Abstract
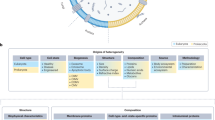
Extracellular vesicles (EVs) are heterogeneous lipid containers with a complex molecular cargo comprising several populations with unique roles in biological processes. These vesicles are closely associated with specific physiological features, which makes them invaluable in the detection and monitoring of various diseases. EVs play a key role in pathophysiological processes by actively triggering genetic or metabolic responses. However, the heterogeneity of their structure and composition hinders their application in medical diagnosis and therapies. This diversity makes it difficult to establish their exact physiological roles, and the functions and composition of different EV (sub)populations. Ensemble averaging approaches currently employed for EV characterization, such as western blotting or ‘omics’ technologies, tend to obscure rather than reveal these heterogeneities. Recent developments in single-vesicle analysis have made it possible to overcome these limitations and have facilitated the development of practical clinical applications. In this review, we discuss the benefits and challenges inherent to the current methods for the analysis of single vesicles and review the contribution of these approaches to the understanding of EV biology. We describe the contributions of these recent technological advances to the characterization and phenotyping of EVs, examination of the role of EVs in cell-to-cell communication pathways and the identification and validation of EVs as disease biomarkers. Finally, we discuss the potential of innovative single-vesicle imaging and analysis methodologies using microfluidic devices, which promise to deliver rapid and effective basic and practical applications for minimally invasive prognosis systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hooke, R. Micrographia: or Some Physiological Descriptions of Minute Bodies Made by Magnifying Glasses. With Observations and Inquiries Thereupon (1665).
Die Altmann, R. Elementarorganismen und ihre Beziehungen zu den Zellen (von Veit, 1890).
van Leeuwenhoek, A. Opera Omnia, seu Arcana Naturae ope exactissimorum Microscopiorum detecta, experimentis variis comprobata, Epistolis ad varios illustres viros.
Wolf, P. The nature and significance of platelet products in human plasma. Br. J. Haematol. 13, 269–288 (1967).
Herr, D. R. et al. Ultrastructural characteristics of DHA-induced pyroptosis. Neuromolecular Med. https://doi.org/10.1007/s12017-019-08586-y (2020).
Patras, L. & Banciu, M. Intercellular crosstalk via extracellular vesicles in tumor milieu as emerging therapies for cancer progression. Curr. Pharm. Des. 25, 1980–2006 (2019).
Andaloussi, S. E. L., Mäger, I., Breakefield, X. O. & Wood, M. J. A. Extracellular vesicles: biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 12, 347–357 (2013).
Buzas, E. I., György, B., Nagy, G., Falus, A. & Gay, S. Emerging role of extracellular vesicles in inflammatory diseases. Nat. Rev. Rheumatol. 10, 356–364 (2014).
Panagopoulou, M. S., Wark, A. W., Birch, D. J. S. & Gregory, C. D. Phenotypic analysis of extracellular vesicles: a review on the applications of fluorescence. J. Extracell. Vesicles 9, 1710020 (2020).
Yáñez-Mó, M. et al. Biological properties of extracellular vesicles and their physiological functions. J. Extracell. Vesicles 4, 27066 (2015).
Van Niel, G., D’Angelo, G. & Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 19, 213–228 (2018).
Nieuwland, R. & Sturk, A. Why do cells release vesicles? Thromb. Res. 125, S49–S51 (2010).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659 (2007).
Anderson, H. C., Mulhall, D. & Garimella, R. Role of extracellular membrane vesicles in the pathogenesis of various diseases, including cancer, renal diseases, atherosclerosis, and arthritis. Lab. Investig. 90, 1549–1557 (2010).
Wiklander, O. P. B., Brennan, M., Lötvall, J., Breakefield, X. O. & Andaloussi, S. E. L. Advances in therapeutic applications of extracellular vesicles. Sci. Transl. Med. 11, 1–16 (2019).
An, T. et al. Exosomes serve as tumour markers for personalized diagnostics owing to their important role in cancer metastasis. J. Extracell. Vesicles 4, 27522 (2015).
Becker, A. et al. Extracellular vesicles in cancer: cell-to-cell mediators of metastasis. Cancer Cell 30, 836–848 (2016).
Zhao, H. et al. Tumor microenvironment derived exosomes pleiotropically modulate cancer cell metabolism. eLife 5, e10250 (2016).
Chiang, C. Y. & Chen, C. Toward characterizing extracellular vesicles at a single-particle level. J. Biomed. Sci. 26, 9 (2019).
Théry, C. et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 7, 1535750 (2018).
Zhang, H. et al. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric-flow field-flow fractionation. Nat. Cell Biol. 20, 332–343 (2018).
Mathieu, M., Martin-Jaular, L., Lavieu, G. & Théry, C. Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat. Cell Biol. 21, 9–17 (2019).
Tai, Y. L., Chen, K. C., Hsieh, J. T. & Shen, T. L. Exosomes in cancer development and clinical applications. Cancer Sci. 109, 2364–2374 (2018).
Willms, E., Cabañas, C., Mäger, I., Wood, M. J. A. & Vader, P. Extracellular vesicle heterogeneity: subpopulations, isolation techniques, and diverse functions in cancer progression. Front. Immunol. 9, 738 (2018).
Kalluri, R. & LeBleu, V. S. The biology, function, and biomedical applications of exosomes. Science 367, eaau6977 (2020).
Van Der Pol, E. et al. Optical and non-optical methods for detection and characterization of microparticles and exosomes. J. Thromb. Haemost. 8, 2596–2607 (2010).
Soung, Y. H., Ford, S., Zhang, V. & Chung, J. Exosomes in cancer diagnostics. Cancers (Basel) 9, 8 (2017).
Puente-Massaguer, E., Lecina, M. & Gòdia, F. Application of advanced quantification techniques in nanoparticle-based vaccine development with the Sf9 cell baculovirus expression system. Vaccine 38, 1849–1859 (2020).
Pick, H., Alves, A. C. & Vogel, H. Single-vesicle assays using liposomes and cell-derived vesicles: from modeling complex membrane processes to synthetic biology and biomedical applications. Chem. Rev. 118, 8598–8654 (2018).
Tkach, M., Kowal, J. & Théry, C. Why the need and how to approach the functional diversity of extracellular vesicles. Philos. Trans. R. Soc. B Biol. Sci. 373, 20160479 (2018).
Goñi, F. M. The basic structure and dynamics of cell membranes: an update of the Singer-Nicolson model. Biochim. Biophys. Acta 1838, 1467–1476 (2014).
