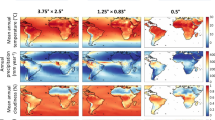
Abstract
Curated global climate data have been generated from climate model outputs for the last 120,000 years, whereas reconstructions going back even further have been lacking due to the high computational cost of climate simulations. Here, we present a statistically-derived global terrestrial climate dataset for every 1,000 years of the last 800,000 years. It is based on a set of linear regressions between 72 existing HadCM3 climate simulations of the last 120,000 years and external forcings consisting of CO2, orbital parameters, and land type. The estimated climatologies were interpolated to 0.5° resolution and bias-corrected using present-day climate. The data compare well with the original HadCM3 simulations and with long-term proxy records. Our dataset includes monthly temperature, precipitation, cloud cover, and 17 bioclimatic variables. In addition, we derived net primary productivity and global biome distributions using the BIOME4 vegetation model. The data are a relevant source for different research areas, such as archaeology or ecology, to study the long-term effect of glacial-interglacial climate cycles for periods beyond the last 120,000 years.
Measurement(s) | temperature • precipitation • cloud cover • net primary productivity |
Technology Type(s) | computational modeling technique |
Factor Type(s) | atmospheric carbon dioxide • orbital parameters • land surface type (ocean, land, land ice) |
Sample Characteristic - Environment | climate system |
Sample Characteristic - Location | Earth (planet) |
Machine-accessible metadata file describing the reported data: https://doi.org/10.6084/m9.figshare.15073479
Similar content being viewed by others
Background & Summary
Studying the ecology and environment throughout past climatic changes often involves environmental reconstructions that are either based on paleoclimate proxies or on paleoclimate simulations. Unfortunately, even by today’s standards, simulating the climate over periods of thousands or hundreds of thousands of years in a continuous way can still be a costly and time-consuming endeavour. Present or future climate simulations are based on comprehensive Global Climate Models (GCMs) that resolve processes at high temporal and spatial resolution, such as those used in the fifth IPCC Assessment Report1. Climate model reconstructions for longer, continuous periods back in time are, therefore, challenging. They have to span a much longer period and are, thus, computationally too expensive. Instead, GCMs provide snapshots for a specific time or short transients in the order of a few thousand years. For longer, transient simulations of tens or hundreds of thousands of years, we rely on simulations from Earth System Models of Intermediate Complexity (EMICs)2,3 but they come at the cost of lower spatial resolution and a simplified representation of the climate system4.
Although there are a some high-resolution paleoclimate data sets readily available for download, for example, WorldClim5, PaleoClim6, or ecoClimate7, their temporal coverage is limited to a few snapshots of key periods in the past, for example, Mid-Holocene (6,000 years before present (BP)) the Last Glacial Maximum (21,000 years BP), or the Last Interglacial Period (130,000 years BP). An exception is PaleoView8 which covers the transient period of the last deglaciation, but this only goes back 21,000 years. Longer, continuous climate data sets of the past, based on HadCM39 snapshots, have become available more recently, for example a Northern Hemisphere data set for the last 60,000 years10 or a bias-corrected, high-resolution terrestrial climate data set of the last 120,000 years11.
Here, we used a linear regression model to extend existing HadCM3 climate simulations of the last 120,000 years to create climate reconstructions of the last 800,000 years. We then applied a bias correction12 of the model output using present-day gridded observational data (CRU TS v. 4.0413) to downscale climate output to a final horizontal resolution of 0.5°11,13. This new data set is complementary to the aforementioned high-resolution terrestrial climate data set of the last 120,000 years11 (which should be preferred for studies of the last glacial cycle, as they are based directly on the GCM output), and it is an extension for readers to explore the climate history for earlier periods of the past.
In this paper, we present annual and monthly mean climatologies for the last 800,000 years in 1,000 year time steps (Table 1). The data set includes air temperature, precipitation, total cloud cover, 17 bioclimatic variables14, as well as biomes and annual and monthly net primary productivity, the latter based on BIOME4 simulations15, run on the debiased climatologies. We validated the long-term climate change signal using time series of various proxy records.
Methods
Our climate reconstructions are based on a set of linear regression models for each of the HadCM3 model grid boxes (N = nlon × nlat = 96 × 73 = 7008). Each linear model predicts a climate variable, which can be either temperature, precipitation, or total cloud cover, i.e., the dependent variable. The independent variables, i.e., the forcing terms, of the model are three orbital parameters, atmospheric CO2, and a surface type mask (land, ocean, or land ice), five variables in total.
Each linear model uses 72 data points given by the HadCM3 snapshots throughout the past 120 thousand years (ka)16,17. These snapshots cover both the Last Glacial Maximum, one of the coldest glacial stages, and the Last Interglacial, one of the warmest interglacial stages during the Middle and Late Pleistocene. By applying available long-term forcing to the solutions of the linear models, we reconstructed the climate for periods before 120 ka. The forcing consists of CO218, interpolated to 1 ka intervals for the last 800 ka, orbital parameters, taken from numerical solution to the Earth’s orbit around the sun19, and surface type masks based on numerical ice-sheet model output2 and a global sea-level record20. At this stage, the reconstructed climate of the last 800,000 years has the same coarse spatial resolution as the underlying HadCM3 snapshots. In a last step, we applied a bias correction (including spatial downscaling) for the terrestrial climate to derive a spatially explicit data set that covers the last 800 ka in 1 ka intervals with a spatial resolution of 0.5° × 0.5°. For each variable, these steps were repeated for each monthly mean climatology (Jan–Dec), as well as for the annual mean values (Table 1).
A comprehensive overview of our approach is shown in Fig. 1 and further details of our experimental setup are given below.
Flowchart showing how the different data sets have been used as input for the different stages of the paleo-climate data generation: A linear regression combines 72 low-resolution (LR) HadCM3 snapshot simulations with the external forcings, i.e., CO2, orbital parameters, and surface type masks (ocean, land, land ice), which provides the basis of the long-term climate reconstructions using long-term forcings and surface type masks. The final bias correction procedure yields the high-resolution (HR), bias-corrected (BC) climate data set for the last 800 ka.
