Abstract
In prokaryotes, protein phosphorylation plays a critical role in regulating a broad spectrum of biological processes and occurs mainly on various amino acids, including serine (S), threonine (T), tyrosine (Y), arginine (R), aspartic acid (D), histidine (H) and cysteine (C) residues of protein substrates. Through literature curation and public database integration, here we reported an updated database of phosphorylation sites (p-sites) in prokaryotes (dbPSP 2.0) that contains 19,296 experimentally identified p-sites in 8,586 proteins from 200 prokaryotic organisms, which belong to 12 phyla of two kingdoms, bacteria and archaea. To carefully annotate these phosphoproteins and p-sites, we integrated the knowledge from 88 publicly available resources that covers 9 aspects, namely, taxonomy annotation, genome annotation, function annotation, transcriptional regulation, sequence and structure information, family and domain annotation, interaction, orthologous information and biological pathway. In contrast to version 1.0 (~30 MB), dbPSP 2.0 contains ~9 GB of data, with a 300-fold increased volume. We anticipate that dbPSP 2.0 can serve as a useful data resource for further investigating phosphorylation events in prokaryotes. dbPSP 2.0 is free for all users to access at: http://dbpsp.biocuckoo.cn.
Similar content being viewed by others
Introduction
As one of the most well-characterized and important post-translational modifications (PTMs), protein phosphorylation plays an essential role in almost all signalling pathways and biological processes, from eukaryotes to prokaryotes1,2,3,4,5. This reversibly dynamic process is precisely modulated by protein kinases (PKs) and protein phosphatases (PPs), which are involved in linking or removing a phosphate group at specific residues of protein substrates1,2,3,4,5. The first eukaryotic phosphoprotein was discovered in 1883 by Olof Hammarsten, a Swedish biochemist, who detected phosphorous in a secreted protein, casein, from milk6. Although later studies demonstrated that many proteins can be phosphorylated in eukaryotes, it was long debated whether protein phosphorylation also exists in prokaryotes until the discovery of isocitrate dehydrogenase in Escherichia coli, the first identified prokaryotic phosphoprotein, in 19797,8. In contrast with eukaryotic phosphorylation, which occurs mainly at specific serine (S), threonine (T) and tyrosine (Y) residues of proteins5, prokaryotic protein phosphorylation can occur at additional types of amino acids, such as arginine (R), aspartic acid (D), histidine (H) and cysteine (C)1,9,10,11,12,13. Given the importance of phosphorylation in the regulation of protein functions11,12,13, the identification of novel phosphorylation sites (p-sites) in proteins is fundamental for understanding the molecular mechanism and regulatory roles of prokaryotic phosphorylation.
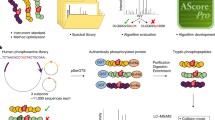
Previously, experimental identification of p-sites with conventional biochemical assays was usually labour intensive, time consuming and expensive and was accomplished in a low-throughput (LTP) manner. The LTP methods mainly included site-directed mutagenesis (SDM) of candidate p-sites14, in vitro kinase assay (IKA) to identify potential kinase-specific p-sites15, detection of p-sites in purified proteins with LTP mass spectrometry (LTP-MS)16, and N-terminal sequencing of phosphopeptides (NSP)17. The quality of p-sites identified in LTP studies is higher, because usually multiple assays were performed, and the biological functions of p-sites were also carefully analyzed. Recently, advances in the development of proteomic techniques using high-throughput MS (HTP-MS) have enabled the large-scale phosphoproteomic identification of p-sites in prokaryotic proteins18,19,20,21. For example, Macek et al. conducted phosphoproteomic profiling to detect 54 phosphoserine (pS), 16 phosphothreonine (pT) and 8 phosphotyrosine (pY) residues of 78 proteins in Bacillus subtilis, as well as 81 pS/pT/pY sites of 79 E. coli phosphoproteins18,19. For arginine phosphorylation, Elsholz et al. systematically identified 121 phosphoarginine (pR) residues in 87 B. subtilis proteins20, whereas Schmidt et al. later quantitatively characterized 134 phosphoproteins with 217 pR sites in B. subtilis21. More recently, Lai et al. detected 159 phosphohistidine (pH) and 69 phosphoaspartic acid (pD) sites of 197 phosphopeptides in nine prokaryotic organisms13. Because an increasing number of LTP and HTP p-site investigations have been reported, the collection, curation, integration and annotation of known phosphoproteins and p-sites in prokaryotes will provide invaluable information for better understanding the host-pathogen interaction and development of antimicrobial agents.
