Abstract
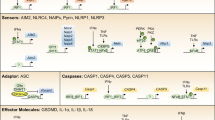
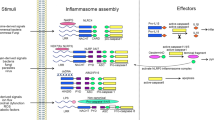
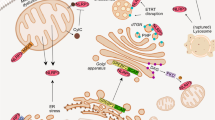
Inflammasomes are multimeric heterogeneous mega-Dalton protein complexes that play key roles in the host innate immune response to infection and sterile insults. Assembly of the inflammasome complex following infection or injury begins with the oligomerization of the upstream inflammasome-forming sensor and proceeds through a multistep process of well-coordinated events and downstream effector functions. Together, these steps enable elegant experimental readouts with which to reliably assess the successful activation of the inflammasome complex and cell death. Here, we describe a comprehensive protocol that details several in vitro (in bone marrow–derived macrophages) and in vivo (in mice) strategies for activating the inflammasome and explain how to subsequently assess multiple downstream effects in parallel to unequivocally establish the activation status of the inflammasome and cell death pathways. Our workflow assesses inflammasome activation via the formation of the apoptosis-associated speck-like protein containing a CARD (ASC) speck; cleavage of caspase-1 and gasdermin D; release of IL-1β, IL-18, caspase-1, and lactate dehydrogenase from the cell; and real-time analysis of cell death by imaging. Analyses take up to ~24 h to complete. Overall, our multifaceted approach provides a comprehensive and consistent protocol for assessing inflammasome activation and cell death.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files. No datasets were generated or analyzed during the current study. All source data underlying the figures are available from the corresponding author upon request.
References
Kesavardhana, S. & Kanneganti, T. D. Mechanisms governing inflammasome activation, assembly and pyroptosis induction. Int. Immunol. 29, 201–210 (2017).
Man, S. M., Karki, R. & Kanneganti, T. D. Molecular mechanisms and functions of pyroptosis, inflammatory caspases and inflammasomes in infectious diseases. Immunol. Rev. 277, 61–75 (2017).
Martinon, F., Burns, K. & Tschopp, J. The inflammasome: a molecular platform triggering activation of inflammatory caspases and processing of proIL-beta. Mol. Cell. 10, 417–426 (2002).
Kanneganti, T. D., Lamkanfi, M. & Nunez, G. Intracellular NOD-like receptors in host defense and disease. Immunity 27, 549–559 (2007).
Liston, A. & Masters, S. L. Homeostasis-altering molecular processes as mechanisms of inflammasome activation. Nat. Rev. Immunol. 17, 208–214 (2017).
Tartey, S. & Kanneganti, T. D. Inflammasomes in the pathophysiology of autoinflammatory syndromes. J. Leukoc. Biol. 107, 379–391 (2020).
Man, S. M. & Kanneganti, T. D. Converging roles of caspases in inflammasome activation, cell death and innate immunity. Nat. Rev. Immunol. 16, 7–21 (2016).
Place D. E. & Kanneganti, T. D. Cell death-mediated cytokine release and its therapeutic implications. J. Exp. Med. 216, 1474-1486 (2019).
Brydges, S. D. et al. Divergence of IL-1, IL-18, and cell death in NLRP3 inflammasomopathies. J. Clin. Invest. 123, 4695–4705 (2013).
Hoffman, H. M., Mueller, J. L., Broide, D. H., Wanderer, A. A. & Kolodner, R. D. Mutation of a new gene encoding a putative pyrin-like protein causes familial cold autoinflammatory syndrome and Muckle-Wells syndrome. Nat. Genet. 29, 301–305 (2001).
Brydges, S. D. et al. Inflammasome-mediated disease animal models reveal roles for innate but not adaptive immunity. Immunity 30, 875–887 (2009).
Sharma, D., Sharma, B. R., Vogel, P. & Kanneganti, T. D. IL-1beta and caspase-1 drive autoinflammatory disease independently of IL-1alpha or caspase-8 in a mouse model of familial mediterranean fever. Am. J. Pathol. 187, 236–244 (2017).
