Abstract
Cancer invasion and metastasis are challenging to study in vivo since they occur deep inside the body over extended time periods. Organotypic 3D culture of fresh tumor tissue enables convenient real-time imaging, genetic and microenvironmental manipulation and molecular analysis. Here, we provide detailed protocols to isolate and culture heterogenous organoids from murine and human primary and metastatic site tumors. The time required to isolate organoids can vary based on the tissue and organ type but typically takes <7 h. We describe a suite of assays that model specific aspects of metastasis, including proliferation, survival, invasion, dissemination and colony formation. We also specify comprehensive protocols for downstream applications of organotypic cultures that will allow users to (i) test the role of specific genes in regulating various cellular processes, (ii) distinguish the contributions of several microenvironmental factors and (iii) test the effects of novel therapeutics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data mentioned in the protocol are included. Any additional information may be provided by the corresponding author upon request. Source data are provided with this paper.
References
Nelson, C. M. & Bissell, M. J. Of extracellular matrix, scaffolds, and signaling: tissue architecture regulates development, homeostasis, and cancer. Annu. Rev. Cell Dev. Biol. 22, 287–309 (2006).
Simian, M. et al. The interplay of matrix metalloproteinases, morphogens and growth factors is necessary for branching of mammary epithelial cells. Development 128, 3117–3131 (2001).
Ewald, A. J. Practical considerations for long-term time-lapse imaging of epithelial morphogenesis in three-dimensional organotypic cultures. Cold Spring Harb. Protoc. 2013, 100–117 (2013).
Nguyen-Ngoc, K. V. et al. ECM microenvironment regulates collective migration and local dissemination in normal and malignant mammary epithelium. Proc. Natl Acad. Sci. U. S. A. 109, E2595–E2604 (2012).
Ewald, A. J. et al. Mammary collective cell migration involves transient loss of epithelial features and individual cell migration within the epithelium. J. Cell Sci. 125, 2638–2654 (2012).
Chen, Q. et al. A temporal requirement for Hippo signaling in mammary gland differentiation, growth, and tumorigenesis. Genes Dev. 28, 432–437 (2014).
Streuli, C. H., Bailey, N. & Bissell, M. J. Control of mammary epithelial differentiation: basement membrane induces tissue-specific gene expression in the absence of cell-cell interaction and morphological polarity. J. Cell Biol. 115, 1383–1395 (1991).
Mroue, R. & Bissell, M. J. Three-dimensional cultures of mouse mammary epithelial cells. Methods Mol. Biol. 945, 221–250 (2013).
Liu, Y. et al. Novel role for Netrins in regulating epithelial behavior during lung branching morphogenesis. Curr. Biol. 14, 897–905 (2004).
Del Moral, P. M. & Warburton, D. Explant culture of mouse embryonic whole lung, isolated epithelium, or mesenchyme under chemically defined conditions as a system to evaluate the molecular mechanism of branching morphogenesis and cellular differentiation. Methods Mol. Biol. 633, 71–79 (2010).
Lee, J. H. et al. Lung stem cell differentiation in mice directed by endothelial cells via a BMP4-NFATc1-thrombospondin-1 axis. Cell 156, 440–455 (2014).
Sato, T. et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology 141, 1762–1772 (2011).
Ootani, A. et al. Sustained in vitro intestinal epithelial culture within a Wnt-dependent stem cell niche. Nat. Med. 15, 701–706 (2009).
Huch, M. et al. Unlimited in vitro expansion of adult bi-potent pancreas progenitors through the Lgr5/R-spondin axis. EMBO J. 32, 2708–2721 (2013).
Ghosh, S. et al. PI3K/mTOR signaling regulates prostatic branching morphogenesis. Dev. Biol. 360, 329–342 (2011).
Drost, J. et al. Organoid culture systems for prostate epithelial and cancer tissue. Nat. Protoc. 11, 347–358 (2016).
Huch, M. et al. In vitro expansion of single Lgr5+ liver stem cells induced by Wnt-driven regeneration. Nature 494, 247–250 (2013).
Barker, N. et al. Lgr5(+ve) stem cells drive self-renewal in the stomach and build long-lived gastric units in vitro. Cell Stem Cell 6, 25–36 (2010).
Steinberg, Z. et al. FGFR2b signaling regulates ex vivo submandibular gland epithelial cell proliferation and branching morphogenesis. Development 132, 1223–1234 (2005).
