Abstract
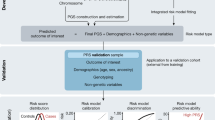
A polygenic score (PGS) or polygenic risk score (PRS) is an estimate of an individual’s genetic liability to a trait or disease, calculated according to their genotype profile and relevant genome-wide association study (GWAS) data. While present PRSs typically explain only a small fraction of trait variance, their correlation with the single largest contributor to phenotypic variation—genetic liability—has led to the routine application of PRSs across biomedical research. Among a range of applications, PRSs are exploited to assess shared etiology between phenotypes, to evaluate the clinical utility of genetic data for complex disease and as part of experimental studies in which, for example, experiments are performed that compare outcomes (e.g., gene expression and cellular response to treatment) between individuals with low and high PRS values. As GWAS sample sizes increase and PRSs become more powerful, PRSs are set to play a key role in research and stratified medicine. However, despite the importance and growing application of PRSs, there are limited guidelines for performing PRS analyses, which can lead to inconsistency between studies and misinterpretation of results. Here, we provide detailed guidelines for performing and interpreting PRS analyses. We outline standard quality control steps, discuss different methods for the calculation of PRSs, provide an introductory online tutorial, highlight common misconceptions relating to PRS results, offer recommendations for best practice and discuss future challenges.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414 (2019).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Yang, J. et al. Common SNPs explain a large proportion of heritability for human height. Nat. Genet. 42, 565–569 (2010).
Dudbridge, F. Power and predictive accuracy of polygenic risk scores. PLoS Genet. 9, e1003348 (2013).
Dudbridge, F. Polygenic epidemiology. Genet. Epidemiol. 40, 268–272 (2016).
Yang, J. et al. GCTA: A tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Palla, L. & Dudbridge, F. A fast method that uses polygenic scores to estimate the variance explained by genome-wide marker panels and the proportion of variants affecting a trait. Am. J. Hum. Genet. 97, 250–259 (2015).
Purcell, S. M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Wray, N. R. et al. Research review: polygenic methods and their application to psychiatric traits. J. Child Psychol. Psychiatry 55, 1068–1087 (2014).
Euesden, J., Lewis, C. M. & O’Reilly, P. F. PRSice: polygenic risk score software. Bioinformatics 31, 1466–1468 (2015).
Choi, S. W. & O’Reilly, P. F. PRSice-2: polygenic risk score software for biobank-scale data. Gigascience 8, giz082 (2019).
Agerbo, E. et al. Polygenic risk score, parental socioeconomic status, family history of psychiatric disorders, and the risk for schizophrenia: a Danish population-based study and meta-analysis. JAMA Psychiatry 72, 635–641 (2015).
Mavaddat, N. et al. Polygenic risk scores for prediction of breast cancer and breast cancer subtypes. Am. J. Hum. Genet. 104, 21–34 (2019).
Mullins, N. et al. Polygenic interactions with environmental adversity in the aetiology of major depressive disorder. Psychol. Med. 46, 759–770 (2016).
Natarajan, P. et al. Polygenic risk score identifies subgroup with higher burden of atherosclerosis and greater relative benefit from statin therapy in the primary prevention setting. Circulation 135, 2091–2101 (2017).
Mak, T. S. H. et al. Polygenic scores via penalized regression on summary statistics. Genet. Epidemiol. 41, 469–480 (2017).
Ge, T. et al. Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nat. Commun. 10, 1–10 (2019).
Speed, D. & Balding, D. J. MultiBLUP: improved SNP-based prediction for complex traits. Genome Res. 24, 1550–1557 (2014).
Zhou, X., Carbonetto, P. & Stephens, M. Polygenic modeling with Bayesian sparse linear mixed models. PLoS Genet. 9, e1003264 (2013).
Shi, J. et al. Winner’s curse correction and variable thresholding improve performance of polygenic risk modeling based on genome-wide association study summary-level data. PLoS Genet. 12, e1006493 (2016).
Lello, L. et al. Accurate genomic prediction of human height. Genetics 210, 477–497 (2018).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Evans, L. M. et al. Comparison of methods that use whole genome data to estimate the heritability and genetic architecture of complex traits. Nat. Genet. 50, 737–745 (2018).
Coleman, J. R. I. et al. Quality control, imputation and analysis of genome-wide genotyping data from the Illumina HumanCoreExome microarray. Brief. Funct. Genomics 15, 298–304 (2016).
Marees, A. T. et al. A tutorial on conducting genome-wide association studies: quality control and statistical analysis. Int. J. Methods Psychiatr. Res. 27, e1608 (2018).
