Abstract
In a clinical contest, it is common to use dedicated phantoms to perform quality assurance test to check the performance of a SPECT system. Some of these phantoms are also used to calibrate the system for dosimetric evaluation of patients undergoing radiometabolic cancer therapy. In this work, a 3D-OSEM reconstructed 177Lu SPECT dataset of a homogeneous cylindrical phantom is described. This dataset was acquired to investigate the variation of the SPECT calibration factor, counts convergence, noise and uniformity by varying the number of subsets and iterations. In particular, the dataset is composed of images reconstructed using five different numbers of subsets and sixteen different numbers of iterations, for a total of 80 different configurations. The dataset is suitable for comparison with other reconstruction algorithms (e.g. FBP, MLEM, etc.) and radionuclides (e.g. technetium, yttrium). In regards to the uniformity issue, the same dataset allows the user to perform radiomic investigations on the influence of the border effect on the reconstructed images.
Measurement(s) | Calibration Factor • coefficient of variation • Texture Descriptor |
Technology Type(s) | Single Photon Emission Computed Tomography • Image Feature |
Similar content being viewed by others
Background & Summary
Single Photon Emission Computed Tomography/Computed Tomography (SPECT/CT) is a common clinical hybrid system allowing the acquisition of anatomical (CT) and functional (SPECT) information of patients undergoing radio-drug administration for cancer diagnosis and/or therapeutic purposes. For instance, molecular radiotherapy based on peptide receptor radionuclide therapy is a well-assessed method for the treatment of neuroendocrine tumors1,2,3,4. Moreover, peptides labelled with Lutetium-177 (177Lu) have an established use in the treatment of this disease5,6,7,8,9,10, and also with theranostic radiopharmaceuticals in metastatic prostate cancer patients11,12,13,14.
Typically, SPECT/CT acquisitions are reconstructed using of commercial software that allow to fuse SPECT and CT acquisition in one single three-dimensional (3D) dataset. However, even if there have been improvements of different imaging modalities for the detection and diagnosis of human diseases, a mandatory step before performing personalized dosimetry is the evaluation of accurate quantitative information from reconstructed images15,16. In particular, for SPECT/CT acquisitions, the recommendations outlined on the Medical Internal Radiation Dose (MIRD) pamphlet No. 2317 and MIRD No. 2618 highlight the importance of the SPECT image reconstruction process based on phantoms and patients, but do not state uniquely defined procedures and protocols. This is also supported by different studies that are focused on different factors influencing the SPECT reconstruction process, for instance attenuation and scatter correction19, collimator type and energy window20, reconstruction parameters and photo-peak choice21.
Moreover, there has been an increase in the diagnostic information within a single study and the possibility of extracting “quantitative” features from both morphological and functional tomographic images. Radiomics22 is the discipline that studies and analyses the features extracted from medical images, in order to generate a predictive model combining the information provided by the “quantitative” features with those of patient outcome. Although many studies address the usefulness of radiomic investigations in a clinical context23,24,25,26, only a few once deal with their use for the quantitative evaluation of SPECT images27.
In this contest, our dataset has a twofold aim: analysing the influence of reconstruction parameters usually used in the SPECT image reconstruction process, and investigating possible relationships between radiomic analysis and reconstruction parameters. For these aims, a homogeneous phantom filled with a known amount of 177Lu was acquired on a SPECT/CT system used in our clinical contest. The original SPECT acquisition was reconstructed using 3D Order Subsets Expectation Maximization (3D-OSEM) algorithm, considering different numbers of subsets and iterations. This dataset was used in a first study28 to analyse the total counts convergence, calibration factors and noise level as a function of the 3D-OSEM reconstruction parameters. Moreover, in a second study29, the same dataset has been analysed using a free downloaded radiomic software30: a statistical investigation of the radiomic feature trends as a function of the reconstruction parameters was performed in order to explore the potential role of radiomics as a new approach for quantitative characterization of 177Lu SPECT images.
