Abstract
Elongation, splicing and polyadenylation are fundamental steps of transcription, and studying their coordination requires simultaneous monitoring of these dynamic processes on one transcript. We recently developed a full-length nascent RNA sequencing method in the model plant Arabidopsis that simultaneously detects RNA polymerase II position, splicing status, polyadenylation site and poly(A) tail length at genome-wide scale. This method allows calculation of the kinetics of cotranscriptional splicing and detects polyadenylated transcripts with unspliced introns retained at specific positions posttranscriptionally. Here we describe a detailed protocol for this method called FLEP-seq (full-length elongating and polyadenylated RNA sequencing) that is applicable to plants. Library production requires as little as one nanogram of nascent RNA (after rRNA/tRNA removal), and either Nanopore or PacBio platforms can be used for sequencing. We also provide a complete bioinformatic pipeline from raw data processing to downstream analysis. The minimum time required for FLEP-seq, including RNA extraction and library preparation, is 36 h. The subsequent long-read sequencing and initial data analysis ranges between 31 and 40 h, depending on the sequencing platform.
Similar content being viewed by others
Introduction
In eukaryotes, transcription by RNA polymerase II (Pol II) involves initiation, elongation and termination. The cotranscriptional processes, including mRNA capping, splicing, cleavage and polyadenylation, turn nascent RNAs into mature mRNAs that are eventually exported to the cytoplasm. Pre-mRNA splicing occurs mainly cotranscriptionally and is tightly coupled with Pol II elongation1,2,3,4,5,6,7,8,9. Many high-throughput sequencing methods have been developed to characterize nascent RNAs at the genome-wide scale quantitatively, and have revealed novel insights into transcriptional regulation by tracking Pol II position at nucleotide resolution as well as the status of splicing10. These methods can be grouped by their strategies for nascent RNA enrichment, including isolating the chromatin fraction (caRNA-seq11,12, Start-seq13, SMIT4), immunoprecipitation of Pol II (NET-seq14, mNET-seq15), nuclear run-on with labeled nucleotide (GRO-seq16, PRO-seq17,18), and metabolic labeling (4sU-seq19, TT-seq20, SLAM-seq21, TimeLapse-seq22). Recent progress in full-length RNA sequencing permitted by single-molecule long-read sequencing platforms (PacBio and Nanopore in particular) has furthered our knowledge on transcriptional regulation by enabling simultaneous detection of elongation and splicing on the same nascent RNA molecule. This has led to the development of methods such as long-read sequencing of nascent RNA4,5,6, nano-COP7,8 and POINT-nano9, which have revealed the divergent landscape of cotranscriptional splicing kinetics in yeast, mouse, fly and human4,5,6,7,8,9(Table 1).
Besides elongation and splicing, next-generation sequencing has also been adopted to characterize poly(A) sites (PAS-seq23, PAT-seq24, DRS25) and poly(A) tail length (Tail-seq26, mTail-seq27, PAL-seq28). Direct measurement on the full-range of poly(A) tail lengths has become feasible because of advances in long-read sequencing techniques and has led to the development of PacBio-based methods such as FLAM-seq29 and PAIso-seq30, or direct RNA sequencing by Nanopore in Arabidopsis31 (Table 1), human32 (Table 1), Caenorhabditis elegans33 and, more recently, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)34.
Compared with these existing single-molecule nascent RNA sequencing methods, our approach is uniquely positioned to study the coordination among elongation, splicing and polyadenylation all together (Table 1). Therefore, we named it FLEP-seq (full-length elongating and polyadenylated RNA sequencing). This difference turned out to be important as we found that many nascent transcripts in the model plant Arabidopsis are fully transcribed yet incompletely spliced, and their retained introns need to be removed before the transcripts can be released into the cytoplasm, a process we refer to as posttranscriptional splicing35. In addition to plants, polyadenylated nascent RNAs are also abundant in animal cells, including embryonic stem cells36, macrophages12, neuronal cells37 and differentiating germline cells38. These chromatin-tethered, intron-containing RNAs can be further spliced in response to various environmental and developmental signals. Thus, posttranscriptional splicing of polyadenylated nascent RNA could serve as a rapid-response layer of posttranscriptional gene regulation in eukaryotes.
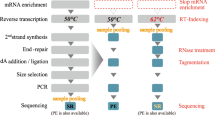
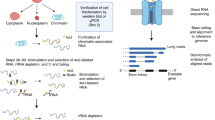
After enrichment of nascent RNAs by purifying the chromatin-associated fraction, to obtain the full-length ones, we ligate a universal DNA adapter to the 3′ end of nascent RNAs (including both the elongating and the polyadenylated fractions) and perform template-switching reverse transcription (RT) reaction using a primer that is complementary to the adapter. After PCR amplification of the full-length cDNAs, the final library can be sequenced on either Nanopore or PacBio (Fig. 1). Compared with the direct RNA sequencing approach, which requires 500–1,000 ng of poly(A)+ RNA7,8,31,32,33, the amplification step in FLEP-seq enabled us to produce comprehensive RNA profiles from as little as 1 ng of nascent RNA (after rRNA and tRNA depletion), thus extending its utility to situations where materials are limited. We also developed a complete bioinformatic pipeline for both Nanopore and PacBio data (Fig. 2). In particular, we developed a software named PolyAcaller to find polyadenylated transcripts and measure poly(A) tail length as described in the ‘Experimental design’ section. FLEP-seq can be applied to reveal cotranscriptional splicing kinetics, the dynamic coordination of splicing, and polyadenylation on multiintron genes, as well as the chromatin retention and the posttranscriptional splicing of introns that are potentially involved in developmental regulation or adaption to environmental signals.
Nascent RNA is purified from the chromatin fraction. After ribosomal RNA (rRNA) and tRNA depletion, a universal DNA adapter is ligated to the 3′ end of RNA. By performing RT with a custom primer complementary to the 3′ adapter and template switching, nascent RNAs are converted to full-length cDNA and amplified by PCR. The resulting double-stranded DNA library can be sequenced on either Nanopore or PacBio platforms. TSS, transcription start site. PAS, polyadenylation signal. RT, reverse transcription. TS, template switching.
Applications of the method
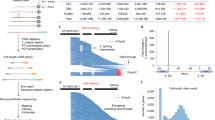
First, our method can accurately estimate the cotranscriptional splicing kinetics for introns genome-wide as previously demonstrated in Arabidopsis35, similar to long-read sequencing of nascent RNA and nano-COP4,5,6,7,8. Capturing the full-length elongating transcripts enables us to study the coordination between splicing and elongation by tracking the distance of Pol II positions relative to the 3′ splice site (3′ SS) of introns, which reveals the splicing kinetics (Fig. 3a).
FLEP-seq captures both the nascent elongating and polyadenylated transcripts, simultaneously detecting the Pol II position, splicing status, polyadenylation sites and poly(A) tail length on single molecules. a–e, With the splicing status, Pol II position and poly(A) information, users can analyze cotranscriptional splicing kinetics (a), posttranscriptional intron retention (b), different PAS isoforms (c), overall landscape of poly(A) tail length (d) and correlation between splicing and poly(A) tail length at gene level (e). For a,b and c, the red line represents the poly(A) tail. Only elongating transcripts are shown in a, while only polyadenylated transcripts are shown in b and c. For d and e, the median of poly(A) tail length and incompletely spliced ratio of genes with more than ten detected poly(A) reads were calculated. The genome-wide distribution of the median poly(A) tail length and the correlation between incompletely spliced ratio and median poly(A) tail length are shown in d and e, respectively. Figure adapted from ref. 35.
In addition, our method can simultaneously track the splicing status of multiple introns on a single transcript on both the elongating and polyadenylated fraction of RNAs. Analysis of the order of splicing among multiple introns could provide insight into the cooperative splicing of neighboring introns as reported in yeast4,5, fly, human5,6,7 and Arabidopsis35. Moreover, the analysis of the splicing status of polyadenylated transcripts can distinguish the cotranscriptionally spliced introns from the posttranscriptionally spliced ones, thus providing a more complex view of intron retention (Fig. 3b).
FLEP-seq can also be used to explore the genome-wide landscape of alternative polyadenylation (Fig. 3c) and poly(A) tail length (Fig. 3d). Furthermore, with the intron-containing yet polyadenylated transcripts in our library, our method can explore the crosstalk between posttranscriptional splicing, poly(A) site usage, and poly(A) tail length (Fig. 3e). Finally, our method also revealed many splicing intermediates, which can be used to estimate splicing reaction rates39,40.
By taking advantage of the comprehensive information on a full-length nascent RNA molecule, FLEP-seq can uniquely reveal the complexity among elongation, splicing, polyadenylation sites and poly(A) tail length. While we developed FLEP-seq to examine the dynamics of nascent RNA, it can be extended to characterize RNAs from other subcellular fractions, such as cytoplasm, nucleoplasm or immunoprecipitated RNA.
Experimental design
General considerations
We organize the procedure of FLEP-seq into four main modules: cell fractionation and RNA preparation (Steps 1–46), double-stranded cDNA library construction (Steps 47–90), long-read sequencing (Steps 91–93) and data analysis (Steps 94–116). Researchers can choose to use one or several of these modules to suit their needs. The first module describes the isolation of chromatin-bound factions (nascent RNA), RNA purification and ribosomal RNA depletion. In the section of cell fractionation, we provide a protocol for Arabidopsis. The following three modules can be generally applied to RNA from any other cell fractions (e.g., cytoplasm, nucleoplasm or immunoprecipitated RNA) and other types of linear RNA of interest (e.g., lncRNA, rRNA, RNA intermediates), not limited to nascent RNA.