Bhatia, V. K. et al. Amphipathic motifs in BAR domains are essential for membrane curvature sensing. EMBO J. 28, 3303–3314 (2009).
Mathiasen, S. et al. Nanoscale high-content analysis using compositional heterogeneities of single proteoliposomes. Nat. Methods 11, 931–934 (2015).
Brett, S. I. et al. Immunoaffinity based methods are superior to kits for purification of prostate derived extracellular vesicles from plasma samples. Prostate 77, 1335–1343 (2017).
Royo, F. et al. Different EV enrichment methods suitable for clinical settings yield different subpopulations of urinary extracellular vesicles from human samples. J. Extracell. Vesicles 5, 29497 (2016).
Ramirez, M. I. et al. Technical challenges of working with extracellular vesicles. Nanoscale 10, 881–906 (2018).
Sódar, B. W. et al. Low-density lipoprotein mimics blood plasma-derived exosomes and microvesicles during isolation and detection. Sci. Rep. 6, 24316 (2016).
Woo, J. R., Sharma, S. & Gimzewski, J. The role of isolation methods on a nanoscale surface structure and its effect on the size of exosomes. J. Circ. Biomark. 5, 11 (2016).
Takahashi, K., Yan, I. K., Kim, C., Kim, J. & Patel, T. Analysis of extracellular RNA by digital PCR. Front. Oncol. 4, 129 (2014).
Liu, Y. & Lu, Q. Extracellular vesicle microRNAs: biomarker discovery in various diseases based on RT-qPCR. Biomark. Med. 9, 791–805 (2015).
Giannopoulou, L., Zavridou, M., Kasimir-Bauer, S. & Lianidou, E. S. Liquid biopsy in ovarian cancer: the potential of circulating miRNAs and exosomes. Transl. Res. 205, 77–91 (2019).
Crocetti, E. Epidemiology of prostate cancer in Europe. Centre for Parliamentary Studies https://ec.europa.eu/jrc/en/publication/epidemiology-prostate-cancer-europe (2015).
Torrano, V. et al. Vesicle-MaNiA: extracellular vesicles in liquid biopsy and cancer. Curr. Opin. Pharmacol. 29, 47–53 (2016).
Heidenreich, A. et al. EAU guidelines on prostate cancer. Part 1: screening, diagnosis, and local treatment with curative intent—update 2013. Eur. Urol. 65, 124–137 (2014).
Humphrey, P. A. Diagnosis of adenocarcinoma in prostate needle biopsy tissue. J. Clin. Pathol. 60, 35–42 (2007).
Shariat, S. F. & Roehrborn, C. G. Using biopsy to detect prostate cancer. Rev. Urol. 10, 262–280 (2008).
Clos-Garcia, M. et al. Metabolic alterations in urine extracellular vesicles are associated to prostate cancer pathogenesis and progression. J. Extracell. Vesicles 7, 1470442 (2018).
Höög, J. L. & Lötvall, J. Diversity of extracellular vesicles in human ejaculates revealed by cryo-electron microscopy. J. Extracell. Vesicles 4, 28680 (2015).
Duijvesz, D. et al. Immuno-based detection of extracellular vesicles in urine as diagnostic marker for prostate cancer. Int. J. Cancer 137, 2869–2878 (2015).
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell Biol. 200, 373–383 (2013).
Raposo, G. & Stahl, P. D. Extracellular vesicles: a new communication paradigm? Nat. Rev. Mol. Cell Biol. 20, 509–510 (2019).
Théry, C., Clayton, A., Amigorena, S. & Raposo, G. Isolation and characterization of exosomes from cell culture supernatants. in Current Protocols in Cell Biology https://doi.org/10.1002/0471143030.cb0322s30 (2006).
Giulietti, M. et al. Exploring small extracellular vesicles for precision medicine in prostate cancer. Front. Oncol. 8, 221 (2018).
Russell, A. E. et al. Biological membranes in EV biogenesis, stability, uptake, and cargo transfer: an ISEV position paper arising from the ISEV membranes and EVs workshop. J. Extracell. Vesicles 8, 1684862 (2019).
Chen, C. et al. Isolation of a novel bacterial strain capable of producing abundant extracellular membrane vesicles carrying a single major cargo protein and analysis of its transport mechanism. Front. Microbiol. 10, 3001 (2020).
Szatanek, R. et al. The methods of choice for extracellular vesicles (EVs) characterization. Int. J. Mol. Sci. 18, 1153 (2017).
Tatischeff, I., Larquet, E., Falcon-Perez, J. M., Turpin, P.-Y. & Kruglik, S. G. Fast characterisation of cell-derived extracellular vesicles by nanoparticles tracking analysis, cryo-electron microscopy, and raman tweezers microspectroscopy. J. Extracell. Vesicles 1, 19179 (2012).
Hyenne, V. et al. Studying the fate of tumor extracellular vesicles at high spatiotemporal resolution using the zebrafish embryo. Dev. Cell 48, 554–572.e7 (2019).
Tian, Q. et al. Nanoparticle counting by microscopic digital detection: selective quantitative analysis of exosomes via surface-anchored nucleic acid amplification. Anal. Chem. 90, 6556–6562 (2018).
Carney, R. P. et al. Multispectral optical tweezers for biochemical fingerprinting of CD9-positive exosome subpopulations. Anal. Chem. 89, 5357–5363 (2017).
Enciso-Martinez, A. et al. Synchronized Rayleigh and Raman scattering for the characterization of single optically trapped extracellular vesicles. Nanomedicine 24, 102109 (2020).
Stremersch, S. et al. Identification of individual exosome-like vesicles by surface enhanced raman spectroscopy. Small 12, 3292–3301 (2016).
Yuana, Y. et al. Cryo-electron microscopy of extracellular vesicles in fresh plasma. J. Extracell. Vesicles 2, 21494 (2013).
Daaboul, G. G. et al. Digital detection of exosomes by interferometric imaging. Sci. Rep. 6, 37246 (2016).
Ridolfi, A. et al. AFM-based high-throughput nanomechanical screening of single extracellular vesicles. Anal. Chem. 92, 10274–10282 (2020).
Kim, S. Y., Khanal, D., Kalionis, B. & Chrzanowski, W. High-fidelity probing of the structure and heterogeneity of extracellular vesicles by resonance-enhanced atomic force microscopy infrared spectroscopy. Nat. Protoc. 14, 576–593 (2019).
