Abstract
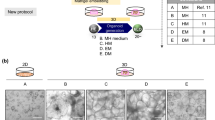
Hepatic stellate cells (HSCs) are nonparenchymal liver cells responsible for extracellular matrix homeostasis and are the main cells involved in the development of liver fibrosis following injury. The lack of reliable sources of HSCs has hence limited the development of complex in vitro systems to model liver diseases and toxicity. Here we describe a protocol to differentiate human induced pluripotent stem cells (iPSCs) into hepatic stellate cells (iPSC-HSCs). The protocol is based on the addition of several growth factors important for liver development sequentially over 12 d. iPSC-HSCs present phenotypic and functional characteristics of primary HSCs and can be expanded or frozen and used to perform high-throughput in vitro studies. We also describe how to coculture iPSC-HSCs with hepatocytes, which self-assemble into three-dimensional (3D) hepatic spheroids. This protocol enables the generation of HSC-like cells for in vitro modeling and drug screening studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The authors declare that all data supporting the findings of this study are available within the paper. Source data are provided with this paper.
References
Hernandez-Gea, V. & Friedman, S. L. Pathogenesis of liver fibrosis. Annu. Rev. Pathol. https://doi.org/10.1146/annurev-pathol-011110-130246 (2011).
Friedman, S. L. Hepatic stellate cells: protean, multifunctional, and enigmatic cells of the liver. Physiol. Rev. https://doi.org/10.1152/physrev.00013.2007 (2008).
Mederacke, I. et al. Fate tracing reveals hepatic stellate cells as dominant contributors to liver fibrosis independent of its aetiology. Nat. Commun. https://doi.org/10.1038/ncomms3823 (2013).
Issa, R. et al. Apoptosis of hepatic stellate cells: involvement in resolution of biliary fibrosis and regulation by soluble growth factors. Gut https://doi.org/10.1136/gut.48.4.548 (2001).
van Grunsven, L. A. 3D in vitro models of liver fibrosis. Adv. Drug Deliv. Rev. https://doi.org/10.1016/j.addr.2017.07.004 (2017).
Lauschke, V. M. et al. Novel 3D culture systems for studies of human liver function and assessments of the hepatotoxicity of drugs and drug candidates. Chem. Res. Toxicol. https://doi.org/10.1021/acs.chemrestox.6b00150 (2016).
Sancho-Bru, P. et al. Genomic and functional characterization of stellate cells isolated from human cirrhotic livers. J. Hepatol. https://doi.org/10.1016/j.jhep.2005.02.035 (2005).
Perea, L., Coll, M. & Sancho-Bru, P. Assessment of liver fibrotic insults in vitro. in Protocols in In Vitro Hepatocyte Research (eds. Vinken, M. & Rogiers, V.) 391–401 (Springer, 2015). https://doi.org/10.1007/978-1-4939-2074-7_30
Xu, L. et al. Human hepatic stellate cell lines, LX-1 and LX-2: new tools for analysis of hepatic fibrosis. Gut https://doi.org/10.1136/gut.2004.042127 (2005).
Herrmann, J., Gressner, A. M. & Weiskirchen, R. Immortal hepatic stellate cell lines: useful tools to study hepatic stellate cell biology and function? J. Cell. Mol. Med. https://doi.org/10.1111/j.1582-4934.2007.00060.x (2007).
Nishio, M. et al. Feeder-free and serum-free production of hepatocytes, cholangiocytes, and their proliferating progenitors from human pluripotent stem cells: application to liver-specific functional and cytotoxic assays. Cell. Rep. https://doi.org/10.1089/cell.2011.0064 (2018).
Takebe, T. et al. Vascularized and functional human liver from an iPSC-derived organ bud transplant. Nature https://doi.org/10.1038/nature12271 (2013).
Sancho-Bru, P. et al. Directed differentiation of murine-induced pluripotent stem cells to functional hepatocyte-like cells. J. Hepatol. https://doi.org/10.1016/j.jhep.2010.06.014 (2011).
Coll, M. et al. Generation of hepatic stellate cells from human pluripotent stem cells enables in vitro modeling of liver fibrosis. Cell Stem Cell https://doi.org/10.1016/j.stem.2018.05.027 (2018).
