Abstract
This data release of 117 healthy community-dwelling adults provides multimodal high-quality neuroimaging and behavioral data for the investigation of brain-behavior relationships. We provide structural MRI, resting-state functional MRI, movie functional MRI, together with questionnaire-based and task-based psychological variables; many of the participants have multiple datasets from retesting over the course of several years. Our dataset is distinguished by utilizing open-source data formats and processing tools (BIDS, FreeSurfer, fMRIPrep, MRIQC), providing data that is thoroughly quality checked, preprocessed to various extents and available in multiple anatomical spaces. A customizable denoising pipeline is provided as open-source code that includes tools for the generation of functional connectivity matrices and initialization of individual difference analyses. Behavioral data include a comprehensive set of psychological assessments on gold-standard instruments encompassing cognitive function, mood and personality, together with exploratory factor analyses. The dataset provides an in-depth, multimodal resource for investigating associations between individual differences, brain structure and function, with a focus on the domains of social cognition and decision-making.
Measurement(s) | Blood Oxygen Level-Dependent Functional MRI • brain • psychological measurement |
Technology Type(s) | functional magnetic resonance imaging • 3 T MRI scanner • behavioral assessments |
Sample Characteristic - Organism | Homo sapiens |
Sample Characteristic - Location | United States of America |
Similar content being viewed by others
Background & Summary
Investigating brain-behavior relationships and their individual differences requires multimodal data that include at least neural data (typically structural or functional magnetic resonance imaging data, structural MRI (sMRI) or functional MRI (fMRI)) together with behavioral data (typically questionnaire-based scores). Traditionally, such data have been acquired in individual studies, often with modest sample sizes, and focused on a specific research question. Several more recent datasets combine neuroimaging and behavioral data in larger samples with broader but shallow coverage of cognitive domains; a few datasets also provide exceptionally dense data with deep phenotyping, but in very small samples1,2. For instance, the UK Biobank3 provides very broad behavioral data together with MRI and genetic data on 500,000 subjects – a resource that has been utilized by over 20,000 researchers to date and has yielded a number of important findings4,5. At the opposite extreme, dense longitudinal rest-state fMRI acquired on a single individual showed that functional brain networks are more fine-grained than originally thought1. For many large databases, sample sizes are now becoming sufficiently large that nonlinear modeling (e.g., with deep learning) is becoming possible to apply to brain-behavior relationships6. However, breadth across variables and large samples typically come at the expense of shallow assessment of most behavioral variables, and often limited quality control over individual neuroimaging data. For instance, cognitive variables like intelligence are assessed with short tasks or questionnaires, rather than gold-standards in the field such as the Wechsler Adult Intelligence Scale7, which may result in poor precision as well as low construct validity. More comprehensive psychological assessment is provided in the Human Connectome Project, (HCP), which also provides high-resolution structural and functional neuroimaging data8 and remains a multimodal resource generating a large number of novel discoveries (see https://www.humanconnectome.org/study/hcp-young-adult/publications). However, across datasets, there is a tradeoff between in-depth and/or quality-checked data, on the one hand, versus sample size and domain breadth, on the other hand. For instance, social cognition is multifaceted and complex. Thus it can only be adequately assessed with a variety of measures to describe individual functioning (and potential individual differences therein). In addition, a major practical limitation for users is that databases generally provide only specific neuroimaging formats and processing steps, which often become outdated or require further conversion and processing before analyses can be applied.
Our dataset lies intermediate in the space of databases outlined above: a medium-sized sample (n = 117) but with exceptional quality control, range of data types, accessibility and ease of usability. Manually quality-checked structural and denoised functional (resting-state and movie) MRI data are organized in BIDS9 and are provided with quality metrics and in multiple processed formats (including individual native space and multiple standard template spaces). We use standardized preprocessing and quality control tools, such as fMRIPrep10, FreeSurfer11, and MRIQC12. Behavioral data encompass comprehensive metrics on intelligence, personality, mood, and social cognition. A subset of the subjects were retested over months-years. All data were collected by the NIMH-funded Caltech Conte Center (http://conte.caltech.edu) and are tailored towards investigations of social cognition and decision-making. The precision with which cognitive processes can be estimated usually demands longer tasks or questionnaires, and/or the extraction of latent factors across multiple observed measures, both of which we provide here. As detailed in Table 1 below, the dataset includes a variety of gold-standard psychological state and trait variables relevant to social decision-making. For example, intelligence provides an important continuous variable that could be used as a covariate in all analyses. Likewise measures of depression, stress, valenced mood and anxiety may be used as covariates or utilized for formulating exclusionary criteria. The dataset also includes trait-level questionnaire-based measures that specifically address social decision-making and behavior. For example, several of our measures are commonly used to assess traits related to autism spectrum disorder (e.g. Empathizing Quotient13, Systematizing Quotient14, and Social Responsiveness Scale15), the social network index has been widely used as a proxy for social connectedness in real life, and the 16PF personality questionnaire provides a fine-grained assessment of personality traits related to social engagement from which the standard “Big-Five” personality factors can be easily derived. Especially notable is the Mayer-Salovey-Caruso Emotional Intelligence Test16, a comprehensive and time-intensive collection of questionnaire-based and task-based measures that index multiple facets of social and emotional ability. Taken together, this array of behavioral measures provides a particularly rich assessment of individual differences relevant to social decision-making, and item-level data availability permits researchers to explore additional structure. We have included an exploratory factor analysis to showcase how the included measures and their loadings on potentially underlying factors can be used to leverage the richness of this new dataset towards novel questions in human cognitive neuroscience.
Three features further distinguish our dataset: (1) we went to considerable lengths to control the quality of surface reconstructions by manual visual inspection and correction of all structural MRI datasets (where necessary); (2) we provide functional neuroimaging data in multiple preprocessed formats and anatomical spaces (including both volumetric and surface data) with open-source processing tools. This not only affords greater flexibility in how the data might be analyzed, but largely obviates the need to conduct further preprocessing or transformations by users — a task that can be complex and require substantial computational cost and time; (3) we provide a customizable denoising pipeline for the analysis of functional connectivity data that includes not only state-of-the-art denoising, but also incorporates the generation of functional connectivity matrices on the parcellated data and initializes analysis workflow for individual difference studies. Taken together, these features aim to provide a dataset that can most easily be used immediately to address scientific questions of interest by neuroscientists, psychologists, and data scientists.
