Abstract
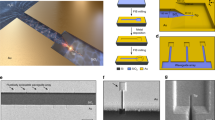
The spatiotemporal aspects of early signaling events during interactions between cells and their environment dictate multiple downstream outcomes. While advances in nanopatterning techniques have allowed the isolation of these signaling events, a major limitation of conventional nanopatterning methods is its dependence on gold (Au) or related materials that plasmonically quench fluorescence and, thus, are incompatible with super-resolution fluorescence microscopy. Here we describe a novel method that integrates nanopatterning with single-molecule resolution fluorescence imaging, thus enabling mechanistic dissection of molecular-scale signaling events in conjunction with nanoscale geometry manipulation. Our method exploits nanofabricated titanium (Ti) whose oxide (TiO2) is a dielectric material with no plasmonic effects. We describe the surface chemistry for decorating specific ligands such as cyclo-RGD (arginine, glycine and aspartate: a ligand for fibronectin-binding integrins) on TiO2 nanoline and nanodot substrates, and demonstrate the ability to perform dual-color super-resolution imaging on these patterns. Ti nanofabrication is similar to other metallic materials like Au, while the functionalization of TiO2 is relatively fast, safe, economical, easy to set up with commonly available reagents, and robust against environmental parameters such as humidity. Fabrication of nanopatterns takes ~2–3 d, preparation for functionalization ~1.5–2 d, and functionalization 3 h, after which cell culture and imaging experiments can be performed. We suggest that this method may facilitate the interrogation of nanoscale geometry and force at single-molecule resolution, and should find ready applications in early detection and interpretation of physiochemical signaling events at the cell membrane in the fields of cell biology, immunology, regenerative medicine, and related fields.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
Data availability
No original code has been used for this paper. The raw data that support the anticipated results are available at figshare: https://doi.org/10.6084/m9.figshare.19337648. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
References
Geiger, B., Spatz, J. P. & Bershadsky, A. D. Environmental sensing through focal adhesions. Nat. Rev. Mol. Cell Biol. 10, 21–33 (2009).
Cukierman, E., Pankov, R., Stevens, D. R. & Yamada, K. M. Taking cell–matrix adhesions to the third dimension. Science 294, 1708–1712 (2001).
Doyle, A. D. & Yamada, K. M. Mechanosensing via cell-matrix adhesions in 3D microenvironments. Exp. Cell Res. 343, 60–66 (2016).
Changede, R., Cai, H., Wind, S. J. & Sheetz, M. P. Integrin nanoclusters can bridge thin matrix fibres to form cell–matrix adhesions. Nat. Mater. 18, 1366–1375 (2019).
Erickson, B. et al. Nanoscale structure of type I collagen fibrils: quantitative measurement of D-spacing. Biotechnol. J. 8, 117–126 (2013).
Früh, S. M., Schoen, I., Ries, J. & Vogel, V. Molecular architecture of native fibronectin fibrils. Nat. Commun. 6, 1–10 (2015).
Mossman, K. D., Campi, G., Groves, J. T. & Dustin, M. L. Altered TCR signaling from geometrically repatterned immunological synapses. Science 310, 1191–1193 (2005).
Lee, K.-H. et al. T cell receptor signaling precedes immunological synapse formation. Science 295, 1539–1542 (2002).
Morrison, S. J. & Kimble, J. Asymmetric and symmetric stem-cell divisions in development and cancer. Nature 441, 1068–1074 (2006).
Moya, I. M. & Halder, G. Hippo–YAP/TAZ signalling in organ regeneration and regenerative medicine. Nat. Rev. Mol. Cell Biol. 20, 211–226 (2019).
Davalos, D. et al. ATP mediates rapid microglial response to local brain injury in vivo. Nat. Neurosci. 8, 752–758 (2005).
Papenfort, K. & Bassler, B. L. Quorum sensing signal–response systems in Gram-negative bacteria. Nat. Rev. Microbiol. 14, 576–588 (2016).
Lindner, M. et al. A fast and simple contact printing approach to generate 2D protein nanopatterns. Front. Chem. 7, 655 (2019).
Huang, Y., Tran, H. & Ober, C. K. High-resolution nanopatterning of free-standing, self-supported helical polypeptide rod brushes via electron beam lithography. ACS Macro Lett. 10, 755–759 (2021).
Han, P. et al. Five piconewtons: the difference between osteogenic and adipogenic fate choice in human mesenchymal stem cells. ACS Nano 13, 11129–11143 (2019).
Killops, K. L. et al. Nanopatterning biomolecules by block copolymer self-assembly. ACS Macro Lett. 1, 758–763 (2012).
