Abstract
We present the newest version of CoryneRegNet, the reference database for corynebacterial regulatory interactions, available at www.exbio.wzw.tum.de/coryneregnet/. The exponential growth of next-generation sequencing data in recent years has allowed a better understanding of bacterial molecular mechanisms. Transcriptional regulation is one of the most important mechanisms for bacterial adaptation and survival. These mechanisms may be understood via an organism’s network of regulatory interactions. Although the Corynebacterium genus is important in medical, veterinary and biotechnological research, little is known concerning the transcriptional regulation of these bacteria. Here, we unravel transcriptional regulatory networks (TRNs) for 224 corynebacterial strains by utilizing genome-scale transfer of TRNs from four model organisms and assigning statistical significance values to all predicted regulations. As a result, the number of corynebacterial strains with TRNs increased twenty times and the back-end and front-end were reimplemented to support new features as well as future database growth. CoryneRegNet 7 is the largest TRN database for the Corynebacterium genus and aids in elucidating transcriptional mechanisms enabling adaptation, survival and infection.
Similar content being viewed by others
Introduction
Next-generation sequencing (NGS) has unraveled the genomic sequence of a multitude of bacterial genomes1. Despite the amount of information, these data do not fully explain how organisms orchestrate their survival on a molecular level. To understand the mechanisms that coordinate an organism’s adaptation to environmental changes, it is crucial to understand how a cell maintains transcription2,3. The main players in the transcriptional regulation of bacterial organisms are transcription factors (TFs). These regulatory proteins recognize transcription factor binding sites (TFBSs) in the upstream region of the respective target genes (TGs), stimulating or repressing their expression4,5,6. Experimental studies such as RNA-Seq7, microarray8, ChIP-chip and ChIP-seq9 have been applied in order to reveal regulatory interactions in a cell. Nevertheless, performing these experiments for all bacterial strains would be labor-intensive and, thus, financially infeasible4,10. As a result, these experimental data are not available for every member of a bacterial genus.
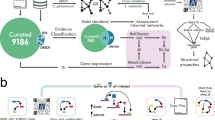
To alleviate this lack of data, genome-scale transfer of TRNs has been applied providing insights into the regulatory mechanisms of bacterial organisms10,11. In this context, a model organism is an organism with the most complete and experimentally validated TRN which can be utilized to predict regulatory interactions in other organisms, called target organisms, with incomplete or less validated TRNs. TRNs are constructed as directed graphs where the nodes represent TFs and their TGs while the criterion to create an edge from a TF to a TG is the regulatory interaction between them6,12,13. Edge labels may then indicate the corresponding TFBSs and/or the type of the regulatory interaction. A reliable method to transfer TRNs from a model organism to taxonomically related target organisms is to consider a regulatory interaction to be conserved between two organisms when the TF, the TG and the TFBS are all conserved4. Figure 1a illustrates a conserved regulatory interaction between two organisms. An example of a TRN transferred from a model organism to a target organism is represented in Fig. 1b. More extensive explanations on genome-scale TRN transfer methods can be found in Baumbach et al.11 and Kiliç et al.14.
Overview of the computational reconstruction of TRNs. (a) General concept of regulatory conservation. (b) TRN transfer scheme: The TRN of a model organism (top) and the predicted TRN of the target organism including all transferred regulations (bottom). In the networks, nodes represent the genes and arrows represent the regulatory interactions.
Both experimentally and computationally reconstructed TRNs are publicly available in databases such as RegulonDB15 for Escherichia coli, EHECRegNet16 for human pathogenic Escherichia coli, TB Portal17 and MTB Network Portal18 for Mycobacterium tuberculosis, DBTBS19 and Subtiwiki20 for Bacillus subtilis, and CoryneRegNet21 for the Corynebacterium genus. RegulonDB15 focuses on detailed and manually curated transcriptional regulation data retrieved from literature for E. coli. MTB Network Portal18 and Subtiwiki20 provide literature-mined transcriptional regulation data on M. tuberculosis and B. subtilis. Abasy Atlas22 is an online collection of regulatory data covering 42 bacteria retrieved from both literature and other online databases. However, there is no resource focusing on corynebacterial gene regulatory networks, and no database that stores predicted TRNs based on evolutionary conservation across a whole collection of model and target organisms. CoryneRegNet has served as the reference database of the genus Corynebacterium since 200623. This genus includes organisms with medical, veterinary and biotechnological relevance24,25,26,27. While the National Center for Biotechnology Information (NCBI) database contains more than 60 corynebacterial species with fully sequenced and annotated genomes, there are TRNs of only eight of these species available in online databases.
