_reads.genomescope.
Canu: genomeSize = 636 m -nanopore-raw.
minimap2: -x map-ont.
purge_dups: −2 -T cutoffs -c PB.base.cov.
BUSCO: -m prot -f -l eukaryota_odb9.
ProtTest: -all-distributions -F -AIC -BIC.
Default parameters were used for Porechop, Pilon, Racon, RepeatMasker, RepeatScout, RepeatScout, Augustus, Genemark, Prothint, Braker, hmmsearch, OrthoFinder, MUSCLE and Gblock.
References
Rondon, S. I. Decoding Phthorimaea operculella (Lepidoptera: Gelechiidae) in the new age of change. J. Integr. Agr. 19, 316–324 (2020).
Trivedi, T. P. & Rajagopal, D. Distribution, biology, ecology and management of potato tuber moth, Phthorimaea operculella (Zeller) (Lepidoptera: Gelechiidae): A review. Int. J. Pest Manage. 38, 279–285 (1992).
Doğramacı, M. & Tingey, W. M. Comparison of insecticide resistance in a North American field population and a laboratory colony of potato tuberworm (Lepidoptera: Gelechiidae). J. Pest Sci. 81, 17 (2008).
Collantes, L. G., Raman, K. V. & Cisneros, F. H. Effect of six synthetic pyrethroids on two populations of potato tuber moth, Phthorimaea operculella (Zeller) (Lepidoptera: Gelechiidae), in Peru. Crop Prot. 5, 355–357 (1986).
Huemer, P. et al. DNA barcode library for European Gelechiidae (Lepidoptera) suggests greatly underestimated species diversity. ZooKeys 921, 141–157 (2020).
Tabuloc, C. et al. Sequencing of Tuta absoluta genome to develop SNP genotyping assays for species identification. J. Pest Sci. 92, 1397–1407 (2019).
Chang, P. E. C. & Metz, M. A. Classification of Tuta absoluta (Meyrick, 1917) (Lepidoptera: Gelechiidae: Gelechinae: Gnorimoschemini) based on cladistic analysis of morphology. P. Entomol. Soc. Wash. 123, 41–54 (2021).
Bergero, R. & Charlesworth, D. The evolution of restricted recombination in sex chromosomes. Trends Ecol. Evol. 24, 94–102 (2009).
Makee, H. & Tafesh, N. Sex chromatin body as a marker of radiation-induced sex chromosome aberrations in the potato tuber moth, Phthorimaea operculella (Lepidoptera: Gelechiidae). J. Pest Sci. 79, 75–82 (2006).
Cao, L. et al. Chromosome-level genome of the peach fruit moth Carposina sasakii (Lepidoptera: Carposinidae) provides a resource for evolutionary studies on moths. Mol. Ecol. Resour. 21, 834–848 (2021).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204 (2017).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Guan, D. et al. Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics 36, 2896–2898 (2020).
Walker, B. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One 9, e112963 (2014).
Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746 (2017).
Zhang, M. et al. Contig orders and orientations on Phthorimaea operculella chromosomes. figshare https://doi.org/10.6084/m9.figshare.21510693.v1 (2022).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21, 351–358 (2005).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 6, 11 (2015).
Zhang, M. et al. Phthorimaea operculella genome repeat sequences annotation. figshare https://doi.org/10.6084/m9.figshare.21431649.v1 (2022).
Zhang, M. et al. Supplementary tables for ‘Chromosomal-level genome assembly of potato tuberworm, Phthorimaea operculella: a pest of solanaceous crops’. figshare https://doi.org/10.6084/m9.figshare.21510738.v1 (2022).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic. Acids Res. 34, 435–439 (2006).
Lomsadze, A., Ter-Hovhannisyan, V., Chernoff, Y. O. & Borodovsky, M. Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic. Acids Res. 33, 6494–506 (2005).
Hoff, J., Lomsadze, A., Borodovsky, M. & Stanke, M. Whole-Genome Annotation with BRAKER. Methods Mol. Biol. 1962, 65–95 (2019).
Zhang, M. et al. Gene prediction for five Lepidopteran genomes. figshare https://doi.org/10.6084/m9.figshare.21431598.v1 (2022).
Waterhouse, R. M. et al. BUSCO applications from quality assessments to gene prediction and phylogenomics. Mol. Biol. Evol. 35, 543–548 (2018).
El-Gebali, S. et al. The Pfam protein families database in 2019. Nucleic. Acids Res. 47, D427–D432 (2019).
UniProt Consortium. UniProt: a worldwide hub of protein knowledge. Nucleic. Acids Res. 47, D506–D515 (2019).
Koonin, E. V. et al. A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol. 5, R7 (2004).
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A. C. & Kanehisa, M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic. Acids Res. 35, W182–185 (Web Server issue) (2007).
Nelson, D. R. The cytochrome p450 homepage. Hum. Genomics 4, 59–65 (2009).
You, Y. et al. Characterization and expression profiling of glutathione S-transferases in the diamondback moth, Plutella xylostella (L.). BMC Genomics 16, 152 (2015).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic. Acids Res. 32, 1792–1797 (2004).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–1165 (2011).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Mitreva, M. et al. The draft genome of the parasitic nematode Trichinella spiralis. Nat. Genet. 43, 228–235 (2011).
Han, M. V., Thomas, G. W., Lugo-Martinez, J. & Hahn, M. W. Estimating gene gain and loss rates in the presence of error in genome assembly and annotation using CAFE 3. Mol. Biol. Evol. 30, 1987–1997 (2013).
Zhang, M. et al. Genbank https://identifiers.org/insdc.gca:GCA_024500475.1 (2022).
NCBI Sequence Read Archive https://www.ncbi.nlm.nih.gov/sra/SRP405340 (2022).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Acknowledgements
This work was supported by the the National Key Research and Development Program of China (2018YFD0200802); the Key Research and Development Program of Zhejiang Province (Grant No. 2019C04007); the National Nature Science Foundation of China (Grant No. 32072432, 32272636).
Author information
Authors and Affiliations
Contributions
Y.G., X.C. and W.Z. conceived the research project. M.Z. and Y.G. collected the samples, R.L. and W.Z. performed the analyses. Y.G., R.L., W.Z., M.Z., B.X., R.N., S.R., J.Z., S.P., S.L. and X.X. wrote and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, M., Cheng, X., Lin, R. et al. Chromosomal-level genome assembly of potato tuberworm, Phthorimaea operculella: a pest of solanaceous crops. Sci Data 9, 748 (2022). https://doi.org/10.1038/s41597-022-01859-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-022-01859-5