Zong, S. et al. Single molecule localization imaging of exosomes using blinking silicon quantum dots. Nanotechnology 29, 065705 (2017).
Filipe, V., Hawe, A. & Jiskoot, W. Critical evaluation of nanoparticle tracking analysis (NTA) by NanoSight for the measurement of nanoparticles and protein aggregates. Pharm. Res. 27, 796–810 (2010).
Bachurski, D. et al. Extracellular vesicle measurements with nanoparticle tracking analysis—an accuracy and repeatability comparison between NanoSight NS300 and ZetaView. J. Extracell. Vesicles 8, 1596016 (2019).
Rikkert, L. G. et al. Cancer-ID: toward identification of cancer by tumor-derived extracellular vesicles in blood. Front. Oncol. 10, 608 (2020).
Smith, Z. J. et al. Single exosome study reveals subpopulations distributed among cell lines with variability related to membrane content. J. Extracell. Vesicles 4, 28533 (2015).
Carney, R. P. et al. Targeting tumor-associated exosomes with integrin-binding peptides. advanced biosystems. Physiol. Behav. 1, 1600038 (2017).
Lee, W. et al. Label-free prostate cancer detection by characterization of extracellular vesicles using Raman spectroscopy. Anal. Chem. 90, 11290–11296 (2018).
Kruglik, S. G. et al. Raman tweezers microspectroscopy of circa 100 nm extracellular vesicles. Nanoscale 11, 1661–1679 (2019).
Dai, Y. et al. Combined morpho-chemical profiling of individual extracellular vesicles and functional nanoparticles without labels. Anal. Chem. 92, 5585–5594 (2020).
Enciso-Martinez, A. et al. Label-free identification and chemical characterisation of single extracellular vesicles and lipoproteins by synchronous Rayleigh and Raman scattering. J. Extracell. Vesicles 9, 1730134 (2020).
Lee, W., Lenferink, A. T. M., Otto, C. & Offerhaus, H. L. Classifying Raman spectra of extracellular vesicles based on convolutional neural networks for prostate cancer detection. J. Raman Spectrosc. 51, 293–300 (2020).
Bryce, D. A., Kitt, J. P. & Harris, J. M. Confocal-Raman microscopy characterization of supported phospholipid bilayers deposited on the interior surfaces of chromatographic silica. J. Am. Chem. Soc. 140, 4071–4078 (2018).
Kitt, J. P., Bryce, D. A., Minteer, S. D. & Harris, J. M. Confocal Raman microscopy for in situ measurement of phospholipid-water partitioning into model phospholipid bilayers within individual chromatographic particles. Anal. Chem. 90, 7048–7055 (2018).
Penders, J. et al. Single particle automated Raman trapping analysis. Nat. Commun. 9, 4256 (2018).
Bour, A. et al. Lipid unsaturation properties govern the sensitivity of membranes to photoinduced oxidative stress. Biophys. J. 116, 910–920 (2019).
Collard, L., Sinjab, F. & Notingher, I. Raman spectroscopy study of curvature-mediated lipid packing and sorting in single lipid vesicles. Biophys. J. 117, 1589–1598 (2019).
Bryce, D. A., Kitt, J. P., Myres, G. J. & Harris, J. M. Confocal Raman microscopy investigation of phospholipid monolayers deposited on nitrile-modified surfaces in porous silica particles. Langmuir 36, 4071–4079 (2020).
Krafft, C. et al. A specific spectral signature of serum and plasma-derived extracellular vesicles for cancer screening. Nanomed. Nanotechnol., Biol. Med. 13, 835–841 (2017).
Gualerzi, A. et al. Raman spectroscopy uncovers biochemical tissue-related features of extracellular vesicles from mesenchymal stromal cells. Sci. Rep. 7, 9820 (2017).
Gualerzi, A. et al. Raman spectroscopy as a quick tool to assess purity of extracellular vesicle preparations and predict their functionality. J. Extracell. Vesicles 8, 1568780 (2019).
Zhang, H., Silva, A. C., Zhang, W., Rutigliano, H. & Zhou, A. Raman spectroscopy characterization extracellular vesicles from bovine placenta and peripheral blood mononuclear cells. PLoS ONE 15, e0235214 (2020).
Morasso, C. F. et al. Raman spectroscopy reveals biochemical differences in plasma derived extracellular vesicles from sporadic amyotrophic lateral sclerosis patients. Nanomedicine 29, 102249 (2020).
Cialla, D., Pollok, S., Steinbrücker, C., Weber, K. & Popp, J. SERS-based detection of biomolecules. Nanophotonics 3, 383–411 (2014).
Lee, C. et al. 3D plasmonic nanobowl platform for the study of exosomes in solution. Nanoscale 7, 9290–9297 (2015).
Fazio, B. et al. SERS detection of biomolecules at physiological pH via aggregation of gold nanorods mediated by optical forces and plasmonic heating. Sci. Rep. 6, 26952 (2016).
Park, J. et al. Exosome classification by pattern analysis of surface-enhanced Raman spectroscopy data for lung cancer diagnosis. Anal. Chem. 89, 6695–6701 (2017).
Rojalin, T., Phong, B., Koster, H. & Carney, R. P. Nanoplasmonic approaches for sensitive detection and molecular characterization of extracellular vesicles. Front. Chem. 7, 729 (2019).
Wang, J., Koo, K. M., Wang, Y. & Trau, M. Engineering state-of-the-art plasmonic nanomaterials for SERS-based clinical liquid biopsy applications. Adv. Sci. 6, 1900730 (2019).
Pramanik, A. et al. Mixed-dimensional heterostructure material-based SERS for trace level identification of breast cancer-derived exosomes. ACS Omega 3, 16602–16611 (2020).
Zabeo, D. et al. Exosomes purified from a single cell type have diverse morphology. J. Extracell. Vesicles 6, 1329476 (2017).
Orlov, I. et al. The integrative role of cryo electron microscopy in molecular and cellular structural biology. Biol. Cell 109, 81–93 (2017).
Dubochet, J. et al. Cryo-electron microscopy of vitrified specimens. Q. Rev. Biophys. 21, 129–228 (1988).
Conde-Vancells, J. et al. Characterization and comprehensive proteome profiling of exosomes secreted by hepatocytes. J. Proteome Res. 7, 5157–5166 (2008).
Zonneveld, M. I. et al. Recovery of extracellular vesicles from human breast milk is influenced by sample collection and vesicle isolation procedures. J. Extracell. Vesicles https://doi.org/10.3402/jev.v3.24215 (2014).
