Abstract
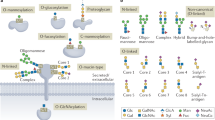
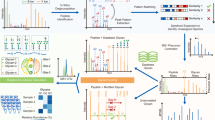
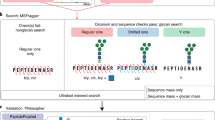
Protein glycosylation is one of the most common protein modifications. A major type of protein glycosylation is O-GalNAcylation, in which GalNAc-type glycans are attached to protein Ser or Thr residues via an O-linked glycosidic bond. O-GalNAcylation is thought to play roles in protein folding, stability, trafficking and protein interactions, and identification of the site-specific O-GalNAc glycoproteome is a crucial step toward understanding the biological significance of the modification. However, lack of suitable methodology, absence of consensus sequon of O-GalNAcylation sites and complex O-GalNAc glycan structures pose analytical challenges. We recently developed a mass spectrometry-based method called extraction of O-linked glycopeptides (EXoO) that enables large-scale mapping of site-specific mucin-type O-GalNAcylation sites. Here we provide a detailed protocol for EXoO, which includes seven stages of: (1) extraction and proteolytic digestion of proteins to peptides, (2) sequential guanidination and de-salting of peptides, (3) enrichment of glycopeptides, (4) solid-phase peptide conjugation and release of O-GalNAc glycopeptides using the OpeRATOR protease, (5) liquid chromatography with tandem mass spectrometry analysis of O-GalNAc glycopeptides, (6) identification of O-GalNAc glycopeptides by database search and (7) quantification of O-GalNAc glycopeptides. Using this protocol, thousands of O-GalNAcylation sites from hundreds of glycoproteins with information regarding site-specific O-GalNAc glycan can be identified and quantified from complex samples. The protocol can be performed by a researcher with basic proteomics skills and takes about 4 d to complete.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Code availability
Extract_oxonium_ion_spectra_intensity.exe script is freely available at GitHub (download at https://github.com/appleTree2017/Extraction-of-O-linked-glycopeptides-EXoO-).
References
Gahmberg, C. G. & Tolvanen, M. Why mammalian cell surface proteins are glycoproteins. Trends Biochem. Sci. 21, 308–311 (1996).
Jang, J. H. & Hanash, S. Profiling of the cell surface proteome. Proteomics 3, 1947–1954 (2003).
Kovtun, Y. V. & Goldmacher, V. S. Cell killing by antibody–drug conjugates. Cancer Lett. 255, 232–240 (2007).
Selvaraj, P. et al. The T lymphocyte glycoprotein CD2 binds the cell surface ligand LFA-3. Nature 326, 400–403 (1987).
Orentas, R. J. et al. Identification of cell surface proteins as potential immunotherapy targets in 12 pediatric cancers. Front. Oncol. 2, 194 (2012).
Posey, A. D. Jr. et al. Engineered CAR T cells targeting the cancer-associated Tn-glycoform of the membrane mucin MUC1 control adenocarcinoma. Immunity 44, 1444–1454 (2016).
Westerlund, B. & Korhonen, T. K. Bacterial proteins binding to the mammalian extracellular matrix. Mol. Microbiol. 9, 687–694 (1993).
Maddon, P. J. et al. The T4 gene encodes the AIDS virus receptor and is expressed in the immune system and the brain. Cell 47, 333–348 (1986).
Kennedy, S. Proteomic profiling from human samples: the body fluid alternative. Toxicol. Lett. 120, 379–384 (2001).
Brockhausen, I. & Stanley, P. O-GalNAc glycans. in Essentials of Glycobiology 3rd edn (eds A. Varki et al.) (Cold Spring Harbor Laboratory Press, 2015).
Tarelli, E., Smith, A. C., Hendry, B. M., Challacombe, S. J. & Pouria, S. Human serum IgA1 is substituted with up to six O-glycans as shown by matrix assisted laser desorption ionisation time-of-flight mass spectrometry. Carbohydr. Res. 339, 2329–2335 (2004).
Arnold, J. N. et al. The glycosylation of human serum IgD and IgE and the accessibility of identified oligomannose structures for interaction with mannan-binding lectin. J. Immunol. 173, 6831–6840 (2004).