Methods
Participants
Adults (enrollment n = 191; 18–50 years old at time of enrollment) were recruited from the Los Angeles area via Craigslist and publicly distributed flyers over the course of the past 8 years. Informed consent was obtained from all subjects prior to participation in accordance with the institutional review board (IRB) at the California Institute of Technology. Subjects were excluded if they had a full-scale IQ below 90, were not fluent in English, had a first-degree relative with schizophrenia or autism spectrum disorder, were currently taking psychotropic medication, had uncorrected vision or hearing impairment, and moderate-severe depression or indication of current suicidality (Beck Depression Inventory–II total = 25+; score of 3 or 4 on item 917). Additional exclusionary criteria included history of any of the following: premature birth, epilepsy, major medical condition, metabolic disorder, chemotherapy or radiation, brain surgery, head injury, eating disorder, neurological condition, psychosis, bipolar disorder, autism, suicide attempt, substance dependence or abuse, alcoholism, color blindness or strabismus.
Following a brief phone screening, 191 individuals came to Caltech for the enrollment visit. The final sample was reduced to 117 individuals due to exclusions and attrition. Information acquired during the enrollment visit resulted in exclusion of 47 based on our inclusion/exclusion criteria, 19 were excluded during MRI safety screening or due to features of MRI testing (6 due to claustrophobia, incompatible tattoos or pregnancy, 12 due to excessive motion during MRI scanning, 1 incidental structural abnormality per expert radiological review) and 8 dropped out of the study following the enrollment visit. The final participant group of 117 adults did not differ from the initial sample in gender (χ2 = 0.305, p = 0.581), age (t(187) = −0.594, mean difference = −0.581, 95% CI [−2.511, 1.349]), ethnicity (χ2 = 0.072, p = 0.789), or race (χ2 = 4.270, p = 0.640) (Fig. 1).
Demographics of Final Sample. Demographics of the final sample (n = 117, inner pie) are compared to demographics of participants who were excluded (outer pie; excluded n = 66; attrition n = 8). Top row: Sex (a) and Race (b) proportions. Middle: Ethnicity proportions (c). Bottom row: Number of participants by highest education level (d) and age grouping (e; green = final sample, gray = excluded/attrition group). Abbreviations: AA, Associates in Arts; BA/BS, Bachelor of Arts/Science; Grad, Graduate degree; HS, high school; Some C, some college.
Structural magnetic resonance imaging
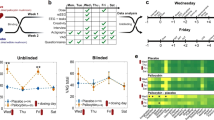
All MRI data were acquired using either a 3 Tesla TIM Trio (2012 to 2017) or an upgraded 3 T Prisma.Fit system (2018 to 2019) (Siemens Medical Solutions, Malvern, PA) with a 32 channel head receive array coil. Stimulus presentation and response capture were performed using an LCD back-projection system and optical response button box (controlled via Psychophysics Toolbox 3). T1w structural imaging was performed in all 117 participants. T2w imaging was added in the second phase of the Conte Center (2018 onwards) and both structural contrasts were acquired in a subset of participants (Fig. 2). Incremental modifications were made over the years to the structural imaging protocol, including a change in spatial resolution from 1 mm to 0.9 mm isotropic, the addition of lipid suppression and a change in T1w pulse sequence from single-echo MP-RAGE to multi-echo MEMP-RAGE, which are summarized in Table 2.
Comparison of functional connectivity (FC) matrices estimated before denoising (top row, a,b) and after denoising (bottom row, c,d) on subjects with two complete resting-state runs (N = 116). On the left (a,c), the lower triangular matrices are the average FC derived from single-band resting-state acquisitions (N = 34), while the upper triangular matrices show the average FC derived from multiband resting-state acquisitions (N = 100). Note that some subjects (N = 18) have both SB and MB scans and therefore contribute to both upper and lower triangles. On the right (b,d), lower triangular matrices are derived from movie fMRI data (N = 57), while upper triangular matrices are derived from multiband resting-state acquisitions (N = 100). We used data in CIFTI format registered to the MNI152NLin2009cAsym space, processed through fMRIPrep and denoised with rsDenoise with the strategy described in48 For each subject, two runs were concatenated before computing the average time series for each of 400 parcels of the Schaefer cortical parcellation83. Parcels are grouped following the 7 resting-state networks defined in the Yeo parcellation84. FC was computed as the pairwise Pearson’s correlation between parcel time series (color scale). For subjects with more than one session available, individual FC matrices are averaged across sessions before averaging them across subjects (so that each subject only contributed once).
Functional magnetic resonance imaging
High-quality BOLD fMRI data with whole-brain coverage were acquired in all subjects. BOLD resting-state and movie-viewing fMRI were acquired using single-band or multi-band 2.5 mm isotropic T2*-weighted EPI, depending on protocol version (Table 2 and Fig. 2). The imaging protocol was refined several times during the first phase of the Conte Center, but remained constant during the second phase following the scanner upgrade from TIM Trio to Prisma.Fit. Multiband acceleration was introduced in protocol version 1.2 and imaging parameters for all versions of the fMRI protocols are summarized in Table 2. Either dual-echo gradient echo imaging or phase-encoding polarity reversed SE-EPI image pairs were acquired for distortion correction immediately before each functional run, with identical slice geometry and EPI echo spacing to the BOLD EPI series.
Resting-state data consisted of two runs of between 400 (session 1p1) and 420 (session 2p2) seconds of resting-state with eyes open and instructions to fixate a white central cross on a black background. Movie viewing fMRI consisted of watching the black-and-white Hitchcock film “Bang! You’re Dead (1954)” and the short animated movie “Partly Cloudy”18. Alfred Hitchcock’s “Bang! You’re Dead (1954)” movie was edited from the original 20 min running time down to 8 min., as in19. Instructions were shown on the screen until the subject pressed a key, followed by 10 s of blank screen with a fixation cross. The movie played for 8-min, followed by 10 s blank screen with fixation cross until the end of scanning. The “Partly Cloudy” movie began after 10 s of rest (black screen; TRs 0–5). The first 10 s of the movie consisted of the opening credits (Disney castle, Pixar logo; 12–20 s), followed by 5 minutes, 14 seconds of the movie (without credits at the end), followed by 10 seconds of rest.
