Abstract
Besides being central for understanding both global biodiversity patterns and associated anthropogenic impacts, species range maps are currently only available for a small subset of global biodiversity. Here, we provide a set of assembled spatial data for terrestrial vascular plants listed at the global IUCN red list. The dataset consists of pre-defined native regions for 47,675 species, density of available native occurrence records for 30,906 species, and standardized, large-scale Maxent predictions for 27,208 species, highlighting environmentally suitable areas within species’ native regions. The data was generated in an automated approach consisting of data scraping and filtering, variable selection, model calibration and model selection. Generated Maxent predictions were validated by comparing a subset to available expert-drawn range maps from IUCN (n = 4,257), as well as by qualitatively inspecting predictions for randomly selected species. We expect this data to serve as a substitute whenever expert-drawn species range maps are not available for conducting large-scale analyses on biodiversity patterns and associated anthropogenic impacts.
Measurement(s) | species distributions |
Technology Type(s) | machine learning |
Sample Characteristic - Organism | Tracheophyta |
Sample Characteristic - Environment | terrestrial natural environment |
Similar content being viewed by others
Background & Summary
Life on Earth is essential to human society as it forms the foundation of present welfare1. The growing human population, modern lifestyles and associated pressures on the planet have already resulted in a significant loss of natural habitat and are threatening biodiversity2,3,4,5,6. Different initiatives promote the protection of biodiversity and aim to halt its loss, such as the UN Sustainable Development Goals7, the Intergovernmental Science-Policy Platform on Biodiversity and Ecosystem Services8 and the International Union for the Conservation of Nature (IUCN). Different decision-support tools can contribute to this by assessing environmental performances of products, strategies and policies2,9,10,11. For the development of such tools, but also for the implementation of global conservation strategies and policies itself, spatial data, e.g. in the form of distribution maps of individual species12, are crucial. However, besides many species remaining undiscovered or undescribed, we still lack spatial information for most of the ones we know13. Consequently, comprehensive and ready-to-use datasets for large-scale analyses are only available for a few vertebrate groups14,15,16. This is concerning, as global conservation strategies and biodiversity impact assessments are limited to these groups, while some hyperdiverse species groups, such as plants, are often not considered17,18.
Here, we provide spatial distribution data for a large fraction of red-listed terrestrial vascular plant species at different levels of spatial detail (Fig. 1), i.e. native regions (n = 47,675), occurrence records (n = 30,906) and modelled range estimates (i.e. a predicted relative environmental suitability19 within native regions; n = 27,208). The workflow included data scraping and filtering, as well as variable selection, model calibration and model selection, aiming for best practice20,21,22 but within the constraints of data limitations and computational feasibility at this scale. Species-specific native regions were retrieved from a scheme specifically developed to challenge the lack of distributional knowledge for plant species23. Available native occurrence records were retrieved from the Global Biodiversity Information Facility (GBIF)24 and subsequently filtered. Range estimates were generated using maximum entropy modelling19,25,26,27, and show where environmentally suitable conditions exist within each species’ native regions (Fig. 2a–d).
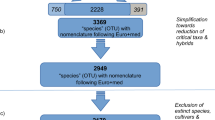
Schematic summary of the dataset. Top: Native region extents were retrieved from Kew’s Plants of the World online. Middle: Occurrence data was retrieved from the Global Biodiversity Information Facility (GBIF)24 and filtered into three different occurrence data types: raw data (blue), presence cells (grey) and thinned data (yellow). Bottom: The different occurrence data types were used in Maxent models to predict relative environmental suitability indices within native regions (i.e. range estimates). Differences between Model 0 and Model 1 to 3. Model 0 was trained to support variable selection using raw data in k-fold cross validated Maxent models (one model for each combination of feature classes, i.e. linear (L), quadratic (Q), hinge (H), product (P) and threshold (T)). The selected variables and each of the three occurrence data types were used to train a set of separate k-fold cross validated Maxent models (one model for each possible combination of feature classes, regularization multipliers and occurrence data type). The overall best performing model was selected for each species based on performance metrics.
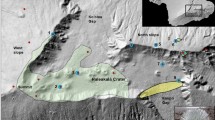
Data examples for randomly selected species and spatial coverage of the dataset. Best performing Maxent prediction, highlighting environmentally suitable conditions within the species native regions (i.e. modelling extent) along retrieved occurrence records (white points) for (a) Amomum pterocarpum, (b) Cedrus libani, (c) Laburnum anagyroides, (d) Megistostegium nodulosum. Performance of the shown predictions indicated by maximum F1-score and the area under the receiver operating characteristics curve for true vs. false positive rate (AUC) and recall vs. precision (AUCPR). Bottom: number of (e) retrieved native regions, (f ) retrieved occurrence records, and (g) generated Maxent predictions across the globe.
The underlying occurrence data is known to be highly spatiotemporally aggregated and variable across administrative borders for some species28,29,30,31. We aimed at counteracting a potential sampling bias by using three differently treated occurrence data types (i.e. different degree of spatial filtering: no filter, presence cells, thinned presence cells), and by dividing occurrence data in equally-sized bins during model calibration32. Up to 96 different models were fitted per species to find optimal variables, model settings and data type. The best prediction was selected for each species based on common performance metrics (i.e. AUC and AUCPR).
However, some predictions will undoubtedly remain flawed by underlying biases. Based on comparisons to expert-drawn range maps available from IUCN (n = 4,257) and qualitative inspection of predictions for randomly selected species, we expect this to mainly influence widespread and common species, and hence, only affect the smallest proportion of global biodiversity33. In addition, the species most vital for assessing anthropogenic impacts or for defining conservation priorities, are more likely to be small-ranged and endemic. Although validating each prediction was not feasible, we found most individually inspected predictions to either offer an improvement compared to elsewhere available data or an acceptable substitute, although at a coarser spatial resolution and less detailed.