Chae, J. J. et al. Gain-of-function Pyrin mutations induce NLRP3 protein-independent interleukin-1beta activation and severe autoinflammation in mice. Immunity 34, 755–768 (2011).
Brenner, M., Ruzicka, T., Plewig, G., Thomas, P. & Herzer, P. Targeted treatment of pyoderma gangrenosum in PAPA (pyogenic arthritis, pyoderma gangrenosum and acne) syndrome with the recombinant human interleukin-1 receptor antagonist anakinra. Br. J. Dermatol. 161, 1199–1201 (2009).
Yu, J. W. et al. Pyrin activates the ASC pyroptosome in response to engagement by autoinflammatory PSTPIP1 mutants. Mol. Cell 28, 214–227 (2007).
Dierselhuis, M. P., Frenkel, J., Wulffraat, N. M. & Boelens, J. J. Anakinra for flares of pyogenic arthritis in PAPA syndrome. Rheumatol. (Oxf.) 44, 406–408 (2005).
Karki, R. & Kanneganti, T. D. Diverging inflammasome signals in tumorigenesis and potential targeting. Nat. Rev. Cancer 19, 197–214 (2019).
Sharma, D. & Kanneganti, T. D. The cell biology of inflammasomes: mechanisms of inflammasome activation and regulation. J. Cell Biol. 213, 617–629 (2016).
Boyden, E. D. & Dietrich, W. F. Nalp1b controls mouse macrophage susceptibility to anthrax lethal toxin. Nat. Genet. 38, 240–244 (2006).
Kanneganti, T. D. et al. Bacterial RNA and small antiviral compounds activate caspase-1 through cryopyrin/Nalp3. Nature 440, 233–236 (2006).
Sutterwala, F. S. et al. Critical role for NALP3/CIAS1/cryopyrin in innate and adaptive immunity through its regulation of caspase-1. Immunity 24, 317–327 (2006).
Mariathasan, S. et al. Cryopyrin activates the inflammasome in response to toxins and ATP. Nature 440, 228–232 (2006).
Martinon, F., Agostini, L., Meylan, E. & Tschopp, J. Identification of bacterial muramyl dipeptide as activator of the NALP3/cryopyrin inflammasome. Curr. Biol. 14, 1929–1934 (2004).
Martinon, F., Petrilli, V., Mayor, A., Tardivel, A. & Tschopp, J. Gout-associated uric acid crystals activate the NALP3 inflammasome. Nature 440, 237–241 (2006).
Franchi, L. et al. Cytosolic flagellin requires Ipaf for activation of caspase-1 and interleukin 1beta in salmonella-infected macrophages. Nat. Immunol. 7, 576–582 (2006).
Miao, E. A. et al. Cytoplasmic flagellin activates caspase-1 and secretion of interleukin 1beta via Ipaf. Nat. Immunol. 7, 569–575 (2006).
Miao, E. A. et al. Innate immune detection of the type III secretion apparatus through the NLRC4 inflammasome. Proc. Natl Acad. Sci. USA 107, 3076–3080 (2010).
Zhao, Y. et al. The NLRC4 inflammasome receptors for bacterial flagellin and type III secretion apparatus. Nature 477, 596–600 (2011).
Fernandes-Alnemri, T., Yu, J. W., Datta, P., Wu, J. & Alnemri, E. S. AIM2 activates the inflammasome and cell death in response to cytoplasmic DNA. Nature 458, 509–513 (2009).
Hornung, V. et al. AIM2 recognizes cytosolic dsDNA and forms a caspase-1-activating inflammasome with ASC. Nature 458, 514–518 (2009).
Burckstummer, T. et al. An orthogonal proteomic-genomic screen identifies AIM2 as a cytoplasmic DNA sensor for the inflammasome. Nat. Immunol. 10, 266–272 (2009).
Roberts, T. L. et al. HIN-200 proteins regulate caspase activation in response to foreign cytoplasmic DNA. Science 323, 1057–1060 (2009).
Chae, J. J. et al. The B30.2 domain of pyrin, the familial Mediterranean fever protein, interacts directly with caspase-1 to modulate IL-1beta production. Proc. Natl Acad. Sci. USA 103, 9982–9987 (2006).