The HadCM3 climate model
HadCM3 is a fully coupled global climate model with an atmospheric component, HadAM3, which has a horizontal resolution of 3.75° × 2.5°, 19 vertical levels, and a time step of 30 minutes. The ocean and sea-ice component of HadCM3 has a horizontal resolution of 1.25° × 1.25° and 20 vertical levels. HadCM3 simulations were run with a prescribed ice-sheet and continental geometry. We used output from the atmospheric component of the 72 available HadCM3 simulations covering the last 120,000 years in 2,000-year intervals from 120,000 to 24,000 years BP and in 1,000-year intervals from 22,000 years BP to present-day16,17.
Surface type mask: Ice-sheet extents, sea level and lakes
As ice sheet extents for the period outside the HadCM3 snapshots, we used model outputs from CLIMBER-2/SICOPOLIS simulations2 for which Northern Hemisphere ice sheet extents and heights are available for the last 800 ka in 1 ka-year intervals. For the more recent period from 122–0 ka, we used the ice sheet configurations from the ICE-6G data set21 (http://www.atmosp.physics.utoronto.ca/ peltier/data.php). Changes in the coast lines affecting the land–sea mask were derived from a global sea-level record20. We overlaid those changes on top of present-day coast lines, taken from the ETOPO1 data set22 (https://ngdc.noaa.gov/mgg/global/global.html), while we preserved inland lakes which were taken from the Global Lakes and Wetlands Database23 (https://www.worldwildlife.org/pages/global-lakes-and-wetlands-database).
Training and test data
We divided the HadCM3 snapshots into a training (80%) and a test data set (20%). The training data set was used to fit the linear model while the test data was used for a comparison to the reconstructed snapshots. For a 80/20 division of the 72 time slices into training and test data, i.e., 58/14, there are \(\left(\begin{array}{c}n\\ k\end{array}\right)=\left(\begin{array}{c}72\\ 58\end{array}\right)\approx 3\times 1{0}^{14}\) possible combinations of snapshots. But instead of randomly dividing the snapshots into the training/test data, we followed an approach with the aim to preserve as much variance as possible in the training data, i.e. maximise the variance of the predictors. This is best illustrated by the phase plots of the parameters, i.e., the predictors (Fig. 2). The training data set covers the edges of each phase plane and thus maximises the phase space covered by the linear regression model. This choice of training data ensured that the linear regression model interpolated within the phase space and did not need to extrapolate for the test data.
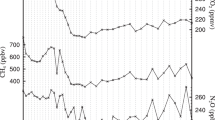
(a) Time series of the four external parameters: CO2 and orbital parameters for the last 800 ka and (b) the associated parameter space as scatter plot matrix (blue dots). The continuous CO2 record is from the EPICA Dome C ice core in Antarctica18. The orbital parameters are numerical solutions for the Earth’s orbit and rotation in terms of eccentricity, precession, and obliquity19. In (b), black lines with black dots represent the total 72 parameter sets. Orange dots highlight the parameter sets of the 58 HadCM3 snapshot simulations which we used as training data (80% of the total 72) for the linear regression model.
The procedure was as follows. We calculated the covariance matrix of the full parameter set (n = 72), Cfull. Then, we randomly created a sample training data set (k = 58) for which we computed the covariance matrix Csample. If the eigenvalues of Csample were larger than the eigenvalues of Cfull, then the training sample data set contained at least as much variance as the full data set and this sample training data set was marked as a candidate for the final training set. After several iterations (N = 10,000), we summed up how many times each time slice had appeared within a candidate training set. We then ranked all time slices according to this number. In the final step, we picked the 80% top-ranked time slices as training data.
Note, the division into training and test data set was made only for the validation of the linear model. For the actual climate reconstructions with the linear regression model, we used all 72 snapshots to make full use of the complete set of available data.
The linear regression model
For each HadCM3 grid box, we fitted a linear regression model to a local series of each climatic variable of interest (temperature, precipitation, or cloud cover) with the following independent variables or forcings: atmospheric CO2 concentrations (as a major greenhouse gas), three variables reflecting the orbital forcing19, and the surface type, which is either ocean, land, or land ice. The orbital parameters are obliquity ε and two combinations of eccentricity e and precession ω: e · sin ω, henceforth referred to as precession index I, and e · cos ω (precession index II), and they are a generally accepted set of orbital forcings24,25. We chose temperature T, precipitation P, and total cloud cover C as dependent variables. The independent variables are given by the normalised forcings.
More formally, let Y(x, t) be a time series of a climate variable in a specific grid box x at time t. Our linear model should explain variations of Y(x, t), ΔY(x, t), around a mean value \(\bar{Y(x,t)}\):
To make the linear model well-conditioned, all independent variables were normalised. The mean was subtracted and the result were then divided by the standard deviation.
Precipitation and total cloud cover are bounded variables which can lead to linear model predictions outside of physically meaningful ranges. A common procedure to prevent these out of bounds predictions is to apply a transformation to the data beforehand. To prevent the linear model from predicting negative precipitation values, we therefore applied a logarithmic transformation to precipitation, which maps values from [0, +∞] to [−∞, +∞]. Thus, in the case of precipitation, the linear regression coefficients predict the response in terms of anomalies in the exponent. For total cloud cover, expressed as fraction between 0 and 1, we choose a logit transformation which maps values from [0, 1] to [−∞, +∞]. Note that the minimum values for precipitation and total cloud cover, as simulated by HadCM3, are never exactly zero (or smaller), except at the poles, 90° N/S, which were excluded for that reason; therefore, the transformations always yield finite values. The decomposition of temperature T, precipitation P, and total cloud cover C, i.e., the ΔY(x, t) on the left hand side of Eq. (1) is:
The linear regression model for each (transformed) anomaly is:
Here, β1 to β5 are the regression coefficients for the respective predictors (see Fig. 3 for maps of β coefficients). Surface type changes are captured by the categorical variable M(x, t) ∈ [ocean, land, land ice]. For example, around coastlines land grid boxes can turn into ocean grid boxes when sea level is high. Similarly, expanding ice sheets turn land grid boxes into ice-covered grid boxes, and the climate variable Y(x, t) may respond to different surface types in different ways. The categorical variables were encoded using Treatment coding. The first level, ocean, was chosen as a reference level and is by definition zero β5(x) = 0. For the other two levels, land and land ice, a different β5(x) was assigned for each—in effect, a different intercept for ΔY(x, t).
We solved the linear model for each transformed variable, applied the extended forcing to generate the 800 ka climate reconstructions, and transformed the resulting data back to its original range according to Eqs. (2–4). At this stage, we have an extended HadCM3 model output of annual and monthly mean for temperature, precipitation, and total cloud cover, still at the original HadCM3 resolution, for the last 800 ka.