Cizmar, P. & Yuana, Y. Detection and characterization of extracellular vesicles by transmission and cryo-transmission electron microscopy. in Extracellular Vesicles: Methods and Protocols (eds. Kuo, W. P. & Shidong, J.) 221–232 (Springer, 2017).
Binnig, G., Quate, F. & Gerber, C. Atomic force microscope. Phys. Rev. Lett. 56, 930–933 (1986).
Sharma, S. et al. Structural-mechanical characterization of nanoparticle exosomes in human saliva, using correlative AFM, FESEM, and force spectroscopy. ACS Nano 4, 1921–1926 (2010).
Parisse, P. et al. Atomic force microscopy analysis of extracellular vesicles. Eur. Biophys. J. 46, 813–820 (2017).
Creasey, R. et al. Atomic force microscopy-based antibody recognition imaging of proteins in the pathological deposits in pseudoexfoliation syndrome. Ultramicroscopy 111, 1055–1061 (2011).
Sebaihi, N., De Boeck, B., Yuana, Y., Nieuwland, R. & Pétry, J. Dimensional characterization of extracellular vesicles using atomic force microscopy. Meas. Sci. Technol. 28, 034006 (2017).
Skliar, M. & Chernyshev, V. S. Imaging of extracellular vesicles by atomic force microscopy. J. Vis. Exp. https://doi.org/10.3791/59254 (2019).
Kim, S. Y., Khanal, D., Tharkar, P., Kalionis, B. & Chrzanowski, W. None of us is the same as all of us: Resolving the heterogeneity of extracellular vesicles using single-vesicle, nanoscale characterization with resonance enhanced atomic force microscope infrared spectroscopy (AFM-IR). Nanoscale Horiz. 3, 430–438 (2018).
Avci, O., Ünlü, N. L., Özkumur, A. Y. & Ünlü, M. S. Interferometric reflectance imaging sensor (IRIS)—a platform technology for multiplexed diagnostics and digital detection. Sensors 15, 17649–17665 (2015).
Trueb, J. T., Avci, O., Sevenler, D., Connor, J. H. & Ünlü, M. S. Robust visualization and discrimination of nanoparticles by interferometric imaging. IEEE J. Sel. Top. Quantum Electron. https://ieeexplore.ieee.org/document/7782781 (2017).
Daaboul, G. G. et al. Enhanced light microscopy visualization of virus particles from Zika virus to filamentous ebolaviruses. PLoS ONE 12, e0179728 (2017).
van der Vlist, E. J., Nolte-’t Hoen, E. N. M., Stoorvogel, W., Arkesteijn, G. J. A. & Wauben, M. H. M. Fluorescent labeling of nano-sized vesicles released by cells and subsequent quantitative and qualitative analysis by high-resolution flow cytometry. Nat. Protoc. 7, 1311–1326 (2012).
Gomes, J. et al. Analytical considerations in nanoscale flow cytometry of extracellular vesicles to achieve data linearity. Thromb. Haemost. 118, 1612–1624 (2018).
Fish, K. N. Total internal reflection fluorescence (TIRF) microscopy. Curr. Protoc. Cytom. https://doi.org/10.1002/0471142956.cy1218s50 (2009).
Kudalkar, E. M., Davis, T. N. & Asbury, C. L. Single-molecule total internal reflection fluorescence microscopy. Cold Spring Harb. Protoc. 2016, pdb.top077800 (2016).
Axelrod, D. Chapter 7: total internal reflection fluorescence microscopy. Methods Cell Biol. 89, 169–221 (2008).
Ha, T. Single-molecule fluorescence resonance energy transfer. Methods 25, 78–86 (2001).
Arluison, V. & Wien, F. RNA Spectroscopy: Methods and Protocols (Springer, 2020).
Cerdán, L. et al. FRET-assisted laser emission in colloidal suspensions of dye-doped latex nanoparticles. Nat. Photonics 6, 621–626 (2012).
Rectenwald, J. et al. A general TR-FRET assay platform for high-throughput screening and characterizing inhibitors of methyl-lysine reader proteins. SLAS Discov. 24, 693–700 (2019).
Maurel, D. et al. Cell-surface protein-protein interaction analysis with time-resolved FRET and snap-tag technologies: application to GPCR oligomerization. Nat. Methods 5, 561–567 (2008).
Dao, T. P. T. et al. Mixing block copolymers with phospholipids at the nanoscale: from hybrid polymer/lipid wormlike micelles to vesicles presenting lipid nanodomains. Langmuir 33, 1705–1715 (2017).
Johnson, J. L. et al. Munc13-4 Is a Rab11-binding protein that regulates Rab11-positive vesicle trafficking and docking at the plasma membrane. J. Biol. Chem. 291, 3423–3438 (2016).
Gayrard, C. & Borghi, N. FRET-based molecular tension microscopy. Methods 94, 33–42 (2016).
Chen, C. et al. Visualization and intracellular dynamic tracking of exosomes and exosomal miRNAs using single molecule localization microscopy. Nanoscale 10, 5154–5162 (2018).
Oleksiuk, O. et al. Single-molecule localization microscopy allows for the analysis of cancer metastasis-specific mirna distribution on the nanoscale. Oncotarget 6, 44745–44757 (2015).
Dabrowska, S. et al. Imaging of extracellular vesicles derived from human bone marrow mesenchymal stem cells using fluorescent and magnetic labels. Int. J. Nanomed. 13, 1653–1664 (2018).
Willig, K. I., Rizzoli, S. O., Westphal, V., Jahn, R. & Hell, S. W. STED microscopy reveals that synaptotagmin remains clustered after synaptic vesicle exocytosis. Nature 440, 935–939 (2006).
Chen, C. et al. Imaging and intracellular tracking of cancer-derived exosomes using single-molecule localization-based super-resolution microscope. ACS Appl. Mater. Interfaces 8, 25825–25833 (2016).
Polanco, J. C., Li, C., Durisic, N., Sullivan, R. & Götz, J. Exosomes taken up by neurons hijack the endosomal pathway to spread to interconnected neurons. Acta Neuropathol. Commun. 6, 10 (2018).
Gustafsson, M. G. L. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Hell, S. W. Toward fluorescence nanoscopy. Nat. Biotechnol. 21, 1347–1355 (2003).
Huang, B. Super-resolution optical microscopy: multiple choices. Curr. Opin. Chem. Biol. 14, 10–14 (2010).