Xu, H. et al. Innate immune sensing of bacterial modifications of Rho GTPases by the pyrin inflammasome. Nature 513, 237–241 (2014).
Karki, R. et al. IRF8 regulates transcription of Naips for NLRC4 inflammasome activation. Cell 173, 920–933 e913 (2018).
Nour, A. M. et al. Anthrax lethal toxin triggers the formation of a membrane-associated inflammasome complex in murine macrophages. Infect. Immun. 77, 1262–1271 (2009).
Jin, T., Curry, J., Smith, P., Jiang, J. & Xiao, T. S. Structure of the NLRP1 caspase recruitment domain suggests potential mechanisms for its association with procaspase-1. Proteins 81, 1266–1270 (2013).
Zhang, L. et al. Cryo-EM structure of the activated NAIP2-NLRC4 inflammasome reveals nucleated polymerization. Science 350, 404–409 (2015).
Datta, D., McClendon, C. L., Jacobson, M. P. & Wells, J. A. Substrate and inhibitor-induced dimerization and cooperativity in caspase-1 but not caspase-3. J. Biol. Chem. 288, 9971–9981 (2013).
Broz, P., von Moltke, J., Jones, J. W., Vance, R. E. & Monack, D. M. Differential requirement for caspase-1 autoproteolysis in pathogen-induced cell death and cytokine processing. Cell Host Microbe 8, 471–483 (2010).
Ramage, P. et al. Expression, refolding, and autocatalytic proteolytic processing of the interleukin-1 beta-converting enzyme precursor. J. Biol. Chem. 270, 9378–9383 (1995).
Kayagaki, N. et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signalling. Nature 526, 666–671 (2015).
Shi, J. et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–665 (2015).
Black, R. A., Kronheim, S. R. & Sleath, P. R. Activation of interleukin-1 beta by a co-induced protease. FEBS Lett. 247, 386–390 (1989).
Kostura, M. J. et al. Identification of a monocyte specific pre-interleukin 1 beta convertase activity. Proc. Natl Acad. Sci. USA 86, 5227–5231 (1989).
Man, S. M. et al. IRGB10 liberates bacterial ligands for sensing by the AIM2 and caspase-11-NLRP3 inflammasomes. Cell 167, 382–396.e317 (2016).
Gurung, P. et al. Toll or interleukin-1 receptor (TIR) domain-containing adaptor inducing interferon-beta (TRIF)-mediated caspase-11 protease production integrates toll-like receptor 4 (TLR4) protein- and Nlrp3 inflammasome-mediated host defense against enteropathogens. J. Biol. Chem. 287, 34474–34483 (2012).
Balakrishnan, A., Karki, R., Berwin, B., Yamamoto, M. & Kanneganti, T. D. Guanylate binding proteins facilitate caspase-11-dependent pyroptosis in response to type 3 secretion system-negative Pseudomonas aeruginosa. Cell Death Discov. 4, 3 (2018).
Malireddi, R. K., Ippagunta, S., Lamkanfi, M. & Kanneganti, T. D. Cutting edge: proteolytic inactivation of poly(ADP-ribose) polymerase 1 by the Nlrp3 and Nlrc4 inflammasomes. J. Immunol. 185, 3127–3130 (2010).
Place, D. E. et al. ASK family kinases are required for optimal NLRP3 inflammasome priming. Am. J. Pathol. 188, 1021–1030 (2018).
Zhu, Q., Man, S. M., Karki, R., Malireddi, R. K. S. & Kanneganti, T. D. Detrimental type I interferon signaling dominates protective AIM2 inflammasome responses during Francisella novicida infection. Cell Rep. 22, 3168–3174 (2018).
Liu, Z. et al. Role of inflammasomes in host defense against Citrobacter rodentium infection. J. Biol. Chem. 287, 16955–16964 (2012).
Kuriakose, T. et al. ZBP1/DAI is an innate sensor of influenza virus triggering the NLRP3 inflammasome and programmed cell death pathways. Sci. Immunol. 1, aag2045 (2016).