Spatial downscaling to 0.5° and bias correction
The CRU TS climate data set
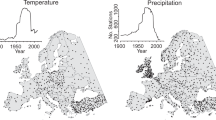
For the bias correction of the extended HadCM3 model output, as predicted by the previously described linear model, we used variables from the CRU TS (Climatic Research Unit Timeseries) data set (v. 4.04)013. Before the bias correction step, the data set has been bi-linearly interpolated from the spatial resolution of 3.75° × 2.5° to the CRU TS resolution of 0.5° × 0.5°. CRU TS v. 4.04 contains monthly time series fields of precipitation, daily maximum and minimum temperatures, cloud cover, and other variables covering all land areas (except Antarctica) for 1901 to 2019. As reference period for the bias correction we chose 1961–1990. We applied the additive “delta” method, which is the most effective bias correction with respect to paleoclimate reconstructions12, to all climate variables predicted by the linear model (subscript LM) to create the final, bias-corrected climate data set \(\widehat{Y}(x,t)\):
The BIOME4 vegetation model
We used the BIOME4 model15 to calculate annual net primary productivity (NPP) and to determine the global distribution of biomes. BIOME4 is a coupled biogeography and biogeochemistry model that simulates competition of different plant functional types (PFTs). It optimises the leaf area of each PFT as a function of NPP. BIOME4 forcing consists of monthly values of temperature, precipitation, and sunshine percentage, as well as values of annual minimum temperature and atmospheric CO2 concentrations. Sunshine percentage was computed using total cloud cover26. For atmospheric CO2 we used the same data set as described earlier18. In its default setup, BIOME4 does not incorporate orbital variations that would affect top-of-atmosphere (TOA) insolation. Instead, it is approximated by a cosine function representative of present-day insolation only. We therefore updated the TOA insolation representation in BIOME4 so that it takes changes in the Earth’s orbit into account27. Further inputs, which were kept constant through time, are water holding capacity and percolation rate.
Data Records
All data records are publicly available as NetCDF files in the project repository in the data directory28.
Monthly mean and annual mean climatologies
The climate variables that are part of our data set are listed in Table 1 and can be downloaded as NetCDF files (Table 2) from the Open Science Framework data repository28. All climate variables are available on a 0.5° × 0.5° resolution, i.e., the same regular grid as the CRU TS v4.04 data set13.
Bioclimatic variables
For ecological applications such as species distribution modelling, bioclimatic variables are more relevant than the actual climate variables because they capture information about annual and seasonal climate conditions as reflected by temperature and precipitation14, for example coldest/warmest or wettest/driest quarter averages of precipitation and temperature. Most of the commonly required bioclimatic variables can be directly derived from monthly mean temperature and precipitation data, and are thus included in our data records (Tables 1, 2). Annual Mean Diurnal Range (BIO2) and Isothermality (BIO3) cannot be calculated from the available climate model data (HadCM3) and are therefore not included. For BIO2, we do not have the monthly minimum and maximum temperature, and BIO3 depends on BIO2 (BIO3 = BIO2/BIO7 × 100).
Net primary productivity and biomes
Our data record contains annual and monthly net primary productivity as well as categorical biome data, both of which were calculated with the BIOME4 vegetation model, using the 800 ka of reconstructed and bias-corrected climate (Tables 1, 2).
Technical Validation
Comparison to original HadCM3 simulations
We validated our reconstruction from the linear model against the original HadCM3 snapshots. As comparison metric, we used R2 values, a goodness of fit estimator that measures the proportion of variance explained by the linear model, and the root mean squared error (RMSE), an estimator of the goodness of the model that measures how far the linear model predictions are from the HadCM3 test data (Fig. 4).
Left panel (a,c,e): Root mean square errors (RMSE) as estimators of the goodness of fit (lower is better) calculated using the test data. Right panel (b,d,f ): R2 values as estimator for the goodness of the model (higher is better) using the training data. Shown are the R2 and RMSEs for (a,b) mean annual temperature, (c,d) mean annual precipitation and (e,f) mean annual total cloud cover. Note, that only the values over land and land ice areas are relevant for the overall quality of the final data product.
Overall, our linear model is a better predictor for temperature than for precipitation and total cloud cover. Temperature responds more directly to local forcings than precipitation and cloud cover, because it is determined by the energy balance of downward and upward longwave and shortwave radiation and turbulent heat fluxes. The downward shortwave radiation depends on incoming solar radiation that is determined by orbital variations, whereas downward longwave radiation is determined by greenhouse gases such as CO2 and water vapour, as well as cloud cover. Large-scale atmospheric circulation changes have a much smaller effect on temperature. The high R2 and low RMSE values in most regions (Fig. 4a,b) mean that temperature is locally well constrained by global CO2 and orbital variations and our linear model captures this effect well.
The matter is more complicated for precipitation and cloud cover. Both variables are directly affected by the hydrological cycle which itself depends on large-scale atmospheric dynamics, such as monsoonal systems in the tropics and subtropics, or mid-latitude storm systems. Local interactions between the atmosphere and the surface, such as evaporation and transpiration over the ocean, or deep convection over the tropics, matter to a lesser extent. Instead, processes and circulation features like moisture transport or the atmospheric Hadley cell dynamics determine the non-local response of precipitation (and cloud cover) to CO2 or orbital variations to a much larger extent. Because of the larger dynamical component of the hydrological cycle, precipitation and cloud cover are much less constrained by the forcing than temperature. As a result, the linear model shows less predictive skill for precipitation and total cloud cover (Fig. 4c–f).
Our reconstruction represented only long-term climatologies of past climate changes, similar to other GCM snapshot of the past11,16. Therefore, the data set does not contain sub-millennial scale variability.
The spatial and temporal covariance of the model output
Our climate reconstructions are based on a pixel-specific linear model, one for each of the HadCM3 grid boxes. By design, spatial autocorrelation is not an issue, which it would be if we were to analyse all points simultaneously. In that case, spatial autocorrelation would invalidate the linear model as the residuals would be spatially autocorrelated.
However, HadCM3 exhibits a certain spatial structure, and such a pixel-by-pixel approach, where spatial grid cells are treated independently, cannot guarantee that this spatial structure, or covariance, is preserved. We can nevertheless show that the reconstructed climate fields exhibit the same spatial structure as the original HadCM3 model output. First of all, the regression coefficient maps (Fig. 3) indicate that the climate response to external forcings is spatially coherent. We calculated this spatial coherence in terms of the spatial covariance matrix (Fig. 5). It consists of the covariance between the time series of any two grid points, i.e., it is a 7008 × 7008 matrix (= 96 × 73 (nlon × nlat) = 7008) of covariances over the 72 snapshots.