Cell fractionation, RNA purification and rRNA depletion
The most critical issue for cell fractionation is the purity of subcellular fractions and the integrity of RNA. To minimize RNA degradation, all buffers, consumables and equipment should be precooled at 4 °C, and steps need to be performed on ice if required. tRNA, RNase inhibitor and a protease inhibitor cocktail are used to prevent the degradation of RNA and proteins at different steps. The cell fractionation protocol we present here can be used for up to 3–4 g of Arabidopsis seedlings. For efficient cell lysis, one should avoid using too much starting material. If necessary, the user can scale up all the buffers accordingly for a large amount of starting material. To monitor the purity of the subcellular fractions, we recommend performing a western blot35 or qRT-PCR41 with protein or RNA from different compartments (Fig. 4). We also recommend running the RNA from all subcellular fractions on an agarose gel to roughly evaluate the RNA integrity and the fractionation efficiency (Fig. 5). To minimize batch-to-batch variation, we recommend processing two samples simultaneously (i.e., wild type and mutant or treatment and control). However, it is not recommended to process more than two samples at a time to avoid prolonged incubation during the subsequent cell fractionation and RNA extraction steps. The purified RNA samples can be safely stored at −80 °C for further processing.
Protein is prepared from total (Total), cytoplasm (Cyto), nucleoplasm (NP1) and chromatin fractions (Cb) and detected with antibodies specific for cytoplasm (UGPase, 51.6 kDa) and chromatin (Histone H3, 17 KDa). Figure adapted from ref. 35.
To assess RNA quality, 5 μl total (Total), cytoplasmic (Cyto), two nucleoplasmic fractions (NP1 and NP2) and chromatin-bound (Cb) fraction were run on 1% (wt/vol) agarose gel with DM2000 Ladder (100–2,000 bp) after RNA extraction. A large amount of tRNA (~100 bp) is present in all fractions as indicated in the figure.
The efficiency of rRNA depletion is particularly important for the quality of the sequencing data. The rRNA is extremely abundant in eukaryotic cells, representing 80–90% RNA in most RNA samples. Based on these considerations, if there are too many rRNA-derived reads in the library, the rRNA depletion should be optimized by adding more input rRNA probe or mixing well when incubating with probe or biotin beads, especially for the cytoplasmic RNA sample or ribosome-enriched fractions.
3′ adapter ligation, RT and cDNA amplification
To capture both the polyadenylated nascent RNA and the elongating transcripts, we ligated a 5′ adenylated, 3′ blocked oligodeoxynucleotide to the 3′ end of RNA after rRNA depletion as the primer binding site of RT. We recommend incubating at 16 °C overnight to improve the ligation efficiency of long RNA substrates (>1 kb). RNase inhibitor is required until the cDNA synthesis. For cDNA synthesis and double-stranded cDNA library amplification, the protocol of PacBio Sequel System to construct a non-size-selected high-quality library is well suited for full-length sequencing. The Clontech SMARTer PCR cDNA Synthesis Kit is used to generate first-strand cDNA. In particular, we use a custom RT primer specific to the 3′ adapter to replace the original primer provided in the kit. After cDNA synthesis, the PCR cycle number for amplification needs to be optimized before large-scale amplification of cDNA. Meanwhile, the quality of library products, including the integrity and the amount of cDNA, can be evaluated by electrophoresis (Fig. 6). After the large-scale PCR with an optimal number of cycles, the double-stranded cDNA product is purified twice using 1× AMPure PB beads.
To demonstrate cDNA distribution, PCR aliquots with various cycle numbers are collected during PCR optimization and run on 1% (wt/vol) agarose gel with DM2000 ladder (L1, 100–2,000 bp) and 1 kb DNA ladder (L2, 1–10 kb). The numbers above the lanes indicate cycle number. In this example, with 1 μl input cDNA, 11–12 cycles are the optimal number of PCR cycles for large-scale amplification, although the smear distribution from 12–16 cycles looks similar.
Full-length library preparation and sequencing
Double-stranded cDNA can be directly used to prepare libraries for both Nanopore and PacBio sequencing. Prior to library construction, Qubit Fluorometer and Agilent 2100 Bioanalyzer are required to monitor the concentration and size distribution of the library (Fig. 7). It is critical to evaluate the quality of DNA fragments and the molar number of DNA molecules of the sample. Too little or too much DNA, or poor quality, can dramatically decrease the library quality and data output. To generate the Oxford Nanopore DNA library, we follow the instructions of DNA Ligation Sequencing Kit (SQK-LSK109) and sequence the library on a MinION for 36 h. The MinION flowcell yields ~14 million reads, and the average read length is ~900 nt (Fig. 8). For the PacBio platform, the library is prepared according to the protocol of SMRTbell Template Prep Kit 1.0-SPv3, and sequenced on a PacBio Sequel II System. The Sequel II SMRT Cell yields ~2 million reads, and the average read length is ~1,200 nt.
Bioanalyzer results after large-scale PCR and DNA library clean-up. a, Sample 1 is an example of a qualified library with high yield and correct size distribution. The majority of the amplified products should be longer than 1 kb. b, Two examples of poor quality. The yield of sample 4 is low. Sample 7 has a broader size distribution with a high proportion of low-molecular-weight products, indicating that the enrichment for long DNA fragments is lost.
a, Total yield of the number of raw reads per hour through the progression of the run. b, Statistics of read length over time throughout the run. Figure adapted from ref. 35.
Data processing and analysis
The data analysis workflow includes several main steps: (a) converting raw signal to DNA sequences (Nanopore) or generating consensus sequences (PacBio), (b) mapping to a reference genome, (c) identifying adapters, (d) finding poly(A) tails, (e) extracting the intron splicing/retention information and the 3′ end position, (f) calculating cotranscriptional splicing kinetics using elongating transcripts, and (g) calculating the incompletely spliced ratio after polyadenylation using poly(A)+ transcripts (Fig. 2).
The basecalling and genome mapping are performed using popular software as described below. As the cDNA can be sequenced from either the 5′ end or the 3′ end, the 3′ adapter is searched on each read from both sides using the local alignment program BLASTn with a small seed size to determine the strand direction of the original RNA, which is key to distinguishing the sense and antisense transcripts. For PacBio data, their official adapter identification software ‘lima’ is integrated in our workflow.
Because FLEP-seq captures both elongating transcripts without poly(A) tails and polyadenylated transcripts, a core component of the analysis workflow is to determine whether a transcript contains a poly(A) tail. To distinguish poly(A) tails from potential short A-stretches in mRNAs, we only use the unmapped portion of the read to perform poly(A) tail analysis. This portion, defined as the sequence between the region mapped to the genome and the 3′ adapter sequence, plus the last ten bases of the region mapping to the genome, and the first five bases of the adapter region, were used as input for our algorithm to find poly(A) tails, and the minimum length for a candidate poly(A) tail was set at 15 nt. While we use a dynamic programming algorithm to find potential poly(A) tails from highly accurate PacBio reads, this problem becomes challenging for nanopore data, as long polymer regions generate a flat electrical signal, and thus cannot be accurately sequenced, meaning that a long poly(A) tail may be recognized as one or a few A’s. Several software tools have been developed to infer the poly(A) length based on the duration time of the measured signal, such as Nanopolish32 and Tailfindr42. However, Nanopolish is designed for data produced from direct RNA sequencing; thus, it is not suitable for a cDNA based method. Tailfindr can indeed be used to analyze poly(A) tail length from regular cDNA Nanopore data. However, it is designed to work with sequences that are known to have a poly(A) tail, such as poly(A)-enriched mRNAs, and it is not optimized to deal with data that contain a large portion of nonpolyadenylated elongating transcripts, such as FLEP-seq data. As a result, Tailfindr has a high false-positive rate when analyzing transcripts that should not have a poly(A) tail, such as the elongating ones (Fig. 9a). To measure the performance of Tailfindr on FLEP-seq data, we separated full-length clean reads into two groups based on the position of their 3′ ends: the reads in group I are located upstream of the last exon and therefore belong to the elongating transcripts that should have little poly(A) signal, while the reads in group II extend downstream of the last exon and thus should include elongating transcripts and polyadenylated transcripts. Tailfindr predicts that 20% of the group I reads contain poly(A) tail, suggesting a high false-positive rate.
a, The percentage of FLEP-seq reads with and without PolyA tail (≥15 nt) identified by Tailfindr or PolyAcaller. The full-length clean reads were classed into two groups. The 3′ end of the type I reads were located upstream of the last exon, and therefore most of them should be elongating transcripts and have no poly(A) tail. The others were type II and included elongating transcripts and polyadenylated transcripts. CB1 and CB2 are two biological replicates of FLEP-seq data35. b, The distribution of poly(A) tail length calculated by PolyAcaller or Tailfindr on Nanopore DNA sequencing data of PCR-amplified eGFP with different (10, 30, 40, 60, 100, 150 nt, indicated by dashed lines) poly(A) lengths35. c, The median poly(A) tail lengths of genes measured by Nanopore are highly consistent with those measured by PacBio using FLEP-seq.
To address this problem, we developed a method called PolyAcaller to calculate the poly(A) tail length. Compared with the 20% from Tailfindr, PolyAcaller estimated that only 2% of the reads in group I have a poly(A) tail, while maintaining a similar performance on group II reads with Tailfindr (Fig. 9a). To further confirm the accuracy of PolyAcaller, we used it to analyze the same Nanopore data of synthetic DNA with different poly(A) tail lengths that were used to benchmark Tailfindr42, and found that the results from PolyAcaller match well with the expected poly(A) tail length, and are also consistent with the results obtained using Tailfindr (Fig. 9b). In addition, we also used FLEP-seq PacBio data to calculate the median poly(A) tail of genes, and the results are highly consistent with those calculated by PolyAcaller from Nanopore data (Person’s r 0.89, Fig. 9c), suggesting that PolyAcaller is highly reliable.