Zheng, M., Karki, R., Vogel, P. & Kanneganti, T.-D. Caspase-6 is a key regulator of innate immunity, inflammasome activation and host defense. Cell 181, 674–687.e13 (2020).
Man, S. M. et al. The transcription factor IRF1 and guanylate-binding proteins target activation of the AIM2 inflammasome by Francisella infection. Nat. Immunol. 16, 467–475 (2015).
Briard, B. et al. Fungal ligands released by innate immune effectors promote inflammasome activation during Aspergillus fumigatus infection. Nat. Microbiol. 4, 316–327 (2019).
Karki, R. et al. Concerted activation of the AIM2 and NLRP3 inflammasomes orchestrates host protection against Aspergillus infection. Cell Host Microbe 17, 357–368 (2015).
van de Veerdonk, F. L. et al. The inflammasome drives protective Th1 and Th17 cellular responses in disseminated candidiasis. Eur. J. Immunol. 41, 2260–2268 (2011).
Malireddi, R. K. S. et al. Innate immune priming in the absence of TAK1 drives RIPK1 kinase activity-independent pyroptosis, apoptosis, necroptosis, and inflammatory disease. J. Exp. Med. 217, e20191644 (2020).
Gurung, P., Sharma, B. R. & Kanneganti, T. D. Distinct role of IL-1beta in instigating disease in Sharpin(cpdm) mice. Sci. Rep. 6, 36634 (2016).
Lukens, J. R. et al. Dietary modulation of the microbiome affects autoinflammatory disease. Nature 516, 246–249 (2014).
Gurung, P., Burton, A. & Kanneganti, T. D. NLRP3 inflammasome plays a redundant role with caspase 8 to promote IL-1beta-mediated osteomyelitis. Proc. Natl Acad. Sci. USA 113, 4452–4457 (2016).
Ippagunta, S. K. et al. Inflammasome-independent role of apoptosis-associated speck-like protein containing a CARD (ASC) in T cell priming is critical for collagen-induced arthritis. J. Biol. Chem. 285, 12454–12462 (2010).
Stienstra, R. et al. Inflammasome is a central player in the induction of obesity and insulin resistance. Proc. Natl Acad. Sci. USA 108, 15324–15329 (2011).
Rhoads, J. P. et al. Oxidized low-density lipoprotein immune complex priming of the Nlrp3 Inflammasome Involves TLR and FcgammaR cooperation and is dependent on CARD9. J. Immunol. 198, 2105–2114 (2017).
Stancu, I. C. et al. Aggregated Tau activates NLRP3-ASC inflammasome exacerbating exogenously seeded and non-exogenously seeded Tau pathology in vivo. Acta Neuropathol. 137, 599–617 (2019).
Saito, T. et al. Single App knock-in mouse models of Alzheimer’s disease. Nat. Neurosci. 17, 661–663 (2014).
Zaki, M. H. et al. The NLRP3 inflammasome protects against loss of epithelial integrity and mortality during experimental colitis. Immunity 32, 379–391 (2010).
Zaki, M. H., Vogel, P., Body-Malapel, M., Lamkanfi, M. & Kanneganti, T. D. IL-18 production downstream of the Nlrp3 inflammasome confers protection against colorectal tumor formation. J. Immunol. 185, 4912–4920 (2010).
Sharma, D. et al. Pyrin inflammasome regulates tight junction integrity to restrict colitis and tumorigenesis. Gastroenterology 154, 948–964 e948 (2018).
Man, S. M. et al. Critical role for the DNA sensor AIM2 in stem cell proliferation and cancer. Cell 162, 45–58 (2015).
Amer, A. et al. Regulation of Legionella phagosome maturation and infection through flagellin and host Ipaf. J. Biol. Chem. 281, 35217–35223 (2006).
Lamkanfi, M. et al. The Nod-like receptor family member Naip5/Birc1e restricts Legionella pneumophila growth independently of caspase-1 activation. J. Immunol. 178, 8022–8027 (2007).
Lamkanfi, M., Malireddi, R. K. & Kanneganti, T. D. Fungal zymosan and mannan activate the cryopyrin inflammasome. J. Biol. Chem. 284, 20574–20581 (2009).