Anderson, C. A. et al. Data quality control in genetic case-control association studies. Nat. Protoc. 5, 1564–1573 (2010).
Speed, D. & Balding, D. J. SumHer better estimates the SNP heritability of complex traits from summary statistics. Nat. Genet. 51, 277–284 (2019).
Han, B. & Eskin, E. Random-effects model aimed at discovering associations in meta-analysis of genome-wide association studies. Am. J. Hum. Genet. 88, 586–598 (2011).
Drepper, U., Miller, S. & Madore, D. md5sum(1): compute/check MD5 message digest. Linux man page (accessed 20 October 2018); https://linux.die.net/man/1/md5sum
National Center for Biotechnology Information. US National Library of Medicine. Data changes that occur between builds. in SNP FAQ Archive. NCBI Help Manual. https://www.ncbi.nlm.nih.gov/books/NBK44467/ (2005).
Hinrichs, A. S. et al. The UCSC Genome Browser Database: update 2006. Nucleic Acids Res. 34, D590–D598 (2006).
Niemi, M. E. K. et al. Common genetic variants contribute to risk of rare severe neurodevelopmental disorders. Nature 562, 268–271 (2018).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Chen, L. M. et al. PRS-on-Spark (PRSoS): a novel, efficient and flexible approach for generating polygenic risk scores. BMC Bioinforma. 19, 295 (2018).
Accounting for sex in the genome. Nat. Med.23, 1243–1243 (2017).
König, I. R. et al. How to include chromosome X in your genome-wide association study. Genet. Epidemiol. 38, 97–103 (2014).
Wray, N. R. et al. Pitfalls of predicting complex traits from SNPs. Nat. Rev. Genet. 14, 507–515 (2013).
Viechtbauer, W. & Cheung, M. W.-L. Outlier and influence diagnostics for meta-analysis. Res. Synth. Methods 1, 112–125 (2010).
Socrates, A. et al. Polygenic risk scores applied to a single cohort reveal pleiotropy among hundreds of human phenotypes. Preprint at https://www.biorxiv.org/content/10.1101/203257v1 (2017).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Vilhjálmsson, B. J. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592 (2015).
Newcombe, P. J. et al. A flexible and parallelizable approach to genome-wide polygenic risk scores. Genet. Epidemiol. 43, 730–741 (2019).
Loh, P.-R. et al. Mixed-model association for biobank-scale datasets. Nat. Genet. 50, 906–908 (2018).
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Privé, F. et al. Efficient analysis of large-scale genome-wide data with two R packages: bigstatsr and bigsnpr. Bioinformatics 34, 2781–2787 (2018).
Márquez‐Luna, C., Loh, P.-R. & Price, A. L. Multiethnic polygenic risk scores improve risk prediction in diverse populations. Genet. Epidemiol. 41, 811–823 (2017).
Clayton, D. Link functions in multi-locus genetic models: implications for testing, prediction, and interpretation. Genet. Epidemiol. 36, 409–418 (2012).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Astle, W. & Balding, D. J. Population structure and cryptic relatedness in genetic association studies. Stat. Sci. 24, 451–471 (2009).
Kong, A. et al. The nature of nurture: effects of parental genotypes. Science 359, 424–428 (2018).
Selzam, S. et al. Comparing within- and between-family polygenic score prediction. Am. J. Hum. Genet. 105, 351–363 (2019).
Price, A. L. et al. New approaches to population stratification in genome-wide association studies. Nat. Rev. Genet. 11, 459–463 (2010).
Cheesman, R. et al. Comparison of adopted and nonadopted individuals reveals gene–environment interplay for education in the UK Biobank. Psychol. Sci. 31, 582–591 (2020).
Mostafavi, H. et al. Variable prediction accuracy of polygenic scores within an ancestry group. eLife 9, e48376 (2020).
Young, A. I. et al. Deconstructing the sources of genotype-phenotype associations in humans. Science 365, 1396–1400 (2019).
Kim, M. S., Patel, K. P., Teng, A. K., Berens, A. J. & Lachance, J. Genetic disease risks can be misestimated across global populations. Genome Biol. 19, 179 (2018).
Martin, A. R. et al. Human demographic history impacts genetic risk prediction across diverse populations. Am. J. Hum. Genet. 100, 635–649 (2017).
Duncan, L. et al. Analysis of polygenic risk score usage and performance in diverse human populations. Nat. Commun. 10, 3328 (2019).