Lastly, sequencing errors of bases at the 3′ end in Nanopore reads may result in inaccurate alignment and, therefore, inaccurate determination of the Pol-II position on the transcript. Thus, we set up a scoring system for measuring the accuracy of 3′ end sequences. We calculated the distance from the start position of the 3′ adapter (for elongating transcripts) or the poly(A) tail (for polyadenylated transcripts) to the end position of the genome mapping region as a 3′ alignment score. We marked reads with a 3′ alignment score >5 as ‘low_accuracy_3end_mapped’ reads in the final output generated by the FLEP-seq analysis pipeline (Step 102). We usually remove ‘low_accuracy_3end_mapped’ reads from analysis concerning the 3′ end of transcripts, such as splicing kinetics and poly(A) site determination. For FLEP-seq PacBio data, the sequence identity is 99.5%; thus, we did not calculate the 3′ alignment score. Previous studies have also reported using RNA Spike-In sequences to estimate the base error rate and Illumina poly(A) RNA-seq data to correct errors in ONT (Oxford Nanopore Technologies) long reads using proofreading software31.
Limitations
Because our method does not include a Pol II immunoprecipitation step, we cannot assume that the 3′ end of the read corresponds to the last nucleotide transcribed by Pol II. Indeed, we observed a high proportion of splicing intermediates with their 3′ end precisely located at the 5′ splicing site of intron35. The precision of the 3′ ends at the 5′ SS can be used to evaluate the degree of RNA fragmentation during experimental manipulation. In addition, incomplete RT can also affect the integrity of 5′ end of the RNA and lead to truncated non-full-length reads. While the use of RT and PCR amplification allows our technique to be used on low amounts of RNA, it is at the cost of other information, such as the various forms of RNA modifications that can be detected by the direct RNA sequencing approach (Table 1).
Materials
Biological materials
-
Arabidopsis seeds from natural accession Columbia (Col-0, ABRC stock no. CS60000), incubated at 4 °C for 2 d stratification and grown on 1/2 Murashige and Skoog plates vertically at 22 °C (16 h light–8 h dark) for 12 d before collection
Reagents
-
Murashige and Skoog basal medium (PhytoTechnology Laboratories, cat. no. M519)
-
Diethyl pyrocarbonate (Sigma-Aldrich, cat. no. D5758)
Caution
Diethyl pyrocarbonate is toxic and sensitive to moisture; storage at 2–8 °C is recommended.
-
Liquid nitrogen
Caution
Liquid nitrogen can cause freeze burns. When handling, please wear personal protective equipment, such as a laboratory coat, cold weather gloves and protective eyewear.
-
Sucrose, molecular biology grade (Sigma-Aldrich, cat. no. S0389)
-
Ficoll PM 400 Type 400 (Sigma-Aldrich, cat. no. F4375)
-
Dextran-40 (Sangon Biotech, cat. no. A600374)
-
Triton X-100, molecular biology grade (Sigma-Aldrich, cat. no. T8787)
-
MgCl2, molecular biology grade, 1 M (Sigma-Aldrich, cat. no. M1028)
-
HEPES, RNase-free, 1 M, pH 7.0–7.6; (Sigma-Aldrich, cat. no. H0887)
-
DTT, powder (Thermo Fisher Scientific, cat. no. R0861)
Caution
DTT may cause respiratory irritation, eye damage and severe skin burns. Avoid breathing, and use only in a well-ventilated area. When handling this reagent, wear an appropriate laboratory coat, gloves and protective eyewear.
-
EDTA-free protease inhibitor cocktail (Roche, cat. no. 4693132001)
-
Baker’s yeast tRNA (Roche, cat. no. 10109495001)
-
Glycerol, molecular biology grade (Sigma-Aldrich, cat. no. G5516)
-
EDTA, RNase-free, 0.5 M, pH 8.0 (Life Technologies, cat. no. AM9260G)
Caution
EDTA may cause respiratory tract, eye damage and severe skin burns. Avoid breathing, and use only in a well-ventilated area. When handling this reagent, wear an appropriate laboratory coat, gloves and protective eyewear.
-
Tris-HCl, RNase-free, 1 M, pH 7.5 (Thermo Fisher Scientific, cat. no. 15567027)
-
TE buffer, pH 7.4 (Sigma-Aldrich, cat. no. 93302-100ML)
-
NaCl, RNase-free, 5 M (Sigma-Aldrich, cat. no. S3014)
-
Histone H3 antibody, 1:5,000 dilution (ABclonal, cat. no. A2348)
-
RNaseOUT recombinant ribonuclease inhibitor, 40 U/μl (Life Technologies, cat. no.10777019)
-
Urea, molecular biology grade (Sigma-Aldrich, cat. no. U5378)
-
Tween 20, molecular biology grade (Sigma-Aldrich, cat. no. P9416)
-
TRIzol LS Reagent (Thermo Fisher Scientific, cat. no. 10296010)
Caution
TRIzol, whose main ingredients are phenol and guanidinium thiocyanate, is light sensitive. Phenol is toxic and corrosive, while guanidinium thiocyanate is irritating. When handling this reagent, wear an appropriate laboratory coat, gloves and protective eyewear.
-
TRIzol reagent (Thermo Fisher Scientific, cat. no. 15596026)
Caution
TRIzol, whose main ingredients are phenol and guanidinium thiocyanate, is light sensitive. Phenol is toxic and corrosive, while guanidinium thiocyanate is irritating. When handling this reagent, wear an appropriate laboratory coat, gloves and protective eyewear.
-
Chloroform, molecular biology grade (Sigma-Aldrich, cat. no. 288306)
Caution
Chloroform is volatile, toxic and irritating. When handling chloroform-containing solutions, wear an appropriate laboratory coat, gloves and protective eyewear. Please dispose of chloroform waste according to institutional regulations.
-
Ethanol, 200 proof, molecular biology grade (Sigma-Aldrich, cat. no. E7023)
Caution
Ethanol is a volatile, flammable and colorless liquid. Keep away from heat, sparks and open flame. Please dispose of ethanol waste according to institutional regulations.
-
Direct-zol RNA Miniprep Plus (Zymo, cat. no. R2070)
-
RNA Clean & Concentrator-5 (Zymo, cat. no. R1015)
-
Agarose (Biowest, cat. no. BY-R0100)
-
DNA gel loading dye, 6× (Thermo Fisher Scientific, cat. no. R0611)
-
DM 2000 DNA marker (CoWin Biosciences, cat. no. CW0632S)
-
1 kb DNA ladder (TransGen Biotech cat. no. BM201-02)
-
SDS-PAGE loading buffer, reducing, 5× (CoWin Biosciences, cat. no. CW0027S)
-
Ultra GelRed, 10,000× (Vazyme, cat. no. GR501)
-
Qubit RNA HS Assay Kit (Thermo Fisher Scientific, cat. no. Q32852)
-
Qubit dsDNA HS Assay Kit (Thermo Fisher Scientific, cat. no. Q32851)
-
Agilent DNA 12000 reagents (Agilent Technologies, cat. no. 50671508)
-
RiboMinus Plant Kit for RNA-Seq (Thermo Fisher Scientific, cat. no. A1083808) or riboPOOL (siTOOLs Biotech) with hydrophilic streptavidin magnetic beads (NEB, cat. no. S1421S)
-
T4 RNA Ligase 2, truncated KQ (NEB, cat. no. M0373S)
-
50% (wt/vol) PEG8000, supplied with T4 RNA Ligase 2, truncated KQ (NEB, cat. no. M0373S)
-
Universal miRNA cloning linker (NEB, cat. no. S1315S; 5′ rAppCTGTAGGCACCATCAAT–NH2 3′)
-
3′ cDNA RT primer (Custom primer, 5′-AAGCAGTGGTATCAACGCAGAGTACATT GATGGTGCCTACAG-3′)
-
SMARTer PCR cDNA Synthesis Kit (TaKaRa, cat. no. 634926)
-
PrimerSTAR GXL DNA polymerase (TaKaRa, cat. no. R050B)
-
Nuclease-free water, supplied with PrimerSTAR GXL DNA Polymerase (TaKaRa, cat. no. R050B)
-
AMPure PB beads (Pacific Biosciences, cat. no. 100-265-900)
-
NEBNext FFPE DNA Repair Mix (NEB, cat. no. M6630S)
-
NEBNext Ultra II End Repair/dA-Tailing Module (NEB, cat. no. E7546S)
-
NEBNext Quick Ligation Module (NEB, cat. no. E6056S)
-
Ligation Sequencing Kit 1D, PM (Nanopore, cat. no. SQK-LSK109)
-
Flow Cell Wash Kit (Nanopore, cat. no. EXP-WSH003)
-
SMRTbell Template Prep Kit 1.0-SPv3 (PacBio, cat. no. 100–991-900)
-
N,N,N′,N′-Tetramethylethylenediamine (Sigma-Aldrich, cat. no. T411019-100ML)
Caution
Handle in a laboratory fume hood.
-
Acrylamide/bis-acrylamide, 30% (wt/vol) solution (Sigma-Aldrich, cat. no. A3574-100ML)
Caution
This reagent is toxic; wear an appropriate laboratory coat and gloves, avoid skin contact and work under a fume hood. Store at 4 °C.
-
Ammonium persulfate (Sigma-Aldrich, cat. no. A3678-100G)
Caution
Prepare fresh and avoid contact and inhalation.
-
Trizma base (Sigma-Aldrich, cat. no. 93362-1KG)
-
SDS solution, 10% (wt/vol) (Thermo Fisher Scientific, cat. no. 15553027)
-
Milk powder (local supermarket)
-
Glycine (Sigma-Aldrich, cat. no. 50046-250G)
-
2-Mercaptoethanol (Sigma-Aldrich, cat. no. M6250-100ML)
-
Bromophenol Blue (Sigma-Aldrich, cat. no. B0126-25G)
-
Methanol (Sigma-Aldrich, cat. no. 34860-1L-R)
Caution
Methanol is volatile and toxic. When handling methanol, work under a fume hood, and wear an appropriate laboratory coat, gloves and protective eyewear. Please dispose of methanol waste according to institutional regulations.