Preprocessing of MRI Data
FreeSurfer segmentation and cortical parcellation
We performed cortical reconstruction and volumetric segmentation of T1w images outside of fMRIPrep with the FreeSurfer image analysis suite (version 7.1.0, http://surfer.nmr.mgh.harvard.edu/)11,20,21,22. In summary, processing included motion correction and averaging23 of volumetric T1w images, removal of non-brain tissue, automated Talairach transformation, segmentation of the subcortical white matter and deep gray matter volumetric structures, intensity normalization24, tessellation of the gray matter-white matter boundary, automated topology correction25,26, and surface deformation following intensity gradients for optimal tissue boundary placement. T1w MP-RAGE data were used for FreeSurfer reconstruction if T1w MEMP-RAGE data from the Phase 2 protocol were unavailable for a given subject. T2w images were passed to FreeSurfer reconstruction where available (n = 59). See Fig. 2 for a full breakdown of T1w and T2w image availability for all subjects.
Standardized MRI preprocessing
Both structural and functional MRI data were minimally preprocessed using fMRIPrep 20.2.110, which is based on Nipype 1.5.127. The processing steps for anatomical and functional MR data are summarized below, with specific software noted in italics. Independent, quality controlled FreeSurfer reconstructions (above) were integrated automatically by the fMRIPrep pipeline. Preprocessing scripts, including the exact parameters used with fMRIPrep and a detailed description of individual steps are provided in the code folder of the OpenNeuro BIDS data release28.
Anatomical data preprocessing
T1-weighted (T1w) structural images were corrected for intensity non-uniformity (N4BiasFieldCorrection, ANTS 2.3.3)29,30 and skull-stripped (antsBrainExtraction.sh, ANTS 2.3.3). Brain tissue was segmented into cerebrospinal fluid (CSF), white-matter (WM) and gray-matter (GM) (fast, FSL 5.0.9)31. Where multiple T1w images were available for a given subject, a robust, registered average was constructed (mri_robust_template, FreeSurfer 6.0.1)20. Brain extracted T1w images were then registered diffeomorphically (antsRegistration, ANTs 2.3.3) to two standard spaces: (1) the ICBM/MNI 152 2009c Nonlinear Asymmetric space used by OpenNeuro (MNI152NLin2009cAsym)32 and (2) the ICBM/MNI 152 Version 6 Nonlinear Asymmetric space used by FSL (MNI152NLin6Asym)33.
Functional data preprocessing
For each of the BOLD runs found per subject (across all tasks and sessions), the following preprocessing was performed. First, a reference volume and its skull-stripped version were generated by aligning and averaging single-band references (SBRefs). Spatial distortion corrections for BOLD EPI data were derived from two spin echo EPI reference images with opposing phase-encoding directions (3dQwarp, AFNI 20160207)34. A distortion-corrected BOLD EPI reference image was constructed and registered to the T1w reference using a boundary-based approach (bbregister, Freesurfer)35. Rigid-body head-motion parameters with respect to the BOLD EPI reference were estimated (mcflirt, FSL 5.0.9)36 before any spatiotemporal filtering. BOLD runs belonging to the single band acquisition sessions were slice-time corrected (3dTshift, AFNI 20160207). The BOLD time series were resampled onto the fsaverage and fsaverage6 standard FreeSurfer surface spaces. The BOLD time series (including slice-timing correction when applied) were resampled onto their original, native space by applying a single, composite transform to correct for head motion and susceptibility distortions. The BOLD time series were resampled into the MNI152NLin2009cAsym standard space. Grayordinate files37 containing 91,000 samples were also generated using the highest-resolution fsaverage as an intermediate standardized surface space. Several physiological confound time series were calculated based on the preprocessed BOLD: framewise displacement (FD), DVARS and three region-wise global signals. FD was computed for each functional run using two definitions: absolute sum of relative motions38 and relative root-mean-square displacement between affine transforms36.
Physiological Denoising of fMRI Data
Physiological noise regressors were extracted using CompCor and are provided for use in alternative physiological denoising approaches, but were not used in the rsDenoise pipeline described below39. Principal components were estimated for the two CompCor variants: temporal (tCompCor) and anatomical (aCompCor). A mask to exclude signal originating in cortex was obtained by eroding the brain mask, ensuring it only contained subcortical structures. Six tCompCor components were then calculated including only the top 5% variable voxels within that subcortical mask. For aCompCor, six components were calculated within the intersection of the subcortical mask and the union of CSF and WM masks calculated in T1w space, after their projection to the native space of each functional run. Framewise displacement38 was calculated for each functional run using the approach implemented by Nipype.
Resting-state and movie fMRI data were further processed with rsDenoise (https://github.com/adolphslab/rsDenoise), a denoising pipeline specifically designed to correct for artifactual influences of non-neuronal fluctuations in signals acquired in the absence of an explicit task. This software was originally developed to study individual differences in intelligence and personality detectable from resting-state fMRI functional connectivity data40,41. The pipeline is based on open-source libraries and frameworks for scientific computing, including SciPy, Numpy, NiLearn, NiBabel, Nipype, Scikit-learn, Pandas and Matplotlib27,42,43,44,45,46,47, and accepts both volumetric data (in NIfTI format) and surface data (GIfTI or CIFTI format) that were minimally preprocessed with either fMRIPrep or the HCP pipelines37. It implements a wide variety of denoising strategies described by previous literature1,48,49,50,51, and works by performing a sequence of operations grouped in seven categories: motion scrubbing, voxel-wise normalization, detrending, tissue regression, global signal regression, motion regression and temporal filtering. In addition to enabling the user to reproduce previously published methods, the software allows testing of new combinations of denoising steps and adding custom functions to the pipeline. The pipeline also offers support for the generation of functional connectivity matrices (as in Fig. 3) and a framework for the prediction of individual differences from functional connectivity features. For the results presented in this work, we adopted a pipeline that reproduces the denoising strategy described in48. There are seven consecutive steps: (1) each voxels’ signal is z-score normalized, (2) using tissue masks, temporal drifts from cerebrospinal fluid (CSF) and white matter (WM) are removed with third-degree Legendre polynomial regressors, (3) CSF and WM mean signals are regressed from gray matter (GM) voxels, (4) rotational and translational realignment parameters and their temporal derivatives are used as explanatory variables in motion regression, (5) signals are low-pass filtered with a Gaussian kernel, (6) temporal drift from gray matter (GM) signal is removed using third-degree Legendre polynomial regressors, and (7) lastly global signal regression (GSR) is performed.