Selzam, S. et al. Predicting educational achievement from DNA. Mol. Psychiatry 22, 267–272 (2017).
Lee, J. J. et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat. Genet. 50, 1112–1121 (2018).
Grotzinger, A. D. et al. Genomic structural equation modelling provides insights into the multivariate genetic architecture of complex traits. Nat. Hum. Behav. 3, 513–525 (2019).
Maier, R. M. et al. Improving genetic prediction by leveraging genetic correlations among human diseases and traits. Nat. Commun. 9, 989 (2018).
Krapohl, E. et al. Multi-polygenic score approach to trait prediction. Mol. Psychiatry 23, 1368–1374 (2018).
Ruderfer, D. M. et al. Polygenic dissection of diagnosis and clinical dimensions of bipolar disorder and schizophrenia. Mol. Psychiatry 19, 1017–1024 (2014).
Bipolar Disorder and Schizophrenia Working Group of the Psychiatric Genomics Consortium. Genomic dissection of bipolar disorder and schizophrenia, including 28 subphenotypes. Cell 173, 1705–1715.e16 (2018).
Turley, P. et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat. Genet. 50, 229–237 (2018).
Rheenen, W. et al. Genetic correlations of polygenic disease traits: from theory to practice. Nat. Rev. Genet. 20, 567–581 (2019).
Visscher, P. M. et al. Statistical power to detect genetic (co)variance of complex traits using SNP data in unrelated samples. PLoS Genet. 10, e1004269 (2014).
Janssens, M. J. J. Co-heritability: its relation to correlated response, linkage, and pleiotropy in cases of polygenic inheritance. Euphytica 28, 601–608 (1979).
Pirinen, M., Donnelly, P. & Spencer, C. C. A. Including known covariates can reduce power to detect genetic effects in case-control studies. Nat. Genet. 44, 848–851 (2012).
Lee, S. H. et al. A better coefficient of determination for genetic profile analysis. Genet. Epidemiol. 36, 214–224 (2012).
Heinzl, H., Waldhör, T. & Mittlböck, M. Careful use of pseudo R-squared measures in epidemiological studies. Stat. Med. 24, 2867–2872 (2005).
Lee, S. H. et al. Estimating missing heritability for disease from genome-wide association studies. Am. J. Hum. Genet. 88, 294–305 (2011).
Won, H.-H. et al. Disproportionate contributions of select genomic compartments and cell types to genetic risk for coronary artery disease. PLoS Genet. 11, e1005622 (2015).
Santoro, M. L. et al. Polygenic risk score analyses of symptoms and treatment response in an antipsychotic-naive first episode of psychosis cohort. Transl. Psychiatry 8, 1–8 (2018).
Power, R. A. et al. Polygenic risk scores for schizophrenia and bipolar disorder predict creativity. Nat. Neurosci. 18, 953–955 (2015).
Mullins, N. et al. GWAS of suicide attempt in psychiatric disorders and association with major depression polygenic risk scores. Am. J. Psychiatry 176, 651–660 (2019).
Khera, A. V. et al. Polygenic prediction of weight and obesity trajectories from birth to adulthood. Cell 177, 587–596.e9 (2019).
Khera, A. V. et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 50, 1219–1224 (2018).
Du Rietz, E. et al. Association of polygenic risk for attention-deficit/hyperactivity disorder with co-occurring traits and disorders. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 3, 635–643 (2018).
Clayton, D. G. Prediction and interaction in complex disease genetics: experience in type 1 diabetes. PLoS Genet. 5, e1000540 (2009).
Dudbridge, F., Pashayan, N. & Yang, J. Predictive accuracy of combined genetic and environmental risk scores. Genet. Epidemiol. 42, 4–19 (2018).
Torkamani, A., Wineinger, N. E. & Topol, E. J. The personal and clinical utility of polygenic risk scores. Nat. Rev. Genet. 19, 581–590 (2018).
Lambert, S. A., Abraham, G. & Inouye, M. Towards clinical utility of polygenic risk scores. Hum. Mol. Genet. 28(R2), R133–R142 (2019).
Gibson, G. On the utilization of polygenic risk scores for therapeutic targeting. PLoS Genet. 15, e1008060 (2019).
Rose, G. Sick individuals and sick populations. Int. J. Epidemiol. 30, 427–432 (2001).
Wynants, L., Collins, G. S. & Van Calster, B. Key steps and common pitfalls in developing and validating risk models. BJOG 124, 423–432 (2017).