-
KH2PO4 (Sigma-Aldrich, cat. no. P9791-500G)
-
Na2HPO4 (Sigma-Aldrich, cat. no. S3264-500G)
-
Prestained protein ladder, 10–180 kDa (Thermo Fisher Scientific, cat. no. 26616)
-
Antibodies against UGPase (Agrisera, cat. no. AS05086; RRID: AB_1031827)
-
Antibodies against histone H3 (ABclonal, cat. no. A2348; RRID: AB_2631273)
-
Anti-rabbit IgG (Sigma-Aldrich, cat. no. A0545-1ML; RRID: AB_257896)
Equipment
-
Miracloth (Millipore, cat. no. 475855-1R)
-
RNase/DNase-free thin wall PCR tubes with flat cap, 0.2 ml (Axygen, cat. no. PCR-02-C)
-
RNase/DNase-free microcentrifuge tubes, 1.5 ml (Axygen, cat. no. MCT-150-C)
-
DNA LoBind microcentrifuge tubes 1.5 ml (Eppendorf, cat. no. 022431021)
-
Pipette tips (Kirgen, cat. no. KG1300, KG1200, KG1100)
-
Refrigerated microcentrifuge 5424R for 1.5 ml microcentrifuge tubes (Eppendorf, cat. no. 5404F1621754)
-
Benchtop centrifuge for 50 ml conical tubes (Thermo Fisher Scientific, cat. no. ST 16R)
-
Mortar and pestle, 100 mm D × 130 ml (GSC International, Inc., cat. no. 4-13021-12)
-
Vortexer (IKA, cat. no. V2S025)
-
Rotator mixer (Kylin-Bell, cat. no. QB328)
-
Magnetic rack (Thermo Fisher Scientific, cat. no. 12321D)
-
3 × 32-well PCR system (Thermo Fisher Scientific, cat. no. 4483636)
-
Mixing and temperature control instruments (Eppendorf, cat. no. ThermoMixer C)
-
NanoDrop One microvolume UV-Vis spectrophotometer (Thermo Fisher Scientific, cat. no. ND-ONE)
-
Qubit 3.0 fluorometer (Life Technologies, cat. no. Q33216)
-
Agilent 2100 Bioanalyzer (Agilent Technologies, cat. no. G2939A)
-
Gel imaging system (Tanon, cat. no. Tanon 3500R)
-
Extra thick blot filter paper (BIO-RAD, cat. no. 1703966)
-
PVDF membrane (Millipore, cat. no. ISEQ00005)
-
Mini-PROTEAN vertical electrophoresis cell, with PowerPac basic power supply (BIO-RAD, cat. no. 1658025FC)
-
Trans-Blot Turbo system (BIO-RAD, cat. no. 1704150EDU)
-
Wide Mini ReadySub-Cell GT Horizontal electrophoresis system (BIO-RAD, cat. no. 1640304)
-
MinION nanopore device (Oxford Nanopore Technologies)
-
Flow Cells -R.9.4.1, Nanopore, FLO-MIN106D (Oxford Nanopore Technologies)
-
MinION computer requirements:
-
Operation system: Windows (7, 8, 10); macOS (Sierra, High Sierra, Mojave); Linux (Ubuntu 16.04 or 18.04)
-
Memory: 16 GB random-access memory
-
CPU: i7 or Xeon with 4+ cores
-
Storage: 1 TB internal SSD
-
Ports: USB3
-
Software
-
MinKNOW (MinION software) (https://community.nanoporetech.com/downloads)
-
Guppy v4.0.11 or above (https://community.nanoporetech.com/downloads)
-
Minimap243 (https://github.com/lh3/minimap2)
-
SAMtools44 (http://www.htslib.org/)
-
BLAST+45 (https://ftp.ncbi.nlm.nih.gov/blast/executables/blast+/LATEST/)
-
Python 3.7 or above, and the following packages:
-
ont_fast5_api (https://github.com/nanoporetech/ont_fast5_api)
-
pandas (https://pandas.pydata.org/)
-
NumPy (https://numpy.org/)
-
Matplotlib (https://matplotlib.org/)
-
Joblib (https://github.com/joblib/joblib)
-
R 3.5.2 or above, and the following packages:
-
Tidyverse (https://www.tidyverse.org/)
-
optparse (https://cran.r-project.org/web/packages/optparse/index.html)
-
Reagent setup
Critical
Take care to avoid RNase contamination when preparing and handling the following reagents.
Honda buffer
Contains 0.44 M sucrose, 1.25% (wt/vol) Ficoll, 2.5% (wt/vol) Dextran T40, 20 mM HEPES (pH 7.4), 10 mM MgCl2, 0.5% (vol/vol) Triton X-100, 1 mM DTT, 1× protease inhibitor cocktail, and 100 ng/μl tRNA. Prepare Honda buffer in a 50 ml RNase/DNase-free tube; one tube per sample. For 50 ml Honda Buffer, mix 7.53 g of sucrose, 0.625 g of Ficoll, 1.25 g of Dextran T40, 1 ml of 1 M HEPES (pH 7.4), 500 μl of 1 M MgCl2, 1.25 ml of 20% (vol/vol) Triton X-100, 50 μl of 1 M DTT, 500 μl of 100× protease inhibitor cocktail and 500 μl of 10 μg/μl tRNA. Prepare this solution with RNase-free reagents. The buffer can be stored at 2–8 °C for up to 1 week. Add tRNA, protease inhibitor cocktail and DTT just before use.
Nucleic resuspension buffer
Contains 50% (vol/vol) glycerol, 0.5 mM EDTA (pH 8.0), 1 mM DTT, 25 mM Tris-HCl (pH 7.5), 100 mM NaCl, 0.4 U/μl RNaseOUT recombinant ribonuclease inhibitor and 100 ng/μl tRNA. For 10 ml nucleic resuspension buffer, mix 5 ml of glycerol, 10 μl of 0.5 M EDTA (pH 8.0), 10 μl of 1 M DTT, 250 μl of 1 M Tris-HCl (pH 7.5), 200 μl of 5 M NaCl, 100 μl of 40 U/μl RNaseOUT recombinant ribonuclease inhibitor and 100 μl of tRNA (10 μg/μl). Prepare this solution with RNase-free reagents. The solution without DTT, tRNA and protease inhibitor cocktail can be stored at −20 °C for months. Add tRNA, protease inhibitor cocktail and DTT just before use.
Washing buffer
Contains 25 mM Tris-HCl (pH 7.5), 300 mM NaCl, 1 M urea, 0.5 mM EDTA (pH 8.0), 1 mM DTT, 1% (vol/vol) Tween 20, 0.4 U/μl RNaseOUT recombinant ribonuclease inhibitor and 200 ng/μl tRNA. For 10 ml washing buffer, mix 250 μl of 1 M Tris-HCl (pH 7.5), 600 μl of 5 M NaCl, 0.6 g urea, 10 μl of 0.5 M EDTA (pH 8.0), 10 μl of 1 M DTT, 100 μl of 40 U/μl RNaseOUT recombinant ribonuclease inhibitor and 100 μl of tRNA (10 μg/μl). Prepare this solution with RNase-free reagents. The solution without DTT, tRNA and protease inhibitor cocktail can be stored at −20 °C for months. Add tRNA, protease inhibitor cocktail and DTT just before use.
tRNA solution
To prepare 10 μg/μl tRNA solution, dissolve 100 mg of yeast tRNA powder in 10 ml RNase-free H2O. Mix by vortexing until fully dissolved, and store at −80 °C for months. Because tRNA is unstable, prepare into aliquots to avoid multiple freeze–thaw cycles.
Protease inhibitor mix (100×)
To prepare protease inhibitor mix (100×), dissolve one tablet of protease inhibitors in 500 μl RNase-free H2O. Mix by vortexing until fully dissolved, and store at −20 °C for months.
DTT solution
To prepare 1 M DTT solution, dissolve 1.55 g of DTT powder in 10 ml RNase-free H2O. Mix by vortexing until fully dissolved. Because DTT in solution is unstable, prepare into aliquots to avoid multiple freeze–thaw cycles and use a fresh solution as much as possible. The stock solution can be stored at −20 °C up to 3 months.
3′ cDNA RT primer (5′-AAGCAGTGGTATCAACGCAGAGTACATTGATGGTGCCCTACAG-3′)
Prepare a 100 μM stock solution by dissolving in TE buffer (pH 7.4). The stock solution can be stored at −20 °C for at least several years. Prepare a 12 μM working solution by diluting with 10 mM Tris pH 7.5, and prepare into 20 μl aliquots. Aliquots can be stored at −20 °C for several years.
Procedure
Cell fractionation
Timing 2 h
Critical
Cell fractionation has been optimized for 12-d-old Arabidopsis seedlings. Protocol optimization may be required if using a different type of starting material.
Critical
RNase-free reagents and equipment (including RNase/DNase-free 1.5 ml and 50 ml tubes) are required. Cool all buffers on ice before use.
-
1
Prepare Honda buffer in a 50 ml RNase/DNase-free tube. Thaw the 10 μg/μl tRNA, 100× protease inhibitor cocktail and 1 M DTT on ice, and add to Honda buffer just before use.
-
2
Precool the centrifuge for 50 ml and 1.5 ml tubes to 4 °C.
-
3
Grind 3–4 g of seedlings into a fine powder with liquid nitrogen using mortar and pestle, and transfer to an ice-cold 50 ml RNase/DNase-free tube with 10 ml Honda buffer.
-
4
After homogenization by gentle vortexing and inverting, filter the homogenate into a new 50 ml RNase/DNase-free tube through two layers of Miracloth.
-
5
Add another 10 ml Honda buffer to the filter to wash the remaining plant material.
-
6
Collect 300 μl filtrate as the total RNA fraction (Total), and add 900 μl TRIzol LS reagent. Vortex to mix thoroughly for 10 s, and put at room temperature (22 °C).
Critical step
If you want to check the purity of cell fractions, retain 50 μl filtrate at 4 °C for western blot analysis as ‘total fraction’.
-
7
Centrifuge the 50 ml tube at 4 °C, 2,000g, for 5 min.
-
8
For RNA extraction, transfer 400 μl supernatant into a new 1.5 ml RNase/DNase-free microcentrifuge tube as the cytoplasmic fraction (Cyto). Discard the remainder of supernatant.