The previous version of CoryneRegNet21 was released in 2012 presenting predicted (transferred) TRNs for eleven corynebacterial strains. The steady increase in corynebacterial genomic sequences contained in public databases allows us to unravel further transcriptional regulatory interactions. In the seventh version of CoryneRegNet, we now present 82,268 regulatory interactions, an increase of more than eleven times compared to the sixth version, as well as 228 TRNs, increasing the number of corynebacterial strains with known TRNs by twenty times. It contains up-to-date regulatory information about the model organisms C. glutamicum ATCC 13032, E. coli K-12, M. tuberculosis H37Rv and B. subtilis 168, and predicted TRNs of 224 target organisms of the Corynebacterium genus. Furthermore, we present an increase of more than seven times the number of corynebacterial species with TRNs available in public databases representing a great improvement for the bacterial gene regulatory network research community.
Results
In this section we present the results of the re-implemented back- and front-end, the updated database content and the predicted TRNs of all fully sequenced and annotated corynebacterial genomes.
Updated database content
In CoryneRegNet 7 we updated the database content by adding new model and target organisms. As in previous versions, TRNs are categorized as either experimentally validated or computationally predicted. The former contains up-to-date TRNs of C. glutamicum ATCC 13032, E. coli K-12, M. tuberculosis H37Rv and B. subtilis 168, the latter contains predicted TRNs of a total of 224 corynebacterial strains. A full list of these strains as well as more details regarding the experimental and predicted databases are presented in Supplementary Table S1. The resulting number of predicted TFs, regulated genes, regulations, binding motifs and profile Hidden Markov Models (HMMs) are presented in Table 1 together with the evolution of the database content throughout previous CoryneRegNet versions.
Novel back- and front-end
In this work, we re-implemented CoryneRegNet’s back- and front-end, allowing the user to browse the database via a modern and easy-to-use web-interface. The new architecture (Fig. 2) is inspired by the Model-View-Controller (MVC) architectural pattern28,29. This modular structure allows components to be modified or replaced, facilitating maintenance and future updates.
Schematic of the CoryneRegNet 7 architecture. The processing layer handles the TRN transfer and data parsing. The database access layer manages any query, update, insertion or deletion in the database. The view is responsible for user interaction through the browser. The controller conducts all the communication and data handling among the other three layers.
CoryneRegNet web-interface
The CoryneRegNet 7 website contains TRNs of 224 corynebacterial target genomes and 4 model organisms. Information about quantities of regulator types, distribution of TFs, distribution of co-regulating TFs and distribution of HMM profile lengths are shown in the statistics page. Those are shown for each database (predicted and experimental) as well as for each organism. Figure 3 represents the statistics page for the experimental database.
Overview of main statistics present in CoryneRegNet. In (a) the pie chart presents the quantities of regulator types in percentages, (b) represents the distribution of the numbers of TFs regulating a gene, (c) presents the distribution of co-regulating TFs, and (d) demonstrates the distribution of HMM profiles lengths.
Through the web-interface, the user is able to browse the TRNs in both table and network format. In table format, the list of regulatory interactions (RIs) provides source, target and operon information (Fig. 4a). The network visualization comes with two different layout options: a gene-centered layout and an operon-centered layout, see Fig. 4b,c, respectively. Both visualizations give access to gene information by clicking on the gene and/or operon of interest. Each network may also be downloaded in.sif file format. Additionally, the user can visualize networks of genes or operons of interest by using the network visualization in the gene information pop-up.
Furthermore, we offer a detailed gene information page which shows gene identifiers linked to NCBI, nucleotide and protein sequences, homologous genes, and regulatory information. This page also allows the user to make additional motif searches in the database. A view of this page showing the putative homologous genes of cg0199 is presented in Fig. 5. The user is provided with information of all genes that are predicted to be homologous to the gene of interest that are present in CoryneRegNet 7.