We want to stress that the presented dataset is generated for the purpose of global spatial screening studies and for building a basis for future, global biodiversity impact assessment models. In concert with powerful, species-specific trait and conservation-related databases, the provided data can benefit future work, such as assessing global extinction probabilities34, effects of terrestrial acidification35, drivers of invasion success36, progress towards reaching global conservation goals37 and act as pre-assessment prior to expert-based range map generation and red list assessments38,39,40,41. With a continuously increasing availability of species occurrence records, the presented dataset can be updated frequently to illustrate the state of knowledge at any time. With more data becoming available, precision is likely to increase in the future.
Methods
Taxonomic scope
A species list containing all terrestrial vascular plants (n = 52,372) of the global IUCN red list was retrieved from IUCN in April 2021, IUCN version 2021-116. We retrieved each species’ accepted name from Plants of the World Online (POWO)42 to facilitate communication to various data portals using the package taxize43 in R44. Plant family, order and class were retrieved from the Integrated Taxonomic Information System45 using the package taxize43 in R. Only species outside the IUCN threat categories “Extinct” and “Extinct in the Wild” were kept, and all species considered as subspecies or varieties according to POWO removed. We attempted to assemble spatial data for each of the remaining 48,144 species.
Native regions
Species-specific native regions (Fig. 1) were retrieved from POWO using a customized web-scraper function (see section Code Availability) and the packages taxize43 and rvest46 in R. The data follows the World Geographical Scheme for Recording Plant Distributions (WGSRPD)23 and includes a continental, country and regional level. Retrieved WGSRPD-regions were matched to its corresponding shapefile at level 4, available from the Biodiversity Information Standards GitHub repository47 and rasterized at 30 arc minutes spatial resolution (approximately 56 km at the equator).
Occurrence records
For species with given native extents in POWO, the maximum number of most recent occurrence points (i.e. 100,000) per native WGSRPD-country was retrieved from the GBIF application programming interface (API) using the package rgbif 48 in R (the equivalent full dataset49 is available at https://doi.org/10.15468/dl.uvd56q). The considered environmental variables have changed tremendously in the past decades50,51 and only cover a limited period of time, i.e. the years 1979–2013 and 2015 respectively (see section Environmental data). Therefore, only records between the years 2000 and 2020 were considered to temporally align occurrence data to both sets of environmental variables as best as possible. If less than 25 records were available for a given species after the year 2000, no temporal filter was set to maximize data retrieval. GBIF records without specified coordinates and with flagged geospatial issues48 were not considered. As such, we expect inaccurate coordinate notations as well as records of specimens preserved in museums or other biodiversity facilities to be typically detected. Only points inside reported native WGSRPD-regions were kept and duplicated records were removed (hereafter: raw data). The number of raw data records was counted per cell (30 arc min.) using the package raster52 in R.
Maxent predictions
We generated spatial predictions within species’ native WGSRPD-regions at 30 arc min. resolution (approximately 56 km at the equator) using maximum entropy modelling (Maxent)19,26,27, for all species with at least 5 raw data records53,54 that were distributed across at least 3 cells, and a native region extent of at least 9 cells. Although an arbitrary threshold, we attempted to allocate computational resources to more meaningful predictions, modelled across larger extents. Maxent is a probability density estimation approach widely used for predicting species distributions based on presence-only data55. Background information, required to fit response curves56, was collected from each cell within each species’ native regions57. For generating models we utilized a high-performance computing infrastructure58 allowing for parallel computations using the Maxent software25 via R packages dismo59 and ENMeval60.
Environmental data
We downloaded all CHELSA bioclimatic variables61,62 (n = 19, see Table 1 for full list) in 30 arc seconds resolution and aggregated, for computational efficiency, to the chosen modelling resolution (30 arc min.) by averaging. CHELSA bioclimatic variables are a set of modelled, biologically relevant, climatic variables based on data collected during the years 1979–201361. In addition, fractions for different natural land cover types, including different types and mosaics of forest, shrubland, grassland and sparse vegetation, (n = 17, see Table 1 for full list) were calculated based on the European Space Agency’s land cover product for the year 2015 in 300 m resolution63. Each land cover class was transformed into a binary raster depicting presence (=1) and absence (=0) of the land cover type. The binary raster was then aggregated to modelling resolution by averaging, resulting in one raster for each land cover class, representing the proportion of land covered by that class per pixel.
Occurrence data types
For some species, several raw data records can be in the same cell at the given spatial resolution (30 arc min.). Although pseudo-replication can inflate model performance (here: during model calibration) and, hence, increases the risk of overfitting, we argue that these occurrence points still contain valid information if they are discrete observations and therefore kept this data. However, we henceforth applied two filters to counteract potential spatial biases, as well as pseudo-replication (Fig. 1). We removed all cell-duplicates from the raw data (hereafter: presence cells), and we applied spatial thinning with a minimum distance of two cells on the presence cells (hereafter: thinned data). Occurrence data was spatially filtered using the R package spThin64.
Model training
A set of Maxent models was fitted for each species using the differently treated occurrence data types. All models were calibrated using k-fold cross validation. The employed occurrence data was partitioned into training and testing bins. For species with only few data points (n < 25), we used k - 1 Jackknife partitioning (k = n)54. For species with more data points (n ≥ 25) we used block partitioning (k = 4) to account for spatial autocorrelation of occurrence points in larger datasets32. This partitioning splits the occurrence data at a longitudinal and latitudinal line, resulting in approximately equally sized bins60.
An initial model (Fig. 1; Model 0) was trained to support the selection of uncorrelated environmental variables using the raw data and all environmental variables (n = 36) for each species. Separate models, one for each possible combination out of all included feature classes (i.e. environmental variables and transformations thereof), were trained. We included linear (l), quadratic (q), product (p), hinge (h) and threshold (t) transformations, resulting in 6 possible combinations (i.e. l, lq, h, lqh, lqhp, and lqhpt). The best performing model was selected based on the corrected Akaike information criterion (AICc)65,66,67. However, if no model performed best in terms of AICc, or if this metric was unavailable for 50% of fitted models, the average testing area under the receiver operating characteristics curve (AUC; see section Technical Validation) during model calibration was used instead. Permutation importance was retrieved for all variables in Model 0. Correlated variables were identified using Spearman’s rank correlation coefficient (ρ) and defined as ρ ≥ | ± 0.7|. In any set of correlated variables, only the variable with the greatest permutation importance was kept.