Spatial covariance matrix of (a) temperature (in units K2), (b) precipitation (in units mm2/a2), and (c) total cloud cover (in units of 12) for HadCM3, our linear regression model (LR model), and the difference between the two. Each value represents a covariance matrix element from a flattened vector with the length of the total number of grid points (n = 7008). The covariance matrices are symmetric and thus are their differences.
For both HadCM3 and the linear model, the covariance matrices are similar in structure and magnitude, and their differences are relatively small (Fig. 5). The covariance matrices are much more similar for temperature than for precipitation and total cloud cover, because the linear regression works better for temperature than for the other two climate fields. Overall, the covariance matrices of the linear model reconstructions are so similar to HadCM3 that we can conclude that the spatial structure is indeed preserved.
For any time series of observations (for example, as shown in Figs. 6, 7), we can assume that data points of that time series are temporally autocorrelated because of possible lag effects that derive from non-equilibrium climate dynamics. However, our reconstructions are based on snapshot simulations that are assumed to be in equilibrium, and any such lags are therefore omitted. The relationship between outputs of snapshots and their forcings can thus be treated as independent data points with no temporal autocorrelation.
(a) Map of the 39 Middle and Late Pleistocene marine sea surface temperature proxies used in this study and their respective time series (b,c). Black dots indicate proxy sea surface temperature while blue lines indicate mean annual temperature as reconstructed for every 1 ka of the last 800 ka. Proxy–derived and model temperature are on the same scale, in). Orange lines are original time series from HadCM3. Grey bars indicate glacial stages. The coefficients for the correlation between the reconstructed temperature (blues lines) and the proxy record (black dots) can be found in Table 3.
(a) Map of the 20 Middle and Late Pleistocene terrestrial climate proxies used in this study and their respective time series (b). Black dots indicate proxy variables (in different units) while blue lines indicate mean annual temperature as reconstructed for every 1 ka of the last 800 ka (in). Orange lines are original time series from HadCM3. Grey bars indicate glacial stages. The coefficients for the correlation between the reconstructed temperature (blues lines) and the proxy record (black dots) can be found in Table 4.
Comparison to marine and terrestrial proxies
We compared the reconstructed climate data with marine proxies (before the downscaling/bias-correction step) as a means of highlighting how well the reconstructed long-term climatologies compare to empirical reconstructions.
Marine sediment cores are valuable archives of past sea surface temperature (SST) records. Because their associated bio-geochemistry is relatively straightforward, marine proxies can be utilised as paleo-thermometers and are thus well suited for a direct proxy–model comparison. For these proxies, we compared model-derived mean annual temperature (MAT) time series directly with proxy-derived SST time series and calculated the correlation between the two. Note that MAT and SST are not the same climatological quantities; SST is the temperature of the ocean surface and has a lower limit of about −1.8°C, the freezing point of saltwater. While we expect MAT and SST to co-vary in low and mid-latitudes, at higher latitudes, seasonal or perennial sea ice cover makes a comparison between both variables problematic.
Although terrestrial proxies are rarely available as direct temperature estimates, we could still calculate the correlation between the model-derived and the proxy-derived time series. However, the interpretation of terrestrial proxies from a climate perspective can be problematic. For example, pollen-based vegetation reconstructions are suggested to be less reliable as climate proxies, particularly for interglacials29. Other land-based proxies such as dust deposits integrate long-term climatic changes over large regions and hence do not necessarily capture climatic effects at their specific location.
For the comparison of our climate reconstructions with proxy reconstruction, we assembled long-term marine SST and terrestrial climate proxy reconstructions (Figs. 6, 7, 8, 9) that cover a period of at least 150 ka during the last 800 ka (Tables 3 and 4).
Scatter plot of the 39 Middle and Late Pleistocene marine proxies (x-axis) used in this study versus reconstructed mean annual temperature (y-axis). The respective correlation coefficient is shown on the top right in each plot and information about each proxy can be found in Table 3. The dashed diagonal line represents the hypothetical 1-1 for a perfect model.
Scatter plot of the 20 Middle and Late Pleistocene terrestrial proxies (x-axis) used in this study versus reconstructed mean annual temperature (y-axis). The respective correlation coefficient is shown on the top right in each plot and information about each proxy can be found in Table 4.
In locations where we have empirical proxies, both on land and over the ocean, our regression-based climate reconstructions match the original HadCM3 simulations well. These reconstructions can therefore be considered as representative of the simulated HadCM3 climate. As a consequence, any differences with respect to the proxies records will persist in our reconstructions and, therefore, needs to be removed.
Model bias
Because the linear model was fitted to match HadCM3 model outputs, the resulting bias of our reconstructions is similar to the HadCM3 model bias. The bias of reconstructed temperature and precipitation with respect to the CRU TS data set has a similar spatial pattern (Fig. 10) and is as large as the bias shown for HadCM3 climate reconstructions of the last 60 ka10. This is also why our long-term reconstructions show the same bias towards the assembled paleo-climate proxy records as the original HadCM3 simulations (Figs. 6, 7). From this, we concluded that this similarity means that our present-day climate reconstruction is of the same quality as the original HadCM3 simulation it is based on. As discussed earlier, the bias was removed from our climate reconstructions using the “delta” method12.
Informed user notice
Our final dataset is just as good as i) the goodness of the applied linear regression model, and ii) the underlying climate model, HadCM3 in our case. How well the linear regression model performs for different variables and different regions can be seen in Fig. 4. It works usually better for temperature and thus for temperature derived bioclimatic variables (see Table 1). For precipitation and total cloud cover, we suggest to carefully assess Fig. 4 if the region of interest is well represented by the statistics-based reconstruction. The actual numbers for the goodness of the fit and the model, R2 and RMSE, can be found in the diagnostic files listed in Table 5.
The quality of the underlying climate model, HadCM3, can be assessed in two ways. First, by looking at the model bias (Fig. 10), and second, by looking how well HadCM3 compares to proxy records that go well back in time and show the longer-term climatic changes happening on glacial–interglacial time scales (Figs. 6, 7, 8, 9). However, most geological proxies are useful for quantitative temperature comparisons, and only few, terrestrial proxies exist for precipitation, and they mostly reflect qualitative changes, e.g., wetter vs. drier periods. For cloud cover no such geological proxies exist.
