Abstract
The development of single-molecule switching (SMS) fluorescence microscopy (also called single-molecule localization microscopy) over the last decade has enabled researchers to image cell biological structures at unprecedented resolution. Using two opposing objectives in a so-called 4Pi geometry doubles the available numerical aperture, and coupling this with interferometric detection has demonstrated 3D resolution down to 10 nm over entire cellular volumes. The aim of this protocol is to enable interested researchers to establish 4Pi-SMS super-resolution microscopy in their laboratories. We describe in detail how to assemble the optomechanical components of a 4Pi-SMS instrument, align its optical beampath and test its performance. The protocol further provides instructions on how to prepare test samples of fluorescent beads, operate this instrument to acquire images of whole cells and analyze the raw image data to reconstruct super-resolution 3D data sets. Furthermore, we provide a troubleshooting guide and present examples of anticipated results. An experienced optical instrument builder will require ~12 months from the start of ordering hardware components to acquiring high-quality biological images.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

























Similar content being viewed by others
Data availability
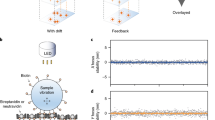
Software and hardware plans are available from online repositories. The CAD file repository (Autodesk Inventor) can be found at https://github.com/4Pi-SMS-consortium/CAD-files. The complete parts list is available at https://github.com/4Pi-SMS-consortium/CAD-files/blob/master/PartsList.xlsx and as Supplementary Table 1. The microscope control software can be downloaded from https://github.com/Gurdon-Super-Res-Lab/Microscope-Control. The raw image files used to create Fig. 25a–c and Fig. 25d–f are available via the Zenodo online repositories: https://zenodo.org/record/3929647#.X6Q-31NKgdU and https://zenodo.org/record/4022827#.X6Q_GVNKgdU, respectively.
Code availability
The data analysis software referenced in this paper is available online at https://github.com/4Pi-SMS-consortium/4Pi-SMS-analysis.
References
Shtengel, G. et al. Interferometric fluorescent super-resolution microscopy resolves 3D cellular ultrastructure. Proc. Natl Acad. Sci. USA 106, 3125–3130 (2009).
Aquino, D. et al. Two-color nanoscopy of three-dimensional volumes by 4Pi detection of stochastically switched fluorophores. Nat. Methods 8, 353–359 (2011).
Wang, G., Hauver, J., Thomas, Z., Darst, S. A. & Pertsinidis, A. Single-molecule real-time 3D imaging of the transcription cycle by modulation interferometry. Cell 167, 1839–1852.e21 (2016).
Huang, F. et al. Ultra-high resolution 3D imaging of whole cells. Cell 166, 1028–1040 (2016).
Del Viso, F. et al. Congenital heart disease genetics uncovers context-dependent organization and function of nucleoporins at cilia. Dev. Cell 38, 478–492 (2016).
Zhang, Y., Lara-Tejero, M., Bewersdorf, J. & Galan, J. E. Visualization and characterization of individual type III protein secretion machines in live bacteria. Proc. Natl Acad. Sci. USA 114, 6098–6103 (2017).
Hwang, J. Y. et al. Dual sensing of physiologic pH and calcium by EFCAB9 regulates sperm motility. Cell 177, 1480–1494.e19 (2019).
Schroeder, L. K. et al. Dynamic nanoscale morphology of the ER surveyed by STED microscopy. J. Cell Biol 218, 83–96 (2019).
Zhang, Y. et al. Nanoscale subcellular architecture revealed by multicolor three-dimensional salvaged fluorescence imaging. Nat. Methods 17, 225–231 (2020).
Karanastasis, A. A. et al. 3D mapping of nanoscale crosslink heterogeneities in microgels. Mater. Horiz. 5, 1130–1136 (2018).
Brown, T. A. et al. Superresolution fluorescence imaging of mitochondrial nucleoids reveals their spatial range, limits, and membrane interaction. Mol. Cell. Biol. 31, 4994–5010 (2011).
Schermelleh, L. et al. Super-resolution microscopy demystified. Nat. Cell Biol. 21, 72–84 (2019).
Barentine, A. E. S. et al. 3D multicolor nanoscopy at 10,000 cells a day. Preprint at https://www.biorxiv.org/content/10.1101/606954v1 (2019).
Booth, M. J. Adaptive optical microscopy: the ongoing quest for a perfect image. Light Sci. Appl. 3, e165 (2014).
Burke, D., Patton, B., Huang, F., Bewersdorf, J. & Booth, M. J. Adaptive optics correction of specimen-induced aberrations in single-molecule switching microscopy. Optica 2, 177–185 (2015).
Van Engelenburg, S. B. et al. Distribution of ESCRT machinery at HIV assembly sites reveals virus scaffolding of ESCRT subunits. Science 343, 653–656 (2014).