In 2015, we developed a new database of phosphorylation sites in prokaryotes (dbPSP) 1.0, which contained 7,391 experimentally identified p-sites, including 2,709 pS, 2,174 pT, 2,187 pY, 142 pR, 84 pD, 90 pH and 5 phosphocysteine (pC) sites, in 3,750 phosphoproteins of 96 prokaryotes22. Compared with the second largest resource, the Phosphorylation Site Database, which curated approximately 1,400 prokaryotic p-sites23, dbPSP 1.0 had a > 4-fold greater data volume. At that time, few annotations were provided, except limited information on p-sites. Due to the large number of prokaryotic p-sites found in recent studies, here we created dbPSP 2.0, which contains 19,296 known p-sites in 8,586 proteins from 200 prokaryotic organisms, through literature curation and public database integration (Fig. 1a, Supplementary Table 1). Furthermore, we carefully annotated these phosphoproteins and p-sites through integrating the knowledge from 88 publicly accessible databases, covering 9 aspects. In contrast with dbPSP 1.0 (~30 MB), this updated database possesses ~9 GB of data, with a 300-fold increased volume. We confirmed that dbPSP 2.0 will be continuously updated and can provide a much more useful resource for exploring protein phosphorylation in prokaryotes.
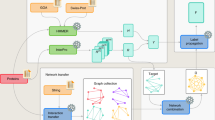
An overview of the dbPSP 2.0 database. (a) Flowchart for creating the database. First, we manually re-curated all entries in version 1.0 to ensure the data quality, searched PubMed to find newly identified p-sites, and integrated known p-sites from other public databases. Then, we mapped all phosphoproteins to public data sources for cross-referencing. In addition to basic information, we further integrated various annotations from 88 public databases that covered 9 aspects: (i) taxonomy annotation, (ii) genome annotation, (iii) function annotation, (iv) transcriptional regulation, (v) sequence and structure information, (vi) family and domain annotation, (vii) interaction, (viii) orthologous information, and (ix) biological pathway. (b) A comparison of the numbers of prokaryotic p-sites in dbPSP 2.0 and in other databases.
Results
dbPSP update
Entries of newly reported p-sites
Compared with version 1.0, version 2.0 contains 11,905 new entries (Fig. 1b). Through literature curation and public database integration, dbPSP 2.0 contains 19,296 non-redundant p-sites on seven different types of amino acid residues in 8,586 substrates from 200 prokaryotic species (Supplementary Table 1). In our dataset, there are 18,576 and 671 p-sites derived from HTP and LTP studies, respectively. The derivation of 96.27% known p-sites from HTP studies indicated the importance and usefulness of MS-based phosphoproteomic profiling for studying prokaryotic phosphorylation. In addition to version 1.0, we also compared dbPSP 2.0 to other existing databases, including the Phosphorylation Site Database23, UniProt24, dbPTM 201925, SysPTM 2.026 and PHOSIDA27, and our database contained a much higher number of known phosphoproteins and p-sites in prokaryotes (Fig. 1b). For each p-site, its corresponding gene name, UniProt accession number, organism, phylum, phosphorylated position, residue type, flanking peptide, data type, experimental method and original reference(s) have been present (Supplementary Table 1).
Distribution of phosphoproteins and p-sites for different residue types and different phyla
In dbPSP 1.0, known p-sites were taken from 96 prokaryotic organisms belonging to 11 phyla, Crenarchaeota, Euryarchaeota, Proteobacteria, Actinobacteria, Firmicutes, Cyanobacteria, Deinococcus-Thermus, Tenericutes, Spirochaetes, Chlamydiae and Thermotogae22. Due to the new data accumulation, known p-sites have been extended to 200 prokaryotic species in 12 phyla by adding a new phylum, Bacteroidetes (Fig. 2a). The distribution of numbers of p-sites among different phyla was analyzed, and it was observed that more p-sites were identified in Proteobacteria and Actinobacteria than in other phyla, with proportions of 27.95% and 23.13%, respectively (Fig. 2a). The Proteobacteria phylum comprises a number of extensively studied microorganisms, such as the most widely used model organism E. coli in microbiological studies7,8, and a human pathogen Shigella flexneri that causes bacillary dysentery mainly in children and results in 14,000 deaths per year28. In Actinobacteria phylum, one of the most notorious species is Mycobacterium tuberculosis, which is the causative agent of tuberculosis (TB) and annually causes 1.5 million deaths29. Due to the high virulence of M. tuberculosis, two related species including the slow-growing Mycobacterium bovis30 and the fast-growing Mycobacterium smegmatis30 were established as models to study mycobacterial physiology. Additionally, we analyzed the distribution of p-sites on different types of amino acid residues and found that pS, pT and pY sites appear more frequently than other types of residues and occupy proportions of 39.67%, 31.55% and 19.87%, respectively (Fig. 2b). Moreover, the distribution of different types of p-sites among the 12 phyla was evaluated (Fig. 2c). The most pR sites were detected in Firmicutes, whereas Proteobacteria had the highest number of pD and pH sites (Fig. 2c). Additional detailed data statistics can be viewed at http://dbpsp.biocuckoo.cn/Statistics.php.