Usage Notes
Examples on how to access the NetCDF files of the reconstructed climate using Python or R are provided in the examples directory of the project repository28.
Code availability
Model code for the linear regression as well as the code for the analysis and visualisation of figures is publicly available in the project repository28. NetCDF files have been processed using cdo30. We used the Python language for most of our scripts with a few bash scripts as wrappers. The workflow for the data generation process is managed by Snakemake31. The linear regression is based on the statsmodels package32. All visualisations are made with matplotlib33 using cartopy34 for maps. Other Python packages used are (in alphabetical order): adjustText35, BeautifulSoup4 (https://www.crummy.com/software/BeautifulSoup/), netCDF436, numpy37, pandas38,39, scipy40, scikit-image41, and tqdm42.
References
Solomon, S. et al. (eds.) IPCC: Climate Change 2007: The Physical Science Basis. Contribution of Working Group I to the Fourth Assessment Report of the Intergovernmental Panel on Climate Change (Cambridge University Press, 2007).
Ganopolski, A. & Calov, R. The role of orbital forcing, carbon dioxide and regolith in 100 kyr glacial cycles. Clim. Past 7, 1415–1425, https://doi.org/10.5194/cp-7-1415-2011 (2011).
Timmermann, A. et al. Modeling Obliquity and CO2 Effects on Southern Hemisphere Climate during the Past 408 ka. J. Climate 27, 1863–1875, https://doi.org/10.1175/JCLI-D-13-00311.1 (2013).
Claussen, M. et al. Earth system models of intermediate complexity: closing the gap in the spectrum of climate system models. Climate Dynamics 18, 579–586, https://doi.org/10.1007/s00382-001-0200-1 (2002).
Hijmans, R. J., Cameron, S. E., Parra, J. L., Jones, P. G. & Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. International Journal of Climatology 25, 1965–1978, https://doi.org/10.1002/joc.1276 (2005).
Brown, J. L., Hill, D. J., Dolan, A. M., Carnaval, A. C. & Haywood, A. M. PaleoClim, high spatial resolution paleoclimate surfaces for global land areas. Scientific Data 5, 180254, https://doi.org/10.1038/sdata.2018.254 (2018).
Lima-Ribeiro, M. S. et al. EcoClimate: a database of climate data from multiple models for past, present, and future for macroecologists and biogeographers. Biodiversity Informatics 10, https://doi.org/10.17161/bi.v10i0.4955 (2015).
Fordham, D. A. et al. PaleoView: a tool for generating continuous climate projections spanning the last 21 000 years at regional and global scales. Ecography 40, 1348–1358, https://doi.org/10.1111/ecog.03031 (2017).
Valdes, P. J. et al. The BRIDGE HadCM3 family of climate models: HadCM3@Bristol v1.0. Geosci. Model Dev. 10, 3715–3743, https://doi.org/10.5194/gmd-10-3715-2017 (2017).
Armstrong, E., Hopcroft, P. O. & Valdes, P. J. A simulated Northern Hemisphere terrestrial climate dataset for the past 60,000 years. Sci Data 6, 1–16, https://doi.org/10.1038/s41597-019-0277-1 (2019).
Beyer, R. M., Krapp, M. & Manica, A. High-resolution terrestrial climate, bioclimate and vegetation for the last 120,000 years. Scientific Data 7, 236, https://doi.org/10.1038/s41597-020-0552-1 (2020).
Beyer, R., Krapp, M. & Manica, A. An empirical evaluation of bias correction methods for palaeoclimate simulations. Climate of the Past 16, 1493–1508, https://doi.org/10.5194/cp-16-1493-2020 (2020).
Harris, I., Osborn, T. J., Jones, P. & Lister, D. Version 4 of the CRU TS monthly high-resolution gridded multivariate climate dataset. Sci Data 7, 1–18, https://doi.org/10.1038/s41597-020-0453-3 (2020).
O’Donnell, M. S. & Ignizio, D. A. Bioclimatic predictors for supporting ecological applications in the conterminous United States. US Geological Survey Data Series 691 (2012).
Kaplan, J. O. et al. Climate change and Arctic ecosystems: 2. Modeling, paleodata-model comparisons, and future projections. J. Geophys. Res. 108, 8171, https://doi.org/10.1029/2002JD002559 (2003).
Singarayer, J. S. & Valdes, P. J. High-latitude climate sensitivity to ice-sheet forcing over the last 120kyr. Quaternary Science Reviews 29, 43–55, https://doi.org/10.1016/j.quascirev.2009.10.011 (2010).
Davies-Barnard, T., Ridgwell, A., Singarayer, J. & Valdes, P. Quantifying the influence of the terrestrial biosphere on glacial–interglacial climate dynamics. Clim. Past 13, 1381–1401, https://doi.org/10.5194/cp-13-1381-2017 (2017).
Bereiter, B. et al. Revision of the EPICA Dome C CO2 record from 800 to 600 kyr before present. Geophys. Res. Lett. 42, 2014GL061957, https://doi.org/10.1002/2014GL061957 (2015).
Berger, A. & Loutre, M. F. Insolation values for the climate of the last 10 million years. Quaternary Science Reviews 10, 297–317, https://doi.org/10.1016/0277-3791(91)90033-Q (1991).
Spratt, R. M. & Lisiecki, L. E. A Late Pleistocene sea level stack. Clim. Past 12, 1079–1092, https://doi.org/10.5194/cp-12-1079-2016 (2016).
Peltier, W. R., Argus, D. F. & Drummond, R. Space geodesy constrains ice age terminal deglaciation: The global ICE-6G_c (VM5a) model. Journal of Geophysical Research: Solid Earth 120, 450–487, https://doi.org/10.1002/2014JB011176 (2014).
Amante, C. & Eakins, B. ETOPO1 1 Arc-Minute Global Relief Model: Procedures, Data Sources and Analysis. National Geophysical Data Center, NOAA NOAA Technical Memorandum NESDIS NGDC-24, https://doi.org/10.7289/V5C8276M (2009).
Lehner, B. & Dőll, P. Development and validation of a global database of lakes, reservoirs and wetlands. Journal of Hydrology 296, 1–22, https://doi.org/10.1016/j.jhydrol.2004.03.028 (2004).