Asahina, K., Zhou, B., Pu, W. T. & Tsukamoto, H. Septum transversum-derived mesothelium gives rise to hepatic stellate cells and perivascular mesenchymal cells in developing mouse liver. Hepatology https://doi.org/10.1002/hep.24119 (2011).
Asahina, K. et al. Mesenchymal origin of hepatic stellate cells, submesothelial cells, and perivascular mesenchymal cells during mouse liver development. Hepatology https://doi.org/10.1002/hep.24119 (2009).
Richter, A. et al. BMP4 promotes EMT and mesodermal commitment in human embryonic stem cells via SLUG and MSX2. Stem Cells https://doi.org/10.1002/stem.1592 (2014).
Evseenko, D. et al. Mapping the first stages of mesoderm commitment during differentiation of human embryonic stem cells. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.1002077107 (2010).
Ng, F. et al. PDGF, TGF-β, and FGF signaling is important for differentiation and growth of mesenchymal stem cells (MSCs): transcriptional profiling can identify markers and signaling pathways important in differentiation of MSCs into adipogenic, chondrogenic, and osteogenic lineages. Blood https://doi.org/10.1182/blood-2007-07-103697 (2008).
Wang, Y., Yu, X., Chen, E. & Li, L. Liver-derived human mesenchymal stem cells: A novel therapeutic source for liver diseases. Stem Cell Res. Ther. https://doi.org/10.1186/s13287-016-0330-3 (2016).
Yamaguchi, T. P., Harpal, K., Henkemeyer, M. & Rossant, J. fgfr-1 is required for embryonic growth and mesodermal patterning during mouse gastrulation. Genes Dev. https://doi.org/10.1101/gad.8.24.3032 (1994).
Dorey, K. & Amaya, E. FGF signalling: diverse roles during early vertebrate embryogenesis. Development https://doi.org/10.1242/dev.037689 (2010).
Schuldiner, M., Yanuka, O., Itskovitz-Eldor, J., Melton, D. & Benvenisty, N. Effects of eight growth factors on the differentiation of cells derived from human embryonic stem cells. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.97.21.11307 (2000).
El Taghdouini, A. et al. In vitro reversion of activated primary human hepatic stellate cells. Fibrogenesis Tissue Repair https://doi.org/10.1186/s13069-015-0031-z (2015).
Senoo, H. et al. Hepatic stellate cell (vitamin A–storing cell) and its relative—past, present and future. Cell Biol. Int. https://doi.org/10.1042/Cbi20100321 (2010).
Ijpenberg, A. et al. Wt1 and retinoic acid signaling are essential for stellate cell development and liver morphogenesis. Dev. Biol. https://doi.org/10.1016/j.ydbio.2007.09.014 (2007).
Chun, Y. S., Byun, K. & Lee, B. Induced pluripotent stem cells and personalized medicine: current progress and future perspectives. Anat. Cell Biol. https://doi.org/10.5115/acb.2011.44.4.245 (2011).
Koui, Y. et al. An in vitro human liver model by iPSC-derived parenchymal and non-parenchymal cells. Stem Cell Rep. https://doi.org/10.1016/j.stemcr.2017.06.010 (2017).
Miyoshi, M. et al. LIM homeobox 2 promotes interaction between human iPS-derived hepatic progenitors and iPS-derived hepatic stellate-like cells. Sci. Rep. https://doi.org/10.1038/s41598-018-37430-9 (2019).
Lee, U. E. & Friedman, S. L. Mechanisms of hepatic fibrogenesis. Best Pract. Res. Clin. Gastroenterol. https://doi.org/10.1016/j.bpg.2011.02.005 (2011).
Dongiovanni, P., Anstee, Q. & Valenti, L. Genetic predisposition in NAFLD and NASH: impact on severity of liver disease and response to treatment. Curr. Pharm. Des. https://doi.org/10.2174/13816128113199990381 (2013).
Ouchi, R. et al. Modeling steatohepatitis in humans with pluripotent stem cell-derived organoids. Cell Metab. https://doi.org/10.1016/j.cmet.2019.05.007 (2019).