The distribution of phosphoproteins and p-sites for different phyla and different residue types in prokaryotes. (a) The distribution of phosphoproteins and p-sites in various phyla. (b) The numbers of different types of phosphoproteins and p-sites in dbPSP 2.0. (c) The distribution of different residue types among different phyla.
Coverage of phosphoproteins in different species
Due to data limitation, here we only calculated the coverage values of phosphoproteins in 50 species with ≥10 phosphorylated substrates (Supplementary Table 2). For each prokaryote, its proteome set was downloaded from UniProt24 by searching the corresponding Proteome ID, e.g., UP000001018 for Sulfolobus acidocaldarius (strain ATCC 33909/DSM 639/JCM 8929/NBRC 15157/NCIMB 11770) (https://www.uniprot.org/proteomes/?query=taxonomy:330779). Then the proportion of phosphoproteins against all protein products were counted, and top 10 species with higher coverage values were shown. From the results, we found that the coverage values of the 10 prokaryotes ranged from 8.47% (Staphylococcus aureus) to 36.06% (S. acidocaldarius) (Fig. 3a). Previously, it was estimated that about 30% of human proteins might be phosphorylated31, and a later study demonstrated that at least 75% of human proteins are phosphorylated in vivo32. Thus, when more and more phosphoproteomic studies are performed for prokaryotes, the coverage values of their phosphoproteins will be undoubtedly increased.
New annotations
Multiple-layer annotation of prokaryotic phosphoproteins
For convenience, dbPSP 2.0 was organized as a phosphoprotein-centred database. To provide an integrative annotation of known phosphoproteins and p-sites, we provided a variety of cross-references to public data sources. For example, gene and protein names were taken mainly from UniProt24, whereas corresponding accession numbers were integrated from UniProt24, Ensembl33, EMBL34, KEGG35 and NCBI GenBank36. Moreover, functional descriptions, protein/nucleotide sequences, and keywords were derived from UniProt24 to provide the basic information for each phosphoprotein entry, while the primary references with PMIDs were provided for each p-site. The gene ontology (GO) annotations in the Gene Ontology resource37 were also included if available. Furthermore, the knowledge from 88 additional public resources, such as ChEMBL38, BacDive39, PDB40, IUPred2A41, InterPro42, BioGRID43, EggNOG 5.044 and Reactome45, was integrated to comprehensively annotate the prokaryotic phosphoproteins. These resources covered 9 aspects, namely, taxonomy annotation, genome annotation, function annotation, transcriptional regulation, sequence and structure information, family and domain annotation, interaction, orthologous information and biological pathway (Fig. 1a). A brief summary of all public resources integrated in dbPSP 2.0 can be accessed at: http://dbpsp.biocuckoo.cn/Links.php. For these resources, the annotation datasets can be downloaded at http://dbpsp.biocuckoo.cn/Download.php.
Dynamic 3D structure details for phosphoproteins
For each phosphoprotein with available 3D structures characterized by X-ray crystallography or NMR spectroscopy, a representative 3D structure was selected for intuitive visualization. Users can select all or specific p-sites for visualizing their locations on protein structures.
HTP p-site classification
In phosphoproteomic studies, phosphopeptides were derived from mass spectrometry spectral datasets, usually with a false discovery rate (FDR) of 0.01 at the peptide-spectrum match (PSM), peptide and protein level for quality control. To pinpoint an exact p-site in a phosphopeptide, a localization probability (LP) score could be calculated by a variety of tools, such as MaxQuant46. LP scores range from 0 to 1, and a higher LP score represents a higher probability of a detected site being a real p-site. Since HTP p-sites were identified from different studies with different confidence, we classified all collected HTP p-sites into four classes based on their LP scores if available, namely, class I (LP > 0.75), class II (LP ≤ 0.75 and >0.5), class III (LP ≤ 0.5 and ≥0.25), and class IV (LP < 0.25), as previously described46. In most of these HTP studies, different reference databases, distinct search engines and/or diverse parameter configurations were adopted for phosphopeptide detection in different organisms. Thus, the aggregation of false positive identifications might result in a considerable higher FPR value in the cumulative dataset. A re-analysis of all raw MS datasets under a unified platform will generate phosphopeptides with much higher quality, although such an effort is not within the scope of dbPSP 2.0, which directly collected known p-sites from published literature.