Janssens, A. C. J. W. & Joyner, M. J. Polygenic risk scores that predict common diseases using millions of single nucleotide polymorphisms: is more, better? Clin. Chem. 65, 609–611 (2019).
Baverstock, K. Polygenic scores: are they a public health hazard? Prog. Biophys. Mol. Biol. 149, 4–8 (2019).
Janssens, A. C. J. W. Validity of polygenic risk scores: are we measuring what we think we are? Hum. Mol. Genet. 28, R143–R150 (2019).
Boyle, E. A., Li, Y. I. & Pritchard, J. K. An expanded view of complex traits: from polygenic to omnigenic. Cell 169, 1177–1186 (2017).
Sniekers, S. et al. Genome-wide association meta-analysis of 78,308 individuals identifies new loci and genes influencing human intelligence. Nat. Genet. 49, 1107–1112 (2017).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Hartwig, F. P. et al. Two-sample Mendelian randomization: avoiding the downsides of a powerful, widely applicable but potentially fallible technique. Int. J. Epidemiol. 45, 1717–1726 (2016).
Krapohl, E. et al. Phenome-wide analysis of genome-wide polygenic scores. Mol. Psychiatry 21, 1188–1193 (2016).
Pingault, J.-B. et al. Using genetic data to strengthen causal inference in observational research. Nat. Rev. Genet. 19, 566–580 (2018).
Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. Ser. B Methodol. 58, 267–288 (1996).
Mak, T. S. H. et al. Polygenic scores for UK Biobank scale data. Preprint at https://www.biorxiv.org/content/10.1101/252270v3 (2018).
Machiela, M. J. et al. Evaluation of polygenic risk scores for predicting breast and prostate cancer risk. Genet. Epidemiol. 35, 506–514 (2011).
Falconer, D. S. Introduction to Quantitative Genetics (Ronald Press, 1960).
Jaffee, S. & Price, T. Gene–environment correlations: a review of the evidence and implications for prevention of mental illness. Mol. Psychiatry 12, 432–442 (2007).
Acknowledgements
We thank the participants in the UK Biobank and the scientists involved in the construction of this resource. We thank Jonathan Coleman and Kylie Glanville for help in management of the UK Biobank resource at King’s College London, and we thank Jack Euesden, Carla Giner-Delgado, Clive Hoggart, Hei Man Wu, Tom Bond, Gerome Breen, Cathryn Lewis, Cecile Janssens and Pak Sham for helpful discussions. This research has been conducted using the UK Biobank Resource under application 18177 (P.F.O.). P.F.O. receives funding from the UK Medical Research Council (MR/N015746/1). S.W.C. is funded by the UK Medical Research Council (MR/N015746/1). This report represents independent research partially funded by the National Institute for Health Research (NIHR) Biomedical Research Centre at South London and Maudsley NHS Foundation Trust and King’s College London. The views expressed are those of the authors and not necessarily those of the NHS, the NIHR or the Department of Health.
Author information
Authors and Affiliations
Contributions
P.F.O. conceived, prepared and wrote the manuscript, with feedback from S.W.C. and T.S.-H.M. S.W.C. performed the analyses and produced the online tutorial, with feedback from P.F.O.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Dorret Boomsma, Brandon Johnson and Anubha Mahajan for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
PRS tutorial: https://choishingwan.github.io/PRS-Tutorial/
GWAS Tutorial: https://github.com/MareesAT/GWA_tutorial
PRSice software: http://PRSice.net
LDpred software: https://github.com/bvilhjal/ldpred
Lassosum software: https://github.com/tshmak/lassosum/blob/master/README.md
The R project for statistical computing: https://www.r-project.org
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Choi, S.W., Mak, T.SH. & O’Reilly, P.F. Tutorial: a guide to performing polygenic risk score analyses. Nat Protoc 15, 2759–2772 (2020). https://doi.org/10.1038/s41596-020-0353-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-0353-1
This article is cited by
-
Appraising associations between signature lipidomic biomarkers and digestive system cancer risk: novel evidences from a prospective cohort study of UK Biobank and Mendelian randomization analyses
Lipids in Health and Disease (2024)
-
Age-dependent genes in adipose stem and precursor cells affect regulation of fat cell differentiation and link aging to obesity via cellular and genetic interactions
Genome Medicine (2024)
-
A study of genetic variants associated with skin traits in the Vietnamese population
BMC Genomics (2024)
-
Recent advances in polygenic scores: translation, equitability, methods and FAIR tools
Genome Medicine (2024)
-
A brief comparison of polygenic risk scores and Mendelian randomisation
BMC Medical Genomics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.