The selected environmental variables were used to train separate models for each of the three differently treated occurrence data types: raw data (Model 1), presence cells (Model 2), and thinned data (Model 3). Model 1 was trained if at least 5 raw data records were available, distributed across at least 3 cells (see above). Model 2 and Model 3 were trained if at least 3 records of the corresponding data type were available to avoid computational failure. Although a smaller sample size, we argue that if those models performed better than Model 1, the threshold of 5 records becomes arbitrary and the assessed performance indicators (see section Technical Validation) more valuable. The same model architecture as in Model 0 was utilized, including model calibration and selection of the best performing model. However, this time, we added five different regularization multipliers (RM; i.e. 1, 2, 3, 5 and 10; based on previous studies68,69,70) to counteract overfitting20,56 and for building simpler, ecologically more relevant, models60. Hence, separate models for each possible combination out of feature classes and RMs were trained (Fig. 1; Model 1–3), resulting in 30 trained models for each data type and up to 90 models per species.
Metadata
Metadata was assembled for all data and includes general information about species (taxonomy and red list status), provided data type (native regions, occurrence records or Maxent prediction), bounding box of native regions, and if relevant, information about the occurrence data (number of raw data records, Moran’s Index71, calculated as a measure of spatial autocorrelation and based on the number of raw occurrence points obtained per cell), and Maxent metadata: training data (filter treatment, number of training data points), thresholds for converting the prediction into binary range maps59, model settings (features, parameters, transformations, regularization multiplier, variables) and out of the box60 model performance, including degree of overfit (DOO) quantified as the difference between calibration and testing AUC during k-fold cross validation70, as well as self-assessed model performance metrics as described in the section Technical Validation.
Data Records
Dataset
The presented dataset is stored in a stable Dryad Digital Repository72 and can be explored at https://plant-ranges.indecol.no. The dataset includes spatial information for 47,675 species at different levels of detail. In total, range estimates (i.e. relative environmental suitability within native regions) have been predicted for 27,208 species using Maxent, for 30,906 species native occurrence records are provided, and for 47,675 species the spatial extent of its native WGSRPD-regions is provided.
All gathered and generated data are stored in netCDF files and can be called by specifying a varname. Spatial predictions are provided in Maxent’s raw as well as default output (i.e. complementary log-log (cloglog) transformed, but see section Usage Notes)27,59,60. The suggested data is stored in folder basic. These netCDF files (default output and raw output) assemble the best performing Maxent prediction (varname: Maxent prediction) for each species selected based on the highest harmonic mean between AUC and AUCPR (see Technical Validation), along with number of occurrence records per cell (varname: Presence cells) and rasterized native WGSRPD-regions (varname: Native region).
The netCDF files in folder advanced contain one Maxent prediction for each occurrence data type (varname: Model 1, Model 2 or Model 3), instead of best performing Maxent prediction (i.e. varname Maxent prediction is not applicable). Number of occurrence records per cell (varname: Presence cells) and rasterized native WGSRPD-regions (varname: Native region) are identical in all netCDF files.
Each band in the netCDF files assembles the mentioned variables for one species. The corresponding bands can be looked up in the metadata (i.e. speciesID). Furthermore, the metadata can be used to select appropriate cut-off thresholds for generating binary range maps, filter models based on species, performance, or desired datatypes, and to lookup the relevant study extent for masking individual predictions (see Usage Notes).
Technical Validation
Maxent predictions
We calculated performance metrics for model 1 to 3 for each species using its corresponding presence cells to validate the Maxent predictions. Receiver operating characteristic curves and the corresponding area under the curve for recall (i.e. true positive rate, sensitivity) versus false positive rate (AUC) as well as precision versus recall (AUCPR) were generated using the packages ROCR73 and PRROC74 in R. Recall was calculated as the fraction of correctly predicted presence cells compared to all presence cells of the reference (Eq. 1), the false positive rate as the fraction of falsely assigned presence cells compared to all true absence cells (Eq. 2), and precision as the fraction of correctly assigned presence cells compared to all predicted presence cells (Eq. 3). In addition, F1-scores (Eq. 4) were calculated as harmonic mean between recall and precision at all possible cut-off thresholds to transform the Maxent prediction into a binary range map. The maximum obtained F1-score indicates how well a potential binary range map performs at equal importance of recall and precision.
AUC and AUCPR are threshold-independent performance measures for binary classifiers. An AUC value of 1 indicates a perfect model, an acceptable AUC value (>0.7)75 indicates the ability to predict many true presences at a low false positive rate, and an AUC value of 0.5 indicates the model performing as good as a random guess. The average AUC obtained across the suggested dataset was 0.95 when comparing predictions to its corresponding presence cells (Table 2), indicating well-performing models for the majority of species. For 26,977 species (99%), at least one Maxent prediction had an AUC value above 0.775.
AUCPR is not affected by true negatives (i.e. true absence) which often dominated our dataset. A higher AUCPR value indicates a relatively higher ability to correctly predict a high proportion of presumably true range while maintaining a high precision compared to a lower AUCPR. However, the AUC and AUCPR values, as well as max. F1-score, described here were calculated based on presence-background data and are highly influenced by class balances. Strictly speaking, both false presences and true absences cannot be determined with presence-only data. Hence, the performance metrics described here can only be used to compare different models for a given species, but not across different species76,77.