Critical step
If you want to check the purity of cell fractions, retain 50 μl supernatant at 4 °C for western blot as ‘cytoplasmic fraction’. Optionally, to minimize the contamination of nuclei or chromatin in the cytoplasmic fraction, high-speed centrifugation (14,000g, 10 min) can be used to pellet the nuclei fraction.
-
9
Add 4 μl RNase inhibitor to the 400 μl supernatant from the previous step, mix well and centrifuge at 4 °C, 14,000g, for 5 min.
-
10
Transfer the supernatant from the previous step to a new 1.5 ml RNase/DNase-free microcentrifuge tube, and centrifuge at 4 °C, 14,000g, for 5 min.
-
11
Transfer 300 μl supernatant from the previous step to a new 1.5 ml RNase/DNase-free microcentrifuge tube, and add 900 μl TRIzol LS reagent. Vortex to mix thoroughly for 10 s, and put at room temperature.
-
12
Resuspend the nuclear pellet from Step 7 with 15 ml Honda buffer. Vortex to resuspend.
-
13
Centrifuge at 4 °C, 2,000g, for 5 min.
-
14
Remove all the supernatant and discard it. Wash the pellet with 10 ml Honda buffer. Vortex to resuspend the pellet.
-
15
Centrifuge at 4 °C, 2,000g, for 5 min.
-
16
Remove all the supernatant and discard it. Resuspend the pellet with 1 ml Honda buffer supplemented with 10 μl RNase inhibitor.
-
17
Transfer the nuclei to a 1.5 ml RNase/DNase-free microcentrifuge tube, and centrifuge at 4 °C, 8,000g, for 1 min.
-
18
Remove the supernatant thoroughly and discard.
-
19
Spin briefly (10 s) to collect the droplets on the walls of tube and remove the supernatant completely.
Critical step
It is important to completely remove the supernatant containing cytoplasmic RNA.
-
20
Weigh the nuclei pellet.
-
21
Add one volume of nuclei resuspension buffer to the pellet according to the weight of the pellet (e.g., 100 mg = 100 μl). Stir the pellet with a pipette tip to mix.
Critical step
Do not mix by pipetting. Because nuclei are sticky, pipetting can cause undesirable sample loss when using it to resuspend nuclei. However, it is important to mix the buffer thoroughly to ensure sufficient lysis.
-
22
Cut 2–3 mm off the tip of a 200 μl pipette tip. Use it to add two volumes (based on the pellet’s weight) of washing buffer to the mixture, and wash the pellet by pipetting up and down 30 times.
Critical step
It is important to cut the end of the pipette tip to preserve the integrity of RNA molecules.
-
23
Centrifuge at 4 °C, 8,000g, for 1 min.
-
24
Transfer the supernatant to a new 1.5 ml RNase/DNase-free microcentrifuge tube as the nucleoplasmic fraction 1 (NP1) without transferring any of the pellet; then remove the remaining supernatant completely and discard.
Critical step
If you want to check the purity of cell fractions, retain 50 μl supernatant for western blot as ‘nucleoplasmic fraction’.
-
25
Add three volumes of TRIzol LS reagent to the supernatant NP1. Vortex to mix thoroughly for 10 s, and put at room temperature.
-
26
For the second wash, resuspend the pellet from Step 24 with one volume of nuclei resuspension buffer. Add one volume of washing buffer, and wash the chromatin by pipetting up and down seven to ten times.
Critical step
It is important to cut the end of the pipette tip to preserve the integrity of RNA molecules.
-
27
Centrifuge at 4 °C, 8,000g, for 1 min.
-
28
Transfer the supernatant to a new 1.5 ml RNase/DNase-free microcentrifuge tube as the nucleoplasmic fraction 2 (NP2) without transferring any of the pellet. Then remove the supernatant thoroughly and discard.
-
29
Add three volumes of TRIzol LS reagent to the supernatant NP2. Vortex to mix thoroughly for 10 s, and put at room temperature.
-
30
The pellet from Step 29 is retained as the chromatin-bound fraction (Cb). Add 1 ml TRIzol to it, vortex to mix thoroughly and keep at room temperature for 10 min.
Critical step
It is important to completely resuspend the pellet in TRIzol at this step to enable complete RNA extraction. If you want to check the purity of cell fractions, retain a small fraction of the chromatin pellet for western blot analysis.
-
31
(Optional) The purity of total (from Step 6), cytoplasmic (from Step 8), nucleoplasmic (from Step 24) and chromatin fractionation (from Step 30) can be evaluated by western blot according to our previous report35 (Fig. 4). For Arabidopsis, antibodies to UGPase (1:1,500) and histone H3 (1:5,000) were used as cytoplasmic and chromatin fraction-specific markers, respectively.
RNA purification from fractions
Timing 2 h
-
32
Add 0.2× volume of chloroform to the RNA tube from Steps 6, 11, 25, 29 and 30, vortex for 10 s, and incubate at room temperature for 5 min.
-
33
Centrifuge at 4 °C, 14,000g, for 10 min.
-
34
Transfer the supernatant to a new 1.5 ml RNase/DNase-free microcentrifuge tube, and add an equal volume of 100% (vol/vol) ethanol. Invert the tube gently to mix.
-
35
Purify the RNA using Direct-zol RNA Miniprep Plus kit with on-column digestion of DNA according to the manufacturer’s instructions.
-
36
Elute RNA into a 1.5 ml RNase/DNase-free microcentrifuge tube with 100 μl RNase-free water.
-
37
Take 1 μl RNA to quantify the concentration with Nanodrop.
Expected results: ~3–4 μg nascent RNA (including rRNA and tRNA) per sample for 3–4 g Arabidopsis seedlings.
-
38
Check the RNA integrity on a 1% (wt/vol) agarose gel. Load 5 μl purified RNA per sample.
Critical step
The 25 S and 18 S rRNA bands are used to assess the quality of RNA. Theoretically, the ratio of 25 S and 18 S rRNA bands of Arabidopsis should be ~2:1. However, it is difficult to get the theoretical ratio in practice. RNA can be used for library construction if the rRNA bands are sharp and intense (Fig. 5). The purity of RNA fractionation can be evaluated by quantitative PCR as described previously40.
Pause point
RNA can be stored at −80 °C for months.
-
39
(Optional) The high throughput and low cost of Illumina sequencing make it a widely accessible tool for exploring differences in gene expression and unspliced intron ratio. For users who are interested in analyzing intron retention using Illumina data, here we list the optional steps to perform Illumina mRNA sequencing and provide the corresponding bioinformatic pipelines as described in Box 1.
Ribosomal RNA and tRNA depletion
Timing At least 3.5 h
Critical
We often use 3–4 μg starting nascent RNA. Theoretically, 10% (wt/wt) RNA is recovered after ribosomal RNA depletion from total RNA. However, the recovery is far less than the theoretical value here owing to the large amount of tRNA in the chromatin-bound RNA fraction.
Critical
We have tried Thermo RiboMinus Plant Kit and riboPOOL probe targeting Pan-plant to remove rRNA. It is necessary to perform two rounds of rRNA depletion with Thermo RiboMinusTM Plant Kit. Only one round is needed for riboPOOL probe. The timing of this section depends on the kit used. The tRNA (~100 bp) is removed with ZYMO RNA Clean & Concentrator-5 kit during RNA concentration, which recovers only RNAs longer than 200 bp.
-
40
Concentrate the RNA following the instructions of the ZYMO RNA Clean & Concentrator-5 kit.
-
41
Elute the RNA with 10 μl RNase-free water.
-
42
Perform rRNA depletion according to the manufacturer’s instructions.
-
43
After rRNA depletion, clean and concentrate the RNA using the ZYMO RNA Clean & Concentrator-5 kit.
-
44
Elute the RNA with 6 μl RNase-free water.
-
45
Check the RNA concentration with Qubit, and assess the recovery after rRNA depletion.
Pause point
RNA can be stored at −80 °C for months.
3′ adapter ligation
Timing overnight
Critical
For ligation, the FLEP-seq protocol can be used with as little as 1 ng RNA as input.
-
46
Add 1 μl 50 pmol 3′ adapter (Universal miRNA cloning linker,) and 5 μl rRNA-depleted RNA to an RNase/DNase-free microtube, and mix by pipetting.
-
47
Incubate at 65 °C for 5 min, and place on ice for at least 1 min.
-
48
Add ligation reagents as follows:
Reagent
Volume (μl)
Final concentration
10× T4 RNA ligase reaction buffer
2
1×
RNaseOUT (40 U/μl)
1
2 U/μl
T4 RNA Ligase 2, truncated K227Q (20 U/μl)
1
1 U/μl
50% (wt/vol) PEG 8000
10
25% (wt/vol)
Total
20
–
-
49
Mix thoroughly by pipetting, and spin briefly in a microcentrifuge.
-
50
Incubate at 16 °C for 10 h.
-
51
Clean and concentrate the RNA using the ZYMO RNA Clean & Concentrator-5 kit.
-
52
Elute the RNA with 6 μl RNase-free water.
Pause point
RNA can be stored at −80 °C for months.
cDNA synthesis
Timing 2 h
-
53
Add 3.5 μl RNA and 1 μl 12 μM 3′ cDNA RT primer to a 0.2 ml RNase/DNase-free tube.
Critical step
The 3′ cDNA RT primer is designed based on the universal miRNA cloning linker to replace the original SMART CDS Primer II A provided with the Takara SMARTer PCR cDNA Synthesis Kit.
-
54
Mix by pipetting, and spin the tubes briefly in a microcentrifuge.
-
55
Incubate the tube at 72 °C for 3 min, and then slowly ramp to 42 °C at 0.1 °C/s, then incubate at 42 °C for 2 min in a PCR instrument (hot lid temperature 105 °C).
-
56
Prepare the following master mix at room temperature in the order shown:
Reagent
Volume (μl)
Final concentration
5× First-strand buffer
2
1×
DTT (100 mM)
0.25
2.5 mM
dNTP mix (10 mM)
1
1 mM
SMARTer II A oligonucleotide (12 μM)
1
1.2 mM
RNase inhibitor (40 U/μl)
0.25
1 U/μl
SMARTScribe reverse transcriptase (100 U/μl)
1
10 U/μl
Total
5.5
–
-
57
Place the master mix at 42 °C for 1 min to bring it up to temperature.