At this point, CoryneRegNet 7 offers the biggest collection of profile HMMs publicly available for the Corynebacterium genus. See the methodology section for an explanation of how those were generated. These profiles and their logos are available for download in the gene information page of genes encoding transcription factors. In addition, the user can utilize the profile HMMs stored in the database to search the upstream region of genes present in CoryneRegNet 7. Two kinds of motif searches are provided: (i) the upstream region of the gene of interest can be searched with HMM profiles of an organism of interest (Fig. 6a) and (ii) the HMM profile of the TF of interest can then be used to identify potential binding sites in the upstream regions of all genes of an organism in the database (Fig. 6b).
Finally, the website provides a comprehensive help page with theoretical and practical explanations of the website content, methodology and navigation, including a broad collection of published literature concerning Corynebacterial transcriptional regulation. This help page can be accessed at www.exbio.wzw.tum.de/coryneregnet/docs&help.htm.
Discussion
In CoryneRegNet version 7, we entirely redesigned the back- and front-end to support the updated database content, new functional features as well as future database growth. As we did in the other versions, we present TRNs for all fully sequenced and annotated corynebacterial genomes available in NCBI (June 2019). Consequently, CoryneRegNet 7 currently offers the biggest knowledge base available regarding TRNs of corynebacterial organisms. Along with our newly-designed web interface, we include an operon network layout and the option to download network views from each organisms network visualization as a file (.sif). This allows users to modify and enrich the network locally and create personalized visualizations based on their own research using third-party software.
We also significantly improved our transfer pipeline in two ways. First, we replaced the use of Position Weight Matrices (PWMs) with profile HMMs in our motif conservation analysis. Profile HMMs enable the modelling of insertions and deletions, greatly improving the detection of remote homologous and model nucleotide dependency as well as length variations in the model’s binding sites30,31,32. TFBSs are considered to have low evolutionary conservation between species11 and this strategy provides more robustness and flexibility when predicting them33. It is an advantage considering that mutations in these sites are expected to occur from one species to another. Second, we added the calculation of p-values, i.e. the likelihood of observing this conservation by chance, for each regulatory interaction which provides important information in interpreting the results.
Even though great progress has been made in the TRN field, there are still a few limitations concerning bacterial TRNs. The transfer of TRNs has been hindered by the limited availability of experimentally validated data on bacterial TRNs which are available for only a few model organisms such as E. coli, B. subitilis and C. glutamicum4,14. TRN transfer from one organism to another largely depends on known regulatory interactions in the model organism as well as genome similarity between model and target organism4,11,13,14. Thus, the more experimental TRN data is available for a greater diversity of bacterial species, the higher the quality of any predicted TRNs. Furthermore, the ability of detecting regulatory interactions acquired by horizontal gene transfer (HGT) becomes relevant4,11,14 considering regulatory interactions related to life-style may not be identified by using only one model organism. Using several model organisms allows us to identify these regulations. Previous studies point out that (i) known virulence determinants in Enterohemorrhagic E. coli are located on mobile genetic elements, which are generally acquired in HGT events16 and (ii) in E. coli neighboring regulators were co-transferred with their TGs in HGT34. Methodologies that support the use of more model organisms together with more high quality experimentally validated TRNs will result in more complete TRNs that will consider lifestyle-related regulatory interactions (e.g. pathogenic or non-pathogenic, free-living or host-associated). A first step in this direction was presented in this study, since we transferred regulatory interactions between organisms of different phyla and presented the joint p-value allowing the researcher to evaluate the degree of conservation of each predicted regulatory interaction.
Methods
Database content update
Genomic data of the 228 organisms used in this work were retrieved from the NCBI database35 in June 2019 (for more details see Supplementary Table S1). TRN data of the model organisms were retrieved from RegulonDB15 for E. coli K-12, Minch et al.36 for M. tuberculosis H37Rv, DBTBS19 for B. subtilis. C. glutamicum ATCC 13032 data from CoryneRegNet 621 were updated with new data from Freyre-González and Tauch (2017)37.