It must be said that the proposed dataset has some limitations. From the point of view of nuclear medicine, it is a part of the acquisitions that must be performed on a SPECT system to implement protocols relating to patient-specific dosimetry. Principally our dataset is suitable to define the calibration factor useful to convert SPECT detected counts in terms of activity, and to assess the level of noise obtainable in the reconstructed SPECT/CT images. Moreover and strictly related with the latter aim, from the point of view of radiomic analysis it is important to point out another issue. In fact, the proposed dataset is a texture analysis of noise in the reconstructed SPECT/CT images, and if this could be a limitation, in a practical nuclear medicine contest it is essential for evaluation of possible non-uniformities that can be introduced by the SPECT system components and tomographic reconstruction process. This latter reason is even more important as recently it was proposed using a uniform phantom for SPECT calibration, within the context of personalized dosimetry for molecular radiotherapy28.
Methods
Phantom characteristics
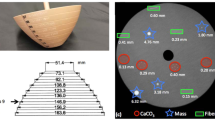
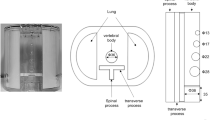
A 6.4 l cylindrical homogeneous phantom (Data Spectrum Corporation, Hillsborough, USA) was used to perform the calibration of a SPECT system. For this purpose, it was filled with a 177Lu concentration of about 0.11 MBq/ml (at the time of SPECT acquisition), mimicking a patient administered with 190 MBq. The accuracy of activity was assessed by means of the clinically available dose calibrator, with an uncertainty of about 5%.
SPECT/CT acquisition
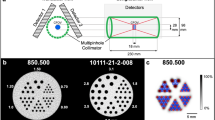
The acquisition was performed by means of a hybrid SPECT/CT system (Discovery NM/CT 670, GE Healthcare, Milwaukee, USA), equipped with two gamma detector heads (9.5 mm NaI(Tl) crystal thickness of 40 cm axial by 54 cm diameter field of view), and an integrated CT component identical to a 16-slice-CT used in diagnostic CT imaging (model: Bright Speed 16, GE Healthcare, Milwaukee, USA). The following SPECT acquisition settings were used: 120 projections with 180° mode detector head, 30 seconds per projection, non-circular step-and-shoot acquisition orbit, 128 × 128 pixel matrix, with 4.42 × 4.42 mm pixel size. For scatter correction, projection data were acquired in three energy windows using a parallel-hole medium energy general purpose (MEGP) collimator: a symmetrical 20% wide energy window was centred at 208 keV177Lu photopeak (energy window: 187.2 keV − 228.8 keV), together with two 8.7% and 11.8% wide adjacent scatter windows, providing the upper and lower scatter windows, respectively. The SPECT acquisition was followed by a CT scan (120 kV, 80mAs, 1.375 pitch, 16 × 1.25 mm collimation, 3.75 slice thickness reconstruction), and the acquired images were reconstructed with filtered back projection algorithm using the default convolution kernel for routine low dose CT (LD-CT) examination of the abdomen.
SPECT/CT reconstruction
SPECT/CT data processing was carried out on a dedicated workstation (Xeleris 3.1108, GE Healthcare, Milwaukee, USA), provided with a software from the same vendor (Dosimetry Toolkit Package, GE Healthcare, Milwaukee, USA). The SPECT images were reconstructed by means of 3D-OSEM algorithm including resolution recovery, scatter correction and attenuation correction (performed by linear attenuation coefficient (μ) maps estimated from the acquired LD-CT). Reconstruction of SPECT images was performed by considering 5, 10, 15, 20 and 30 subsets, with a number of iterations from 1 to 7 with an incremental step of 1, and from 10 to 50 with an incremental step of 5. Hence, a total of 80 combinations have been considered. No pre- and post-reconstruction filters were used. At the end of the reconstruction process, the software re-bins the CT matrix to the SPECT one giving a 256 × 256 pixel matrix dataset, with a 2.21 mm isotropic voxel. In Fig. 1 some 3D-OSEM reconstructed SPECT images of the cylindrical phantom considered are reported. These reconstructions are provided in a higher number with respect to our previous published studies and also the methods are an expanded version of the ones described in our related work28
Data Records
A public figshare collection called “2021_MezzengaEmilio_Collection1”31 was created. It is structured into eight folders, named and organized as follow:
-
“2021_EM_Original_CT_Data”: the folder contains 108 DICOM files related to the original CT sequence of the phantom, named as: “CTxxx.dcm”, with xxx = 001, 002, …, 108.