Factor Analysis. (a) Spearman rank-order correlations between each pair of variables. Variables are ordered according to the four-factor varimax-rotated solution, with dark outline boxing each factor grouping. (b) Four-factor varimax-rotated solution based on data from 144 participants. Maximal absolute loadings of task scores onto each of the four factors, leading to the interpretation of the factors we give in the text. Lighter colors indicate flipped scale interpretation (negative loadings).
Behavioral assessment
Assessment of cognitive and behavioral functioning was conducted using the 12 standardized psychological instruments described in Table 1. These instruments were administered by one trained research assistant (T.A.), and the majority of data were collected on one day. Demographic and behavioral data are curated in a comma-separated value (CSV) file, accompanied by a data dictionary explaining all variables28. The dataset includes summary scores and item-wise responses. Descriptive group statistics of the summary scores from all behavioral measures are provided in Table 3.
Descriptive group statistics of the summary scores from all behavioral measures are provided in Table 3. When available, participants’ scores were converted to standardized scores using published norms that account for demographic factors relevant to each measure (per the publisher). The WASI52, WASI-II53, and SRS-215 norms are age-specific. STAI54 and MSCEIT16 norms are specific for age and sex. Table 3 presents 95% confidence intervals for the difference from the expected mean (e.g. participant T-score minus 50) based on 1000 bootstrapped samples. The 95% confidence intervals indicate that our cohort had elevated IQ scores, with elevated emotion perception but lower emotion management (MSCEIT) scores than the published normative sample. On average, personality traits (16PF) reported in our sample indicated elevations in liveliness, sensitivity, vigilance, abstractedness, openness to change, and self-reliance, with reduced evidence of warmth, rule-consciousness and tension. The 95% confidence intervals for SRS-2 and STAI trait anxiety difference scores included zero, but our cohort reported notably low levels of state anxiety. Additionally, for the tests with standardized scores we examined the number of participants who scored more than 1.5 standard deviations above or below the normative mean (i.e. within the range of clinical significance). After applying measure-wise Bonferroni adjustment, the frequency of participants with clinically-significant scores was not greater than expected by chance for any measure.
In addition to the psychological variables from specific tasks, we also provide an example use case of the rich psychological data in an exploratory factor analysis based on all of the behavioral measures available in all of the subjects (Note: MSCEIT and SRS-2 were not included as they were not available for all participants and STAI state was not included due to high correlation with STAI trait; Fig. 4). We conducted exploratory factor analysis on all subjects with complete datasets, which were 144 Conte Center participants, of which the 117 whose imaging data are presented here were a proper subset. Due to non-normal distribution of multiple measures, Spearman rank-order coefficients were used for all correlations in the factor analysis (see Fig. 4). The number of factors was estimated in R55 using the following methods (processing packages are shown in italics): Horn’s Parallel Analysis56 (paran); Cattell’s Scree Optimal Coordinate Index57 (nFactors); CNG scree test58 (nFactors); Zoski and Jurs’ multiple regression b coefficient59 (nFactors); the Minimum Average Partial (MAP) test, both the original60 and revised61 versions (paramap); and the Very Simple Structure criterion (vss). All tests, with the exception of Horn’s Parallel Analysis, consistently predicted three to four factors. Based on these estimates, four factors were retained. The R code for estimating the optimal number of factors and generating rotated and unrotated solutions for 3- and 4-factor models, as well as all data files related to this analysis are provided at https://github.com/adolphslab/ConteDataRelease/blob/main/FactorAnalysis/Factor_Analysis.R.
An example abbreviated BIDS directory structure for one subject showing the range of imaging and auxiliary data types available for multiple protocol variants. Briefly, the main data records consist of: (i) structural MRI (raw T1w and T2w images; manually edited segmented and parcellated cortical data), (ii) resting-state fMRI (raw, preprocessed, denoised, available in 3 anatomical spaces), (iii) movie fMRI (raw, preprocessed, available in 3 anatomical spaces), (iv) physiological data to accompany the fMRI datasets An overview of all the MRI data available across the entire subject sample is provided in Fig. 5.
Specifying a three-factor and four-factor solution, factor analysis was conducted in R using maximum likelihood estimation, with varimax rotation and without rotation (fa), and factor scores were generated with the Bartlett formula. Figure 4 shows factor loadings for the four-factor varimax-rotated solution. Factor loadings for the rotated and unrotated solutions were highly congruent (rc = 0.99 for factors 1 and 2 and rc = 0.91 for factors 3 and 4). Factor 1 is associated with negative emotionality, including elevations in anxiety, depression, stress, negative affect, and emotional instability, as well as lowered empathy. Factor 2 reflects cognitive flexibility, with elevations on cognitive ability and openness to change, and a negative association with rule consciousness. Factor 3 relates to elevated levels of social engagement. Factor 4 reflects cognitive rigidity. It is noteworthy that lowest factor loadings were for two social measures (SNI People in Network and 16PF Sensitivity), suggesting that while these factors account for some shared variance in social skills, they are unlikely to mask unique individual variations in social functioning. Individual scores across these 4 factors are provided for all our 117 subjects as part of this data release28; however, this is only one illustrative approach to factor analysis and should not preclude exploration using alternative methods.
Data Records
The data types described below are available on the OpenNeuro data sharing platform28. The dataset follows the Brain Imaging Data Structure (BIDS version 1.6.0)9 which organizes the imaging data using a simple folder structure with nested files, each with standardized file naming conventions and accompanying JSON and TSV format metadata. T1w and T2w structural images were irreversibly deidentified using a customization of pydeface (https://github.com/jmtyszka/voxface). An example of the data structure and variety of data types available for subjects is given in Fig. 5. Note that events TSV files are empty placeholders for BIDS validation in the absence of response behavior for passive movie viewing and resting-state series.
Availability of structural (sMRI, a) and functional (fMRI, b) runs for each subject and session (i.e., protocol version). Note that not all subject ID labels are shown for clarity. Key: Cyan = one run, Blue = two runs. See Table 2 for full pulse sequence parameter details.