Multi-alignment (MSA) of orthologs
Here, potential orthologues of known phosphoproteins were obtained from Clusters of Orthologous Groups of proteins (COG)47. For each orthologous group, all protein sequences were multi-aligned using MUSCLE48, and a conservation ratio was calculated for the sequences containing the same types of phosphorylatable residues against all sequences in the group. The distribution of the conservation ratio ranged from 0 to 1 was illustrated for all p-sites in the orthologous groups (Fig. 3b), and we only detected 227 p-sites with a conservation ratio > 0.9 (Supplementary Table 3). These highly conserved p-sites might be useful for the investigation of conserved functions of phosphorylation in prokaryotes.
Browse lists and detailed phosphoprotein information page
dbPSP 2.0 was developed with a user-friendly website interface, and multiple browse and search options were implemented to conveniently query the data. Here, we chose B. subtilis ClpP, an ATP-dependent Clp protease proteolytic subunit, as an example to introduce the usage of dbPSP 2.0. Two browse options, ‘Browse by phyla’ (Fig. 4a) and ‘Browse by residue types’ (Fig. 4b), are accessible to browse the data. In the option ‘Browse by phyla’, 12 representative diagrams for all phyla are listed. The user can click the phylum to link the taxonomic category of the given phylum (Fig. 4a). The user can select ‘Bacillus subtilis (strain 168)’ to retrieve a list of phosphoproteins in a tabular format with ‘dbPSP ID’, ‘UniProt Accession’, ‘Gene Name’, ‘Protein Name’ and ‘Organism’ (Fig. 4a). In the option ‘Browse by residue types’, the user can choose one of the 7 residue types to browse all phosphoproteins with the given phosphorylation residue type. For example, by clicking the diagram of arginine, all proteins with pR sites are listed (Fig. 4b). Through selecting ‘PP04832’, the dbPSP ID of ClpP (Fig. 4a,b), the detailed phosphoprotein page for ClpP, is displayed (Fig. 4c,d). For a brief overview, the dbPSP ID, protein/gene names, organism, and dynamic structure details are presented (Fig. 4c). The ‘Sites’ part provides mainly detailed information on p-sites, and the original peptide and primary reference can be shown by clicking the ‘View’ button of each p-site (Fig. 4c). To access additional information on the phosphoprotein, users can click the label ‘Annotation’ on the left menu and select the interesting aspect to access the corresponding resources (Fig. 4d). For each resource, the annotation details are presented on a new page after clicking the ‘More’ icon (Fig. 4d). In addition to the browse options, multiple search options, including ‘Substrate Search’, ‘Peptide Search’, ‘Advanced Search’, ‘Batch Search’ and ‘BLAST Search’, were also developed for users to easily access the database.
Sequence preferences of different types of p-sites
Due to the limited number of pC sites, here we only analyzed the sequence preferences of pS, pT, pY, pR, pD and pH sites by using pLogo49 for bacteria and archaea (Fig. 5). For prokaryotic pS, pT and pY sites, we also compared their sequence preferences to those of eukaryotic phosphorylation, including 382,105 pS, 123,247 pT and 59,824 pY sites by integrating two previously developed databases, dbPAF50 and dbPPT51. For pS and pT sites in archaea, R or lysine (K) residues most frequently occur at the +1 position, with a lesser extent at the +2 position (Fig. 5a). In bacteria, K residues are over-represented at the −1 position for pS sites, whereas S, D, glycine (G) and proline (P) are enriched at the −2, −1, +1 and +2 positions for pT sites, respectively (Fig. 5a). For pY sites, S residues frequently appear at the +1 position for eukaryotic phosphorylation, whereas K residues preferentially appear at the −2 position for bacteria and the −1 and −2 positions for archaea (Fig. 5a). For prokaryotic pD sites, methionine (M) and P residues are over-represented at the +3 and +4 positions around p-sites in bacteria but not archaea (Fig. 5b). For pH sites, S residues preferentially appear at the −1 position for bacteria (Fig. 5b). Due to data limitation, the sequence preference of pR sites in only bacteria was analyzed, and asparagine (N) residues are enriched at the −1 position (Fig. 5b).
Analyses of sequence preferences for p-sites in prokaryotes with pLogo49. (a) For pS, pT and pY residues, comparisons of sequence preferences among bacteria, archaea and eukaryotes are shown. (b) The sequence preferences of pD, pH and pR sites in bacteria and archaea. Due to data limitations, pR sites in only bacteria were analyzed.