Gross, O. Measuring the inflammasome. Methods Mol. Biol. 844, 199–222 (2012).
Marim, F. M., Silveira, T. N., Lima, D. S. Jr. & Zamboni, D. S. A method for generation of bone marrow-derived macrophages from cryopreserved mouse bone marrow cells. PLoS One 5, e15263 (2010).
Tran, T. A. T. et al. Whole blood assay as a model for in vitro evaluation of inflammasome activation and subsequent caspase-mediated interleukin-1 beta release. PLoS One 14, e0214999 (2019).
Vrentas, C. E. et al. Inflammasomes in livestock and wildlife: insights into the intersection of pathogens and natural host species. Vet. Immunol. Immunopathol. 201, 49–56 (2018).
Pelegrin, P., Barroso-Gutierrez, C. & Surprenant, A. P2X7 receptor differentially couples to distinct release pathways for IL-1beta in mouse macrophage. J. Immunol. 180, 7147–7157 (2008).
Yu, J. W. et al. Cryopyrin and pyrin activate caspase-1, but not NF-kappaB, via ASC oligomerization. Cell Death Differ. 13, 236–249 (2006).
Lamkanfi, M. et al. Targeted peptidecentric proteomics reveals caspase-7 as a substrate of the caspase-1 inflammasomes. Mol. Cell. Proteom. 7, 2350–2363 (2008).
Malireddi, R. K. S. et al. TAK1 restricts spontaneous NLRP3 activation and cell death to control myeloid proliferation. J. Exp. Med. 215, 1023–1034 (2018).
Malireddi, R. K. S., Kesavardhana, S. & Kanneganti, T. D. ZBP1 and TAK1: master regulators of NLRP3 inflammasome/pyroptosis, apoptosis, and necroptosis (PAN-optosis). Front. Cell. Infect. Microbiol. 9, 406 (2019).
Van Opdenbosch, N. et al. Caspase-1 engagement and TLR-induced c-FLIP expression suppress ASC/caspase-8-dependent apoptosis by inflammasome sensors NLRP1b and NLRC4. Cell Rep. 21, 3427–3444 (2017).
Gurung, P. et al. FADD and caspase-8 mediate priming and activation of the canonical and noncanonical Nlrp3 inflammasomes. J. Immunol. 192, 1835–1846 (2014).
Christgen, S., et al Identification of the PANoptosome: a molecular platform triggering pyroptosis, apoptosis, and necroptosis (PANoptosis). Front. Cell. Infect. Microbiol. 10, 237 (2020).
Samir, P., Malireddi, R.K.S., Kanneganti, T.-D. The PANoptosome: A deadly protein complex driving pyroptosis, apoptosis, and necroptosis (PANoptosis). Front. Cell. Infect. Microbiol. 10, 238 (2020).
Kesavardhana, S. et al. The Zα2 domain of ZBP1 is a molecular switch regulating influenza-induced PANoptosis and perinatal lethality during development. J. Biol. Chem. 295, 8325–8330 (2020).
Karki, R. et al. Interferon regulatory factor 1 regulates PANoptosis to prevent colorectal cancer. JCI Insight 5, e136720 (2020).
Thornberry, N. A. et al. A novel heterodimeric cysteine protease is required for interleukin-1 beta processing in monocytes. Nature 356, 768–774 (1992).
Cerretti, D. P. et al. Molecular cloning of the interleukin-1 beta converting enzyme. Science 256, 97–100 (1992).
Boucher, D., Chan, A., Ross, C. & Schroder, K. Quantifying caspase-1 activity in murine macrophages. Methods Mol. Biol. 1725, 163–176 (2018).
Boucher, D., Duclos, C. & Denault, J. B. General in vitro caspase assay procedures. Methods Mol. Biol. 1133, 3–39 (2014).
Kaushal, V., Herzog, C., Haun, R. S. & Kaushal, G. P. Caspase protocols in mice. Methods Mol. Biol. 1133, 141–154 (2014).
Swacha, P., Gekara, N. O. & Erttmann, S. F. Biochemical and microscopic analysis of inflammasome complex formation. Methods Enzymol. 625, 287–298 (2019).