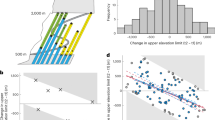
Therefore, we evaluated the Maxent predictions by comparison to available expert-based range maps, as an additional evaluation dataset32. Expert-based range maps were retrieved from IUCN, if available (hereafter: reference ranges). Only reference ranges that were labelled as “native” and “extant (resident)” or “probably extant (resident)” were considered. For 4,257 species of our Maxent predictions, range maps were available at IUCN. These species were unevenly distributed in space (Fig. 3a), across IUCN red list categories (Fig. 3d) as well as the plant classes dicots (Magnoliopsida, n = 3,480), monocots (Liliopsida, n = 731), ferns (Polypodiopsida, n = 27), conifers (Pinopsida, n = 17), and lycopods (Lycopodiopsida, n = 2). Reference ranges were used to calculate the above described performance measures (i.e. max. F1-score, AUC and AUCPR). However, this time we dealt, presumably, with actual presences and absences of the given species, making the performance metrics comparable across species76. Maxent predictions for species classified as “data-deficient” (DD) obtained the lowest, and predictions for species classified as “near-threatened” (NT), “vulnerable” (VU) and “endangered” (EN) the highest AUC values (Fig. 3d). However, these differences were marginal and all average values consistently high across different IUCN categories (mean AUC: 0.9; Table 2) and across the globe (Fig. 3b). Although AUC is a strong indication of model performance75, the predictions seem to rarely accommodate both a high recall and a high precision (represented in either max. F1-score or AUCPR value) when compared to reference ranges. However, we found a large variation and no clear trend in AUCPR values for species across different threat-level categories (Fig. 3d), and although the average AUCPR was lowest for species native to parts of central Africa, India and south-eastern Asia (Fig. 3c), we expect these values to be of little explanatory power due to the limited sample sizes in these regions (Fig. 3a). Moreover, AUCPR seems to increase with increasing data availability (Fig. 3d). We assume that low data coverage in sparsely populated areas influenced modelling performance for some, primarily widespread, species, highlighting that sometimes more spatially distributed occurrence data is required for making expert-alike range maps78.
Performance metrics for the suggested Maxent predictions. (a) Number of reference range maps available used for calculating performance metrics. Average values for species native to the corresponding regions of area under the receiver operating characteristics curve for (b) true vs. false positive rate (AUC) and (c) recall vs. precision (AUCPR). (d) Mean and standard deviation of AUC (blue) and AUCPR (yellow) per rounded log-transformed number of raw occurrence data points (left) and for species in different IUCN red list categories (right), i.e. data-deficient (DD), least concern (LC), near-threatened (NT), vulnerable (VU), endangered (EN) and critically endangered (CR). Significant differences across IUCN categories in d are indicated by different letters in bars for AUC (white text) and AUCPR (black text).
Furthermore, based on a qualitative assessment of predictions for twelve randomly selected species, we expect uncertainties due to differences in data availability across administrative borders as well as for highly naturalized species. For instance, the clustered occurrence records for Cedrus libani in Lebanon (Fig. 2b) resulted in less precise data than elsewhere available for this species79, while the prediction for Laburnum anagyroides (Fig. 2c) was affected by naturalized occurrence records outside its native origin80 but still within its native WGSRPD-regions. However, this will be most problematic for abundant, widespread, and naturalized species, and hence only relevant for the smallest fraction of global biodiversity33. In addition, the predictions for more vulnerable species, presumably small-ranged or endemic, seem to perform better than species in the lowest red list category (i.e. least concern (LC)) in terms of AUC when compared to reference ranges (Fig. 3d).
In fact, the remaining randomly selected predictions were either consistent with point data (e.g. Terminalia macrostachya81), reflected the current knowledge of elsewhere available data, although at a coarser spatial resolution and less detailed (e.g. Mammillaria grahamii82), or offered an improvement compared to previously unavailable spatial data (e.g. Eucalyptus elliptica83, Megistostegium nodulosum84 (Fig. 2d), Memecylon elegantulum85, Psidium salutare86,87, Siparuna conica88,89, Trisetaria dufourei90). However, the prediction of Pyracantha angustifolia was difficult to evaluate due to poorly understood range dynamics91, highlighting the need for more data for vascular plant species.
We want to stress that our predictions indicate environmentally suitable conditions even if isolated from known species occurrence locations. For instance, Amomum pterocarpum seems to be restricted to southern India and Sri Lanka92,93 while our prediction indicates environmentally suitable conditions in north-eastern India (Fig. 2a), which in fact, supports a possible observation nearby94. We further detected several expert-based range maps with a substantial mismatch to our data, confirming that some of the expert-based data may be too conservative95 (e.g. Magnolia pugana)96. However, we also found expert-based ranges being smaller (e.g. Vallesia glabra or Tetraclinis articulata)97,98 than predicted environmental suitability indicates, or being incorrectly georeferenced (e.g. Corylus cornuta)99. Hence, besides highlighting mismatches to expert-based range maps, we expect this dataset to be of sufficient quality to serve as time- and cost-efficient range map substitutes and pre-assessed range estimates for currently unmapped species.
External data
The retrieved native WGSRPD-regions are provided by POWO under a CC BY 3.0 license (https://creativecommons.org/licenses/by/3.0/) and have been checked for consistency to assure proper workflow of data retrieval from POWO and feature matching to the WGSRPD level 4 shapefile. However, the data provider, POWO, cannot warrant the quality or accuracy of the WGSRPD data42. In addition, other data (e.g. ecoregions100) may ecologically be more relevant than administrative boundaries. However, WGSRPD offers the most detailed data on species’ native origins available on a large-scale, to the best of our knowledge. An attempt in matching native WGSRPD-regions to ecoregions was discontinued after loss of information due to incompatible geographical boundaries. Hence, we consider the utilized WGSRPD-regions, currently, as the best compromise between level of detail and availability of data on species’ native origins. Furthermore, spatial inaccuracies and biases in the occurrence data retrieved from GBIF were counteracted by the implemented filtering steps, the coarse spatial resolution, by avoiding non-native occurrence records and the model calibration techniques. However, any unforeseen misclassified or misreported records may flaw predictions for individual species. In addition, data retrieval via GBIF’s API was limited to 100,000 occurrence records per request. We extended this limit by sending one request per native country for each species, and hence, expect this issue to be irrelevant for our study. We further want to stress that most of the generated predictions have not been validated individually, and that some predictions may be erroneous either due to data limitations or simply because digitally stored data can contain minor but crucial blunders. For instance, in terms of nomenclature, the red-listed species Cotoneaster cambricus is endemic to Wales101, but also seems to be a synonym for a widespread species according to POWO42. Consequently, either our spatial prediction or the expert-based range for this species is incorrect.