Hess, S. T., Girirajan, T. P. K. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Nienhaus, K. & Nienhaus, G. U. Where do we stand with super-resolution optical microscopy? J. Mol. Biol. 428, 308–322 (2016).
Bachmann, M., Fiederling, F. & Bastmeyer, M. Practical limitations of superresolution imaging due to conventional sample preparation revealed by a direct comparison of CLSM, SIM and dSTORM. J. Microsc. 262, 306–315 (2016).
Witters, D., Knez, K., Ceyssens, F., Puers, R. & Lammertyn, J. Digital microfluidics-enabled single-molecule detection by printing and sealing single magnetic beads in femtoliter droplets. Lab Chip 13, 2047–2054 (2013).
Gao, W., Li, X., Zeng, L. & Peng, T. Rapid isothermal detection assay: a probe amplification method for the detection of nucleic acids. Diagn. Microbiol. Infect. Dis. 60, 133–141 (2008).
Jia, S. et al. Emerging technologies in extracellular vesicle-based molecular diagnostics. Expert Rev. Mol. Diagn. 14, 307–321 (2014).
Chen, W. W. et al. BEAMing and droplet digital PCR analysis of mutant IDH1 mRNA in glioma patient serum and cerebrospinal fluid extracellular vesicles. Mol. Ther. Nucleic Acids 2, e109 (2013).
Worst, T. S. et al. miR-10a-5p and miR-29b-3p as extracellular vesicle-associated prostate cancer detection markers. Cancers (Basel) 12, 43 (2020).
Takahashi, K. et al. Circulating extracellular vesicle-encapsulated HULC is a potential biomarker for human pancreatic cancer. Cancer Sci. 111, 98–111 (2020).
Liu, C. et al. Single-exosome-counting immunoassays for cancer diagnostics. Nano Lett. 18, 4226–4232 (2018).
Diefenbach, R. J., Lee, J. H. & Rizos, H. Monitoring melanoma using circulating free DNA. Am. J. Clin. Dermatol. 20, 1–12 (2019).
Kong, L., Lee, C., Earhart, C. M., Cordovez, B. & Chan, J. W. A nanotweezer system for evanescent wave excited surface enhanced Raman spectroscopy (SERS) of single nanoparticles. Opt. Express 23, 6793 (2015).
Zong, S. et al. Facile detection of tumor-derived exosomes using magnetic nanobeads and SERS nanoprobes. Anal. Methods 8, 5001–5008 (2016).
Lee, C., Carney, R., Lam, K. & Chan, J. W. SERS analysis of selectively captured exosomes using an integrin-specific peptide ligand. J. Raman Spectrosc. 48, 1771–1776 (2017).
Tian, Y. F., Ning, C. F., He, F., Yin, B. C. & Ye, B. C. Highly sensitive detection of exosomes by SERS using gold nanostar@Raman reporter@nanoshell structures modified with a bivalent cholesterol-labeled DNA anchor. Analyst 143, 4915–4922 (2018).
Zhang, W. et al. Enabling sensitive phenotypic profiling of cancer-derived small extracellular vesicles using surface-enhanced raman spectroscopy nanotags. ACS Sens. 5, 764–771 (2020).
Schie, I. W. et al. High-throughput screening raman spectroscopy platform for label-free cellomics. Anal. Chem. 90, 2023–2030 (2018).
Xiong, Q. et al. Magnetic nanochain integrated microfluidic biochips. Nat. Commun. 9, 1743 (2018).
Beekman, P. et al. Immuno-capture of extracellular vesicles for individual multi-modal characterization using AFM, SEM and Raman spectroscopy. Lab Chip 19, 2526–2536 (2019).
Rüger, J., Mondol, A. S., Schie, I. W., Popp, J. & Krafft, C. High-throughput screening Raman microspectroscopy for assessment of drug-induced changes in diatom cells. Analyst 144, 4488–4492 (2019).
Noble, J. M. et al. Direct comparison of optical and electron microscopy methods for structural characterization of extracellular vesicles. J. Struct. Biol. https://doi.org/10.1016/j.jsb.2020.107474 (2020).
Lian, H., He, S., Chen, C. & Yan, X. Flow cytometric analysis of nanoscale biological particles and organelles. Annu. Rev. Anal. Chem. 12, 389–409 (2019).
Chukhchin, D. G., Bolotova, K., Sinelnikov, I., Churilov, D. & Novozhilov, E. Exosomes in the phloem and xylem of woody plants. Planta 251, 12 (2020).
Plaut, J. S. et al. Quantitative atomic force microscopy provides new insight into matrix vesicle mineralization. Arch. Biochem. Biophys. 667, 14–21 (2019).
Arraud, N. et al. Extracellular vesicles from blood plasma: determination of their morphology, size, phenotype and concentration. J. Thromb. Haemost. 12, 614–627 (2014).
Bevers, E. M., Comfurius, P. & Zwaal, R. F. A. Changes in membrane phospholipid distribution during platelet activation. Biochim. Biophys. Acta 736, 57–66 (1983).
Fadok, V. A. et al. Exposure of phosphatidylserine on the surface of apoptotic lymphocytes triggers specific recognition and removal by macrophages. J. Immunol. 148, 2207–2216 (1992).
Zwaal, R. F. A. & Schroit, A. J. Pathophysiologic implications of membrane phospholipid asymmetry in blood cells. J. Am. Soc. Hematol. 89, 333–340 (1997).
Biró, É. et al. Human cell-derived microparticles promote thrombus formation in vivo in a tissue factor-dependent manner. J. Thromb. Haemost. 1, 2561–2568 (2003).
Morel, O., Jesel, L., Freyssinet, J. M. & Toti, F. Cellular mechanisms underlying the formation of circulating microparticles. Arterioscler. Thromb. Vasc. Biol. 31, 15–26 (2011).
Emelyanov, A. et al. Cryo-electron microscopy of extracellular vesicles from cerebrospinal fluid. PLoS ONE 15, e0227949 (2020).
Yekula, A. et al. Large and small extracellular vesicles released by glioma cells in vitro and in vivo. J. Extracell. Vesicles 9, 1689784 (2020).
Thane, K. E., Davis, A. M. & Hoffman, A. M. Improved methods for fluorescent labeling and detection of single extracellular vesicles using nanoparticle tracking analysis. Sci. Rep. 9, 12295 (2019).
Dragovic, R. A. et al. Isolation of syncytiotrophoblast microvesicles and exosomes and their characterisation by multicolour flow cytometry and fluorescence nanoparticle tracking analysis. Methods 87, 64–74 (2015).