Araya-Melo, P. A., Crucifix, M. & Bounceur, N. Global sensitivity analysis of the Indian monsoon during the Pleistocene. Clim. Past 11, 45–61, https://doi.org/10.5194/cp-11-45-2015 (2015).
Lord, N. S. et al. Emulation of long-term changes in global climate: application to the late Pliocene and future. Climate of the Past 13, 1539–1571, https://doi.org/10.5194/cp-13-1539-2017 (2017).
Hoyt, D. V. Percent of Possible Sunshine and the Total Cloud Cover. Monthly Weather Review 105, 648–652, 10.1175/1520-0493(1977)105<0648:POPSAT>2.0.CO;2 (1977).
Berger, A. L. Long-term variations of daily insolation and Quaternary climatic changes. J. Atm. Sci. 35, 2362–2367 (1978).
Krapp, M. Terrestrial climate of the last 800,000 years, Open Science Framework, https://doi.org/10.17605/OSF.IO/8N43X (2021).
Herzschuh, U. et al. Glacial legacies on interglacial vegetation at the Pliocene-Pleistocene transition in NE Asia. Nature Communications 7, 11967, https://doi.org/10.1038/ncomms11967 (2016).
Schulzweida, U. CDO User Guide. Zenodo https://doi.org/10.5281/zenodo.3539275 (2019).
Kőster, J. & Rahmann, S. Snakemake—a scalable bioinformatics workflow engine. Bioinformatics 28, 2520–2522, https://doi.org/10.1093/bioinformatics/bts480 (2012).
Seabold, S. & Perktold, J. Statsmodels: Econometric and statistical modeling with python. In 9th Python in Science Conference (2010).
Hunter, J. D. Matplotlib: A 2D graphics environment. Computing In Science & Engineering 9, 90–95, https://doi.org/10.1109/MCSE.2007.55 (2007).
Met Office. Cartopy: a cartographic python library with a Matplotlib interface (2010–2015).
Flyamer, I. et al. Phlya/adjustText: 0.8 beta. Zenodo https://doi.org/10.5281/zenodo.3924114 (2020).
Whitaker, J. et al. Unidata/netcdf4-python: version 1.5.5 release. Zenodo https://doi.org/10.5281/zenodo.4308773 (2020).
Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362, https://doi.org/10.1038/s41586-020-2649-2 (2020).
Reback, J. et al. pandas-dev/pandas: Pandas 1.0.3. Zenodo https://doi.org/10.5281/zenodo.3715232 (2020).
McKinney, W. Data Structures for Statistical Computing in Python. In Walt, S. v. d. & Millman, J. (eds.) Proceedings of the 9th Python in Science Conference, 56–61, https://doi.org/10.25080/Majora-92bf1922-00a (2010).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nature Methods 17, 261–272, https://doi.org/10.1038/s41592-019-0686-2 (2020).
Walt, Svd et al. scikit-image: image processing in Python. PeerJ 2, e453, https://doi.org/10.7717/peerj.453 (2014).
da Costa-Luis, C. et al. tqdm: A fast, Extensible Progress Bar for Python and CLI. Zenodo https://doi.org/10.5281/zenodo.4531988 (2021).
Past Interglacials Working Group of PAGES. Interglacials of the last 800,000 years. Rev. Geophys. 54, 2015RG000482, https://doi.org/10.1002/2015RG000482 (2016).
Schaefer, G. et al. Planktic foraminiferal and sea surface temperature record during the last 1 Myr across the Subtropical Front, Southwest Pacific. Marine Micropaleontology 54, 191–212, https://doi.org/10.1016/j.marmicro.2004.12.001 (2005).
Ruddiman, W. F., Raymo, M. E., Martinson, D. G., Clement, B. M. & Backman, J. Pleistocene evolution: Northern hemisphere ice sheets and North Atlantic Ocean. Paleoceanography and Paleoclimatology 4, 353–412, https://doi.org/10.1029/PA004i004p00353 (1989).
Nürnberg, D., Müller, A. & Schneider, R. R. Paleo-sea surface temperature calculations in the equatorial east Atlantic from Mg/Ca ratios in planktic foraminifera: A comparison to sea surface temperature estimates from U37K’, oxygen isotopes, and foraminiferal transfer function. Paleoceanography and Paleoclimatology 15, 124–134, https://doi.org/10.1029/1999PA000370 (2000).
Horikawa, K., Murayama, M., Minagawa, M., Kato, Y. & Sagawa, T. Latitudinal and downcore (0–750 ka) changes in n-alkane chain lengths in the eastern equatorial Pacific. Quaternary Research 73, 573–582, https://doi.org/10.1016/j.yqres.2010.01.001 (2010).
Martrat, B. et al. Four Climate Cycles of Recurring Deep and Surface Water Destabilizations on the Iberian Margin. Science 317, 502–507, https://doi.org/10.1126/science.1139994 (2007).
Rincón-Martínez, D. & Leduc, G. Sea surface temperature calculated from alkenones for the last 285 ka with high-reolution Holocene of sediment core MD02-2529, Panama Basin. PANGAEA https://doi.org/10.1594/PANGAEA.777473 (2012).
Rodrigues, T., Voelker, A. H. L., Grimalt, J. O., Abrantes, F. & Naughton, F. Iberian Margin sea surface temperature during MIS 15 to 9 (580–300 ka): Glacial suborbital variability versus interglacial stability. Paleoceanography 26, PA1204, https://doi.org/10.1029/2010PA001927 (2011).
Hayward, B. W. et al. Planktic foraminifera-based sea-surface temperature record in the Tasman Sea and history of the Subtropical Front around New Zealand, over the last one million years. Marine Micropaleontology 82–83, 13–27, https://doi.org/10.1016/j.marmicro.2011.10.003 (2012).
Russon, T. et al. Inter-hemispheric asymmetry in the early Pleistocene Pacific warm pool. Geophysical Research Letters 37, https://doi.org/10.1029/2010GL043191 (2010).
Bard, E., Rostek, F. & Sonzogni, C. Interhemispheric synchrony of the last deglaciation inferred from alkenone palaeothermometry. Nature 385, 707–710, https://doi.org/10.1038/385707a0 (1997).
Rostek, F. et al. Reconstructing sea surface temperature and salinity using $\delta{18}O$ and alkenone records. Nature 364, 319, https://doi.org/10.1038/364319a0 (1993).