-
58
Add 5.5 μl master mix to each tube from Step 55, mix by pipetting and spin the tubes briefly in a microcentrifuge.
Critical step
It is critical to add the master mix immediately after Step 56. You can prepare the master mix during the incubation at 72 °C.
-
59
Incubate the tubes at 42 °C for 90 min, and terminate the reaction at 70 °C for 10 min in a thermal cycler (hot lid temperature 105 °C).
Pause point
cDNA can be stored at 4 °C overnight or at −80 °C for months.
PCR cycle optimization
Timing 3.5 h
Critical
To minimize the PCR bias resulting from overamplification, it is highly recommended to perform PCR cycle optimization to determine the best cycle number for large scale PCR, especially for low-input RNA. Use 1 μl first-strand cDNA for PCR cycle optimization.
-
60
Prepare PCR master mix in a 0.2 ml RNase/DNase-free tube as follows:
Reagent
Volume (μl)
Final concentration
5× PrimeSTAR GXL buffer
10
1×
First-strand cDNA
1
–
dNTP mix (2.5 mM each)
4
0.2 mM each
5′ PCR primer IIA (12 μM)
1
0.24 μM
Nuclease-free water
33
–
PrimeSTAR GXL DNA polymerase (1.25 U/μl)
1
0.025 U/μl
Total
50
–
Critical step
To check for possible PCR contamination, a tube of negative control without cDNA template should be also prepared for PCR and run using the maximum number of PCR cycles.
-
61
Run the PCR with the following conditions:
Cycle no.
Denature
Anneal
Extend
1
98 °C, 30 s
2–11 (10 cycles)
98 °C, 10 s
65 °C, 15 s
68 °C, 10 min
12
68 °C, 5 min
-
62
Transfer 5 μl PCR product to a tube with 1 μl 6× DNA loading after ten cycles, and label the tube ‘10’.
-
63
Return the tube with the remaining 45 μl to the PCR instrument, and run two cycles with following conditions (hot lid temperature 105 °C).
Cycle no.
Denature
Anneal
Extend
1–2 (2 cycles)
98 °C, 10 s
65 °C, 15 s
68 °C, 10 min
-
64
Transfer 5 μl PCR product to a tube with 1 μl 6× DNA loading after two cycles, and label the tube ‘12’.
-
65
Repeat Steps 64 and 65 for ‘14’ and ‘16’ cycles.
-
66
Run the four aliquots on a 1% (wt/vol) agarose gel, and determine the optimal number of PCR cycles.
Expected result: the optimal number of cycles is dependent on the RNA input. Typically, 3–4 μg input nascent RNA requires 11–13 cycles of PCR amplification. Lower-input RNA may require more cycles (Fig. 6).
Pause point
Unused cDNA (backups) can be stored at −80 °C for months.
Large-scale PCR
Timing 3–4 h
-
67
Set up 8× 50 μl PCR reactions with the first-strand cDNA as input.
-
68
Prepare a PCR master mix in 1.5 ml DNase-free tube as follows:
Reagent
Volume (μl)
Final concentration
5× PrimeSTAR GXL buffer
80
1×
First-strand cDNA
8
–
dNTP mix (2.5 mM each)
32
0.2 mM each
5′ PCR primer IIA (12 μM)
8
0.24 μM
Nuclease-free water
263
–
PrimeSTAR GXL DNA polymerase (1.25 U/μl)
8
0.025 U/μl
Total
400
–
-
69
Prepare aliquot of 50 μl PCR master mix into eight PCR DNase-free tubes, and perform PCR on a PCR thermal cycler using the optimal cycle (N) determined at Step 67.
Critical step
To check for possible PCR contamination, a tube containing a negative control without cDNA template should also be prepared for PCR.
-
70
Perform PCR using the following settings (hot lid temperature 105 °C):
Cycle no.
Denature
Anneal
Extend
1
98 °C, 30 s
2 − (N + 1)
98 °C, 10 s
65 °C, 15 s
68 °C, 10 min
N + 2
68 °C, 5 min
N indicates the optimal cycle determined by PCR cycle optimization.
Pause point
DNA can be stored at 4 °C for days and at −20 °C for months.
DNA library clean-up and quality control
Timing 2 h
-
71
Let the AMPure PB beads warm to room temperature for 30 min.
-
72
Pool all the PCR reactions from Step 71 (excluding the negative control) into a 1.5 ml DNA LoBind microcentrifuge tube, and warm to room temperature.
-
73
Add 1× volume beads into the PCR solution, mix well by pipetting and incubate in a rotator mixer for 10 min at room temperature.
-
74
Spin briefly in a microcentrifuge to collect the beads.
-
75
Place the tube on a magnetic rack, and keep until the solution appears completely clear. Remove the supernatant carefully by pipetting without disturbing the bead pellet.
-
76
Keep the tube on the magnetic rack, and wash the beads with 1 ml freshly prepared 70% (vol/vol) ethanol. Do not disturb the bead pellet.
Critical step
70% ethanol is hygroscopic and should be freshly prepared or stored in a tightly capped polypropylene tube for no more than 3 d.
-
77
After 30 s, aspirate and discard the 70% (vol/vol) ethanol.
-
78
Repeat Steps 76 and 77.
-
79
Spin the tube briefly in a microcentrifuge to collect the residual ethanol, and place back on the magnetic rack. Pipette off any remaining ethanol.
-
80
Air-dry the beads for 1 min, and check for any remaining droplets in the tube.
-
81
Add 100 μl nuclease-free water to the beads, and tap the tube to mix the pellet until all the beads are uniformly resuspended.
Critical step
Do not pipet to mix. It is hard to resuspend the beads if they are overdried and stick to the tube. Using a DNA LoBind microcentrifuge tube can reduce the loss of DNA.
-
82
Incubate the mix at room temperature for 2 min.
-
83
Spin down briefly to pellet the beads, and place the tube on the magnetic rack until the solution is clear.
-
84
Transfer the supernatant to a new 1.5 ml DNA LoBind microcentrifuge tube, without disturbing the pellet.
-
85
Perform a second round of AMPure PB beads purification with 1× volume beads as in Steps 74–85, and elute the beads with 40 μl nuclease-free water.
-
86
Check the DNA concentration and quality with Qubit and Agilent 2100 Bioanalyzer.
Expected results: this step should yield 5–30 ng/μl DNA in 40 μl water. The integrity of the library is important, as DNA of poor quality (e.g., highly fragmented) cannot be used for full-length sequencing. For example, the size of most transcripts in Arabidopsis should be larger than 1 kb (Fig. 7).
Pause point
DNA can be stored at 4 °C for days and at -20 °C for months.
-
87
Either proceed to Oxford Nanopore DNA library construction and sequencing (option A) or PacBio sequencing (option B).
-
(A)
Oxford Nanopore DNA library construction and sequencing
Timing 2–3 h for library preparation and 36 h for sequencing
-
(i)
Prepare the DNA sequencing library, including end-repair/A-tailing with NEB End repair/ dA-tailing Module reagents and Nanopore adapter ligation, and flow cell loading with the DNA ligation sequencing kit (SQK-LSK109) following the manufacturer’s instructions. 200 fmol input DNA is required. The online tool NEBioCalculator (https://nebiocalculator.neb.com/#!/dsdnaamt) is recommended to convert dsDNA mass to moles of dsDNA.
Critical step
Make sure the DNA meets the quantity and quality requirements specified by the manufacturers. Using too little or too much DNA will affect your library preparation. The average length of DNA molecules can be obtained from the Agilent 2100 Bioanalyzer.
-
(ii)
Check the quality of MinION flow cell to ensure it has enough functional pores for a good sequencing run using the MinKNOW software following the manufacturer’s instructions.
-
(iii)
Prime and load the library to a MinION flow cell according to the manufacturer’s instructions.
-
(iv)
Sequence the library on the MinION using the MinKNOW software according to the manufacturer’s instructions. The output format option must be set to include FAST5 if the user wants to measure poly(A) tail length. The run length can be set as 36 h, or you can set a longer time and stop it manually. The default bias voltage is −180 mV, which is suitable for a new flow cell. However, if the flow cell is reused or a stopped sequencing process is rerun, the bias voltage needs to be adjusted based on the total previous runtime of the flow cell as per the manufacturer’s instructions. It is recommended that real-time basecalling be disabled during sequencing to save time and perform basecalling with Guppy after sequencing as described in Step 88A(ii).
Critical step
Ensure the computer has sufficient space to meet the data storage requirements. If low pore occupancy or low sequencing yield are obtained or less data per library is required, wash the flow cell with Flow Cell Wash Kit (EXP-WSH003) following the manufacturer’s instructions and run a second library or store the flow cell at 4 °C for later use.
-
(v)
After the sequencing is complete, flush and store the flow cell following the manufacturer’s instructions.
Critical step
If less data per library is required, wash the flow cell with Flow Cell Wash Kit (EXP-WSH003) following the manufacturer’s instructions and run a second library or store the flow cell at 4 °C for later use.
-
(i)
-
(B)
PacBio sequencing
Timing 2 d
-
(i)
Prepare the library according to the instructions of SMRTbell Template Prep Kit 1.0-SPv3 starting from the step of DNA damage repair. The recommended input DNA (from Step 87) is 160–500 ng for this step.
-
(ii)
Sequence the final library on a PacBio Sequel II System according to the operations guide of the Sequel II System.
-
(i)
-
(A)
-
88
Analyze sequencing data following either option A (Nanopore) or option B (PacBio).