In order to predict TRNs for the 224 corynebacterial strains, we extended the transfer methodology described by Baumbach and collaborators4. First, TF binding profiles were generated for every TF of the model organisms. For this purpose, binding sites of each TF were collected, aligned with Clustal Omega38 and binding profiles were generated using HMM-build from the HMMER package39. Second, we performed all-vs-all protein BLAST40 search and selected the best bidirectional BLAST hits (BBHs) using a cutoff of 10−10 to predict homologous proteins. The upstream regions (−560, +20) of all genes and operons in the analysis were identified. Third, the upstream regions of all homologous TGs in the target organisms were scanned using HMMER30 to predict conserved TFBSs. Figure 1a illustrates this approach. The HMM profiles of the conserved TFs were applied to the upstream regions of the potentially regulated TGs by using HMMER’s default parameters, which corresponds to a p-value of ~10−5 41. Genes with an intergenic distance of less than 50 base pairs were considered to be part of the same operon and predicted regulatory interactions to the first gene were extended to the operon4. The role of a predicted regulatory interaction is inherited from the model regulatory interaction used in the transfer. Finally, profile HMMs were generated for predicted TFs as described above for model TFs. The interaction p-value was obtained by applying Tippet’s method42. The R package Metap43 was used to calculate the joint p-value of the p-values obtained in the homology and motif searches. These steps are summarized in Fig. 7.
Implementation of CoryneRegNet 7
The CoryneRegNet7 back-end was developed using Java and the database was implemented using the PostgreSQL44 management system for relational databases. Hibernate45 was used for object-relational mapping between PostgreSQL and Java. The Spring framework46 was used to implement a Model-View-Controller (MVC) architectural pattern28,29. The new front-end was developed using HTML547, CSS348, Bootstrap49 and JavaScript. We used vis.js50 to implement the network visualization and graphs in the statistics page were drawn using D351. The website was tested and runs on Google Chrome and Mozilla Firefox. Furthermore, the screen resolution was optimized for tablet (1024 × 768), laptop (1920 × 1080) and desktop (2560 × 1440) resolutions.
Data availability
All data generated in this work is provided to the research community free of charge as comma separated values (.csv format) via the figshare repository52 and in CoryneRegNet’s download section (http://www.exbio.wzw.tum.de/coryneregnet/processToDownalod.htm).
Code availability
CoryneRegNet7 code is available on GitHub: https://github.com/baumbachlab/CoryneRegNet7.
References
Haft, D. H. et al. RefSeq: an update on prokaryotic genome annotation and curation. Nucleic Acids Res. 46, D851–D860 (2018).
Röttger, R., Rückert, U., Taubert, J. & Baumbach, J. How little do we actually know? On the size of gene regulatory networks. IEEE/ACM Trans. Comput. Biol. Bioinform. 9, 1293–1300 (2012).
Park, J. & Wang, H. H. Systematic and synthetic approaches to rewire regulatory networks. Curr Opin Syst Biol 8, 90–96 (2018).
Baumbach, J., Rahmann, S. & Tauch, A. Reliable transfer of transcriptional gene regulatory networks between taxonomically related organisms. BMC Syst. Biol. 3, 8 (2009).
Voordeckers, K., Pougach, K. & Verstrepen, K. J. How do regulatory networks evolve and expand throughout evolution? Curr. Opin. Biotechnol. 34, 180–188 (2015).
Hao, T. et al. The Genome-Scale Integrated Networks in Microorganisms. Front. Microbiol. 9, 296 (2018).
Nagalakshmi, U. et al. The transcriptional landscape of the yeast genome defined by RNA sequencing. Science 320, 1344–1349 (2008).
Schena, M., Shalon, D., Davis, R. W. & Brown, P. O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270, 467–470 (1995).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Leyn, S. A. et al. Comparative genomics and evolution of transcriptional regulons in Proteobacteria. Microb Genom 2, e000061 (2016).
Baumbach, J. On the power and limits of evolutionary conservation—unraveling bacterial gene regulatory networks. Nucleic Acids Res. 38, 7877–7884 (2010).
Babu, M. M., Lang, B. & Aravind, L. Methods to Reconstruct and Compare Transcriptional Regulatory Networks. Methods in Molecular Biology 541, 163–180 (2009).
Thompson, D., Regev, A. & Roy, S. Comparative analysis of gene regulatory networks: from network reconstruction to evolution. Annu. Rev. Cell Dev. Biol. 31, 399–428 (2015).
Kılıç, S. & Erill, I. Assessment of transfer methods for comparative genomics of regulatory networks in bacteria. BMC Bioinformatics 17(Suppl 8), 277 (2016).