Talley, S. et al. A caspase-1 biosensor to monitor the progression of inflammation in vivo. J. Immunol. 203, 2497–2507 (2019).
Poreba, M., Strózyk, A., Salvesen, G. S. & Drag, M. Caspase substrates and inhibitors. Cold Spring Harb. Perspect. Biol. 5, a008680 (2013).
Sester, D. P. et al. Assessment of inflammasome formation by flow cytometry. Curr. Protoc. Immunol. 114, 14.40.11–14.40.29 (2016).
Mazanek, Z. & Sohn, J. Tracking the polymerization of DNA sensors and inflammasomes using FRET. Methods Enzymol. 625, 87–94 (2019).
Xia, S., Ruan, J. & Wu, H. Monitoring gasdermin pore formation in vitro. Methods Enzymol. 625, 95–107 (2019).
Sarhan, J. et al. Caspase-8 induces cleavage of gasdermin D to elicit pyroptosis during Yersinia infection. Proc. Natl Acad. Sci. USA 115, E10888–E10897 (2018).
Orning, P. et al. Pathogen blockade of TAK1 triggers caspase-8-dependent cleavage of gasdermin D and cell death. Science 362, 1064–1069 (2018).
Broz, P. & Monack, D. M. Measuring inflammasome activation in response to bacterial infection. Methods Mol. Biol. 1040, 65–84 (2013).
Nagata, K. et al. Generation of App knock-in mice reveals deletion mutations protective against Alzheimer’s disease-like pathology. Nat. Commun. 9, 1800 (2018).
Hoffmann, E., Neumann, G., Kawaoka, Y., Hobom, G. & Webster, R. G. A DNA transfection system for generation of influenza A virus from eight plasmids. Proc. Natl Acad. Sci. USA 97, 6108–6113 (2000).
Nierman, W. C. et al. Genomic sequence of the pathogenic and allergenic filamentous fungus Aspergillus fumigatus. Nature 438, 1151–1156 (2005).
Kuehne, S. A. et al. Importance of toxin A, toxin B, and CDT in virulence of an epidemic Clostridium difficile strain. J. Infect. Dis. 209, 83–86 (2014).
Brand, D. D., Latham, K. A. & Rosloniec, E. F. Collagen-induced arthritis. Nat. Protoc. 2, 1269–1275 (2007).
Liu, Z. et al. DOCK2 confers immunity and intestinal colonization resistance to Citrobacter rodentium infection. Sci. Rep. 6, 27814 (2016).
Franchi, L. et al. Critical role for Ipaf in Pseudomonas aeruginosa–induced caspase-1 activation. Eur. J. Immunol. 37, 3030–3039 (2007).
Werner, J. L. et al. Requisite role for the dectin-1 beta-glucan receptor in pulmonary defense against Aspergillus fumigatus. J. Immunol. 182, 4938–4946 (2009).
Christ, A. et al. Western diet triggers NLRP3-dependent innate immune reprogramming. Cell 172, 162–175.e114 (2018).
Cardona, A. E., Huang, D., Sasse, M. E. & Ransohoff, R. M. Isolation of murine microglial cells for RNA analysis or flow cytometry. Nat. Protoc. 1, 1947–1951 (2006).
He, Y., Taylor, N. & Bhattacharya, A. Isolation and culture of astrocytes from postnatal and adult mouse brains. Methods Mol. Biol. 1938, 37–47 (2019).
Weischenfeldt, J. & Porse, B. Bone marrow-derived macrophages (BMM): isolation and applications. Cold Spring Harb. Protoc. 2008, pdb.prot5080 (2008).
Roney, K. Bone marrow-derived dendritic cells. Methods Mol. Biol. 1031, 71–76 (2013).
Link, A. J. & LaBaer, J. Trichloroacetic acid (TCA) precipitation of proteins. Cold Spring Harb. Protoc. 2011, 993–994 (2011).
Kusumbe, A. P., Ramasamy, S. K., Starsichova, A. & Adams, R. H. Sample preparation for high-resolution 3D confocal imaging of mouse skeletal tissue. Nat. Protoc. 10, 1904–1914 (2015).