Takasato, M. et al. Kidney organoids from human iPS cells contain multiple lineages and model human nephrogenesis. Nature 536, 238 (2016).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Dekkers, J. F. et al. A functional CFTR assay using primary cystic fibrosis intestinal organoids. Nat. Med. 19, 939–945 (2013).
Gabriel, E. et al. Recent Zika virus isolates induce premature differentiation of neural progenitors in human brain organoids. Cell Stem Cell 20, 397–406.e5 (2017).
Qian, X., Nguyen, H. N., Jacob, F., Song, H. & Ming, G. L. Using brain organoids to understand Zika virus-induced microcephaly. Development 144, 952–957 (2017).
Cheung, K. J., Gabrielson, E., Werb, Z. & Ewald, A. J. Collective invasion in breast cancer requires a conserved basal epithelial program. Cell 155, 1639–1651 (2013).
Cheung, K. J. et al. Polyclonal breast cancer metastases arise from collective dissemination of keratin 14-expressing tumor cell clusters. Proc. Natl Acad. Sci. U. S. A. 113, E854–E863 (2016).
Tiriac, H. et al. Organoid profiling identifies common responders to chemotherapy in pancreatic cancer. Cancer Discov. 8, 1112–1129 (2018).
Tuveson, D. & Clevers, H. Cancer modeling meets human organoid technology. Science 364, 952–955 (2019).
Neal, J. T. et al. Organoid modeling of the tumor immune microenvironment. Cell 175, 1972–1988.e16 (2018).
Padmanaban, V. et al. E-cadherin is required for metastasis in multiple models of breast cancer. Nature 573, 439–444 (2019).
Drost, J. & Clevers, H. Organoids in cancer research. Nat. Rev. Cancer 18, 407–418 (2018).
Vlachogiannis, G. et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science 359, 920–926 (2018).
Lee, S. H. et al. Tumor evolution and drug response in patient-derived organoid models of bladder cancer. Cell 173, 515–528.e17 (2018).
Shamir, E. R. & Ewald, A. J. Three-dimensional organotypic culture: experimental models of mammalian biology and disease. Nat. Rev. Mol. Cell Biol. 15, 647–664 (2014).
Sirka, O. K., Shamir, E. R. & Ewald, A. J. Myoepithelial cells are a dynamic barrier to epithelial dissemination. J. Cell Biol. 217, 368–2281 (2018).
Georgess, D. et al. Twist1-induced epithelial dissemination requires Prkd1 signaling. Cancer Res. 80, 204–218 (2020).
Clevers, H. Modeling development and disease with organoids. Cell 165, 1586–1597 (2016).
Huang, L. et al. Ductal pancreatic cancer modeling and drug screening using human pluripotent stem cell- and patient-derived tumor organoids. Nat. Med. 21, 1364–1371 (2015).
Takasato, M., Er, P. X., Chiu, H. S. & Little, M. H. Generation of kidney organoids from human pluripotent stem cells. Nat. Protoc. 11, 1681–1692 (2016).
Tammela, T. et al. A Wnt-producing niche drives proliferative potential and progression in lung adenocarcinoma. Nature 545, 355–359 (2017).
Kalabis, J. et al. Isolation and characterization of mouse and human esophageal epithelial cells in 3D organotypic culture. Nat. Protoc. 7, 235–246 (2012).
Zhang, X., Bush, K. T. & Nigam, S. K. In vitro culture of embryonic kidney rudiments and isolated ureteric buds. Methods Mol. Biol. 886, 13–21 (2012).
Lancaster, M. A. & Knoblich, J. A. Generation of cerebral organoids from human pluripotent stem cells. Nat. Protoc. 9, 2329–2340 (2014).
Unbekandt, M. & Davies, J. A. Dissociation of embryonic kidneys followed by reaggregation allows the formation of renal tissues. Kidney Int. 77, 407–416 (2010).
Debnath, J., Muthuswamy, S. K. & Brugge, J. S. Morphogenesis and oncogenesis of MCF-10A mammary epithelial acini grown in three-dimensional basement membrane cultures. Methods 30, 256–268 (2003).
Takebe, T. et al. Vascularized and functional human liver from an iPSC-derived organ bud transplant. Nature 499, 481–484 (2013).
Shamir, E. R. et al. Twist1-induced dissemination preserves epithelial identity and requires E-cadherin. J. Cell Biol. 204, 839–856 (2014).
Nuciforo, S. et al. Organoid models of human liver cancers derived from tumor needle biopsies. Cell Rep. 24, 1363–1376 (2018).