Farahani, R. M. & Xaymardan, M. Platelet-derived growth factor receptor alpha as a marker of mesenchymal stem cells in development and stem cell biology. Stem Cells Int. https://doi.org/10.1155/2015/362753 (2015).
Ullah, I., Subbarao, R. B. & Rho, G. J. Human mesenchymal stem cells—current trends and future prospective. Biosci. Rep. https://doi.org/10.1042/BSR20150025 (2015).
Kumar, M. et al. A fully defined pluripotent stem cell derived multi-liver-cell model for steatohepatitis and fibrosis. Preprint at bioRxiv https://doi.org/10.1101/2020.09.03.280883 (2020).
Ordovás, L. et al. Efficient recombinase-mediated cassette exchange in hPSCs to study the hepatocyte lineage reveals AAVS1 locus-mediated transgene inhibition. Stem Cell Rep. https://doi.org/10.1016/j.stemcr.2015.09.004 (2015).
Leite, S. B. et al. Novel human hepatic organoid model enables testing of drug-induced liver fibrosis in vitro. Biomaterials https://doi.org/10.1016/j.biomaterials.2015.11.026 (2016).
Acknowledgements
This work was performed in Center Esther Koplowitz and Vrije Universiteit Brussel. We thank the Advanced Optical Microscopy Unit from the Scientific and Technological Centers of the University of Barcelona for their support with the confocal image technique. We are indebted to the Cytomics Unit of the Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS) for flow cytometry analysis. Figures were created using BioRender.com. P.S.-B. is supported by the Fondo de Investigación Sanitaria Carlos III, cofinanced by the Fondo Europeo de Desarrollo Regional (FEDER), Unión Europea, ‘Una manera de hacer Europa’ (FIS PI20/00765, PI17/00673), DTS18/00088, COST Action H2020 PRO-EURO-DILI-NET CA17112 and Miguel Servet (CPII16/00041) to P.S.-B., PFIS (FI18/00215) to R.M. and APIF to J.V. MC FIS (PI18/00862). I.M. is supported by an FWO–V post-doctoral fellowship (12N5419 N) and S.V. is supported by an FWO–V postdoctoral fellowship (1243121N). L.A.v.G is supported by FWO projects G042719N, FWO-SBO-S001121 ‘iPSC-LiMiC’ and a ‘Wetenschappelijk Fonds Willy Gepts’ grant from the Vrije Universiteit Brussel. C.M.V. is supported by FWO-G0D9917N; IWT-140045, HILIM-3D; FWO-SBO-S001121, iPSC-LiMiC and the European Union’s Horizon 2020 research and innovation program under grant agreement 681002 (EU-ToxRisk).
Author information
Authors and Affiliations
Contributions
J.V. and R.A.M. equally contributed to the optimization of expansion and freezing processes, characterization of the cells and drafting the manuscript. I.M. and A.S. performed the 3D spheroid experiments, and S.V. and. A.S. carried out parallel iPSC differentiations using a different iPSC line (not included in the manuscript). M.C. participated in the differentiation protocol design and reviewed the manuscript. S.A., T.R.-T., B.A.-B., C.M.-S. and D.B., participated in the differentiation protocol process. C.M.V. contributed to the design of the IPSCs differentiation protocol and reviewed the manuscript. L.A.v.G. guided the iPSC and spheroid studies in Brussels and critically reviewed the manuscript. P.S.-B. designed and supervised the study and critically reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
M.C., C.M.V. and P.S.-B. have a patent application PCT/EP2016/079464, and L.A.v.G. has a patent application PCT/EP2015/062551.
Additional information
Peer review information Nature Protocols thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Coll, M. et al. Cell Stem Cell 23, 101–113.e7 (2018): https://doi.org/10.1016/j.stem.2018.05.027
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 4
Statistical source data.
Rights and permissions
About this article
Cite this article
Vallverdú, J., Martínez García de la Torre, R.A., Mannaerts, I. et al. Directed differentiation of human induced pluripotent stem cells to hepatic stellate cells. Nat Protoc 16, 2542–2563 (2021). https://doi.org/10.1038/s41596-021-00509-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00509-1
This article is cited by
-
The Current Proceedings of PSC-Based Liver Fibrosis Therapy
Stem Cell Reviews and Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.