Koifman, N., Biran, I., Aharon, A., Brenner, B. & Talmon, Y. A direct-imaging cryo-EM study of shedding extracellular vesicles from leukemic monocytes. J. Struct. Biol. 198, 177–185 (2017).
LeClaire, M., Gimzewski, J. & Sharma, S. A review of the biomechanical properties of single extracellular vesicles. Nano Sel. https://doi.org/10.1002/nano.202000129 (2020).
Royo, F. et al. Differences in the metabolite composition and mechanical properties of extracellular vesicles secreted by hepatic cellular models. J. Extracell. Vesicles 8, 1575678 (2019).
Logozzi, M. et al. Microenvironmental pH and exosome levels interplay in human cancer cell lines of different histotypes. Cancers (Basel) 10, 370 (2018).
Royo, F. et al. Transcriptomic profiling of urine extracellular vesicles reveals alterations of CDH3 in prostate cancer. Oncotarget 7, 6835–6846 (2016).
Federici, C. et al. Exosome release and low pH belong to a framework of resistance of human melanoma cells to cisplatin. PLoS ONE 9, e88193 (2014).
Oosthuyzen, W. et al. Quantification of human urinary exosomes by nanoparticle tracking analysis. J. Physiol. 591, 5833–5842 (2013).
Logozzi, M. et al. Increased PSA expression on prostate cancer exosomes in in vitro condition and in cancer patients. Cancer Lett. 403, 318–329 (2017).
Logozzi, M., Spugnini, E., Mizzoni, D., Di Raimo, R. & Fais, S. Extracellular acidity and increased exosome release as key phenotypes of malignant tumors. Cancer Metastasis Rev. 38, 93–101 (2019).
Calorini, L., Peppicelli, S. & Bianchini, F. Extracellular acidity as favouring factor of tumor progression and metastatic dissemination. Exp. Oncol. 34, 79–84 (2012).
Huber, V. et al. Cancer acidity: an ultimate frontier of tumor immune escape and a novel target of immunomodulation. Semin. Cancer Biol. 43, 74–89 (2017).
Padda, R. S. et al. Nanoscale flow cytometry to distinguish subpopulations of prostate extracellular vesicles in patient plasma. Prostate 79, 592–603 (2019).
Xian, Y., Zhou, M., Han, S., Yang, R. & Wang, Y. A FRET biosensor reveals free zinc deficiency in diabetic beta-cell vesicles. Chin. Chem. Lett. 31, 468–472 (2020).
Nguyen, D. B. et al. Characterization of microvesicles released from human red blood cells. Cell. Physiol. Biochem. 38, 1085–1099 (2016).
Polanco, J. C., Scicluna, B. J., Hill, A. F. & Götz, J. Extracellular vesicles isolated from the brains of rTg4510 mice seed tau protein aggregation in a threshold-dependent manner. J. Biol. Chem. 291, 12445–12466 (2016).
Wang, Y. et al. The release and trans-synaptic transmission of Tau via exosomes. Mol. Neurodegener. 12, 5 (2017).
Yang, J. E. et al. Complexity and ultrastructure of infectious extracellular vesicles from cells infected by non-enveloped virus. Sci. Rep. 10, 7939 (2020).
Santos, M. F. et al. VAMP-associated protein-A and oxysterol-binding protein–related protein 3 promote the entry of late endosomes into the nucleoplasmic reticulum. J. Biol. Chem. 293, 13834–13848 (2018).
Mannavola, F. et al. Tumor-derived exosomes promote the in vitro osteotropism of melanoma cells by activating the SDF-1/CXCR4/CXCR7 axis. J. Transl. Med. 17, 230 (2019).
Sorkin, R. et al. Nanomechanics of extracellular vesicles reveals vesiculation pathways. Small 14, e1801650 (2018).
Vorselen, D. et al. The fluid membrane determines mechanics of erythrocyte extracellular vesicles and is softened in hereditary spherocytosis. Nat. Commun. 9, 4960 (2018).
Böcking, T., Upadhyayula, S., Rapoport, I., Capraro, B. R. & Kirchhausen, T. Reconstitution of clathrin coat disassembly for fluorescence microscopy and single-molecule analysis. Methods Mol. Biol. 1847, 121–146 (2018).
Mattheyses, A. L., Atkinson, C. E. & Simon, S. M. Imaging single endocytic events reveals diversity in clathrin, dynamin, and vesicle dynamics. Traffic 12, 1394–1406 (2011).
Van Lengerich, B., Rawle, R. J., Bendix, P. M. & Boxer, S. G. Individual vesicle fusion events mediated by lipid-anchored DNA. Biophys. J. 105, 409–419 (2013).
Mattie, S., Kazmirchuk, T., Mui, J., Vali, H. & Brett, C. L. Visualization of SNARE-mediated organelle membrane hemifusion by electron microscopy. Methods Mol. Biol. 1860, 361–377 (2019).
Hu, Y., Tian, Z. & Diao, J. Single-molecule fluorescence measurement of SNARE-mediated vesicle fusion. in SNAREs: Methods and Protocols (ed. Fratti, R.) 335–344 (2019).
Lin, C. C. et al. Control of membrane gaps by synaptotagmin-Ca 2+ measured with a novel membrane distance ruler. Nat. Commun. 5, 5859 (2014).
Stratton, B. S. et al. Cholesterol increases the openness of SNARE-mediated flickering fusion pores. Biophys. J. 110, 1538–1550 (2016).
Cao, H. et al. In vivo real-time imaging of extracellular vesicles in liver regeneration via aggregation-induced emission luminogens. ACS Nano 13, 3522–3533 (2019).
Lai, C. P. et al. Dynamic biodistribution of extracellular vesicles in vivo using a multimodal imaging reporter. ACS Nano 8, 483–494 (2014).
Gangadaran, P., Hong, C. M. & Ahn, B. C. Current perspectives on in vivo noninvasive tracking of extracellular vesicles with molecular imaging. Biomed Res. Int. 2017, (2017).
Lai, C. P., Tannous, B. A. & Breakefield, X. O. Noninvasive in vivo monitoring of extracellular vesicles. in. Methods Mol. Biol. 1098, 249–258 (2014).
Van Der Vos, K. E. et al. Directly visualized glioblastoma-derived extracellular vesicles transfer RNA to microglia/macrophages in the brain. Neuro. Oncol. 18, 58–69 (2016).