Technical Validation
Quality Control of Automated Cortical and Subcortical Reconstructions
Freesurfer supports visual inspection and manual corrections of automatic reconstruction to the initial and final brain masks, white and gray matter delineation and specification of white matter bias correction control points. All initial tissue constructions were visually inspected and manually corrected as necessary by a team of eight trained editors (DK, DAK, TR, ZE, DL, SL, WZ, JMT). Training included i) prior training through Freesurfer course material and ii) expert-guided learning of manual interventions (http://surfer.nmr.mgh.harvard.edu/fswiki/CourseDescription). Editors were randomly assigned to edit 10–15 scans. The most common issues that needed correction included: 1) inclusion of non-brain tissue (e.g., dura, skull, sinus blood) in the grey matter (pial surface), 2) incomplete temporal pole reconstruction, 3) white matter surface inaccuracies in ventral temporal regions. Manual edits were applied as outlined in detail by the FreeSurfer documentation (http://surfer.nmr.mgh.harvard.edu/fswiki/Tutorials) and respective reconstruction steps were run as implemented by the pipeline. Resulting next round reconstructions were again visually inspected and edited where necessary. An example of the impact of editing the brain mask on the pial surface in an individual subject is shown in Fig. 6 (top) with the surface displacement caused by editing, averaged over all subjects, shown in Fig. 6 (bottom).
Example impact of manual brain mask editing on pial surface estimation. Prior to correction (a), the automatically estimated pial surface extended into the sagittal sinus (arrows). Deletion of voxels from the brain mask (b, heatmap color scale overlay) restored the pial surface to its edited position (arrows). (c) Cortical regions requiring pial surface editing. The number of subjects with pial surface displacement following editing of greater than 1 mm is shown overlaid on the partially inflated fsaverage pial surface. Overall, pial editing is concentrated in medial temporal, ventromedial frontal and lateral temporal cortices, consistent with areas prone to local susceptibility effects resulting in boundary inaccuracies.
Quality control of fMRIPrep reports
fMRIPrep provides visual quality assessment reports per subject allowing a thorough visual assessment of processing quality. Three raters (D.K., J.M.T., P.G.) each visually inspected about one third of all reports, using previously agreed-upon criteria with regards to i) visual artifacts, ii) registration/transformation errors, iii) brain tissue segmentation and iv) quality of susceptibility distortion correction. We used a threshold intended to be conservative for gross errors, yet not specific to minor inaccuracies. We provide the three-tiered ratings (1, major issues; 2, minor issues; 3, no obvious issues) in a CSV file (fmriprep_output_manualQA.csv)28.
Image quality control metrics for bold fmri
Detailed image quality metrics (IQMs) were calculated for all structural and functional imaging series using MRIQC (v0.15, Stanford Center for Reproducible Neuroscience)62 and full reports are included in this data release. Two example IQMs for the fMRI series, frame wise displacement (FD) and temporal signal-to-noise ratio (tSNR), are reported in more detail here.
Framewise displacement
Rigid body head motion was characterized using the framewise displacement (FD) metric defined in63. FD was computed with and without linear low-pass filtering (LPF) (Butterworth filter, order 5, f < 0.2 Hz) of the individual motion parameter time series calculated by MRIQC. LPF minimizes high frequency respiratory contamination in FD timeseries following arguments made in64,65,66,67. Filtered FD distributions for the three fMRI experiments (“Bang, You’re Dead!”, “Partly Cloudy” and resting-state) are shown in Fig. 7. Note that a very small number of subjects show rare relatively large motion spikes at times during the scan, as expected in a larger sample. All motion is fully characterized in the combination of fMRIPrep and MRIQC reports of this data release28.
Head motion measured by framewise displacement (FD). (a) Raw and (b) low-pass filtered (LPF) temporal mean FD for all task and resting-state fMRI runs. The scatter plots compare typical (temporal median) and upper range (temporal 95th percentile) FD for all fMRI runs. (c) Temporal median LPF FD by task (“Bang, You’re Dead!”, “Partly Cloudy”) and resting-state, and (d) by MRI protocol version.
Temporal SNR
Temporal signal-to-noise ratio (tSNR) was calculated by MRIQC for each fMRI series. Raw tSNR estimates were normalized to voxel volume and EPI repetition time (TR) to allow comparison between sequence variants with different multiband acceleration factors and spatial resolutions (Fig. 8).
Mean, whole-brain temporal SNR normalized to repetition time and voxel volume for comparison between sequence protocol variants. Multiband protocol variants consistently perform between 2.5 and 3 times better than the single band variant (core1p1) in terms of tSNR efficiency. (a) Mid-coronal sections of the normalized tSNR efficiency (raw tSNR calculated by the MRIQC pipeline, adjusted for voxel volume and TR) averaged over all available subjects for the initial single-band protocol (core1p1) and (b) second phase multi-band T2*-weighted EPI protocol (core2p2) demonstrating the increase in normalized tSNR efficiency offered by multiband acquisitions despite the reduction in spatiotemporal resolution from 3.0 mm and 2.5 s to 2.5 mm and 0.7 s. (c) Normalized tSNR distributions within the brain, showing an approximately three-fold increase in mean normalized tSNR with the core2p2 protocol. (d) Whole-brain averaged normalized tSNR distributions for each task and protocol version.
Usage Notes
Limitation and opportunities of an in-depth sample of small size
As compared to other multimodal data releases such as the HCP or UK BioBank, the sample size of the present release is small. It is by now well known that small sample sizes severely limit the statistical reliability of conclusions that can be drawn about individual differences using neuroimaging data68,69,70, in line with a general upwards correction for the statistical reliability of correlations between datasets71. Generalization of findings regarding individual differences is thus limited in our dataset, although the details will vary depending on the exact question asked and method used72. As we have recommended previously70, we encourage the use of a predictive framework (using cross-validation within the dataset and/or replication to other, independent datasets), permutation-based statistical evaluation, and where feasible pre-registration in order to minimize the risk of false positive findings. A recent example based on a subset of the present dataset (prior to its further processing and release) illustrates that valuable negative findings, as well as estimates of sample sizes required for future studies, can be derived from this dataset73. We would anticipate that the present data release may be more valuable for adding cautionary notes and power estimates to the literature than for strong demonstrations of positive findings.