Usage Notes
All data handling, modelling and visualization was done using R version 4.0.344 in RStudio version 1.4.1103102. Handling of all spatial data was done using the R packages raster, rgdal, maptools, rgeos and sp52,103,104,105,106. A showcase for opening the different data types for individual species, is available at https://github.com/jannebor/plant_range_estimates. Although functionality of the code may be given at newer, or older, versions, we expect the best user-experience using the versions specified in this descriptor.
Maxent predictions are given as raw and cloglog transformed output. These outputs are related monotonically, meaning that the performance metrics described in this study, as well as a potential binary range map (excluding prevalence dependent thresholds), will be identical for both raw and cloglog output56. For users mostly interested in qualitative analyses, both predictions can simply be interpreted as indices of environmental suitability20. However, due to rescaling, the exact interpretation and appearance of each output differs. In general, Maxent’s output interpretation depends on the underlying data, and differs, in our case between Model 1 (raw data including pseudo-replicates = abundance) compared to Model 2 and 3 (presence), but gives an estimate of the abundance, or presence, of the species in relation to the true modelled quantity (either abundance or presence). Maxent’s raw output reflects the exponential Maxent model itself, and can be interpreted as a relative occurrence (or presence) rate summing up to 120. The raw output does not rely on any assumptions20, however, it may not perform well in visualizing actual differences in suitability107. Being rescaled on a more common range from 0 to 1, the cloglog transformation compresses extreme values, and hence facilities visualization and comparison amongst predictions27. It can, arguably, be interpreted as a relative probability of presence under certain assumptions27. However, as these assumptions are rarely met, we strongly discourage users from this interpretation and suggest interpreting the cloglog output values as an estimate of relative environmental suitability20 instead.
We further suggest using Maxent predictions with an AUC below 0.7 only in exceptions, and in large-scale studies. In general, our predictions may overestimate true range extents of endemic species and underestimate ranges of widespread species. However, in worst case, the entire native WGSPRD-regions are outlined as being environmentally suitable, which may be acceptable in some cases, but not in others.
In addition, Model 1 has been fitted with the suggested minimum number of records for generating meaningful distributions models53,54, but Model 2 and 3 were in some cases trained with less records. Whether this low sample size as well as its implied uncertainty is acceptable or not will differ between users and applications and needs to be considered.
The full data, including Maxent predictions (cloglog transformed), underlying occurrence records, native regions and corresponding metadata, can be explored at https://plant-ranges.indecol.no. Here, the predictions based on individual models (Model 1 to 3) as well as a suggested (i.e. best performing) prediction highlight environmentally suitable conditions, if available for the selected species. Predictions can potentially be transformed into a map indicating where the species is most certainly found, as required for local management and conservation actions95, or into a conservative range map, best suited for analysing global patterns108 and highlighting where a species is certainly absent109. However, the choice of an appropriate cut-off threshold is highly application specific. We outlined “potential range maps” in the data explorer for illustrational purposes only and based on the best performing prediction. We applied different cut-off thresholds to represent different levels of confidence using the R package dismo59. The threshold at which there was no omission (possibly suitable), the threshold at which the F1-score is highest (probably suitable) and presence cells (presence).
Code availability
All data and code is available without restrictions under the terms of a Creative Commons Zero (CC0) waiver (https://creativecommons.org/share-your-work/public-domain/cc0/). R code for retrieving and filtering data from POWO and GBIF, and for generating and evaluating Maxent models is available on GitHub (https://github.com/jannebor/plant_range_estimates). Any further requests can be directed to the corresponding author.
References
Millennium Ecosystem Assessment. Ecosystems and Human Well-being: Biodiversity Synthesis. (World Resources Institute, 2005).
Moran, D. & Kanemoto, K. Identifying species threat hotspots from global supply chains. Nat. Ecol. Evol. 1, 0023 (2017).
Newbold, T. Future effects of climate and land-use change on terrestrial vertebrate community diversity under different scenarios. Proc. R. Soc. B Biol. Sci. 285, 20180792 (2018).
Newbold, T. et al. Global effects of land use on local terrestrial biodiversity. Nature 520, 45–50 (2015).
Newbold, T. et al. Has land use pushed terrestrial biodiversity beyond the planetary boundary? A global assessment. Science (80-.). 353, 288–291 (2016).
Verones, F., Moran, D., Stadler, K., Kanemoto, K. & Wood, R. Resource footprints and their ecosystem consequences. Sci. Rep. 7, 40743 (2017).
United Nations. Transforming our World: the 2030 Agenda for Sustainable Development. A/RES/70/1 (United Nations, 2015).
Díaz, S. et al. Pervasive human-driven decline of life on Earth points to the need for transformative change. Science (80-.). 366, eaax3100 (2019).
Lenzen, M. et al. International trade drives biodiversity threats in developing nations. Nature 486, 109–112 (2012).
Hellweg, S. & Milà i Canals, L. Emerging approaches, challenges and opportunities in life cycle assessment. Science (80-.). 344, 1109–1113 (2014).
Chaudhary, A. & Brooks, T. M. National Consumption and Global Trade Impacts on Biodiversity. World Dev. 121, 178–187 (2019).
Pereira, H. M., Ziv, G. & Miranda, M. Countryside Species-Area Relationship as a Valid Alternative to the Matrix-Calibrated Species-Area Model. Conserv. Biol. 28, 874–876 (2014).
Lomolino, M. V & Heaney, L. R. Frontiers of Biogeography: New Directions in the Geography of Nature. (Sinauer Associates Inc. Publishers, 2004).
World Wildlife Fund. WildFinder: Online database of species distributions. http://www.worldwildlife.org/WildFinder (2006).
BirdLife International. IUCN Red List for birds. http://www.birdlife.org (2019).
IUCN. The IUCN Red List of Threatened Species. Version 2021-1 https://www.iucnredlist.org (2021).
Curran, M. et al. Toward Meaningful End Points of Biodiversity in Life Cycle Assessment. Environ. Sci. Technol. 45, 70–79 (2011).