Application of dbPSP
After the publication of dbPSP 1.0, it has been visited more than 180,000 times and has served as a highly useful resource for studying prokaryotic phosphorylation50,52,53,54,55,56. For example, Garcia-Garcia et al. re-analyzed the phosphoproteomic datasets in dbPSP and found that phosphoproteins are essential for the regulation of the cell cycle and DNA-mediated processes in bacteria52. With the help of dbPSP, Venkat et al. experimentally validated that phosphorylation of S280 decreases the enzyme activity of malate dehydrogenase (MDH) in E. coli53. Additionally, Lin et al. utilized p-site information in dbPSP to analyze phosphoproteomic data and dissected the dynamic alteration of phosphorylation in various phosphoproteins during antibiotic treatment and resistance54. Moreover, Hasan et al. adopted pS and pT sites in dbPSP as training datasets and developed a useful tool, Microbial Phosphorylation Site predictor (MPSite), for predicting microbial p-sites55. In addition, the phosphorylation data of representative prokaryotes from dbPSP was utilized for kinase motif enrichment analysis, and the results demonstrated that most eukaryotic phosphorylation motifs could not be recovered in prokaryotes56.
In dbPSP 2.0, we collected and curated newly identified p-sites in prokaryotic phosphoproteins, which could present more complete information on phosphorylation in prokaryotes. Furthermore, dbPSP 2.0 has rich annotations for phosphoproteins and p-sites, which is critical for exploring the function and mechanism of phosphorylation events. In addition, the MSA results of orthologues were provided in this database and will be important for discovering conserved functional p-sites in prokaryote cells. Based on previous studies, dbPSP could work as a well-curated data resource of prokaryotic phosphoproteins to provide helpful support for phosphoproteomic analysis, tool development, and the investigation of prokaryotic phosphorylation events. We anticipate that the updated dbPSP 2.0 could be a comprehensive data resource for better understanding the importance of protein phosphorylation in prokaryotes.
Discussion
Protein phosphorylation is one of most well-studied PTMs and is reported to be involved in regulating numerous cellular processes in prokaryotic cells8,57. In 2015, we collected 7,391 known p-sites of 3,750 proteins in 96 prokaryotes from published literature and developed dbPSP 1.022 to contain these datasets. Due to the accumulation of phosphorylation information, here we released dbPSP 2.0 by adding 11,905 new entries to include newly discovered phosphoproteins and p-sites in prokaryotes. Furthermore, the rich annotations derived from 88 public databases were integrated. In total, dbPSP 2.0 contained 19,296 known p-sites in 8,586 phosphoproteins and occupied the size of ~9 GB, with a 300-fold increase compared to that of version 1.0.
In this study, to cover the diverse biological roles of prokaryotic phosphoproteins, we included multiple-layer knowledge from other databases to comprehensively annotate phosphoproteins. For example, the prokaryotic ClpP enzyme plays an important role in modulating various biological processes, such as cellular stress response, pathogenesis and homeostasis58. Inhibiting the function of ClpP was reported to affect the infectivity and virulence of microbial pathogens59. Moreover, the arginine phosphorylation of ClpP was essential for maintaining its function20,21,60. As shown in Fig. 6, the B. subtilis protease ClpP is annotated as a serine peptidase and participates in eliminating damaged proteins during heat shock, and its activity can be repressed by CtsR as well as by 20,697 compounds. Meanwhile, ClpP might interact with 9 partners and self-assemble in hexameric ring structures (Fig. 6). In particular, we found nearly 15,700 records from 6 orthologous databases to demonstrate that ClpP is a highly conserved subunit in prokaryotes, and the results are consistent with previous studies. In addition, the functional domain and p-site information of ClpP were also provided. In dbPSP 2.0, the curated data resources of p-sites and phosphoproteins as well as annotation information are downloadable at http://dbpsp.biocuckoo.cn/Download.php.
In summary, the dbPSP 2.0 database will be continuously maintained and updated when new p-sites in prokaryotes are identified. In addition to adding additional annotations from other public databases, we will further develop computational tools for the prediction of prokaryotic p-sites. We anticipate that this database can provide helpful support for better understanding the regulatory mechanisms and functions of phosphorylation in prokaryotes.