-
“2021_EM_Original_SPECT_Data”: the folder contains three DICOM files related to the SPECT acquisition. In particular, the triple window (TW) file (“TW.dcm”), the lower scatter window (LScW) file (“LScW.dcm”), and the higher scatter window (HScW) file (“HScW.dcm”). The TW file contains all three acquisition-windows (i.e. photopeak window, high and low scatter window), while LScW and HScW contain the lower and higher scatter acquisition window, respectively.
-
“2021_EM_Isotropic_CT_Data”: the folder contains 182 DICOM files related to the reconstructed CT datasets named as: “IsotropicCT001_CTyyy.dcm”, with yyy = 001, 002, …, 182.
-
“2021_EM_Reconstructed_SPECT_Subset_##” (with ## = 05, 10, 15, 20, 30): each folder (a total of five folders) contains 16 DICOM files. The files are named as “IsotropicNM##SzzI_DS.dcm”, with ## referring to the numbers of subsets used in the SPECT 3D-OSEM reconstruction process, and with zz = 01, 02, 03, 04, 05, 06, 07, 10, 15, 20, 25, 30, 35, 40, 45, 50 referring to the numbers of iterations used in the SPECT 3D-OSEM reconstruction process.
The collection is publicly available at: https://doi.org/10.6084/m9.figshare.c.5468097
Technical Validation
The starting dataset was acquired in compliance with the international protocols related to calibration/acquisition of SPECT images. Today the SPECT system is clinically used in our institute for diagnosis and therapies for different cancer patients.
The starting dataset has been reconstructed and prepared to enable the user on performing evaluations by means of MATLAB (The Mathworks, Inc., MA, US) based on 3D-OSEM total counts convergence, SPECT calibration factor, noise evaluation and radiomic analysis versus the reconstruction parameters used (i.e. number of subsets and iterations).
On regard the nuclear medicine contest, a cylindrical volume of interest (VOI) can be drawn inside the phantom and coaxially with the same, fixing its perimeter at a minimum distance of 3 cm from the inner walls of the phantom. The total, mean and standard deviation of voxel counts can be computed inside the chosen VOI and they can be plotted as a function of the 3D-OSEM reconstruction parameters. In particular, the total voxel counts versus the number of iterations, fixing the number of subsets, can be plotted to evaluate when the 3D-OSEM has reached convergence of total counts inside the chosen VOI. Moreover, normalizing the total counts by the SPECT acquisition time (30 minutes in the acquired dataset) and the phantom’s activity concentration, the SPECT calibration factor related to the acquired phantom can be computed as a function of the 3D-OSEM reconstruction parameters.
Computing the average (μ) and standard deviation (σ) of voxel counts in the same VOI, the coefficient of variation (COV, the percentage ratio between σ and μ) can be computed and plotted as a function of the 3D-OSEM reconstruction parameters, evaluating the level of noise present in the reconstructed images.
Considering the radiomic analysis, the proposed dataset can be used to evaluate the dependence of radiomic features with respect to the VOIs used inside the phantom and their location versus the 3D-OSEM reconstruction parameters.
Usage Notes
The datasets proposed could be used to create a dedicated analysis protocol on the same phantom filled by means of different radionuclides with the aim to:
-
investigate the total SPECT counts convergence varying the reconstruction parameters of the algorithm implemented on the reconstruction workstation;
-
evaluate the noise level in the reconstructed SPECT images as a function of numbers of subsets and iterations;
-
use different radiomic software in order to investigate the variation of radiomic matrices and related features for different volumes of interest, and related locations inside the phantom. On this regard, the method used for this radiomic analysis are described in our related work29.