Ricklefs, F. L. et al. Imaging flow cytometry facilitates multiparametric characterization of extracellular vesicles in malignant brain tumours. J. Extracell. Vesicles 8, 1588555 (2019).
Lai, C. P. et al. Visualization and tracking of tumour extracellular vesicle delivery and RNA translation using multiplexed reporters. Nat. Commun. 6, 7029 (2015).
Verweij, F. J., Hyenne, V., Van Niel, G. & Goetz, J. G. Extracellular vesicles: catching the light in zebrafish. Trends Cell Biol. 29, 770–776 (2019).
Kobayashi-Sun, J. et al. Uptake of osteoblast-derived extracellular vesicles promotes the differentiation of osteoclasts in the zebrafish scale. Commun. Biol. 3, 190 (2020).
Sung, B. H. et al. A live cell reporter of exosome secretion and uptake reveals pathfinding behavior of migrating cells. Nat. Commun. 11, 2092 (2020).
Clos-Garcia, M. et al. Gut microbiome and serum metabolome analyses identify molecular biomarkers and altered glutamate metabolism in fibromyalgia. EBioMedicine 46, 499–511 (2019).
Roman-Canal, B. et al. EV-associated miRNAs from peritoneal lavage are a source of biomarkers in endometrial cancer. Cancers (Basel). 11, 839 (2019).
Tian, Y. et al. Protein profiling and sizing of extracellular vesicles from colorectal cancer patients via flow cytometry. ACS Nano 12, 671–680 (2018).
Clos-Garcia, M. et al. Integrative analysis of fecal metagenomics and metabolomics in colorectal cancer. Cancers (Basel) 12, 1142 (2020).
Royo, F. & Falcon-Perez, J. M. Liver extracellular vesicles in health and disease. J. Extracell. Vesicles https://doi.org/10.3402/jev.v1i0.18825 (2012).
He, D. et al. Total internal reflection-based single-vesicle in situ quantitative and stoichiometric analysis of tumor-derived exosomal microRNAs for diagnosis and treatment monitoring. Theranostics 9, 4494–4507 (2019).
Murakami, Y. et al. Comprehensive miRNA expression analysis in peripheral blood can diagnose liver disease. PLoS ONE 7, e48366 (2012).
Pang, B. et al. Extracellular vesicles: the next generation of biomarkers for liquid biopsy-based prostate cancer diagnosis. Theranostics 10, 2309–2326 (2020).
Vlaeminck-Guillem, V. Extracellular vesicles in prostate cancer carcinogenesis, diagnosis, and management. Front. Oncol. 8, 222 (2018).
Mateo, L., Guitart-Pla, O., Duran-Frigola, M. & Aloy, P. Exploring the OncoGenomic Landscape of cancer. Genome Med. 10, 61 (2018).
Joncas, F. H. et al. Plasma extracellular vesicles as phenotypic biomarkers in prostate cancer patients. Prostate 79, 1767–1776 (2019).
Carlsson, J. et al. Validation of suitable endogenous control genes for expression studies of miRNA in prostate cancer tissues. Cancer Genet. Cytogenet. 202, 71–75 (2010).
Schaefer, A. et al. Suitable reference genes for relative quantification of miRNA expression in prostate cancer. Exp. Mol. Med. 42, 749–758 (2010).
Haka, A. S. et al. Diagnosing breast cancer by using Raman spectroscopy. Proc. Natl Acad. Sci. USA 102, 12371–12376 (2005).
Haka, A. S. et al. Diagnosing breast cancer using Raman spectroscopy: prospective analysis. J. Biomed. Opt. 14, 054023 (2009).
Notarangelo, M. et al. Ultrasensitive detection of cancer biomarkers by nickel-based isolation of polydisperse extracellular vesicles from blood. EBioMedicine 43, 114–126 (2019).
Melo, S. A. et al. Glypican1 identifies cancer exosomes and facilitates early detection of cancer. Nature 523, 177–182 (2015).
Biggs, C. N. et al. Prostate extracellular vesicles in patient plasma as a liquid biopsy platform for prostate cancer using nanoscale flow cytometry. Oncotarget 7, 8839–8849 (2016).
Cannon, D. M., Winograd, J. N. & Ewing, A. G. Quantitative chemical analysis of single cells. Annu. Rev. Biophys. Biomol. Struct. 29, 239–263 (2000).
Li, X., Dunevall, J. & Ewing, A. G. Quantitative chemical measurements of vesicular transmitters with electrochemical cytometry. Acc. Chem. Res. 49, 2347–2354 (2016).
Li, X., Dunevall, J., Ren, L. & Ewing, A. G. Mechanistic aspects of vesicle opening during analysis with vesicle impact electrochemical cytometry. Anal. Chem. 89, 9416–9423 (2017).
Ranjbari, E. et al. Direct measurement of total vesicular catecholamine content with electrochemical microwell arrays. Anal. Chem. 92, 11325–11331 (2020).
Dunevall, J., Majdi, S., Larsson, A. & Ewing, A. Vesicle impact electrochemical cytometry compared to amperometric exocytosis measurements. Curr. Opin. Electrochem 5, 85–91 (2017).
Li, X., Majdi, S., Dunevall, J., Fathali, H. & Ewing, A. G. Quantitative measurements of transmitters in vesicles one at a time in single cell cytoplasm with nano-tip electrodes. Angew. Chem. Int. Ed. Engl. 54, 11978–11982 (2015).
Li, X., Dunevall, J. & Ewing, A. G. Electrochemical quantification of transmitter concentration in single nanoscale vesicles isolated from PC12 cells. Faraday Discuss 210, 353–364 (2018).
Ren, L. et al. Zinc regulates chemical-transmitter storage in nanometer vesicles and exocytosis dynamics as measured by amperometry. Angew. Chem. Int. Ed. Engl. 56, 4970–4975 (2017).
Li, X., Dunevall, J. & Ewing, A. G. Using single-cell amperometry to reveal how cisplatin treatment modulates the release of catecholamine transmitters during exocytosis. Angew. Chem. 128, 9187–9190 (2016).
Zupanc, J., Bas, E. & Erdogmus, D. Analysis of lipid vesicle populations from microscopy video sequences. 2010 Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. EMBC’10 5050–5053 https://doi.org/10.1109/IEMBS.2010.5626223 (2010)
Barriere, H. & Lukacs, G. L. Analysis of endocytic trafficking by single-cell fluorescence ratio imaging. Curr. Protoc. Cell Biol. https://doi.org/10.1002/0471143030.cb1513s40 (2008)
Chen, T. et al. Microwave biosensor dedicated to the dielectric spectroscopy of a single alive biological cell in its culture medium. in Microwave Symposium Digest (IMS), 2013 IEEE MTT-S International (2013).