Woods, J. S. et al. Ecosystem quality in LCIA: status quo, harmonization, and suggestions for the way forward. Int. J. Life Cycle Assess. 23, 1995–2006 (2018).
Phillips, S. J., Anderson, R. P. & Schapire, R. E. Maximum entropy modeling of species geographic distributions. Ecol. Modell. 190, 231–259 (2006).
Merow, C., Smith, M. J. & Silander, J. A. A practical guide to MaxEnt for modeling species’ distributions: what it does, and why inputs and settings matter. Ecography (Cop.). 36, 1058–1069 (2013).
Araújo, M. B. et al. Standards for distribution models in biodiversity assessments. Sci. Adv. 5, eaat4858 (2019).
Zurell, D. et al. A standard protocol for reporting species distribution models. Ecography (Cop.). 43, 1261–1277 (2020).
Brummitt, R. K., Pando, F., Hollis, S. & Brummitt, N. A. World Geographical Scheme for Recording Plant Distributions. International Working Group on Taxonomic Databases (TDWG) https://www.tdwg.org/standards/wgsrpd/ (2001).
GBIF. The Global Biodiversity Information Facility: What is GBIF? https://www.gbif.org/what-is-gbif (2021).
Phillips, S. J., Dudík, M. & Schapire, R. E. Maxent software for modeling species niches and distributions (Version 3.4.0). http://biodiversityinformatics.amnh.org/open_source/maxent/ (2016).
Phillips, S. J., Dudík, M. & Schapire, R. E. A maximum entropy approach to species distribution modeling. Proc. Twenty-first Int. Conf. Mach. Learn. 655–662 (2004).
Phillips, S. J., Anderson, R. P., Dudík, M., Schapire, R. E. & Blair, M. E. Opening the black box: an open-source release of Maxent. Ecography (Cop.). 40, 887–893 (2017).
Reddy, S. & Dávalos, L. M. Geographical sampling bias and its implications for conservation priorities in Africa. J. Biogeogr. 30, 1719–1727 (2003).
Hortal, J., Jiménez-Valverde, A., Gómez, J. F., Lobo, J. M. & Baselga, A. Historical bias in biodiversity inventories affects the observed environmental niche of the species. Oikos 117, 847–858 (2008).
Isaac, N. J. B. & Pocock, M. J. O. Bias and information in biological records. Biol. J. Linn. Soc. 115, 522–531 (2015).
Feeley, K. J. & Silman, M. R. Keep collecting: accurate species distribution modelling requires more collections than previously thought. Divers. Distrib. 17, 1132–1140 (2011).
Radosavljevic, A. & Anderson, R. P. Making better Maxent models of species distributions: complexity, overfitting and evaluation. J. Biogeogr. 41, 629–643 (2014).
ter Steege, H. et al. Hyperdominance in the Amazonian Tree Flora. Science (80-.). 342, 1243092 (2013).
Kuipers, K. J. J., Hellweg, S. & Verones, F. Potential Consequences of Regional Species Loss for Global Species Richness: A Quantitative Approach for Estimating Global Extinction Probabilities. Environ. Sci. Technol. 53, 4728–4738 (2019).
Gade, A. L., Hauschild, M. Z. & Laurent, A. Globally differentiated effect factors for characterising terrestrial acidification in life cycle impact assessment. Sci. Total Environ. 761, 143280 (2021).
Géron, C. et al. Urban alien plants in temperate oceanic regions of Europe originate from warmer native ranges. Biol. Invasions 23, 1765–1779 (2021).
Mair, L. et al. A metric for spatially explicit contributions to science-based species targets. Nat. Ecol. Evol. 5, 836–844 (2021).
Bachman, S., Moat, J., Hill, A., de la Torre, J. & Scott, B. Supporting Red List threat assessments with GeoCAT: geospatial conservation assessment tool. Zookeys 150, 117–126 (2011).
Cardoso, P. red - an R package to facilitate species red list assessments according to the IUCN criteria. Biodivers. Data J. 5, e20530 (2017).
Lee, C. K. F., Keith, D. A., Nicholson, E. & Murray, N. J. Redlistr: tools for the IUCN Red Lists of ecosystems and threatened species in R. Ecography (Cop.). 42, 1050–1055 (2019).
Bachman, S., Walker, B., Barrios, S., Copeland, A. & Moat, J. Rapid Least Concern: towards automating Red List assessments. Biodivers. Data J. 8 (2020).
POWO. Plants of the World Online. Facilitated by the Royal Botanic Gardens, Kew. http://www.plantsoftheworldonline.org/ (2021).
Chamberlain, S. et al. taxize: Taxonomic information from around the web. R package version 0.9.98. https://github.com/ropensci/taxize (2020).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria https://www.r-project.org/ (2021).
ITIS. Integrated Taxonomic Information System. https://www.itis.gov/ (2021).
Wickham, H. rvest: Easily Harvest (Scrape) Web Pages. R package version 0.3.5. https://cran.r-project.org/package=rvest (2019).
Desmet, P. & Page, R. WGSRPD. GitHub repository https://github.com/tdwg/wgsrpd (2018).
Chamberlain, S. et al. rgbif: Interface to the Global Biodiversity Information Facility API. R package version 3.6.0. https://cran.r-project.org/package=rgbif (2021).
GBIF. GBIF Occurrence Download. https://doi.org/10.15468/dl.uvd56q (2021).
Winkler, K., Fuchs, R., Rounsevell, M. & Herold, M. Global land use changes are four times greater than previously estimated. Nat. Commun. 12, 2501 (2021).
Sippel, S., Meinshausen, N., Fischer, E. M., Székely, E. & Knutti, R. Climate change now detectable from any single day of weather at global scale. Nat. Clim. Chang. 10, 35–41 (2020).
Hijmans, R. J. raster: Geographic Data Analysis and Modeling. R package version 3.0-7. https://cran.r-project.org/package=raster (2019).
Hernandez, P. A., Graham, C. H., Master, L. L. & Albert, D. L. The effect of sample size and species characteristics on performance of different species distribution modeling methods. Ecography (Cop.). 29, 773–785 (2006).