Santos-Zavaleta, A. et al. RegulonDB v 10.5: tackling challenges to unify classic and high throughput knowledge of gene regulation in E. coli K-12. Nucleic Acids Res. 47, D212–D220 (2019).
Pauling, J. et al. On the trail of EHEC/EAEC–unraveling the gene regulatory networks of human pathogenic Escherichia coli bacteria. Integr. Biol. 4, 728–733 (2012).
Galagan, J. E. et al. TB database 2010: overview and update. Tuberculosis 90, 225–235 (2010).
Turkarslan, S. et al. A comprehensive map of genome-wide gene regulation in Mycobacterium tuberculosis. Sci Data 2, 150010 (2015).
Sierro, N., Makita, Y., de Hoon, M. & Nakai, K. DBTBS: a database of transcriptional regulation in Bacillus subtilis containing upstream intergenic conservation information. Nucleic Acids Res. 36, D93–6 (2008).
Zhu, B. & Stülke, J. SubtiWiki in 2018: from genes and proteins to functional network annotation of the model organism Bacillus subtilis. Nucleic Acids Res. 46, D743–D748 (2018).
Pauling, J., Röttger, R., Tauch, A., Azevedo, V. & Baumbach, J. CoryneRegNet 6.0–Updated database content, new analysis methods and novel features focusing on community demands. Nucleic Acids Res. 40, D610–4 (2012).
Ibarra-Arellano, M. A., Campos-González, A. I., Treviño-Quintanilla, L. G., Tauch, A. & Freyre-González, J. A. Abasy Atlas: a comprehensive inventory of systems, global network properties and systems-level elements across bacteria. Database 2016, 1–16 (2016).
Baumbach, J., Brinkrolf, K., Czaja, L. F., Rahmann, S. & Tauch, A. CoryneRegNet: an ontology-based data warehouse of corynebacterial transcription factors and regulatory networks. BMC Genomics 7, 24 (2006).
Jamal, S. B., Tiwari, S., Silva, A. & Azevedo, V. Pathogenesis of Corynebacterium diphtheriae and available vaccines; an overview. Glob. J. Infect. Dis. Clin. Res 3, 20–24 (2017).
Viana, M. V. C. et al. Comparative genomic analysis between Corynebacterium pseudotuberculosis strains isolated from buffalo. PLoS One 12, e0176347 (2017).
Parise, D. et al. First genome sequencing and comparative analyses of Corynebacterium pseudotuberculosis strains from Mexico. Stand. Genomic Sci. 13, 21 (2018).
Baltz, R. H., Demain, A. L. & Davies, J. E. Manual of Industrial Microbiology and Biotechnology. (American Society for Microbiology Press, 2010).
Pope, K. G. & Krasner, S. A cookbook for using the model-view-controller user interface paradigm in Smalltalk-80. Journal of Object-Oriented Programming 1 (1988).
Gupta, P. & Govil, M. C. MVC Design Pattern for the multi framework distributed applications using XML, spring and struts framework. International Journal on Computer Science and Engineering 2, 1047–1051 (2010).
Wheeler, T. J. & Eddy, S. R. nhmmer: DNA homology search with profile HMMs. Bioinformatics 29, 2487–2489 (2013).
Madera, M. & Gough, J. A comparison of profile hidden Markov model procedures for remote homology detection. Nucleic Acids Res. 30, 4321–4328 (2002).
Delorenzi, M. & Speed, T. An HMM model for coiled-coil domains and a comparison with PSSM-based predictions. Bioinformatics 18, 617–625 (2002).
Riva, A. The MAPPER2 Database: a multi-genome catalog of putative transcription factor binding sites. Nucleic Acids Res. 40, D155–61 (2012).
Price, M. N., Dehal, P. S. & Arkin, A. P. Horizontal gene transfer and the evolution of transcriptional regulation in Escherichia coli. Genome Biol. 9, R4 (2008).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 47, D23–D28 (2019).
Minch, K. J. et al. The DNA-binding network of Mycobacterium tuberculosis. Nat. Commun. 6, 5829 (2015).
Freyre-González, J. A. & Tauch, A. Functional architecture and global properties of the Corynebacterium glutamicum regulatory network: Novel insights from a dataset with a high genomic coverage. J. Biotechnol. 257, 199–210 (2017).