Nonetheless, the dataset is distinguished by its in-depth and comprehensive psychological and behavioral assessments, especially in the domain of social cognition and decision making. We note that the factor analysis that we also provide (Fig. 4), while of interest in its own right, in no way precludes more fine-grained investigation of the original individual variables. Indeed, we would recommend that the factors be considered as broader covariates in analyses that wish to isolate variance in a specific individual behavioral variable more selectively. The in-depth behavior data together with high-quality neuroimaging data provides a powerful platform to discover new brain-behavior relationships even with our modest sample size, since the measurement error of the variables is no less important than the sample size. However, we would expect such positive discoveries to be relatively constrained, ideally driven by specific pre-registered hypotheses. One possible research program could thus consist of an initial discovery study in a large-sample database, such as the UK Biobank, followed with a hypothesis-driven replication of the finding in our database—where the relevant variables are provided both with greater precision and, for the behavioral data, likely greater validity. The breadth of psychological characterization in our data release (see Table 1) provides further opportunities for comparison with other databases, where related cognitive variables are estimated from less detailed assessments. Applications of “far replicability”74 could be extended to databases in clinical populations (e.g., of participants with psychiatric diagnoses of depression, anxiety, autism, schizophrenia, and other disorders that impact social cognition and decision-making).
Our data release is also distinguished by providing multiple data formats and degrees of preprocessing. This affords the opportunity to test results, for instance, against variations in denoising decisions in an accessible and straightforward manner, as a further check on the robustness of findings to variations in typically complex processing pipelines, a well-known source of variation in the results obtained75,76. The denoising code we are co-releasing, in particular, allows researchers to explore a range of processing pipelines with substantial flexibility. Taken together, the internal processing flexibility enabled by this data release, together with the above recommendations to interface the present data with other datasets that purport to measure similar variables, should aim to maximize the meaningful generalizability of findings.
Note on the informed usage notes and quality control (QC)
We highlight below some processing and quality-specific aspects regarding the MRI data of this release.
We have used a combination of manual (human) and automated quality inspection of both structural (human: manual visual inspection and editing of FreeSurfer outputs; automated: MRIQC) and functional MRI data (human: manual visual inspection and resulting QC rating data; automated: MRIQC; see fMRIPrepQC_ratings.csv). We provide the outputs of our careful QC with the actual data resulting from it. It is the responsibility of the end-users to use the information available depending on their intended use of the data and study-specific QC criteria. For example, rigor and attention to minor surface reconstruction errors might be less strict for studies that aim to use cortical reconstruction outputs only for surface-based registration. In contrast, for a specific volumetric study (e.g., cortical thickness analysis), one might be less lenient. Note that given in vivo data (as well as the current possible imaging resolution) there is no clear “ground truth” for anatomical tissue segmentations, beyond consensus in human judgment of the images. In addition, remaining image quality aspects due to factors such as motion and regional susceptibility effects (e.g., in inferior temporal brain regions) cannot be eliminated post hoc and result in residual imprecision in individual data. These and other intrinsic measurement errors in our dataset require users to apply expert judgment in how they use the data release to answer specific scientific questions of interest.
For example, caution should be applied when using functional data in orbital frontal regions and data processed with fMRIPrep. As of the submission date of this paper, there is a known issue with susceptibility distortion correction (SDC) using spin echo fieldmaps as implemented in fMRIPrep. fMRIPrep currently uses AFNI’s 3dQwarp function to implement distortion correction, which can produce suboptimal SDC outputs in some subjects (see https://github.com/nipreps/fmriprep/issues/2210). While issues such as this are not a result of our specific data, they can be serious issues that require knowledge about the limitations inherent to MRI and established processing tools, an ongoing set of issues actively discussed among expert users.
Code availability
We used containerized versions of fMRIPrep 20.2.1 and MRIQC for data preprocessing and quality control. Example calling scripts for fMRIPrep, jupyter lab notebooks for figure recreation and R code for the example factor analysis are provided at https://github.com/adolphslab/ConteDataRelease.
The code to reproduce resting-state and movie analyses are provided at https://github.com/adolphslab/rsDenoise. As outlined in detail in the source, this codebase can easily be adapted to run many different configurations of denoising decisions on the data.
References
Poldrack, R. A. et al. Long-term neural and physiological phenotyping of a single human. Nat. Commun. 6, 8885 (2015).
Gordon, E. M. et al. Precision Functional Mapping of Individual Human Brains. Neuron 95, 791–807.e7 (2017).
Sudlow, C. et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Spreng, R. N. et al. The default network of the human brain is associated with perceived social isolation. Nat. Commun. 11, 6393 (2020).
Elliott, L. T. et al. Genome-wide association studies of brain imaging phenotypes in UK Biobank. Nature 562, 210–216 (2018).
Zabihi, M. et al. Non-linearity matters: a deep learning solution to generalization of hidden brain patterns across population cohorts. https://doi.org/10.1101/2021.03.10.434856.
Wechsler, D. Wechsler Adult Intelligence Scale - 4th edition. (Pearson, 2008).
Van Essen, D. C. et al. The WU-Minn Human Connectome Project: An overview. Neuroimage 80, 62–79 (2013).
Gorgolewski, K. J. et al. The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Sci Data 3, 160044 (2016).
Esteban, O. et al. fMRIPrep: a robust preprocessing pipeline for functional MRI. Nat. Methods 16, 111–116 (2019).
Fischl, B. FreeSurfer. Neuroimage 62, 774–781 (2012).
Esteban, O. et al. MRIQC: Advancing the automatic prediction of image quality in MRI from unseen sites. PLoS One 12, e0184661 (2017).
Baron-Cohen, S. & Wheelwright, S. The empathy quotient: an investigation of adults with Asperger syndrome or high functioning autism, and normal sex differences. J. Autism Dev. Disord. 34, 163–175 (2004).
Baron-Cohen, S., Richler, J., Bisarya, D., Gurunathan, N. & Wheelwright, S. The systemizing quotient: an investigation of adults with Asperger syndrome or high-functioning autism, and normal sex differences. Philos. Trans. R. Soc. Lond. B Biol. Sci. 358, 361–374 (2003).
Constantino, J. N. & Gruber, C. P. Social Responsiveness Scale Second Edition (SRS-2). (Western Psychological Services, 2012).
Mayer, J., Salovey, P. & Caruso, D. Mayer-Salovey-Caruso Emotional Intelligence Test Manual. (Multi-Health Systems, 2002).
Beck, A. T., Steer, R. A. & Brown, G. K. BDI-II, Beck Depression Inventory: Manual. (Psychological Corporation, 1996).
Reher, P. S. Partly Cloudy [Motion Picture] (Pixar Animation Studios and Walt Disney Pictures (2009).
Naci, L., Cusack, R., Anello, M. & Owen, A. M. A common neural code for similar conscious experiences in different individuals. PNAS Proceedings of the National Academy of Sciences of the United States of America, 111(39), 14277–14282 (2014).