Pearson, R. G., Raxworthy, C. J., Nakamura, M. & Townsend Peterson, A. Predicting species distributions from small numbers of occurrence records: a test case using cryptic geckos in Madagascar. J. Biogeogr. 34, 102–117 (2006).
Phillips, S. J. & Dudík, M. Modeling of species distributions with Maxent: new extensions and a comprehensive evaluation. Ecography (Cop.). 31, 161–175 (2008).
Elith, J. et al. A statistical explanation of MaxEnt for ecologists. Divers. Distrib. 17, 43–57 (2011).
Anderson, R. P. & Raza, A. The effect of the extent of the study region on GIS models of species geographic distributions and estimates of niche evolution: preliminary tests with montane rodents (genus Nephelomys) in Venezuela. J. Biogeogr. 37, 1378–1393 (2010).
Själander, M., Jahre, M., Tufte, G. & Reissmann, N. EPIC: An Energy-Efficient, High-Performance GPGPU Computing Research Infrastructure. arXiv 1–4 (2019).
Hijmans, R. J., Phillips, S., Leathwick, J. & Elith, J. dismo: Species Distribution Modeling. R package version 1.1-4. https://cran.r-project.org/package=dismo (2017).
Muscarella, R. et al. ENMeval: An R package for conducting spatially independent evaluations and estimating optimal model complexity for Maxent ecological niche models. Methods Ecol. Evol. 5, 1198–1205 (2014).
Karger, D. N. et al. Climatologies at high resolution for the earth’s land surface areas. Sci. Data 4, 170122 (2017).
Karger, D. N. et al. Data from: Climatologies at high resolution for the earth’s land surface areas. Dryad, Dataset https://doi.org/10.5061/dryad.kd1d4 (2018).
ESA. Land Cover CCI Product User Guide Version 2. Tech. Rep. http://maps.elie.ucl.ac.be/CCI/viewer/download.php (2017).
Aiello-Lammens, M. E., Boria, R. A., Radosavljevic, A., Vilela, B. & Anderson, R. P. spThin: an R package for spatial thinning of species occurrence records for use in ecological niche models. Ecography (Cop.). 38, 541–545 (2015).
Akaike, H. Information Theory and an Extension of the Maximum Likelihood Principle. in 2nd International Symposium on Information Theory (eds. Petrov, B. N. & Csaki, F.) 267–281 (Akademia Kiado, 1973).
Hurvich, C. M. & Tsai, C.-L. Regression and time series model selection in small samples. Biometrika 76, 297–307 (1989).
Sugiura, N. Further analysts of the data by akaike’ s information criterion and the finite corrections. Commun. Stat. - Theory Methods 7, 13–26 (1978).
Morales, N. S., Fernández, I. C. & Baca-González, V. MaxEnt’s parameter configuration and small samples: are we paying attention to recommendations? A systematic review. PeerJ 5, e3093 (2017).
Shcheglovitova, M. & Anderson, R. P. Estimating optimal complexity for ecological niche models: A jackknife approach for species with small sample sizes. Ecol. Modell. 269, 9–17 (2013).
Warren, D. L. & Seifert, S. N. Ecological niche modeling in Maxent: the importance of model complexity and the performance of model selection criteria. Ecol. Appl. 21, 335–342 (2011).
Moran, P. A. P. Notes on Continuous Stochastic Phenomena. Biometrika 37, 17 (1950).
Borgelt, J., Sicacha-Parada, J., Skarpaas, O. & Verones, F. Native range estimates for red-listed vascular plants. Dryad, Dataset https://doi.org/10.5061/dryad.qbzkh18h9 (2022).
Sing, T., Sander, O., Beerenwinkel, N. & Lengauer, T. ROCR: visualizing classifier performance in R. Bioinformatics 21, 3940–3941 (2005).
Grau, J., Grosse, I. & Keilwagen, J. PRROC: computing and visualizing precision-recall and receiver operating characteristic curves in R. Bioinformatics 31, 2595–2597 (2015).
Hosmer, D. W., Lemeshow, S. & Sturdivant, R. X. Applied Logistic Regression. The Statistician 45 (Wiley, 2013).
Lobo, J. M., Jiménez-Valverde, A. & Real, R. AUC: a misleading measure of the performance of predictive distribution models. Glob. Ecol. Biogeogr. 17, 145–151 (2008).
Sofaer, H. R., Hoeting, J. A. & Jarnevich, C. S. The area under the precision‐recall curve as a performance metric for rare binary events. Methods Ecol. Evol. 10, 565–577 (2019).
Meyer, C., Weigelt, P. & Kreft, H. Multidimensional biases, gaps and uncertainties in global plant occurrence information. Ecol. Lett. 19, 992–1006 (2016).
Caudullo, G., Welk, E. & San-Miguel-Ayanz, J. Chorological maps for the main European woody species. Data Br. 12, 662–666 (2017).
Rivers, M. C. Laburnum anagyroides. The IUCN Red List of Threatened Species 2017: e.T79919483A79919650 https://doi.org/10.2305/IUCN.UK.2017-3.RLTS.T79919483A79919650.en (2017).
Botanic Gardens Conservation International Group & IUCN SSC Global Tree Specialist. Terminalia macrostachya. The IUCN Red List of Threatened Species 2019: e.T150118895A150118897 https://doi.org/10.2305/IUCN.UK.2019-3.RLTS.T150118895A150118897.en (2019).
Heil, K., Terry, M. & Corral-Díaz, R. Mammillaria grahamii (amended version of 2013 assessment). The IUCN Red List of Threatened Species 2017: e.T152723A121546147 https://doi.org/10.2305/IUCN.UK.2017-3.RLTS.T152723A121546147.en (2017).
Brooker, M. & Kleinig, D. Field Guide to Eucalypts. (Bloomings Books, 2006).
Koopman, M. M. A synopsis of the Malagasy endemic genus Megistostegium Hochr. (Hibisceae, Malvaceae). Adansonia 33, 101–113 (2011).