Chen, W., Dubuc, D. & Grenier, K. Parametric study of a microwave sensor dedicated to the dielectric spectroscopy of single particles and biological cells. in 2015 European Microwave Conference (EuMC 2015) 829–832 (2015).
Chen, W., Dubuc, D. & Grenier, K. Impact of sensor metal thickness on microwave spectroscopy sensitivity for individual particles and biological cells analysis. in 2016 IEEE Topical Conference on Biomedical Wireless Technologies, Networks, and Sensing Systems (BioWireleSS) 81–83 (2016).
Zhang, M. et al. Electrically controlled tunable broadband interferometric dielectric spectroscopy: groundwork for single cell analysis. in 2019 49th European Microwave Conference (EuMC) 650–653 (2019).
Cui, Y. et al. Analyzing single giant unilamellar vesicles with a slotline-based RF nanometer sensor. IEEE Trans. Microw. Theory Tech. 64, 1339–1347 (2016).
Wu, M. et al. Isolation of exosomes from whole blood by integrating acoustics and microfluidics. Proc. Natl Acad. Sci. USA 114, 10584–10589 (2017).
Ku, A. et al. Acoustic enrichment of extracellular vesicles from biological fluids. Anal. Chem. 90, 8011–8019 (2018).
Ku, A. et al. A urinary extracellular vesicle microRNA biomarker discovery pipeline; from automated extracellular vesicle enrichment by acoustic trapping to microRNA sequencing. PLoS ONE 14, e0217507 (2019).
Carter, E. P. et al. Visualizing Ebolavirus particles using single-particle interferometric reflectance imaging sensor (SP-IRIS). in Ebolaviruses: Methods and Protocols (eds. Groseth, A. & Hoenen, T.) 373–393 (Springer, 2017).
Ünlü, N. L., Kanik, F. E., Seymour, E., Connor, J. H. & Ünlü, M. S. DNA-directed antibody immobilization for robust protein microarrays: application to single particle detection DNA-directed antibody immobilization. in Biosensors and Biodetection: Methods and Protocols (eds. Rasooly, A. & Prickril, B.) 187–206 (Springer, 2017).
Akagi, T. et al. On-chip immunoelectrophoresis of extracellular vesicles released from human breast cancer cells. PLoS ONE 10, e0123603 (2015).
Friedrich, R. et al. A nano flow cytometer for single lipid vesicle analysis. Lab Chip 17, 830–841 (2017).
Obeid, S. et al. NanoBioAnalytical characterization of extracellular vesicles in 75-nm nanofiltered human plasma for transfusion: a tool to improve transfusion safety. Nanomedicine 20, 10197 (2019).
Yokota, S. et al. Extracellular vesicles nanoarray technology: immobilization of individual extracellular vesicles on nanopatterned polyethylene glycol-lipid conjugate brushes. PLoS ONE 14, e0224091 (2019).
Ji, Y. et al. Multiplexed profiling of single-cell extracellular vesicles secretion. Proc. Natl Acad. Sci. USA 116, 5979–5984 (2019).
Bai, Y. et al. Absolute quantification and analysis of extracellular vesicle lncRNAs from the peripheral blood of patients with lung cancer based on multi-colour fluorescence chip-based digital PCR. Biosens. Bioelectron. 142, 111523 (2019).
Akagi, T., Kato, K., Hanamura, N., Kobayashi, M. & Ichiki, T. Evaluation of desialylation effect on zeta potential of extracellular vesicles secreted from human prostate cancer cells by on-chip microcapillary electrophoresis. Jpn. J. Appl. Phys. 53, 06JL01 (2014).
Weber, A., Wehmeyer, J. C., Schmidt, V., Lichtenberg, A. & Akhyari, P. Rapid fluorescence-based characterization of single extracellular vesicles in human blood with nanoparticle-tracking analysis. J. Vis. Exp. https://doi.org/10.3791/58731 (2019).
Marku, A. et al. The LRRK2 N-terminal domain influences vesicle trafficking: impact of the E193K variant. Sci. Rep. 10, 3799 (2020).
Acknowledgements
The authors of this review were supported by funds from the European Union’s Horizon 2020 research and innovation programme under grant agreement no. 860303. We thank MINECO for the TenTaCles (Spanish Excellence Network in Exosomes) and the Severo Ochoa Excellence Accreditation (SEV-2016-0644). This project has received funding from the Spanish Ministry of Economy and Competitiveness MINECO (RTI2018-094969-B-I00).
Author information
Authors and Affiliations
Contributions
All of the authors wrote, edited and discussed this review.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Paolo Bergese, Cees Otto and Frederik Johannes Verweij for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Kruglik, S. G. et al. Nanoscale 11, 1661–1679 (2019): https://doi.org/10.1039/C8NR04677H
Royo, F. et al. J. Extracell. Vesicles 8, 1575678 (2019): https://doi.org/10.1080/20013078.2019.1575678
Tatischeff, I., Larquet, E., Falcon-Perez, J. M., Turpin, P.-Y. & Kruglik, S. G. J. Extracell. Vesicles 1, 19179 (2012): https://doi.org/10.3402/jev.v1i0.19179
Rights and permissions
About this article
Cite this article
Bordanaba-Florit, G., Royo, F., Kruglik, S.G. et al. Using single-vesicle technologies to unravel the heterogeneity of extracellular vesicles. Nat Protoc 16, 3163–3185 (2021). https://doi.org/10.1038/s41596-021-00551-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00551-z
This article is cited by
-
Immunophenotype profile by flow cytometry reveals different subtypes of extracellular vesicles in porcine seminal plasma
Cell Communication and Signaling (2024)
-
Exploring capabilities of elemental mass spectrometry for determination of metal and biomolecules in extracellular vesicles
Analytical and Bioanalytical Chemistry (2024)
-
Engineering of Cell Derived-Nanovesicle as an Alternative to Exosome Therapy
Tissue Engineering and Regenerative Medicine (2024)
-
Therapeutic potential of MSCs and MSC-derived extracellular vesicles in immune thrombocytopenia
Stem Cell Research & Therapy (2023)
-
Advanced extracellular vesicle bioinformatic nanomaterials: from enrichment, decoding to clinical diagnostics
Journal of Nanobiotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.