Reuter, M., Schmansky, N. J., Rosas, H. D. & Fischl, B. Within-subject template estimation for unbiased longitudinal image analysis. Neuroimage 61, 1402–1418 (2012).
Desikan, R. S. et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 31, 968–980 (2006).
Destrieux, C., Fischl, B., Dale, A. & Halgren, E. Automatic parcellation of human cortical gyri and sulci using standard anatomical nomenclature. Neuroimage 53, 1–15 (2010).
Reuter, M., Rosas, H. D. & Fischl, B. Highly accurate inverse consistent registration: a robust approach. Neuroimage 53, 1181–1196 (2010).
Sled, J. G. & Bruce Pike, G. Understanding intensity non-uniformity in MRI. Medical Image Computing and Computer-Assisted Intervention — MICCAI’98 614–622, https://doi.org/10.1007/bfb0056247 (1998).
Fischl, B. et al. Automatic segmentation of the structures in the human brain. NeuroImage 13, 118 (2001).
Segonne, F., Pacheco, J. & Fischl, B. Geometrically Accurate Topology-Correction of Cortical Surfaces Using Nonseparating Loops. IEEE Transactions on Medical Imaging 26, 518–529 (2007).
Gorgolewski, K. et al. Nipype: a flexible, lightweight and extensible neuroimaging data processing framework in python. Front. Neuroinform. 5, 13 (2011).
Kliemann, D. et al. Caltech Conte Center, a multimodal data resource for exploring social cognition and decision-making, OpenNeuro, https://openneuro.org/datasets/ds003798 (2021).
Tustison, N. J. et al. N4ITK: improved N3 bias correction. IEEE Trans. Med. Imaging 29, 1310–1320 (2010).
Avants, B.B., Tustison, N. J., Johnson, H. J. Advanced Normalization Tools (ANTs) at https://github.com/ANTsX/ANTs.
Zhang, Y., Brady, M. & Smith, S. Segmentation of brain MR images through a hidden Markov random field model and the expectation-maximization algorithm. IEEE Trans. Med. Imaging 20, 45–57 (2001).
Fonov, V. S., Evans, A. C., McKinstry, R. C., Almli, C. R. & Collins, D. L. Unbiased nonlinear average age-appropriate brain templates from birth to adulthood. Neuroimage Supplement 1, S102 (2009).
Evans, A. C., Janke, A. L., Collins, D. L. & Baillet, S. Brain templates and atlases. Neuroimage 62, 911–922 (2012).
Cox, R. W. & Hyde, J. S. Software tools for analysis and visualization of fMRI data. NMR Biomed. 10, 171–178 (1997).
Greve, D. N. & Fischl, B. Accurate and robust brain image alignment using boundary-based registration. Neuroimage 48, 63–72 (2009).
Jenkinson, M., Bannister, P., Brady, M. & Smith, S. Improved Optimization for the Robust and Accurate Linear Registration and Motion Correction of Brain Images. Neuroimage 17, 825–841 (2002).
Glasser, M. F. et al. The minimal preprocessing pipelines for the Human Connectome Project. Neuroimage 80, 105–124 (2013).
Power, J. D. et al. Methods to detect, characterize, and remove motion artifact in resting state fMRI. Neuroimage 84, 320–341 (2014).
Behzadi, Y., Restom, K., Liau, J. & Liu, T. T. A component based noise correction method (CompCor) for BOLD and perfusion based fMRI. Neuroimage 37, 90–101 (2007).
Dubois, J., Galdi, P., Han, Y., Paul, L. K. & Adolphs, R. Resting-state functional brain connectivity best predicts the personality dimension of openness to experience. Personal Neurosci 1 (2018).
Dubois, J., Galdi, P., Paul, L. K. & Adolphs, R. A distributed brain network predicts general intelligence from resting-state human neuroimaging data. Philos. Trans. R. Soc. Lond. B Biol. Sci. 373 (2018).
Virtanen, P. et al. Author Correction: SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 352 (2020).
Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362 (2020).
Abraham, A. et al. Machine learning for neuroimaging with scikit-learn. Front. Neuroinform. 8, 14 (2014).
Pedregosa, F. et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
McKinney, W. Data structures for statistical computing in python. in Proceedings of the 9th Python in Science Conference 445, 51–56 (Austin, TX, 2010).
Hunter, J. D. Matplotlib: A 2D Graphics Environment. Computing in Science & Engineering 9, 90–95 (2007).
Finn, E. S. et al. Functional connectome fingerprinting: identifying individuals using patterns of brain connectivity. Nat. Neurosci. 18, 1664–1671 (2015).
Satterthwaite, T. D. et al. An improved framework for confound regression and filtering for control of motion artifact in the preprocessing of resting-state functional connectivity data. Neuroimage 64, 240–256 (2013).
Siegel, J. S. et al. Data Quality Influences Observed Links Between Functional Connectivity and Behavior. Cereb. Cortex 27, 4492–4502 (2017).
Kong, R. et al. Spatial Topography of Individual-Specific Cortical Networks Predicts Human Cognition, Personality, and Emotion. Cerebral Cortex 29, 2533–2551 (2019).
Wechsler, D. WASI: Wechsler abbreviated scale of intelligence. (Psychological Corporation, 1999).
Wechsler, D. Wechsler Abbreviated Scale of Intelligence - Second Edition. (Psychological Corporation, 2011).
Spielberger, C. D. Manual for the State-trait Anxiety Inventory (form Y) (‘self-evaluation Questionnaire’). (Consulting Psychologists Press, 1983).
R Core Team. R: A Language and Environment for Statistical Computing. at https://www.R-project.org/ (R Foundation for Statistical Computing, 2017).
Çokluk, Ö. & Koçak, D. Using Horn’s Parallel Analysis Method in Exploratory Factor Analysis for Determining the Number of Factors. Educational Sciences: Theory and Practice 16, 537–551 (2016).
Cattell, R. B. The Scree Test For The Number Of Factors. Multivariate Behav. Res. 1, 245–276 (1966).
Gorsuch, R. L. & Nelson, J. CNG scree test: an objective procedure for determining the number of factors. (1981).
Zoski, K. & Jurs, S. Using multiple regression to determine the number of factors to retain in factor analysis. Multiple Linear Regression Viewpoints 20, 5–9 (1993).