Stutz, A., Horvath, G. L., Monks, B. G. & Latz, E. ASC speck formation as a readout for inflammasome activation. Methods Mol. Biol. 1040, 91–101 (2013).
Simpson, R. J. Homogenization of mammalian tissue. Cold Spring Harb. Protoc. 2010, pdb.prot5455 (2010).
Zondag, H. A. [Determination and diagnostic significance of the lactate dehydrogenase isoenzymes]. Jaarb. Kankeronderz. Kankerbestrijd. Ned. [Yearb. Cancer Res. Fight Cancer Neth.] 14, 327–329 (1964).
Samir, P. et al. DDX3X acts as a live-or-die checkpoint in stressed cells by regulating NLRP3 inflammasome. Nature 573, 590–594 (2019).
Sharma, D., Malik, A., Guy, C., Vogel, P. & Kanneganti, T. D. TNF/TNFR axis promotes pyrin inflammasome activation and distinctly modulates pyrin inflammasomopathy. J. Clin. Invest. 129, 150–162 (2019).
Fink, S. L., Bergsbaken, T. & Cookson, B. T. Anthrax lethal toxin and Salmonella elicit the common cell death pathway of caspase-1-dependent pyroptosis via distinct mechanisms. Proc. Natl Acad. Sci. USA 105, 4312–4317 (2008).
Wickliffe, K. E., Leppla, S. H. & Moayeri, M. Killing of macrophages by anthrax lethal toxin: involvement of the N-end rule pathway. Cell Microbiol. 10, 1352–1362 (2008).
Karki, R., Lee, E., Sharma, B. R., Banoth, B. & Kanneganti, T. D. IRF8 regulates Gram-negative bacteria-mediated NLRP3 inflammasome activation and cell death. J. Immunol. 204, 2514–2522 (2020).
Wellington, M., Koselny, K., Sutterwala, F. S. & Krysan, D. J. Candida albicans triggers NLRP3-mediated pyroptosis in macrophages. Eukaryot. Cell 13, 329–340 (2014).
Acknowledgements
We thank members of the Kanneganti lab, both past and present, for their comments and suggestions. Work from our laboratory was supported by the US National Institutes of Health (AI101935, AI124346, AR056296, and CA253095 to T.-D.K.) and the American Lebanese Syrian Associated Charities (to T.-D.K.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
R.E.T., R.K.S.M., and T.-D.K. conceptualized the manuscript. R.E.T. and R.K.S.M. wrote the first draft. All authors reviewed and approved the final draft of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Andrea Dorfleutner, Florian Schmidt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Zheng, M., Karki, R., Vogel, P. & Kanneganti, T.-D. Cell 181, 674–687.e13 (2020): https://www.cell.com/cell/fulltext/S0092-8674(20)30333-0
Samir, P. et al. Nature 573, 590–594 (2019): https://www.nature.com/articles/s41586-019-1551-2
Man, S. M. et al. Cell 167, 382–396.e17 (2016): https://www.cell.com/cell/fulltext/S0092-8674(16)31245-4
Supplementary information
Rights and permissions
About this article
Cite this article
Tweedell, R.E., Malireddi, R.K.S. & Kanneganti, TD. A comprehensive guide to studying inflammasome activation and cell death. Nat Protoc 15, 3284–3333 (2020). https://doi.org/10.1038/s41596-020-0374-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-0374-9
This article is cited by
-
Pnpt1 mediates NLRP3 inflammasome activation by MAVS and metabolic reprogramming in macrophages
Cellular & Molecular Immunology (2023)
-
Integrated NLRP3, AIM2, NLRC4, Pyrin inflammasome activation and assembly drive PANoptosis
Cellular & Molecular Immunology (2023)
-
Inflammasome activity is controlled by ZBTB16-dependent SUMOylation of ASC
Nature Communications (2023)
-
PML at mitochondria-associated membranes governs a trimeric complex with NLRP3 and P2X7R that modulates the tumor immune microenvironment
Cell Death & Differentiation (2023)
-
Role of pyroptosis in inflammation and cancer
Cellular & Molecular Immunology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.