Nguyen-Ngoc, K. V. et al. 3D culture assays of murine mammary branching morphogenesis and epithelial invasion. Methods Mol. Biol. 1189, 135–162 (2015).
Guy, C. T., Cardiff, R. D. & Muller, W. J. Induction of mammary tumors by expression of polyomavirus middle T oncogene: a transgenic mouse model for metastatic disease. Mol. Cell. Biol. 12, 954–961 (1992).
Green, J. E. et al. The C3(1)/SV40 T-antigen transgenic mouse model of mammary cancer: ductal epithelial cell targeting with multistage progression to carcinoma. Oncogene 19, 1020–1027 (2000).
Dhanasekaran, R. et al. MYC and Twist1 cooperate to drive metastasis by eliciting crosstalk between cancer and innate immunity. Elife 9, e50731 (2020).
Nguyen-Ngoc, K. V. & Ewald, A. J. Mammary ductal elongation and myoepithelial migration are regulated by the composition of the extracellular matrix. J. Microsc. 251, 212–223 (2013).
Acknowledgements
We thank members of the Ewald laboratory for helpful comments on the manuscript and for sharing organoid yield information (data points in Fig. 2f). We also thank the Cooperative Human Tissue Network (CHTN) for providing patient samples used in this study. We thank Jin Zhu for assistance with FACS experiments. K.J.C. was supported by the Burroughs Welcome Fund Career Award for Medical Scientists 1013355.01. D.G. was supported by a Postdoctoral Fellowship Grant from the Susan G. Komen Foundation (PDF15332336). A.J.E. received support for this project through grants from The Breast Cancer Research Foundation (BCRF-18-048), the Metastatic Breast Cancer Network, Twisted Pink, Hope Scarves, Theresa’s Research Foundation and the National Institutes of Health/National Cancer Institute (U01CA217846, U01CA212007, U54CA2101732 and 3P30CA006973).
Author information
Authors and Affiliations
Contributions
V.P., K.J.C., E.M.G, N.M.N., A.K.F., E.H., D.G. and A.J.E. contributed to the development and optimization of protocols described in this manuscript. W.M. and P.T.T. provided valuable advice on adaptations of these methods to different model systems. V.P., E.M.G. and A.J.E. wrote the manuscript with useful input from all authors.
Corresponding author
Ethics declarations
Competing interests
A.J.E. and K.J.C. have a patent related to the use of keratin-14 as a biomarker for invasive cancer cell populations. A.J.E. and V.P. have a patent related to the use of antibodies for cancer therapy. A.J.E.’s spouse is an employee of Immunocore.
Additional information
Peer review information Nature Protocols thanks Johan de Rooij and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Padmanaban, V. et al. Nature 573, 439–444 (2019): https://doi.org/10.1038/s41586-019-1526-3
Cheung, K. et al. Proc. Natl Acad. Sci. USA 113, E854–E863 (2016): https://doi.org/10.1073/pnas.1508541113
Cheung, K. et al. Cell 155, 1639–1651 (2013): https://doi.org/10.1016/j.cell.2013.11.029
Extended Data
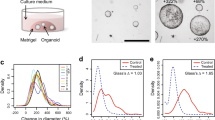
Extended Data Fig. 1 Variables that affect organoid yield from mammary human tumor organoids.
a, Organoid yield increased as the protocol was optimized during the course of the study. b, Variations in organoid yield based on modifications to the protocol. Median with SD is represented.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 7
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Padmanaban, V., Grasset, E.M., Neumann, N.M. et al. Organotypic culture assays for murine and human primary and metastatic-site tumors. Nat Protoc 15, 2413–2442 (2020). https://doi.org/10.1038/s41596-020-0335-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-0335-3
This article is cited by
-
Exploring the interaction between extracellular matrix components in a 3D organoid disease model to replicate the pathophysiology of breast cancer
Journal of Experimental & Clinical Cancer Research (2023)
-
Triple negative breast tumors contain heterogeneous cancer cells expressing distinct KRAS-dependent collective and disseminative invasion programs
Oncogene (2023)
-
Role of adipose tissue-derived cytokines in the progression of inflammatory breast cancer in patients with obesity
Lipids in Health and Disease (2022)
-
Leucyl-tRNA synthetase is a tumour suppressor in breast cancer and regulates codon-dependent translation dynamics
Nature Cell Biology (2022)
-
Morphological screening of mesenchymal mammary tumor organoids to identify drugs that reverse epithelial-mesenchymal transition
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.