Methods
Data collection and update
In dbPSP 1.0, we manually collected 7,391 p-sites in 3,750 non-redundant prokaryotic phosphoproteins from the literature22. In this study, the phosphorylation events in prokaryotes newly reported since 2014 were considered and collected. To obtain known p-sites from the literature, we searched the PubMed database with multiple general keywords, such as ‘bacteria phosphoproteomics’, ‘archaea phosphorylation’, ‘archaebacteria phospho-site’. All the retrieved 39,997 articles were manually curated to collect the experimentally identified prokaryotic p-sites, and collected p-sites were then mapped to protein sequences obtained from UniProt (release 2019_05)24 (Fig. 1a). We also integrated the prokaryotic p-sites from other public databases, with 1,400, 427, 419, 345 and 317 p-sites from Phosphorylation Site Database23, UniProt24, dbPTM 201925, SysPTM 2.026 and PHOSIDA27, respectively (Fig. 1b). These datasets were cross-checked with our manually collected dataset and then integrated into the dbPSP 2.0 database.
Structure data collection and prediction
The 3D structures of phosphoproteins for intuitive visualization were obtained from the PDB40 if available. A JavaScript molecular visualization library, 3Dmol.js61, was used to support the dynamic structure chart in the browser interface. In addition, the probabilities of disordered binding regions and disorder propensity values were predicted by using ANCHOR241 and IUPred241, respectively. The details are provided on the phosphoprotein page.
Web interface construction
HTML, PHP and JavaScript were applied to develop the web interface as the front-end. The MySQL server was applied to manage the data as the back-end. The backlog and cache data will be cleared regularly, and the dbPSP database will be maintained and optimized continuously.
Data availability
All the collected phosphoproteins, p-sites and various annotations are freely available at http://dbpsp.biocuckoo.cn/Download.php. For convenience, phosphorylation datasets can be downloaded in three data types, including the total dataset, the phylum-specific datasets, and the residue-specific datasets The datasets of phosphoproteins in prokaryotes have been uploaded to figshare62, https://doi.org/10.6084/m9.figshare.11436879. The annotation datasets were classified by their functional categories, and users can choose the corresponding options based on their own purposes. All data sets in dbPSP are made available under a Creative Commons CC 3.0 BY license (https://creativecommons.org/licenses/by/3.0/cn/).
Code availability
The source code of dbPSP 2.0 database has been uploaded to GitHub: https://github.com/BioCUCKOO/dbPSP2.0.
References
Mijakovic, I., Grangeasse, C. & Turgay, K. Exploring the diversity of protein modifications: special bacterial phosphorylation systems. FEMS Microbiol Rev 40, 398–417 (2016).
Esser, D. et al. Protein phosphorylation and its role in archaeal signal transduction. FEMS Microbiol Rev 40, 625–647 (2016).
Stock, A. M., Robinson, V. L. & Goudreau, P. N. Two-component signal transduction. Annu Rev Biochem 69, 183–215 (2000).
Moglich, A. Signal transduction in photoreceptor histidine kinases. Protein Sci 28, 1923–1946 (2019).
Guo, Y. et al. iEKPD 2.0: an update with rich annotations for eukaryotic protein kinases, protein phosphatases and proteins containing phosphoprotein-binding domains. Nucleic Acids Res 47, D344–D350 (2019).
Tagliabracci, V. S., Pinna, L. A. & Dixon, J. E. Secreted protein kinases. Trends in biochemical sciences 38, 121–130 (2013).
Garnak, M. & Reeves, H. C. Phosphorylation of Isocitrate dehydrogenase of Escherichia coli. Science 203, 1111–1112 (1979).
Cozzone, A. J. Protein phosphorylation in prokaryotes. Annu Rev Microbiol 42, 97–125 (1988).
Matthews, H. R. Protein kinases and phosphatases that act on histidine, lysine, or arginine residues in eukaryotic proteins: a possible regulator of the mitogen-activated protein kinase cascade. Pharmacol Ther 67, 323–350 (1995).
Khoury, G. A., Baliban, R. C. & Floudas, C. A. Proteome-wide post-translational modification statistics: frequency analysis and curation of the swiss-prot database. Sci Rep 1, 90 (2011).
Trentini, D. B. et al. Arginine phosphorylation marks proteins for degradation by a Clp protease. Nature 539, 48–53 (2016).
Fuhs, S. R. & Hunter, T. pHisphorylation: the emergence of histidine phosphorylation as a reversible regulatory modification. Curr Opin Cell Biol 45, 8–16 (2017).
Lai, S. J. et al. Site-specific His/Asp phosphoproteomic analysis of prokaryotes reveals putative targets for drug resistance. BMC Microbiol 17, 123 (2017).
Kitanishi, K. et al. Identification and functional and spectral characterization of a globin-coupled histidine kinase from Anaeromyxobacter sp. Fw109-5. J Biol Chem 286, 35522–35534 (2011).