-
(A)
Nanopore data analysis pipeline
Timing 36 h (basecalling) + 4 h (downstream analysis)
-
(i)
Download the scripts used below from https://github.com/ZhaiLab-SUSTech/FLEPSeq/tree/master/script. The main functionalities of these scripts are described in Table 2. Download the TAIR10 genome sequence from TAIR (https://www.arabidopsis.org) and the genome annotation file from https://github.com/ZhaiLab-SUSTech/FLEPSeq/tree/master/genome_lib. These annotation information are extracted from published ARAPORT11 annotation file (https://www.arabidopsis.org/download_files/Genes/Araport11_genome_release/Araport11_GFF3_genes_transposons.201606.gff.gz)46.
Table 2 Description of scripts in the FLEP-seq bioinformatic pipeline -
(ii)
(Optional) Nanopore basecalling.
Basecalling is the most time-consuming step in Nanopore data analysis, and thus is usually performed during sequencing by MinKNOW. However, the Guppy software is updated frequently, and the latest version tends to yield the highest accuracy. The researchers can perform basecalling after sequencing using the command below:
$ guppy_basecaller -i raw_fast5_dir -s out_fast5_dir -c dna_r9.4.1_450bps_hac.cfg --recursive --fast5_out --disable_pings --qscore_filtering --device "cuda:all:100%"
-
(iii)
Convert FASTQ files to FASTA format.
$ python fastqdir2fasta.py --indir path/to/fastq_pass --out all.fasta
-
(iv)
Use minimap2 to map reads to reference genome.
$ minimap2 -ax splice --secondary=no genome.fasta all.fasta -t 10 > tmp.sam
Critical step
For organisms with short introns, such as Arabidopsis, it might be better to use the parameter ‘-G’ to set the max intron length, for example, ‘-G 12000’.
$ samtools sort -o mapped.bam tmp.sam $ samtools index mapped.bam $ rm tmp.sam
-
(v)
(Optional) Remove reads derived from rRNAs, tRNAs, snoRNAs, snRNA as well as mitochondrial and chloroplast RNAs.
$ python filter_rRNA_bam.py --inbam mapped.bam --inbed genome_lib/rRNAtRNAetc.bed --out clean.bam $ samtools index clean.bam
-
(vi)
Find 3′ adapter in reads.
$ python adapterFinder.py --inbam clean.bam --inseq all.fasta --out adapter.result.txt --threads 36
-
(vii)
Identify poly(A) tails, and estimate their length.
This script requires the fast5 files generated by MinKNOW or Guppy to identify poly(A). These files are stored in the ‘fast5_pass’ (for MinKNOW) or ‘workspace’ subdirectory (for Guppy). This script also requires the sequencing_summary.txt file generated by MinKNOW or Guppy to obtain the fast5 file name of each read. A demo sequencing_summary.txt file is stored in https://github.com/ZhaiLab-SUSTech/FLEPSeq/blob/master/demo_data/out_fast5_dir/sequencing_summary.test.txt.
$ python PolyAcaller.py --inadapter adapter.result.txt --summary sequencing_summary.txt --fast5dir fast5_dir --out polyA_tail.result.txt --threads 36
-
(viii)
Extract read information.
This script will produce a table containing intron retention information and the 3′ end position of transcripts.
$ python extract_read_info.py --inbam clean.bam –inbed genome_lib/exon_intron_pos.repr.bed --out read.intron_retention.txt
-
(ix)
Merge the above analysis results.
This script will also mark each full-length read as one of the following categories: (a) elongating transcripts, (b) polyadenylated transcripts, (c) splicing intermediates, (d) elongating transcripts with a lower mapping accuracy at 3′ end (might be due to sequencing error), (e) reads missing 5′ end (probably due to incomplete RT).
$ Rscript merge_read_info.R --type Nanopore --inreadinfo read.intron_retention.txt --inadapter adapter.result.txt --inpolya polyA_tail.result.txt --out read.info.txt
-
(x)
Analyze splicing kinetics.
The 3′ end of elongating transcripts captured by FLEP-seq represents the position of RNA Pol II. Additionally, the splicing status of each intron in a transcript can also be recovered by full-length sequencing. The intron splicing status and the transcribed distance between the 3′ splicing site of the intron and the 3′ end of the read are extracted by ‘prepare_data_for_splice_kinetics.py’. The transcribed distances past the 3′ splicing site are binned into 20 nt windows, and the ratio of spliced introns in each transcribed distance bin is calculated by ‘plot_intron_splicing_kinetics.R’ to estimate cotranscriptional splicing kinetics.
$ python prepare_data_for_splice_kinetics.py --inreadinfo read.info.txt --inbed genome_lib/exon_intron_pos.repr.bed --out read.intron.pos.splicing.txt $ Rscript plot_intron_splicing_kinetics.R --inrelpos read.intron.pos.splicing.txt --inreadinfo read.info.txt --inintron genome_lib/select_introns.txt --out read.splicing_kinetics.txt --pdf read.splicing_kinetics.pdf
-
(xi)
Calculate the intron retention ratio of polyadenylated transcripts.
$ Rscript cal_polya_transcript_ir.R --inrelpos read.intron.pos.splicing.txt --inreadinfo read.info.txt --outrna mRNA.incompletely_spliced_ratio.txt --outintron intron.unspliced_ratio.txt
-
(i)
-
(B)
PacBio data analysis pipeline
Timing 30 h (generating circular consensus sequences) + 1 h (downstream analysis)
-
(i)
Generate highly accurate single-molecule consensus reads using CCS.
This is the most time-consuming step in PacBio data analysis. For the data we used (~12 M subreads), it took ~30 h to generate consensus sequences (Hifi-reads) on Intel Xeon 6140 CPU at 2.3 GHz.
$ ccs --num-threads 36 --min-rq 0.9 --report-file ccs.report input.subreads.bam ccs.bam
-
(ii)
Remove the adapter sequence using lima.
$ echo ‘>primer_3p\nAAGCAGTGGTATCAACGCAGAGTACATTGATGGTGCCTACAG\n>primer_5p\nAAGCAGTGGTATCAACGCAGAGTACATGGG\n' > primer.fasta $ lima -j 36 ccs.bam primer.fasta lima.bam --isoseq --peek-guess $ python lima_bam2fasta.py --infile lima.primer_3p--primer_5p.bam --out all.fasta
-
(iii)
Use minimap2 to map reads to reference genome, and remove rRNA-derived reads. This is identical to Step 87A(iv–v) in the Nanopore pipeline.
-
(iv)
Identify poly(A) tails, and estimate their lengths.
$ python pacbio_find_polyA.py --inbam clean.bam --inseq all.fasta --out polyA_tail.result.txt
-
(v)
Extract read information.
This is identical to Step 88A(viii) in the Nanopore pipeline.
-
(vi)
Merge the above analysis results.
$ Rscript merge_read_info.R --type PacBio --inreadinfo read.intron_retention.txt --inpolya polyA_tail.result.txt --out read.info.txt
-
(vii)
Analyze splicing kinetics, and calculate the intron retention ratio of polyadenylated transcripts as described in Step 88A(x–xi) in the Nanopore pipeline.
-
(viii)
(Optional) Visualize the alignment using our in-house jupyter notebook alignment_visualization.ipynb.
-
(i)
-
(A)
Troubleshooting
Troubleshooting advice can be found in Table 3.
Timing
RNA purification and 3′ adapter ligation
-
Steps 1–31, cell fractionation: 2 h
-
Steps 32–39, RNA purification from fractions: 2 h
-
Steps 40–45, ribosomal RNA and tRNA depletion: at least 3.5 h
-
Steps 46–52, 3′ adapter ligation: overnight
Full-length library preparation and sequencing
-
Steps 53–59, cDNA synthesis: 2 h
-
Steps 60–66, PCR cycle optimization: 3.5 h
-
Steps 67–70, large-scale PCR: 3–4 h
-
Steps 71–86, DNA library clean-up and quality control: 2 h
-
Step 87A, Oxford Nanopore DNA library construction and sequencing: 38–39 h
-
Step 87B, PacBio DNA library construction and sequencing: 2 d
Data analysis
-
Step 88A, Nanopore data analysis: 36 h (optional, basecalling) + 4 h (downstream analysis)
-
Step 88B, PacBio data analysis: 30 h (generating CCS reads) + 1 h (downstream analysis)
-
Box 1, Illumina data analysis: 6 h
Anticipated results
With this protocol, researchers can expect to construct a full-length cDNA library of chromatin-bound nascent RNA from Arabidopsis tissues and perform data analysis on both polyadenylated and nonpolyadenylated transcripts. The purity of the subcellular fractions can be monitored by western blotting (Fig. 4). The integrity and quality of RNA fractions can be evaluated by gel electrophoresis (Fig. 5) and Nanodrop. Typically, 3–4 μg nascent RNA (including rRNA and tRNA) can be obtained from 4 g of 12-d-old Arabidopsis seedlings and subsequently requires 11–13 cycles of PCR amplification; lower amount of input RNA may require a higher number of PCR cycles (Fig. 6). After the final library clean-up and elution, one can expect a yield of 5–30 ng/μl DNA in 40 μl water. The integrity of library is important; DNA of poor quality (e.g., highly fragmented) cannot be used for full-length sequencing. For example, the size of most transcripts in Arabidopsis should be >1 kb as evaluated by Agilent 2100 (Fig. 7).
A typical Nanopore FLEP-seq library in Arabidopsis, e.g., the CB1 or CB2 samples that we have published previously31, can yield 7–12 million raw reads, >95% of which can be assigned to the genome. After removing rRNA, chloroplast, mitochondria and noncoding transcripts, ~70% of raw reads can be mapped to protein coding gene. After further filtering, a library from Arabidopsis seedlings can yield ~3 million full-length clean reads, which covers ~20,000 protein-coding genes. After removing splicing intermediate and reads with low accuracy of 3′ end mapping, we obtained more than 1 million elongating transcripts and 0.5 million polyadenylated transcripts.
Both RNA integrity and sequencing depth are critical for data analysis. RNA degradation during experimental manipulation could lead to the failure of library construction (Fig. 7). rRNA contamination and library complexity can also affect the sequencing depth. The possible reasons for poor data quality and corresponding solutions are discussed in the Troubleshooting section (Table 3).
Data availability
The data used in this study were previously published35 and deposited at NCBI under the accession number PRJNA591665.