Sasaki, H., Bothner, B., Dell, A. & Fukuda, M. Carbohydrate structure of erythropoietin expressed in Chinese hamster ovary cells by a human erythropoietin cDNA. J. Biol. Chem. 262, 12059–12076 (1987).
Yang, W. et al. Glycoform analysis of recombinant and human immunodeficiency virus envelope protein gp120 via higher energy collisional dissociation and spectral-aligning strategy. Anal. Chem. 86, 6959–6967 (2014).
Hang, H. C. & Bertozzi, C. R. The chemistry and biology of mucin-type O-linked glycosylation. Bioorg. Med. Chem. 13, 5021–5034 (2005).
Yang, W., Ao, M., Hu, Y., Li, Q. K. & Zhang, H. Mapping the O‐glycoproteome using site‐specific extraction of O‐linked glycopeptides (EXoO). Mol. Syst. Biol. 14, e8486 (2018).
Steentoft, C. et al. Mining the O-glycoproteome using zinc-finger nuclease-glycoengineered SimpleCell lines. Nat. Methods 8, 977–982 (2011).
Woo, C. M., Iavarone, A. T., Spiciarich, D. R., Palaniappan, K. K. & Bertozzi, C. R. Isotope-targeted glycoproteomics (IsoTaG): a mass-independent platform for intact N- and O-glycopeptide discovery and analysis. Nat. Methods 12, 561–567 (2015).
Nilsson, J. et al. Enrichment of glycopeptides for glycan structure and attachment site identification. Nat. Methods 6, 809–811 (2009).
Medzihradszky, K. F., Kaasik, K. & Chalkley, R. J. Tissue-specific glycosylation at the glycopeptide level. Mol. Cell Proteomics 14, 2103–2110 (2015).
Hoffmann, M., Marx, K., Reichl, U., Wuhrer, M. & Rapp, E. Site-specific O-glycosylation analysis of human blood plasma proteins. Mol. Cell Proteomics 15, 624–641 (2016).
Mechref, Y. Use of CID/ETD mass spectrometry to analyze glycopeptides. Curr. Protoc. Protein Sci. Chapter 12, Unit 12 11 11–11 (2012).
Steentoft, C. et al. Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J. 32, 1478–1488 (2013).
Tabak, L. A. The role of mucin-type O-glycans in eukaryotic development. Semin. Cell Dev. Biol. 21, 616–621 (2010).
Kufe, D. W. Mucins in cancer: function, prognosis and therapy. Nat. Rev. Cancer 9, 874–885 (2009).
Sun, S. et al. Comprehensive analysis of protein glycosylation by solid-phase extraction of N-linked glycans and glycosite-containing peptides. Nat. Biotechnol. 34, 84 (2016).
Zhang, H., Li, X. J., Martin, D. B. & Aebersold, R. Identification and quantification of N-linked glycoproteins using hydrazide chemistry, stable isotope labeling and mass spectrometry. Nat. Biotechnol. 21, 660–666 (2003).
Yang, S., Hu, Y. W., Sokoll, L. & Zhang, H. Simultaneous quantification of N- and O-glycans using a solid-phase method. Nat. Protoc. 12, 1229–1244 (2017).
Huang, J. et al. Highly efficient release of glycopeptides from hydrazide beads by hydroxylamine assisted PNGase F deglycosylation for N-glycoproteome analysis. Anal. Chem. 87, 10199–10204 (2015).
Yang, S. et al. Deciphering protein O-glycosylation: solid-phase chemoenzymatic cleavage and enrichment. Anal. Chem. 90, 8261–8269 (2018).
Yang, W. et al. Comparison of enrichment methods for intact N-and O-linked glycopeptides using strong anion exchange and hydrophilic interaction liquid chromatography. Anal. Chem. 89, 11193–11197 (2017).
Zhu, Z., Su, X., Clark, D. F., Go, E. P. & Desaire, H. Characterizing O-linked glycopeptides by electron transfer dissociation: fragmentation rules and applications in data analysis. Anal. Chem. 85, 8403–8411 (2013).
Good, D. M., Wirtala, M., McAlister, G. C. & Coon, J. J. Performance characteristics of electron transfer dissociation mass spectrometry. Mol. Cell Proteomics 6, 1942–1951 (2007).