M, A. et al. Activation of integrin function by nanopatterned adhesive interfaces. Chemphyschem 5, 383–388 (2004).
Cai, H. et al. Full control of ligand positioning reveals spatial thresholds for T cell receptor triggering. Nat. Nanotechnol. 13, 610–617 (2018).
Lohm€, T. et al. Supported membranes embedded with fixed arrays of gold nanoparticles. Nano Lett. 11, 37 (2011).
Cai, H. et al. Molecular occupancy of nanodot arrays. ACS Nano 10, 4173–4183 (2016).
Chighizola F, M. et al. The glycocalyx affects force loading-dependent mechanotransductive topography sensing at the nanoscale. Preprint at https://www.biorxiv.org/content/10.1101/2021.03.02.433591v2 (2021).
Schulte, C. et al. Conversion of nanoscale topographical information of cluster-assembled zirconia surfaces into mechanotransductive events promotes neuronal differentiation. J. Nanobiotechnol. 14, 1–24 (2016).
Elias, C. N., Lima, J. H. C., Valiev, R. & Meyers, M. A. Biomedical applications of titanium and its alloys. JOM 60, 46–49 (2008). 2008 603.
Paschalis, E. I. et al. In vitro and in vivo assessment of titanium surface modification for coloring the backplate of the Boston keratoprosthesis. Invest. Ophthalmol. Vis. Sci. 54, 3863–3873 (2013).
Sidambe, A. T. & Oh, J. K. Biocompatibility of advanced manufactured titanium implants—a review. Materials 7, 8168–8188 (2014).
Quinn, R. K. & Armstrong, N. R. Electrochemical and surface analytical characterization of titanium and titanium hydride thin film electrode oxidation. J. Electrochem. Soc. 125, 1790–1796 (1978).
Ponader, S. et al. Effects of topographical surface modifications of electron beam melted Ti-6Al-4V titanium on human fetal osteoblasts. J. Biomed. Mater. Res. A 84, 1111–1119 (2008).
Haslauer, C. M. et al. In vitro biocompatibility of titanium alloy discs made using direct metal fabrication. Med. Eng. Phys. 32, 645–652 (2010).
Guasch, J. et al. Synthesis of binary nanopatterns on hydrogels for initiating cellular responses. Chem. Mater. 28, 1806–1815 (2016).
Ziental, D. et al. Titanium dioxide nanoparticles: prospects and applications in medicine. Nanomaterials 10, 387 (2020).
Rajh, T., Dimitrijevic, N. M. & Rozhkova, E. A. Titanium dioxide nanoparticles in advanced imaging and nanotherapeutics. Methods Mol. Biol. 726, 63–75 (2011).
Rechenmacher, F. et al. A molecular toolkit for the functionalization of titanium-based biomaterials that selectively control integrin-mediated cell adhesion. Chemistry 19, 9218–9223 (2013).
Svensson, F. G., Daniel, G., Tai, C.-W., Seisenbaeva, G. A. & Kessler, V. G. Titanium phosphonate oxo-alkoxide ‘clusters’: solution stability and facile hydrolytic transformation into nano titania. RSC Adv. 10, 6873–6883 (2020).
Eriksson, A. I. K., Edwards, K., Hagfeldt, A. & Hernández, V. A. Physicochemical characterization of phosphopeptide/titanium dioxide interactions employing the quartz crystal microbalance technique. J. Phys. Chem. B 117, 2019–2025 (2013).
Glass, R., Möller, M. & Spatz, J. P. Block copolymer micelle nanolithography. Nanotechnology 14, 1153 (2003).
Harris, J. M. Poly (ethylene glycol) Chemistry: Biotechnical and Biomedical Applications (Springer Science & Business Media, 1992).
Jain, A., Liu, R., Xiang, Y. K. & Ha, T. Single-molecule pull-down for studying protein interactions. Nat. Protoc. 7, 445–452 (2012).
Blümmel, J. et al. Protein repellent properties of covalently attached PEG coatings on nanostructured SiO2-based interfaces. Biomaterials 28, 4739–4747 (2007).
Zhu, B., Eurell, T., Gunawan, R. & Leckband, D. Chain-length dependence of the protein and cell resistance of oligo(ethylene glycol)-terminated self-assembled monolayers on gold. J. Biomed. Mater. Res. 56, 406–416 (2001).
Cai, H. & Wind, S. J. Improved glass surface passivation for single-molecule nanoarrays. Langmuir 32, 10034–10041 (2016).