Code availability
All software used in this study is described in detail in the Equipment section. Custom Python and R code used for bioinformatics analysis have been deposited in a GitHub repository (https://github.com/ZhaiLab-SUSTech/FLEPSeq).
References
Herzel, L., Ottoz, D. S. M., Alpert, T. & Neugebauer, K. M. Splicing and transcription touch base: co-transcriptional spliceosome assembly and function. Nat. Rev. Mol. Cell Biol. 18, 637–650 (2017).
Naftelberg, S., Schor, I. E., Ast, G. & Kornblihtt, A. R. Regulation of alternative splicing through coupling with transcription and chromatin structure. Annu. Rev. Biochem. 84, 165–198 (2015).
Bentley, D. L. Coupling mRNA processing with transcription in time and space. Nat. Rev. Genet. 15, 163–175 (2014).
Carrillo Oesterreich, F. et al. Splicing of nascent RNA coincides with intron exit from RNA Polymerase II. Cell 165, 372–381 (2016).
Herzel, L., Straube, K. & Neugebauer, K. M. Long-read sequencing of nascent RNA reveals coupling among RNA processing events. Genome Res. 28, 1008–1019 (2018).
Reimer, K. A., Mimoso, C. A., Adelman, K. & Neugebauer, K. M. Co-transcriptional splicing regulates 3’ end cleavage during mammalian erythropoiesis. Mol. Cell https://doi.org/10.1016/j.molcel.2020.12.018 (2021).
Drexler, H. L., Choquet, K. & Churchman, L. S. Splicing kinetics and coordination revealed by direct nascent RNA sequencing through nanopores. Mol. Cell 77, 985–998.e988 (2020).
Drexler, H. L. et al. Revealing nascent RNA processing dynamics with nano-COP. Nat. Protoc. https://doi.org/10.1038/s41596-020-00469-y (2021).
Sousa-Luis, R. et al. POINT technology illuminates the processing of polymerase-associated intact nascent transcripts. Mol. Cell 81, 1935–1950.e6 (2021).
Wissink, E. M., Vihervaara, A., Tippens, N. D. & Lis, J. T. Nascent RNA analyses: tracking transcription and its regulation. Nat. Rev. Genet. 20, 705–723 (2019).
Mayer, A. et al. Native elongating transcript sequencing reveals human transcriptional activity at nucleotide resolution. Cell 161, 541–554 (2015).
Bhatt, DevM. et al. Transcript dynamics of proinflammatory genes revealed by sequence analysis of subcellular RNA fractions. Cell 150, 279–290 (2012).
Nechaev, S. et al. Global analysis of short RNAs reveals widespread promoter-proximal stalling and arrest of Pol II in Drosophila. Science 327, 335 (2010).
Churchman, L. S. & Weissman, J. S. Nascent transcript sequencing visualizes transcription at nucleotide resolution. Nature 469, 368–373 (2011).
Nojima, T. et al. Mammalian NET-Seq reveals genome-wide nascent transcription coupled to RNA processing. Cell 161, 526–540 (2015).
Core, L. J., Waterfall, J. J. & Lis, J. T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845 (2008).
Kwak, H., Fuda, N. J., Core, L. J. & Lis, J. T. Precise maps of RNA polymerase reveal how promoters direct initiation and pausing. Science 339, 950–953 (2013).
Judd, J. et al. A rapid, sensitive, scalable method for Precision Run-On sequencing (PRO-seq). Preprint at bioRxiv https://doi.org/10.1101/2020.05.18.102277 (2020).
Rabani, M. et al. Metabolic labeling of RNA uncovers principles of RNA production and degradation dynamics in mammalian cells. Nat. Biotechnol. 29, 436–442 (2011).
Schwalb, B. et al. TT-seq maps the human transient transcriptome. Science 352, 1225 (2016).
Herzog, V. A. et al. Thiol-linked alkylation of RNA to assess expression dynamics. Nat. Methods 14, 1198–1204 (2017).
Schofield, J. A., Duffy, E. E., Kiefer, L., Sullivan, M. C. & Simon, M. D. TimeLapse-seq: adding a temporal dimension to RNA sequencing through nucleoside recoding. Nat. Methods 15, 221–225 (2018).
Shepard, P. J. et al. Complex and dynamic landscape of RNA polyadenylation revealed by PAS-Seq. RNA 17, 761–772 (2011).
Harrison, P. F. et al. PAT-seq: a method to study the integration of 3’-UTR dynamics with gene expression in the eukaryotic transcriptome. RNA 21, 1502–1510 (2015).
Ozsolak, F. et al. Direct RNA sequencing. Nature 461, 814–818 (2009).
Chang, H., Lim, J., Ha, M. & Kim, V. N. TAIL-seq: genome-wide determination of poly(A) tail length and 3′ end modifications. Mol. Cell 53 (2014).
Lim, J., Lee, M., Son, A., Chang, H. & Kim, V. N. mTAIL-seq reveals dynamic poly(A) tail regulation in oocyte-to-embryo development. Genes Dev. 30, 1671–1682 (2016).
Subtelny, A. O., Eichhorn, S. W., Chen, G. R., Sive, H. & Bartel, D. P. Poly(A)-tail profiling reveals an embryonic switch in translational control. Nature 508, 66–71 (2014).
Legnini, I., Alles, J., Karaiskos, N., Ayoub, S. & Rajewsky, N. FLAM-seq: full-length mRNA sequencing reveals principles of poly(A) tail length control. Nat. Methods 16, 879–886 (2019).
Liu, Y., Nie, H., Liu, H. & Lu, F. Poly(A) inclusive RNA isoform sequencing (PAIso−seq) reveals wide-spread non-adenosine residues within RNA poly(A) tails. Nat. Commun. 10, 5292 (2019).
Parker, M. T. et al. Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification. eLife 9, e49658 (2020).
Workman, R. E. et al. Nanopore native RNA sequencing of a human poly(A) transcriptome. Nat. Methods 16, 1297–1305 (2019).
Roach, N. P. et al. The full-length transcriptome of C. elegans using direct RNA sequencing. Genome Res 30, 299–312 (2020).
Kim, D. et al. The architecture of SARS-CoV-2 transcriptome. Cell 181, 914–921.e910 (2020).
Jia, J. et al. Post-transcriptional splicing of nascent RNA contributes to widespread intron retention in plants. Nat. Plants 6, 780–788 (2020).
Boutz, P. L., Bhutkar, A. & Sharp, P. A. Detained introns are a novel, widespread class of post-transcriptionally spliced introns. Genes Dev. 29, 63–80 (2015).
Mauger, O., Lemoine, F. & Scheiffele, P. Targeted intron retention and excision for rapid gene regulation in response to neuronal activity. Neuron 92, 1266–1278 (2016).
Naro, C. et al. An orchestrated intron retention program in meiosis controls timely usage of transcripts during germ cell differentiation. Dev. Cell 41, 82–93.e84 (2017).
Gahura, O. et al. Prp45 affects Prp22 partition in spliceosomal complexes and splicing efficiency of non-consensus substrates. J. Cell. Biochem. 106, 139–151 (2009).
Siatecka, M., Reyes, J. L. & Konarska, M. M. Functional interactions of Prp8 with both splice sites at the spliceosomal catalytic center. Genes Dev. 13, 1983–1993 (1999).
Zhu, D. et al. The features and regulation of co-transcriptional splicing in Arabidopsis. Mol. Plant 13, 278–294 (2020).
Krause, M. et al. tailfindr: alignment-free poly(A) length measurement for Oxford Nanopore RNA and DNA sequencing. RNA 25, 1229–1241 (2019).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Cheng, C.-Y. et al. Araport11: a complete reannotation of the Arabidopsis thaliana reference genome. Plant J. 89, 789–804 (2017).
Acknowledgements
We are grateful for the useful comments and edits suggested by the anonymous reviewers. The group of J.Z. is supported by a Stable Support Plan Program of Shenzhen Natural Science Fund Grant (20200925153345004), a National Key R&D Program of China Grant (2019YFA0903903), the Program for Guangdong Introducing Innovative and Entrepreneurial Teams (2016ZT06S172), the Shenzhen Sci-Tech Fund (KYTDPT20181011104005) and the Key Laboratory of Molecular Design for Plant Cell Factory of Guangdong Higher Education Institutes (2019KSYS006).
Author information
Authors and Affiliations
Contributions
Y.L., J.J. and J.Z. conceived and designed the study. Y.L., J.J. and X.J. developed the method and performed the experiments. J.J., W.M. and J.Z. designed the computational pipeline and analyzed the data. J.Z. conceived and oversaw the study. All authors wrote and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Zongwei Cai, Shona Murphy, Yongsheng Shi and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Jia et al. Nat. Plants 6, 780–788 (2020): https://doi.org/10.1038/s41477-020-0688-1
Key data used in this protocol
Jia et al. Nat. Plants 6, 780–788 (2020): https://doi.org/10.1038/s41477-020-0688-1
Rights and permissions
About this article
Cite this article
Long, Y., Jia, J., Mo, W. et al. FLEP-seq: simultaneous detection of RNA polymerase II position, splicing status, polyadenylation site and poly(A) tail length at genome-wide scale by single-molecule nascent RNA sequencing. Nat Protoc 16, 4355–4381 (2021). https://doi.org/10.1038/s41596-021-00581-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00581-7
This article is cited by
-
Stress responses of plants through transcriptome plasticity by mRNA alternative polyadenylation
Molecular Horticulture (2023)
-
Remodeling of maternal mRNA through poly(A) tail orchestrates human oocyte-to-embryo transition
Nature Structural & Molecular Biology (2023)
-
Short poly(A) tails are protected from deadenylation by the LARP1–PABP complex
Nature Structural & Molecular Biology (2023)
-
Single-molecule targeted accessibility and methylation sequencing of centromeres, telomeres and rDNAs in Arabidopsis
Nature Plants (2023)
-
Comprehensive analysis of mRNA poly(A) tails by PAIso-seq2
Science China Life Sciences (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.