Yadav, G. S., Ravala, S. K., Malhotra, N. & Chakraborti, P. K. Phosphorylation Modulates Catalytic Activity of Mycobacterial Sirtuins. Front Microbiol 7, 677 (2016).
Villarino, A. et al. Proteomic identification of M. tuberculosis protein kinase substrates: PknB recruits GarA, a FHA domain-containing protein, through activation loop-mediated interactions. J Mol Biol 350, 953–963 (2005).
Forest, K. T., Dunham, S. A., Koomey, M. & Tainer, J. A. Crystallographic structure reveals phosphorylated pilin from Neisseria: phosphoserine sites modify type IV pilus surface chemistry and fibre morphology. Mol Microbiol 31, 743–752 (1999).
Macek, B. et al. The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis. Mol Cell Proteomics 6, 697–707 (2007).
Macek, B. et al. Phosphoproteome analysis of E. coli reveals evolutionary conservation of bacterial Ser/Thr/Tyr phosphorylation. Mol Cell Proteomics 7, 299–307 (2008).
Elsholz, A. K. et al. Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis. Proc Natl Acad Sci USA 109, 7451–7456 (2012).
Schmidt, A. et al. Quantitative phosphoproteomics reveals the role of protein arginine phosphorylation in the bacterial stress response. Mol Cell Proteomics 13, 537–550 (2014).
Pan, Z. et al. dbPSP: a curated database for protein phosphorylation sites in prokaryotes. Database (Oxford) 2015, bav031 (2015).
Wurgler-Murphy, S. M., King, D. M. & Kennelly, P. J. The Phosphorylation Site Database: A guide to the serine-, threonine-, and/or tyrosine-phosphorylated proteins in prokaryotic organisms. Proteomics 4, 1562–1570 (2004).
UniProt, C. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res 47, D506–D515 (2019).
Huang, K. Y. et al. dbPTM in 2019: exploring disease association and cross-talk of post-translational modifications. Nucleic Acids Res 47, D298–D308 (2019).
Li, J. et al. SysPTM 2.0: an updated systematic resource for post-translational modification. Database (Oxford) 2014, bau025 (2014).
Gnad, F., Gunawardena, J. & Mann, M. PHOSIDA 2011: the posttranslational modification database. Nucleic Acids Res 39, D253–260 (2011).
Standish, A. J. et al. Unprecedented Abundance of Protein Tyrosine Phosphorylation Modulates Shigella flexneri Virulence. J Mol Biol 428, 4197–4208 (2016).
de Keijzer, J. et al. Mechanisms of Phenotypic Rifampicin Tolerance in Mycobacterium tuberculosis Beijing Genotype Strain B0/W148 Revealed by Proteomics. J Proteome Res 15, 1194–1204 (2016).
Nakedi, K. C., Nel, A. J., Garnett, S., Blackburn, J. M. & Soares, N. C. Comparative Ser/Thr/Tyr phosphoproteomics between two mycobacterial species: the fast growing Mycobacterium smegmatis and the slow growing Mycobacterium bovis BCG. Front Microbiol 6, 237 (2015).
Cohen, P. The origins of protein phosphorylation. Nat Cell Biol 4, E127–130 (2002).
Sharma, K. et al. Ultradeep human phosphoproteome reveals a distinct regulatory nature of Tyr and Ser/Thr-based signaling. Cell Rep 8, 1583–1594 (2014).
Kersey, P. J. et al. Ensembl Genomes 2018: an integrated omics infrastructure for non-vertebrate species. Nucleic Acids Res 46, D802–D808 (2018).
Madeira, F., Madhusoodanan, N., Lee, J., Tivey, A. R. N. & Lopez, R. Using EMBL-EBI Services via Web Interface and Programmatically via Web Services. Curr Protoc Bioinformatics 66, e74 (2019).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44, D457–462 (2016).
Sayers, E. W. et al. GenBank. Nucleic Acids Res 48, D84–D86 (2019).
The Gene Ontology, C. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res 47, D330–D338 (2019).
Gaulton, A. et al. The ChEMBL database in 2017. Nucleic Acids Res 45, D945–D954 (2017).
Reimer, L. C. et al. BacDive in 2019: bacterial phenotypic data for High-throughput biodiversity analysis. Nucleic Acids Res 47, D631–D636 (2019).
Burley, S. K. et al. RCSB Protein Data Bank: biological macromolecular structures enabling research and education in fundamental biology, biomedicine, biotechnology and energy. Nucleic Acids Res 47, D464–D474 (2019).