Darula, Z., Sherman, J. & Medzihradszky, K. F. How to dig deeper? Improved enrichment methods for mucin core-1 type glycopeptides. Mol. Cell Proteomics 11, O111 016774 (2012).
Mulagapati, S., Koppolu, V. & Raju, T. S. Decoding of O-Linked glycosylation by mass spectrometry. Biochemistry 56, 1218–1226 (2017).
Jensen, P. H., Kolarich, D. & Packer, N. H. Mucin-type O-glycosylation-putting the pieces together. FEBS J. 277, 81–94 (2010).
Malaker, S. A. et al. The mucin-selective protease StcE enables molecular and functional analysis of human cancer-associated mucins. Proc. Natl Acad. Sci. USA 116, 7278–7287 (2019).
Khoo, K. H. Advances toward mapping the full extent of protein site-specific O-GalNAc glycosylation that better reflects underlying glycomic complexity. Curr. Opin. Struct. Biol. 56, 146–154 (2019).
Moran, S. & Cattran, D. C. Recent advances in risk prediction, therapeutics and pathogenesis of IgA nephropathy. Minerva Med. 110, 439–449 (2019).
Zhu, F., Li, D. & Chen, K. Structures and functions of invertebrate glycosylation. Open Biol. 9, 180232 (2019).
Haltom, A. R. & Jafar-Nejad, H. O-Linked glycans in drosophila development: overview in Glycoscience: Biology and Medicine (eds Taniguchi, N. et al.) 809–815 (Springer Japan, 2015).
Sutton, C. W., O’Neill, J. A. & Cottrell, J. S. Site-specific characterization of glycoprotein carbohydrates by exoglycosidase digestion and laser desorption mass spectrometry. Anal. Biochem. 218, 34–46 (1994).
Muller, F., Fischer, L., Chen, Z. A., Auchynnikava, T. & Rappsilber, J. On the reproducibility of label-free quantitative cross-linking/mass spectrometry. J. Am. Soc. Mass Spectrom. 29, 405–412 (2018).
Tabb, D. L. et al. Repeatability and reproducibility in proteomic identifications by liquid chromatography-tandem mass spectrometry. J. Proteome Res. 9, 761–776 (2010).
Zhou, J. Y. et al. Quality assessments of long-term quantitative proteomic analysis of breast cancer xenograft tissues. J. Proteome Res. 16, 4523–4530 (2017).
Wu, Z. L. et al. Imaging specific cellular glycan structures using glycosyltransferases via click chemistry. Glycobiology 28, 69–79 (2018).
Dennis, G. Jr et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 4, P3 (2003).
Vizcaino, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Acknowledgements
This work was supported by the National Institutes of Health, the National Cancer Institute, the Early Detection Research Network (EDRN, U01CA152813), the Clinical Proteomic Tumor Analysis Consortium (CPTAC, U24CA210985) and amfAR, the Foundation for AIDS Research on Bringing Bioengineers to Cure HIV (Grant amfAR 109551‐61‐RGRL).
Author information
Authors and Affiliations
Contributions
W.Y. and H.Z. conceived the research. W.Y., A.S. and Y.X. conducted the experiments and validated the protocol. W.Y., Y.X. and M.A. conducted data analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Yang, W. et al. Mol. Syst. Biol. 14, e8486 (2018): https://www.embopress.org/doi/full/10.15252/msb.20188486.
Supplementary information
Supplementary Data 1
Reproducibility of the EXoO protocol. O-GalNAc glycopeptides were isolated four times from the same 50 µl of human serum. O-GalNAc glycopeptides from one of the isolations were subjected to the same LC–MS/MS analysis for four times to determine the reproducibility of LC–MS/MS analysis. As a result, a total of eight LC–MS/MS runs were conducted, including four isolation repeats and four LC–MS/MS repeats of one of the isolations.
Supplementary Data 2
Quantitative performance of the EXoO protocol.
Rights and permissions
About this article
Cite this article
Yang, W., Song, A., Ao, M. et al. Large-scale site-specific mapping of the O-GalNAc glycoproteome. Nat Protoc 15, 2589–2610 (2020). https://doi.org/10.1038/s41596-020-0345-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-0345-1
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.