World Conservation Monitoring Centre. Memecylon elegantulum. The IUCN Red List of Threatened Species 1998: e.T32597A9713234 https://doi.org/10.2305/IUCN.UK.1998.RLTS.T32597A9713234.en (1998).
Landrum, L. R. A revision of the Psidium salutare complex (Myrtaceae). SIDA, Contrib. to Bot. 20, 1449–1469 (2003).
Tropical Plants Database. Ken Fern. tropical.theferns.info https://tropical.theferns.info/viewtropical.php?id=Psidium+salutare (2021).
Bernal, R., Gradstein, S. R. & Celis, M. Siparuna conica S.S.Renner & Hausner. Catálogo de plantas y líquenes de Colombia http://catalogoplantasdecolombia.unal.edu.co (2015).
Renner, S. S. & Hausner, G. New Species of Siparuna (Monimiaceae) II. Seven New Species from Ecuador and Colombia. Missouri Bot. Gard. Press 6, 103–116 (1996).
Melendo, M., Giménez, E., Cano, E., Mercado, F. G. & Valle, F. The endemic flora in the south of the Iberian Peninsula: taxonomic composition, biological spectrum, pollination, reproductive mode and dispersal. Flora - Morphol. Distrib. Funct. Ecol. Plants 198, 260–276 (2003).
Chari, L. D., Martin, G. D., Steenhuisen, S.-L., Adams, L. D. & Clark, V. R. Biology of Invasive Plants 1. Pyracantha angustifolia (Franch.) C.K. Schneid. Invasive Plant Sci. Manag. 13, 120–142 (2020).
Sasidharan, N. Amomum pterocarpum Thwaites. India Biodiversity Portal https://indiabiodiversity.org/species/show/258864#habitat-and-distribution (2013).
Contu, S. Amomum pterocarpum. The IUCN Red List of Threatened Species 2013: e.T44393013A44450020 https://doi.org/10.2305/IUCN.UK.2013-1.RLTS.T44393013A44450020.en (2013).
Babyrose Devi, N., Das, A. & Singh, P. Amomum Pterocarpum (Zingiberaceae): a new record in the flora of Manipur. Int. J. Adv. Res. 6, 546–549 (2018).
Jetz, W., Sekercioglu, C. H. & Watson, J. E. M. Ecological correlates and conservation implications of overestimating species geographic ranges. Conserv. Biol. 22, 110–9 (2008).
Gibbs, D. & Khela, S. Magnolia pugana. The IUCN Red List of Threatened Species 2014: e.T194806A2363344 https://doi.org/10.2305/IUCN.UK.2014-1.RLTS.T194806A2363344.en (2014).
Sayer, C. Vallesia glabra. The IUCN Red List of Threatened Species 2015: e.T62543A72668627 https://doi.org/10.2305/IUCN.UK.2015-2.RLTS.T62543A72668627.en (2015).
Sánchez Gómez, P., Stevens, D., Fennane, M., Gardner, M. & Thomas, P. Tetraclinis articulata. The IUCN Red List of Threatened Species 2011: e.T30318A9534227 https://doi.org/10.2305/IUCN.UK.2011-2.RLTS.T30318A9534227.en (2011).
Stritch, L., Roy, S., Shaw, K. & Wilson, B. Corylus cornuta (errata version published in 2017). The IUCN Red List of Threatened Species 2016: e.T194448A115337731 https://doi.org/10.2305/IUCN.UK.2016-1.RLTS.T194448A2336319.en (2016).
Olson, D. M. et al. Terrestrial ecoregions of the world: A new map of life on Earth. Bioscience 51, 933–938 (2001).
Rivers, M. C. Cotoneaster cambricus. The IUCN Red List of Threatened Species 2017: e.T102827479A102827485 https://doi.org/10.2305/IUCN.UK.2017-3.RLTS.T102827479A102827485.en (2017).
RStudio Team. RStudio: Integrated Development Environment for R. RStudio, PBC, Boston, MA http://www.rstudio.com/ (2021).
Bivand, R., Keitt, T. & Rowlingson, B. rgdal: Bindings for the ‘Geospatial’ Data Abstraction Library. https://cran.r-project.org/package=rgdal (2019).
Bivand, R. & Lewin-Koh, N. maptools: Tools for Handling Spatial Objects. R package version 0.9-5. https://cran.r-project.org/package=maptools/ (2019).
Bivand, R. & Rundel, C. rgeos: Interface to Geometry Engine - Open Source (‘GEOS’). R package version 0.5-1. https://cran.r-project.org/package=rgeos (2019).
Bivand, R. S., Pebesma, E. & Gómez-Rubio, V. Applied Spatial Data Analysis with R. (Springer New York, 2013).
Phillips, S. J. & Elith, J. POC plots: calibrating species distribution models with presence-only data. Ecology 91, 2476–2484 (2010).
Hurlbert, A. H. & Jetz, W. Species richness, hotspots, and the scale dependence of range maps in ecology and conservation. Proc. Natl. Acad. Sci. 104, 13384–13389 (2007).
Jetz, W., McPherson, J. M. & Guralnick, R. P. Integrating biodiversity distribution knowledge: toward a global map of life. Trends Ecol. Evol. 27, 151–159 (2012).
Acknowledgements
We want to thank Radek Lonka and the IndEcol Digital Lab for facilitating the use of the high-performance computing infrastructure and hosting the online application. This study is part of the Transforming Citizen Science for Biodiversity project hosted by the Digital Transformation initiative of the Norwegian University of Science and Technology.
Author information
Authors and Affiliations
Contributions
J.B. was responsible for study design, methodologies, code writing, code execution, and writing the manuscript. J.S.P. contributed to methods for technical validation of the data and writing the manuscript. O.S. contributed to methodologies, interpretation of the data, and writing the manuscript. F.V. contributed to study design, interpreting the results, and writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Borgelt, J., Sicacha-Parada, J., Skarpaas, O. et al. Native range estimates for red-listed vascular plants. Sci Data 9, 117 (2022). https://doi.org/10.1038/s41597-022-01233-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-022-01233-5