Persson, F. et al. Lipid-based passivation in nanofluidics. Nano Lett. 12, 2260–2265 (2012).
Patterson, G. H. & Lippincott-Schwartz, J. A photoactivatable GFP for selective photolabeling of proteins and cells. Science 297, 1873–1877 (2002).
Durisic, N., Laparra-Cuervo, L., Sandoval-Álvarez, Á., Borbely, J. S. & Lakadamyali, M. Single-molecule evaluation of fluorescent protein photoactivation efficiency using an in vivo nanotemplate. Nat. Methods 11, 156–162 (2014).
Yuwono, A. H. et al. Diblock copolymer templated nanohybrid thin films of highly ordered TiO2 nanoparticle arrays in PMMA matrix. Chem. Mater. 18, 5876–5889 (2006).
Polleux, J. et al. Benzyl alcohol and block copolymer micellar lithography: a versatile route to assembling gold and in situ generated titania nanoparticles into uniform binary nanoarrays. ACS Nano 5, 6355–6364 (2011).
Whelan, D. R. & Bell, T. D. M. Super-resolution single-molecule localization microscopy: tricks of the trade. J. Phys. Chem. Lett. 6, 374–382 (2015).
Khater, I. M., Nabi, I. R. & Hamarneh, G. A review of super-resolution single-molecule localization microscopy cluster analysis and quantification methods. Patterns 1, 100038 (2020).
Ovesný, M., Křížek, P., Borkovec, J., Švindrych, Z. & Hagen, G. M. ThunderSTORM: a comprehensive ImageJ plug-in for PALM and STORM data analysis and super-resolution imaging. Bioinformatics 30, 2389–2390 (2014).
Changede, R., Xu, X., Margadant, F. & Sheetz, M. P. Nascent integrin adhesions form on all matrix rigidities after integrin activation. Dev. Cell 35, 614–621 (2015).
Schnitzbauer, J., Strauss, M. T., Schlichthaerle, T., Schueder, F. & Jungmann, R. Super-resolution microscopy with DNA-PAINT. Nat. Protoc. 12, 1198–1228 (2017).
Wolter, S. et al. rapidSTORM: accurate, fast open-source software for localization microscopy. Nat. Methods 9, 1040–1041 (2012).
Nehme, E., Weiss, L. E., Michaeli, T. & Shechtman, Y. Deep-STORM: super-resolution single-molecule microscopy by deep learning. Optica 5, 458–464 (2018).
Vogel, V. & Sheetz, M. P. Mechanical forces matter in health and disease: from cancer to tissue engineering. Nanotechnology 5, 233–303 (2010).
Vogel, V. & Sheetz, M. Local force and geometry sensing regulate cell functions. Nat. Rev. Mol. Cell Biol. 7, 265–275 (2006).
Zhan, T., Rindtorff, N. & Boutros, M. Wnt signaling in cancer. Oncogene 36, 1461–1473 (2016).
Sanchez-Vega, F. et al. Oncogenic signaling pathways in The Cancer Genome Atlas. Cell 173, 321–337.e10 (2018).
Pan, L. et al. Higher-order clustering of the transmembrane anchor of DR5 drives signaling. Cell 176, 1477–1489.e14 (2019).
L Berger, R. M. et al. Nanoscale FasL organization on DNA origami to decipher apoptosis signal activation in cells. Small 17, 2101678 (2021).
Aggarwal, B. B. Signalling pathways of the TNF superfamily: a double-edged sword. Nat. Rev. Immunol. 3, 745–756 (2003).
Wang, H., Luo, X. & Leighton, J. Extracellular matrix and integrins in embryonic stem cell differentiation. Biochem. Insights 8, 15–21 (2015).
Pathan-Chhatbar, S. et al. Direct Regulation of the T Cell Antigen Receptor’s Activity by Cholesterol. Front. Cell Dev. Biol. 8, 1728 (2021).
Goyette, J., Nieves, D. J., Ma, Y. & Gaus, K. How does T cell receptor clustering impact on signal transduction? J. Cell Sci. 132, jcs226423 (2019).
Nassereddine, A. et al. Ligand nanocluster array enables artificial-intelligence-based detection of hidden features in T-cell architecture. Nano Lett. 21, 5606–5613 (2021).
Li, M. & Yu, Y. Innate immune receptor clustering and its role in immune regulation. J. Cell Sci. 134, jcs249318 (2021).
Inoue, M. & Shinohara, M. L. Clustering of pattern recognition receptors for fungal detection. PLoS Pathog. 10, e1003873 (2014).