Caley, T. et al. High-latitude obliquity as a dominant forcing in the Agulhas current system. Clim. Past 7, 1285–1296, https://doi.org/10.5194/cp-7-1285-2011 (2011).
Pahnke, K., Zahn, R., Elderfield, H. & Schulz, M. 340,000-Year Centennial-Scale Marine Record of Southern Hemisphere Climatic Oscillation. Science 301, 948–952, https://doi.org/10.1126/science.1084451 (2003).
Garidel–Thoron, T. d. et al. A multiproxy assessment of the western equatorial Pacific hydrography during the last 30 kyr. Paleoceanography 22, https://doi.org/10.1029/2006PA001269 (2005).
Liu, Z., Altabet, M. A. & Herbert, T. D. Glacial-interglacial modulation of eastern tropical North Pacific denitrification over the last 1.8-Myr. Geophysical Research Letters 32, https://doi.org/10.1029/2005GL024439 (2005).
Yamamoto, M., Yamamuro, M. & Tanaka, Y. The California current system during the last 136,000 years: response of the North Pacific High to precessional forcing. Quaternary Science Reviews 26, 405–414, https://doi.org/10.1016/j.quascirev.2006.07.014 (2007).
Herbert, T. D. Collapse of the California Current During Glacial Maxima Linked to Climate Change on Land. Science 293, 71–76, https://doi.org/10.1126/science.1059209 (2001).
Schefuβ, E., Damsté, J. S. S. & Jansen, J. H. F. Forcing of tropical Atlantic sea surface temperatures during the mid-Pleistocene transition. Paleoceanography 19, https://doi.org/10.1029/2003PA000892 (2004).
Etourneau, J., Martinez, P., Blanz, T. & Schneider, R. Pliocene–Pleistocene variability of upwelling activity, productivity, and nutrient cycling in the Benguela region. Geology 37, 871–874, https://doi.org/10.1130/G25733A.1 (2009).
McClymont, E. L., Rosell-Melé, A., Giraudeau, J., Pierre, C. & Lloyd, J. M. Alkenone and coccolith records of the mid-Pleistocene in the south-east Atlantic: Implications for the U37K’ index and South African climate. Quaternary Science Reviews 24, 1559–1572, https://doi.org/10.1016/j.quascirev.2004.06.024 (2005).
Martínez–Garcia, A. et al. Links between iron supply, marine productivity, sea surface temperature, and CO2 over the last 1.1 Ma. Paleoceanography 24, https://doi.org/10.1029/2008PA001657 (2009).
Crundwell, M., Scott, G., Naish, T. & Carter, L. Glacial–interglacial ocean climate variability from planktonic foraminifera during the Mid-Pleistocene transition in the temperate Southwest Pacific, ODP Site 1123. Palaeogeography, Palaeoclimatology, Palaeoecology 260, 202–229, https://doi.org/10.1016/j.palaeo.2007.08.023 (2008).
Hayward, B. W. et al. The effect of submerged plateaux on Pleistocene gyral circulation and sea-surface temperatures in the Southwest Pacific. Global and Planetary Change 63, 309–316, https://doi.org/10.1016/j.gloplacha.2008.07.003 (2008).
Li, L. et al. A 4-Ma record of thermal evolution in the tropical western Pacific and its implications on climate change. Earth and Planetary Science Letters 309, 10–20, https://doi.org/10.1016/j.epsl.2011.04.016 (2011).
Herbert, T. D., Peterson, L. C., Lawrence, K. T. & Liu, Z. Tropical Ocean Temperatures Over the Past 3.5 Million Years. Science 328, 1530–1534, https://doi.org/10.1126/science.1185435 (2010).
Nürnberg, D. & Groeneveld, J. Pleistocene variability of the Subtropical Convergence at East Tasman Plateau: Evidence from planktonic foraminiferal Mg/Ca (ODP Site 1172 A). Geochemistry, Geophysics, Geosystems 7, https://doi.org/10.1029/2005GC000984 (2006).
Dyez, K. A., Ravelo, A. C. & Mix, A. C. Evaluating drivers of Pleistocene eastern tropical Pacific sea surface temperature. Paleoceanography 31, 2015PA002873, https://doi.org/10.1002/2015PA002873 (2016).
Alonso-Garcia, M. et al. Ocean circulation, ice sheet growth and interhemispheric coupling of millennial climate variability during the mid-Pleistocene (ca 800–400ka). Quaternary Science Reviews 30, 3234–3247, https://doi.org/10.1016/j.quascirev.2011.08.005 (2011).
Medina-Elizalde, M. & W Lea, D. The Mid-Pleistocene Transition in the Tropical Pacific. Science 310, 1009–12, https://doi.org/10.1126/science.1115933 (2005).
Liu, Z. Pleistocene climate evolution in the eastern Pacific and implications for the orbital theory of climate change. Ph.D., Brown University, United States – Rhode Island (2004).
Dyez, K. A. & Ravelo, A. C. Late Pleistocene tropical Pacific temperature sensitivity to radiative greenhouse gas forcing. Geological Society of America 41, 23–26, https://doi.org/10.1130/G33425.1 (2013).
Martínez-Garcia, A., Rosell-Melé, A., McClymont, E. L., Gersonde, R. & Haug, G. H. Subpolar Link to the Emergence of the Modern Equatorial Pacific Cold Tongue. Science 328, 1550–1553, https://doi.org/10.1126/science.1184480 (2010).
Lawrence, K. T., Herbert, T. D., Brown, C. M., Raymo, M. E. & Haywood, A. M. High-amplitude variations in North Atlantic sea surface temperature during the early Pliocene warm period. Paleoceanography 24, PA2218, https://doi.org/10.1029/2008PA001669 (2009).
Schmidt, M. W., Vautravers, M. J. & Spero, H. J. Western Caribbean sea surface temperatures during the late Quaternary. Geochemistry Geophysics Geosystems 7, https://doi.org/10.1029/2005GC000957 (2006).
Ho, S. L. et al. Sea surface temperature variability in the Pacific sector of the Southern Ocean over the past 700 kyr. Paleoceanography 27, https://doi.org/10.1029/2012PA002317 (2012).
Tierney, J. E., deMenocal, P. B. & Zander, P. D. A climatic context for the out-of-Africa migration. The Geological Society of America 45, 1023–1026, https://doi.org/10.1130/G39457.1 (2017).
Beck, J. W. et al. A 550,000-year record of East Asian monsoon rainfall from 10Be in loess. Science 360, 877–881, https://doi.org/10.1126/science.aam5825 (2018).