Velicer, W. F. Determining the number of components from the matrix of partial correlations. Psychometrika 41, 321–327 (1976).
Velicer, W. F., Eaton, C. A. & Fava, J. L. In Problems and Solutions in Human Assessment: Honoring Douglas N. Jackson at Seventy (eds. Goffin, R. D. & Helmes, E.) 41–71, https://doi.org/10.1007/978-1-4615-4397-8_3 (Springer US, 2000).
Esteban, O. et al. Crowdsourced MRI quality metrics and expert quality annotations for training of humans and machines. Sci Data 6, 30 (2019).
Power, J. D., Barnes, K. A., Snyder, A. Z., Schlaggar, B. L. & Petersen, S. E. Spurious but systematic correlations in functional connectivity MRI networks arise from subject motion. Neuroimage 59, 2142–2154 (2012).
Fair, D. A. et al. Correction of respiratory artifacts in MRI head motion estimates. Neuroimage 208, 116400 (2020).
Gratton, C. et al. Removal of high frequency contamination from motion estimates in single-band fMRI saves data without biasing functional connectivity. Neuroimage 116866, https://doi.org/10.1016/j.neuroimage.2020.116866 (2020).
Power, J. D. et al. Distinctions among real and apparent respiratory motions in human fMRI data. Neuroimage 201, 116041 (2019).
Williams, J. C. & Van Snellenberg, J. X. Motion denoising of multiband resting state functional connectivity MRI data: An improved volume censoring method. bioRxiv 860635, https://doi.org/10.1101/860635 (2019).
Button, K. S. et al. Power failure: why small sample size undermines the reliability of neuroscience. Nat. Rev. Neurosci. 14, 365–376 (2013).
Poldrack, R. A. et al. Scanning the horizon: towards transparent and reproducible neuroimaging research. Nat. Rev. Neurosci. 18, 115–126 (2017).
Dubois, J. & Adolphs, R. Building a Science of Individual Differences from fMRI. Trends Cogn. Sci. 20, 425–443 (2016).
Schönbrodt, F. D. & Perugini, M. At what sample size do correlations stabilize? J. Res. Pers. 47, 609–612 (2013).
Grady, C. L., Rieck, J. R., Nichol, D., Rodrigue, K. M. & Kennedy, K. M. Influence of sample size and analytic approach on stability and interpretation of brain-behavior correlations in task-related fMRI data. Hum. Brain Mapp. 42, 204–219 (2021).
Lin, C. et al. No strong evidence that social network index is associated with gray matter volume from a data-driven investigation. Cortex 125, 307–317 (2020).
Nichols, T. E. et al. Best practices in data analysis and sharing in neuroimaging using MRI. Nat. Neurosci. 20, 299–303 (2017).
Kennedy, D. N. et al. Everything Matters: The ReproNim Perspective on Reproducible Neuroimaging. Front. Neuroinform. 13, 1 (2019).
Bhagwat, N. et al. Understanding the impact of preprocessing pipelines on neuroimaging cortical surface analyses. Gigascience 10 (2021).
Cohen, S., Kamarck, T. & Mermelstein, R. A global measure of perceived stress. J. Health Soc. Behav. 24, 385–396 (1983).
Cohen, S. & Williamson, G. In The Social Psychology of Health (eds. Spacapan, S. & Oskamp, S.) (Sage, 1988).
Watson, D., Clark, L. A. & Carey, G. Positive and negative affectivity and their relation to anxiety and depressive disorders. J. Abnorm. Psychol. 97, 346–353 (1988).
Cattell, R. B., Eber, H. W. & Tatsuoka, M. M. Handbook for the Sixteen Personality Factor Questionnaire (16PF). (Institute for Personality and Ability Testing, 1970).
Russell, M. T. & Karol, D. L. The 16PF fifth edition administrator’s manual: with updated norms. 3rd Edition. (Institute for Personality and Ability Testing, 2002).
Cohen, S., Doyle, W. J., Skoner, D. P., Rabin, B. S. & Gwaltney, J. M. Jr. Social ties and susceptibility to the common cold. JAMA 277, 1940–1944 (1997).
Schaefer, A. et al. Local-Global Parcellation of the Human Cerebral Cortex from Intrinsic Functional Connectivity MRI. Cereb. Cortex 28, 3095–3114 (2018).
Yeo, B. T. T. et al. The organization of the human cerebral cortex estimated by intrinsic functional connectivity. J. Neurophysiol. 106, 1125–1165 (2011).
Acknowledgements
Conceptualization (DK, RA, RN, LKP, JMT), Data Curation (DK, PG, DAK, TR, AZE, DL, SL, WZ, RN, JMT), Formal analysis (DK, TA, PG, RLY, LKP, JMT), Funding Acquisition (RA, LKP, JMT), Data Collection (DK, TA, RN, JMT), Protocol/Task Implementation (TA, JMT, LKP), Recruitment & Enrollment (TA, LKP), Project administration & Supervision (DK, RA, RN, LKP, JMT), Writing of the manuscript (DK, RA, TA, PG, DAK, TR, AZE, DL, SL, WZ, RLY, RN, LKP, JMT). Given the complex set of contributions from many authors, we provide a figure in the supplementary material (Figure S1) to summarize their contributions. Contributions reflect the organization of the Caltech Conte Center, with R.A. as director, R.N. as staff responsible for NIH-compliant data-sharing, L.K.P. as PI of a Psychological Assessment Core, and J.M.T. as PI of a Neuroimaging Core. Other notable contributions were essentially all in-person subject testing by T.A., and generation of the denoising code we are sharing here (rsDenoise) by P.G.
Author information
Authors and Affiliations
Contributions
Given the complex set of contributions from many authors, we provide the figure below (Figure S1) to summarize their contributions. Contributions reflect the organization of the Caltech Conte Center, with R.A. as director, R.N. as staff responsible for NIH-compliant data-sharing, L.K.P. as PI of a Psychological Assessment Core, and J.M.T. as PI of a Neuroimaging Core. Other notable contributions were essentially all in-person subject testing by T.A. and generation of the denoising code we are sharing here (rsDenoise) by P.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files associated with this article.
About this article
Cite this article
Kliemann, D., Adolphs, R., Armstrong, T. et al. Caltech Conte Center, a multimodal data resource for exploring social cognition and decision-making. Sci Data 9, 138 (2022). https://doi.org/10.1038/s41597-022-01171-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-022-01171-2