Code availability
The software used for the SPECT/CT data reconstructions was the commercial Dosimetry Toolkit Package (GE Healthcare, Milwaukee, USA). No custom codes have been designed and used for this study.
References
Dash, A., Chakraborty, S., Pillai, M. R. & Knapp, F. F. Jr. Peptide receptor radionuclide therapy: an overview. Cancer Biother Radiopharm. 30, 47–71 (2015).
Molina Trinidad, E. M. & Salas Casas, A. Somatostatine analogs, how biomarkers in the diagnostic and treatment for cancer and others damages. Int J Pharm Sci Rev Res. 27, 31–46 (2014).
Otte, A. et al. Yttrium-90-labelled somatostatin-analogue for cancer treatment. Lancet. 351, 417–8 (1998).
Marincek, N. et al. Somatostatin-based radiotherapy with [90Y-DOTA]-TOC in neuroendocrine tumors: long-term outcome of phase I dose escalation study. J Transl Med. 15, 11–17 (2013).
Paganelli, G. et al. 177 Lu-Dota-octreotate radionuclide therapy of advanced gastrointestinal neuroendocrine tumors: results from a phase II study. Eur J Nucl Med Mol Imaging. 41, 1845–1851 (2014).
Romer, A. et al. Somatostatin-based radiopeptide therapy with [177Lu-DOTA]-TOC versus [90Y-DOTA]-TOC in neuroendocrine tumours. Eur J Nucl Med Mol Imaging. 41, 214–222 (2014).
Ezzidin, S. et al. Outcome of peptide receptor radionuclide therapy with 177Lu-octreotate in advanced grade ½ pancreatic neuroendocrine tumours. Eur J Nucl Med Mol Imaging. 41, 925–933 (2014).
van der Zwan, W. et al. GEPNETs update: Radionuclide therapy in neuroendocrine tumors. Eur J Endocrinol. 172, R1–8 (2015).
Ilan, E. et al. Dose response of pancreatic neuroendocrine tumors treated with peptide receptor radionuclide therapy using 177Lu-DOTATATE. J Nucl Med. 56, 177–82 (2015).
Bodei, L. et al. Peptide receptor radionuclide therapy with 177Lu-Dotatate: the IEO phase I-II study. Eur J Nucl Med Mol Imaging. 38, 2125–35 (2011).
Schäfer, M. et al. Preclinical evaluation of a tailor-made DOTA-conjugated PSMA inhibitor with optimized linker moiety for imaging and endoradiotherapy of prostate cancer. J Nucl Med. 56, 914–20 (2015).
Afshar-Oromieh, A. et al. The novel theranostic PSMA ligand PSMA-617 in the diagnosis of prostate cancer by PET/CT: biodistribution in humans, radiation dosimetry and first evaluation of tumor lesions. J Nucl Med. 56, 1697–705 (2015).
Kratochwil, C. et al. [177Lu]Lutetium-labelled PSMA ligand-induced remission in a patient with metastatic prostate cancer. Eur J Nucl Med Mol Imaging. 42, 987–8 (2015).
Pillai, M. R. A. & Knapp, F. F. Lu-177 labeled therapeutics: 177Lu-PSMA is set to redefine prostate cancer treatment. Curr Radiopharm. 9, 6–7 (2016).
Ritt, P., Vija, H., Hornegger, J. & Torsten, K. Absolute quantification in SPECT. Eur J Nucl Med Imaging. 38(Suppl 1), S69–77 (2011).
Sgouros, G. & Hobbs, R. F. Dosimetry for radiopharmaceutical therapy. Semin Nucl Med. 44, 172–8 (2014).