Kathayat, G. et al. Indian monsoon variability on millennial-orbital timescales. Scientific Reports 6, 24374, https://doi.org/10.1038/srep24374 (2016).
Guo, Z. T., Berger, A., Yin, Q. Z. & Qin, L. Strong asymmetry of hemispheric climates during MIS-13 inferred from correlating China loess and Antarctica ice records. Clim. Past 5, 21–31, https://doi.org/10.5194/cp-5-21-2009 (2009).
Carolin, S. A. et al. Northern Borneo stalagmite records reveal West Pacific hydroclimate across MIS 5 and 6. Earth and Planetary Science Letters 439, 182–193, https://doi.org/10.1016/j.epsl.2016.01.028 (2016).
Waldmann, N., Torfstein, A. & Stein, M. Northward intrusions of low- and mid-latitude storms across the Saharo-Arabian belt during past interglacials. Geology 38, 567–570, https://doi.org/10.1130/G30654.1 (2010).
Landwehr, J. M., Sharp, W. D., Coplen, T. B., Ludwig, K. R. & Winograd, I. J. The Chronology for the δ18O Record from Devils Hole, Nevada, Extended Into the Mid-Holocene. Tech. Rep., US Geological Survey (2011).
Stoykova, D. A., Shopov, Y. Y., Garbeva, D., Tsankov, L. T. & Yonge, C. J. Origin of the climatic cycles from orbital to sub-annual scales. Journal of Atmospheric and Solar-Terrestrial Physics 70, 293–302, https://doi.org/10.1016/j.jastp.2007.08.018 (2008).
Jouzel, J. et al. Orbital and Millennial Antarctic Climate Variability over the Past 800,000 Years. Science 317, 793–796, https://doi.org/10.1126/science.1141038 (2007).
Cheng, H. et al. The climatic cyclicity in semiarid-arid central Asia over the past 500,000 years. Geophysical Research Letters 39, https://doi.org/10.1029/2011GL050202 (2012).
Prokopenko, A. A., Hinnov, L. A., Williams, D. F. & Kuzmin, M. I. Orbital forcing of continental climate during the Pleistocene: a complete astronomically tuned climatic record from Lake Baikal, SE Siberia. Quaternary Science Reviews 25, 3431–3457, https://doi.org/10.1016/j.quascirev.2006.10.002 (2006).
Melles, M. et al. 2.8 Million Years of Arctic Climate Change from Lake El’gygytgyn, NE Russia. Science 337, 315–320, https://doi.org/10.1126/science.1222135 (2012).
Vaks, A., Bar-Matthews, M., Matthews, A., Ayalon, A. & Frumkin, A. Middle-Late Quaternary paleoclimate of northern margins of the Saharan-Arabian Desert: reconstruction from speleothems of Negev Desert, Israel. Quaternary Science Reviews 29, 2647–2662, https://doi.org/10.1016/j.quascirev.2010.06.014 (2010).
Bar-Matthews, M., Ayalon, A., Gilmour, M., Matthews, A. & Hawkesworth, C. J. Sea–land oxygen isotopic relationships from planktonic foraminifera and speleothems in the Eastern Mediterranean region and their implication for paleorainfall during interglacial intervals. Geochimica et Cosmochimica Acta 67, 3181–3199, https://doi.org/10.1016/S0016-7037(02)01031-1 (2003).
Cheng, H. et al. The Asian monsoon over the past 640,000 years and ice age terminations. Nature 534, 640–646, https://doi.org/10.1038/nature18591 (2016).
Tzedakis, P. C., Hooghiemstra, H. & Pälike, H. The last 1.35 million years at Tenaghi Philippon: revised chronostratigraphy and long-term vegetation trends. Quaternary Science Reviews 25, 3416–3430, https://doi.org/10.1016/j.quascirev.2006.09.002 (2006).
Vaks, A. et al. Paleoclimate and location of the border between Mediterranean climate region and the Saharo–Arabian Desert as revealed by speleothems from the northern Negev Desert, Israel. Earth and Planetary Science Letters 249, 384–399, https://doi.org/10.1016/j.epsl.2006.07.009 (2006).
K. Thomas, E. et al. Heterodynes dominate precipitation isotopes in the East Asian monsoon region, reflecting interaction of multiple climate factors. Earth and Planetary Science Letters 455, https://doi.org/10.1016/j.epsl.2016.09.044 (2016).
Hao, Q. et al. Delayed build-up of Arctic ice sheets during 400,000-year minima in insolation variability. Nature 490, 393–396, https://doi.org/10.1038/nature11493 (2012).
Acknowledgements
MK and RB were supported by an ERC Consolidator Grant to AM (Local Adaptation 647787). MK was also supported by the NZ Ministry for Business, Innovation and Employment through the Antarctic Science Platform (ANTA1801). We thank Eric Wolff, Max Holloway, Peter Hopcroft, Chris Brierley, and Michel Crucifix for commenting on early versions of this manuscript. We would also like to thank Matheus Lima-Ribeiro and our other reviewer for their useful comments.
Author information
Authors and Affiliations
Contributions
A.M. and M.K. devised the project. M.K. devised and implemented the linear regression model with input from R.B. and S.L.E. P.J.V. provided additional HadCM3 snapshot simulations. M.K. and A.M. wrote a first draft of the paper which was improved by input from all other authors. M.K. performed the analysis and prepared the figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files associated with this article.
About this article
Cite this article
Krapp, M., Beyer, R.M., Edmundson, S.L. et al. A statistics-based reconstruction of high-resolution global terrestrial climate for the last 800,000 years. Sci Data 8, 228 (2021). https://doi.org/10.1038/s41597-021-01009-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-021-01009-3
This article is cited by
-
The Neanderthal niche space of Western Eurasia 145 ka to 30 ka ago
Scientific Reports (2024)
-
Longstanding behavioural stability in West Africa extends to the Middle Pleistocene at Bargny, coastal Senegal
Nature Ecology & Evolution (2023)
-
Worldwide Late Pleistocene and Early Holocene population declines in extant megafauna are associated with Homo sapiens expansion rather than climate change
Nature Communications (2023)
-
A spatiotemporally explicit paleoenvironmental framework for the Middle Stone Age of eastern Africa
Scientific Reports (2022)
-
Climatic windows for human migration out of Africa in the past 300,000 years
Nature Communications (2021)