Gay, N. J., Symmons, M. F., Gangloff, M. & Bryant, C. E. Assembly and localization of Toll-like receptor signalling complexes. Nat. Rev. Immunol. 14, 546–558 (2014).
Ma, Z. et al. Membrane nanodomains modulate formin condensation for actin remodeling in Arabidopsis innate immune responses. Plant Cell 34, 374–394 (2022).
Yap, L., Tay, H. G., Nguyen, M. T. X., Tjin, M. S. & Tryggvason, K. Laminins in cellular differentiation. Trends Cell Biol. 29, 987–1000 (2019).
Ivetic, A., Green, H. L. H. & Hart, S. J. L-selectin: a major regulator of leukocyte adhesion, migration and signaling. Front. Immunol. 10, 1068 (2019).
Mattila, P. E., Green, C. E., Schaff, U., Simon, S. I. & Walcheck, B. Cytoskeletal interactions regulate inducible L-selectin clustering. Am. J. Physiol. Cell Physiol. 289, C323–C332 (2005).
Liu, S. & Kiick, K. Architecture effects on L-selectin shedding induced by polypeptide-based multivalent ligands. Polym. Chem. 2, 1513–1522 (2011).
Li, W. et al. Large-scale topographical screen for investigation of physical neural-guidance cues. Sci. Rep. 5, 1–8 (2015).
Sun, Y. et al. Self-organizing circuit assembly through spatiotemporally coordinated neuronal migration within geometric constraints. PLoS ONE 6, e28156 (2011).
Umeshima, H. et al. Local traction force in the proximal leading process triggers nuclear translocation during neuronal migration. Neurosci. Res. 142, 38–48 (2019).
Martinez-Garay, I. Molecular mechanisms of cadherin function during cortical migration. Front. Cell Dev. Biol. 8, 964 (2020).
Shikanai, M., Nakajima, K. & Kawauchi, T. N-cadherin regulates radial glial fiber-dependent migration of cortical locomoting neurons. Commun. Integr. Biol. 4, 326 (2011).
Fujioka, T. et al. β1 integrin signaling promotes neuronal migration along vascular scaffolds in the post-stroke brain. EBioMedicine 16, 195–203 (2017).
Chronopoulos, A. et al. Syndecan-4 tunes cell mechanics by activating the kindlin–integrin–RhoA pathway. Nat. Mater. 19, 669–678 (2020).
Couchman, J. R. Syndecans: proteoglycan regulators of cell-surface microdomains? Nat. Rev. Mol. Cell Biol. 4, 926–938 (2003).
Manon-Jensen, T., Itoh, Y. & Couchman, J. R. Proteoglycans in health and disease: the multiple roles of syndecan shedding. FEBS J. 277, 3876–3889 (2010).
Sarker, F. A., Prior, V. G., Bax, S. & O’Neill, G. M. Forcing a growth factor response—tissue-stiffness modulation of integrin signaling and crosstalk with growth factor receptors. J. Cell Sci. 133, jcs242461 (2020).
Schwartz, M. A. & Ginsberg, M. H. Networks and crosstalk: integrin signalling spreads. Nat. Cell Biol. 4, E65–E68 (2002).
Nair, A., Chauhan, P., Saha, B. & Kubatzky, K. F. Conceptual evolution of cell signaling. Int. J. Mol. Sci. 20, 3292 (2019).
Alberts, B. et al. General Principles of Cell Communication. In Molecular Biology of the Cell (Garland Science, 2002).
Cai, D. et al. Mechanical feedback through E-cadherin promotes direction sensing during collective cell migration. Cell 157, 1146 (2014).
Dan, L., Jian, D., Na, L. & Xiaozhong, W. Crosstalk between EGFR and integrin affects invasion and proliferation of gastric cancer cell line, SGC7901. Onco Targets Ther. 5, 271 (2012).
Wee, P. & Wang, Z. Epidermal growth factor receptor cell proliferation signaling pathways. Cancers 9, 52 (2017).
Cherkouk, C. et al. Controlled immobilization of His-tagged proteins for protein–ligand interaction experiments using Ni2+-NTA layer on glass surfaces. Clin. Hemorheol. Microcirc. 61, 523–539 (2015).
Biswas, K. H. et al. E-cadherin junction formation involves an active kinetic nucleation process. Proc. Natl Acad. Sci. USA 112, 10932–10937 (2015).
Zhang, Y., Ge, C., Zhu, C. & Salaita, K. DNA-based digital tension probes reveal integrin forces during early cell adhesion. Nat. Commun. 5, 1–10 (2014).