Dewaraja, Y. K. et al. MIRD Pamphlet No. 23: Quantitative SPECT for Patient-Specific 3-Dimensional Dosimetry in Internal Radionuclide Therapy. J Nucl Med. 53, 1310–25 (2012).
Ljungberg, M., Konijnenberg, M. W., Eckerman, K. F., Dewaraja, Y. K. & Sjögreen-Gleisner, K. MIRD Pamphlet No. 26: Joint EANM/MIRD Guidelines for Quantitative 177Lu SPECT Applied for Dosimetry of Radiopharmaceutical Therapy. J Nucl Med. 57, 151–62 (2016).
Beauregard, J. M., Hofman, M. S., Pereira, J. M., Eu, P. & Hicks, R. J. Quantitative 177Lu SPECT (QSPECT) imaging using a commercially available SPECT/CT system. Cancer Imaging 11, 56–66 (2011).
de Nijs, R., Lagerburg, V., Klausen, T. L. & Holm, S. Improving quantitative dosimetry in 177Lu_DOTATATE SPECT by energy window-based scatter corrections. Nucl Med Comm 35, 522–33 (2014).
Sanders, J. C., Kuwert, T., Hornegger, J. & Ritt, P. Quantitative SPECT/CT imaging of 177Lu with in vivo validation in patients undergoing peptide receptor radionuclide therapy. Mol Immaging Biol. 17, 585–93 (2015).
Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: images are more than pictures, they are data. Radiology 278, 563–577 (2015).
Ganeshan, B., Panayiotou, E., Burnand, K., Dizdarevic, S. & Miles, K. Tumour heterogeneity in non-small cell lung carcinoma assessed by CT texture analysis: A potential marker of survival. Eur Radiol. 22, 796–802 (2012).
Liu, B. et al. Fully automatic and segmentation-robust classification of breast tumors based on local texture analysis of ultrasound images. Pattern Recognit. 43, 280–298 (2010).
Wibmer, A. et al. Haralick texture analysis of prostate MRI: utility for differentiating non-cancerous prostate from prostate cancer and differentiating prostate cancers with different Gleason scores. Eur Radiol. 25, 2840–2850 (2015).
Yu, H. et al. Automated Radiation Targeting in Head-and-Neck Cancer Using Region-Based Texture Analysis of PET and CT Images. Int J Radiat Oncol Biol Phys. 75, 618–625 (2009).
Nichols, K. J., Di Filippo, F. P. & Palestro, C. J. Texture analysis for automated evaluation of Jaszczack phantom SPECT system tests. Medical Physics. 46, 262–272 (2018).
Mezzenga, E. et al. Quantitative accuracy of 177Lu SPECT imaging for molecular radiotherapy. PLoS ONE 12, e0182888 (2017).
Sarnelli, A. et al. Texture analysis in 177Lu SPECT phantom images: Statistical assessment of uniformity requirements using texture features. PLoS ONE 14, e0218814 (2019).
Fang, Y.H. et al. Development and evaluation of an open-source software package “CGITA” for quantifying tumor heterogeneity with molecular images. Biomed. Res. Int. (2014).
Mezzenga, E. 2021_MezzengaEmilio_Collection1, figshare https://doi.org/10.6084/m9.figshare.c.5468097.v1 (2021).
Acknowledgements
We are particularly grateful to Paola Chiaramida (Product Clinical Specialist in Molecular Imaging, GE Healthcare) for her support and valuable suggestions related to SPECT system and Xeleris workstation.
Author information
Authors and Affiliations
Contributions
E.M. study conceptualisation, collected the raw images, performed image reconstruction, wrote the paper. F.P. image collection uploading and structuring, revised the paper. E.L. image consistency analysis, revised the paper. M.L.B. collected the raw images, revised the paper. A.S. designed the study, wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mezzenga, E., Piccinini, F., Loi, E. et al. Reconstructed SPECT images of 177Lu homogeneous cylindrical phantom used for calibration and texture analysis. Sci Data 9, 412 (2022). https://doi.org/10.1038/s41597-022-01535-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-022-01535-8