Sievers, F. et al. Fast, scalable generation of high‐quality protein multiple sequence alignments using Clustal Omega. Molecular Systems Biology 7, 539 (2011).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Research 39, W29–W37 (2011).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Baumbach, J., Brinkrolf, K., Wittkop, T., Tauch, A. & Rahmann, S. CoryneRegNet 2: An Integrative Bioinformatics Approach for Reconstruction and Comparison of Transcriptional Regulatory Networks in Prokaryotes. J. Integr. Bioinform. 3, 1–13 (2006).
Heard, N. A. & Rubin-Delanchy, P. Choosing between methods of combining p-values. Biometrika 105, 239–246 (2018).
Dewey, M. metap: meta-analysis of significance values. R package version 1.1 (2019).
PostgreSQL Team. PostgreSQL: The World’s Most Advanced Open Source Relational Database, version 9.3, https://www.postgresql.org/ (2019).
Hibernate Team. Hibernate Object/Relational Mapping, version 4.3, https://hibernate.org/orm/ (2019).
Spring Framework Team. Spring Framework, version 4.1, https://spring.io/ (2019).
HTML Team. HTML Standard, version 5, https://html.spec.whatwg.org/multipage/ (2019).
CSS Team. CSS Snapshot 2018, version3, https://www.w3.org/TR/css3-roadmap/ (2019).
Otto, M., Thornton, J. & Bootstrap contributors. Bootstrap Introduction, version 4.0, https://getbootstrap.com/docs/4.0/getting-started/introduction/ (2019).
Vis.js Team. Vis.js, community edition, https://visjs.org/ (2019).
Bostock, M., Ogievetsky, V. & Heer, J. D3: Data-Driven Documents. IEEE Trans. Vis. Comput. Graph. 17, 2301–2309 (2011).
Tauch, A. et al. CoryneRegNet 7 - The reference database and online analysis platform for corynebacterial gene regulatory networks. figshare https://doi.org/10.6084/m9.figshare.c.4720991 (2020).
Baumbach, J. et al. CoryneRegNet 3.0—An interactive systems biology platform for the analysis of gene regulatory networks in corynebacteria and Escherichia coli. Journal of Biotechnology 129, 279–289 (2007).
Baumbach, J. CoryneRegNet 4.0–A reference database for corynebacterial gene regulatory networks. BMC Bioinformatics 8, 429 (2007).
Acknowledgements
JB is grateful for support from H2020 grant RepoTrial (nr. 777111) and his VILLUM Young Investigator grant (nr. 13154). MP received support from CNPq (nr. 201336/2018-9), and DP from CAPES (nr. 88887.364607/2019-00) for their work at TUM in Germany. MP’s work was also supported by the German Research Foundation (under SFB924). Contributions by J.P. were funded by the Bavarian State Ministry of Science and the Arts in the framework of the Center Digitisation.Bavaria (ZD.B, Zentrum Digitalisierung.Bayern). V.A. is grateful for support from his CNPq Research Productivity grant (nr. 305093/2015-0), CNPq Universal grant (nr. 405233/2016-7) and FAPEMIG grant (nr. APQ 02600-17). This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Finance Code 001.
Author information
Authors and Affiliations
Contributions
M.P. and D.P. developed the new version of CoryneRegNet and wrote the manuscript. J.B. and V.A. designed the project. J.B., J.P., V.A., A.T. and R.K. reviewed the manuscript. M.P., D.P., J.P. and J.B. revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Parise, M.T.D., Parise, D., Kato, R.B. et al. CoryneRegNet 7, the reference database and analysis platform for corynebacterial gene regulatory networks. Sci Data 7, 142 (2020). https://doi.org/10.1038/s41597-020-0484-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-020-0484-9
This article is cited by
-
A genome-reduced Corynebacterium glutamicum derivative discloses a hidden pathway relevant for 1,2-propanediol production
Microbial Cell Factories (2024)
-
A 4-hydroxybenzoate 3-hydroxylase mutant enables 4-amino-3-hydroxybenzoic acid production from glucose in Corynebacterium glutamicum
Microbial Cell Factories (2023)
-
Competition and evolutionary selection among core regulatory motifs in gene expression control
Nature Communications (2023)
-
A manually curated compendium of expression profiles for the microbial cell factory Corynebacterium glutamicum
Scientific Data (2022)