Meszaros, B., Erdos, G. & Dosztanyi, Z. IUPred2A: context-dependent prediction of protein disorder as a function of redox state and protein binding. Nucleic Acids Res 46, W329–W337 (2018).
Mitchell, A. L. et al. InterPro in 2019: improving coverage, classification and access to protein sequence annotations. Nucleic Acids Res 47, D351–D360 (2019).
Oughtred, R. et al. The BioGRID interaction database: 2019 update. Nucleic Acids Res 47, D529–D541 (2019).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res 47, D309–D314 (2019).
Jupe, S. et al. Interleukins and their signaling pathways in the Reactome biological pathway database. J Allergy Clin Immunol 141, 1411–1416 (2018).
Humphrey, S. J. et al. Dynamic adipocyte phosphoproteome reveals that Akt directly regulates mTORC2. Cell metabolism 17, 1009–1020 (2013).
Galperin, M. Y., Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Expanded microbial genome coverage and improved protein family annotation in the COG database. Nucleic Acids Res 43, D261–269 (2015).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32, 1792–1797 (2004).
O’Shea, J. P. et al. pLogo: a probabilistic approach to visualizing sequence motifs. Nat Methods 10, 1211–1212 (2013).
Ullah, S. et al. dbPAF: an integrative database of protein phosphorylation in animals and fungi. Sci Rep 6, 23534 (2016).
Cheng, H. et al. dbPPT: a comprehensive database of protein phosphorylation in plants. Database (Oxford) 2014, bau121 (2014).
Garcia-Garcia, T. et al. Role of Protein Phosphorylation in the Regulation of Cell Cycle and DNA-Related Processes in Bacteria. Front Microbiol 7, 184 (2016).
Venkat, S. et al. Genetically Incorporating Two Distinct Post-translational Modifications into One Protein Simultaneously. ACS Synth Biol 7, 689–695 (2018).
Lin, M. H. et al. A New Tool to Reveal Bacterial Signaling Mechanisms in Antibiotic Treatment and Resistance. Mol Cell Proteomics 17, 2496–2507 (2018).
Hasan, M. M., Rashid, M. M., Khatun, M. S. & Kurata, H. Computational identification of microbial phosphorylation sites by the enhanced characteristics of sequence information. Sci Rep 9, 8258 (2019).
Bradley, D. & Beltrao, P. Evolution of protein kinase substrate recognition at the active site. PLoS Biol 17, e3000341 (2019).
Bourret, R. B., Borkovich, K. A. & Simon, M. I. Signal transduction pathways involving protein phosphorylation in prokaryotes. Annu Rev Biochem 60, 401–441 (1991).
Vahidi, S. et al. Reversible inhibition of the ClpP protease via an N-terminal conformational switch. Proc Natl Acad Sci USA 115, E6447–E6456 (2018).
Bhandari, V. et al. The Role of ClpP Protease in Bacterial Pathogenesis and Human Diseases. ACS Chem Biol 13, 1413–1425 (2018).
Trentini, D. B., Fuhrmann, J., Mechtler, K. & Clausen, T. Chasing Phosphoarginine Proteins: Development of a Selective Enrichment Method Using a Phosphatase Trap. Mol Cell Proteomics 13, 1953–1964 (2014).
Rego, N. & Koes, D. 3Dmol.js: molecular visualization with WebGL. Bioinformatics 31, 1322–1324 (2015).
Shi, Y. et al. dbPSP 2.0, an updated database of protein phosphorylation sites in prokaryotes. Figshare https://doi.org/10.6084/m9.figshare.11436879 (2020).
Acknowledgements
This work was supported by grants from the Special Project on Precision Medicine under the National Key R&D Program (2017YFC0906600 and 2018YFC0910500), Natural Science Foundation of China (81701567, 31930021, 31970633 and 31671360), China Postdoctoral Science Foundation (2018M642816 and 2019T120648), Fundamental Research Funds for the Central Universities (2017KFXKJC001 and 2019kfyRCPY043), Changjiang Scholars Program of China, and program for HUST Academic Frontier Youth Team. The manuscript has been edited by American Journal Experts (AJE) prior to submission.
Author information
Authors and Affiliations
Contributions
Y.X. and D.P. conceived and supervised this study. Y.S. and Y.Z. collected known p-sites, integrated various data resources and developed this updated database. S.L., C.W., J.Z. and H.X. participated in processing data resources. Y.X., D.P. and Y.S. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shi, Y., Zhang, Y., Lin, S. et al. dbPSP 2.0, an updated database of protein phosphorylation sites in prokaryotes. Sci Data 7, 164 (2020). https://doi.org/10.1038/s41597-020-0506-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-020-0506-7