Brockman, J. M. et al. Live-cell super-resolved PAINT imaging of piconewton cellular traction forces. Nat. Methods 17, 1018–1024 (2020).
WC, L., CH, Y., S, T. & JT, G. Supported membrane formation, characterization, functionalization, and patterning for application in biological science and technology. Curr. Protoc. Chem. Biol. 2, 235–269 (2010).
Pfaff, M. et al. Selective recognition of cyclic RGD peptides of NMR defined conformation by alpha IIb beta 3, alpha V beta 3, and alpha 5 beta 1 integrins. J. Biol. Chem. 269, 20233–20238 (1994).
Ferhan, A. R. et al. Solvent-assisted preparation of supported lipid bilayers. Nat. Protoc. 14, 2091–2118 (2019).
Ulman, A. Ultrathin Organic Films: From Langmuir-Blodgett to Self Assembly (Academic Press, 1991).
Harris, L. J. et al. The three-dimensional structure of an intact monoclonal antibody for canine lymphoma. Nature 360, 369–372 (1992).
Xia, S. et al. Nanoscale architecture of the cortical actin cytoskeleton in embryonic stem cells. Cell Rep. 28, 1251–1267.e7 (2019).
van de Linde, S. et al. Direct stochastic optical reconstruction microscopy with standard fluorescent probes. Nat. Protoc. 6, 991–1009 (2011).
Cai, Y. et al. Nonmuscle myosin IIA-dependent force inhibits cell spreading and drives F-actin flow. Biophys. J. 91, 3907–3920 (2006).
Rust, M., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Gauthier, N. C., Fardin, M. A., Roca-Cusachs, P. & Sheetz, M. P. Temporary increase in plasma membrane tension coordinates the activation of exocytosis and contraction during cell spreading. Proc. Natl Acad. Sci. USA 108, 14467–14472 (2011).
Hu, W., Sarveswaran, K., Lieberman, M. & Bernstein, G. H. Sub-10 nm electron beam lithography using cold development of poly(methylmethacrylate). J. Vac. Sci. Technol. B 22, 1711–1716 (2004).
Shroff, H., White, H. & Betzig, E. Photoactivated localization microscopy (PALM) of adhesion complexes. Curr. Protoc. Cell Biol. 58, 4–21 (2013).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S. T., Girirajan, T. P. K. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Giannone, G. et al. Periodic lamellipodial contractions correlate with rearward actin waves. Cell 116, 431–443 (2004).
Acknowledgements
We thank G. Grenci (Nano and Microfabrication Core, MBI, Singapore) for useful discussions for cleaning nanopatterned coverslips, D. Pitta de Araujo (Science Communication Core, MBI, Singapore) for helpful suggestions for illustrations and A. Wong (Science Communication Core, MBI, Singapore) for his comments on the manuscript text. We thank the Michael W. Davidson group, The Florida State University, Tallahassee, FL, USA for the mPA-GFP-paxillin DNA construct. This work was supported by intramural funds from the Mechanobiology Institute. K.J. was supported by Mechanobiology Institute Graduate Scholarship. M.P.S. received National Institutes of Health (NIH) grant support related to this project (no. RO1-GM113022). K.J. and P.K. acknowledge funding support from Ministry of Education Academic Research Fund Tier2 (MOE2019-T2-2-014). This work was performed, in part, at the Center for Nanoscale Materials, a U.S. Department of Energy Office of Science User Facility, and supported by the US Department of Energy, Office of Science, under contract no. DE-AC02-06CH11357.
Author information
Authors and Affiliations
Contributions
R.C. developed the functionalization of Ti substrates. K.J. and R.C. designed the experiments. K.J. performed experiments, analyzed and summarized the data, illustrations and figures. H.C. and X.Z. developed Ti nanofabrication processes. The manuscript was prepared with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Nicholas Kurniawan, Cas van der Putten, Carsten Schulte and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Changede, R. et al. Nat. Mater. 18, 1366–1375 (2019): https://doi.org/10.1038/s41563-019-0460-y
Extended data
Extended Data Fig. 1 TIRF calibration.
Fluorescence images of a cell expressing paxillin-EGFP imaged in an epifluorescence and a TIRF mode.
Supplementary information
Supplementary Information
Supplementary Method 1.
Rights and permissions
About this article
Cite this article
Jain, K., Kanchanawong, P., Sheetz, M.P. et al. Ligand functionalization of titanium nanopattern enables the analysis of cell–ligand interactions by super-resolution microscopy. Nat Protoc 17, 2275–2306 (2022). https://doi.org/10.1038/s41596-022-00717-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-022-00717-3
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.