Buttler, C. A. et al. Single molecule fate of HIV-1 envelope reveals late-stage viral lattice incorporation. Nat. Commun. 9, 1861 (2018).
Liu, S. & Huang, F. Enhanced 4Pi single-molecule localization microscopy with coherent pupil based localization. Commun. Biol. 3, 220 (2020).
Li, Y. et al. Accurate 4Pi single-molecule localization using an experimental PSF model. Opt. Lett. 45, 3765–3768 (2020).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Wang, Y. et al. Localization events-based sample drift correction for localization microscopy with redundant cross-correlation algorithm. Opt. Express 22, 15982–15991 (2014).
Li, Y. et al. Real-time 3D single-molecule localization using experimental point spread functions. Nat. Methods 15, 367–369 (2018).
Thevathasan, J. V. et al. Nuclear pores as versatile reference standards for quantitative superresolution microscopy. Nat. Methods 16, 1045–1053 (2019).
Antonello, J., Wang, J., He, C., Phillips, M. & Booth, M. Interferometric calibration of a deformable mirror. Zenodo. 19 March 2020 (accessed 21 November 2020) https://zenodo.org/record/3714951#.X6RM2lNKgdU
Acknowledgements
We thank Fang Huang for advice, discussions and original code for the analysis software; Jacopo Antonello for advice, discussions and help with setting up the DMs; Chao He for assembling the DM calibration tool; David Miguel Susano Pinto for help and discussions with the analysis code; Andrew Barentine and Zach Marin for help and discussion about the analysis code; and David Baddeley for imaging advice, helpful discussions and help with the analysis code. J.W., M.A.P. and I.M.D. were supported by John Fell Fund award 141/144 and Wellcome Trust awards 105605/Z/14/Z and 107457/Z/15/Z. E.S.A. was supported by Wellcome Trust awards 095927/B/11/Z and 203285/Z/16/Z. G.S. was supported by Wellcome Trust awards 095927/B/11/Z and 203144/Z/16/Z. Y.Z., K.H., M.D.L. and J.B. were supported by Wellcome Trust awards 095927/A/11/Z and 203285/B/16/Z and National Institutes of Health (NIH) award R01 GM118486. K.H. was additionally supported by NIH award T32EB019941. R.D. was supported by an award from the Engelhorn Foundation. J.R. and Y.L. were supported by European Research Council award ERC CoG-724489, funding from the EMBL and the 4D Nucleome/4DN NIH Common Fund award U01 EB021223. Y.L. was additionally supported by the EMBL Interdisciplinary Postdoc Programme (EIPOD) under Marie Curie Actions COFUND and a start-up grant from the Southern University of Science and Technology, China. M.J.B. was supported by European Research Council award AdOMIS 695140 and Wellcome Trust award 203285/C/16/Z.
Author information
Authors and Affiliations
Contributions
Hardware development: J.B., M.J.B., Y.Z., M.A.P., J.W., Y.L., E.S.A. and G.S.; software development: Y.Z., Y.L. and E.S.A.; specimen/imaging protocols: Y.Z. and M.D.L.; alignment protocols: E.S.A., G.S., J.W., Y.L., Y.Z. and K.H.; index matching protocol: R.D., J.R. and Y.L.; project supervision: J.B., M.J.B., J.R. and I.M.D.; writing and editing of the manuscript: all authors.
Corresponding authors
Ethics declarations
Competing interests
J.B. has financial interests in Bruker Corp. and Hamamatsu Photonics. J.B. is co-inventor of a US patent application (US20170251191A1) related to the 4Pi-SMS system and image analysis used in this work. Y.Z. and J.B. have filed a US patent application about the salvaged fluorescence multicolor imaging method described in this work.
Additional information
Peer review information Nature Protocols thanks Ilaria Testa and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Huang, F. et al. Cell 166, 1028–1040 (2016): https://doi.org/10.1016/j.cell.2016.06.016
Zhang, Y. et al. Nat. Methods 17, 225–231 (2020): https://doi.org/10.1038/s41592-019-0676-4
Zhang, Y. et al. Proc. Natl Acad. Sci. USA 114, 6098–6103 (2017): https://doi.org/10.1073/pnas.1705823114
Supplementary information
Supplementary Information
Supplementary Methods and Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Wang, J., Allgeyer, E.S., Sirinakis, G. et al. Implementation of a 4Pi-SMS super-resolution microscope. Nat Protoc 16, 677–727 (2021). https://doi.org/10.1038/s41596-020-00428-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-00428-7
This article is cited by
-
Vectorial adaptive optics
eLight (2023)
-
Three-dimensional single particle tracking using 4π self-interference of temporally phase-shifted fluorescence
Light: Science & Applications (2023)
-
Mastering lanthanide energy states for next-gen photonic innovation
Science China Chemistry (2023)
-
Dynamics and functions of E-cadherin complexes in epithelial cell and tissue morphogenesis
Marine Life Science & Technology (2023)
-
Global fitting for high-accuracy multi-channel single-molecule localization
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.