Abstract
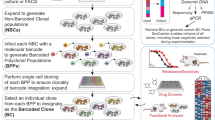
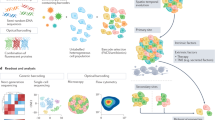
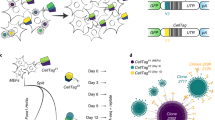
Embedded viral barcoding in combination with high-throughput sequencing is a powerful technology with which to track single-cell clones. It can provide clonal-level insights into cellular proliferation, development, differentiation, migration, and treatment efficacy. Here, we present a detailed protocol for a viral barcoding procedure that includes the creation of barcode libraries, the viral delivery of barcodes, the recovery of barcodes, and the computational analysis of barcode sequencing data. The entire procedure can be completed within a few weeks. This barcoding method requires cells to be susceptible to viral transduction. It provides high sensitivity and throughput, and enables precise quantification of cellular progeny. It is cost efficient and does not require any advanced skills. It can also be easily adapted to many types of applications, including both in vitro and in vivo experiments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
A sample dataset has been deposited in Figshare: https://doi.org/10.35092/yhjc.11374446. This dataset was used to generate Figs. 4 and 5.
Code availability
The Python scripts have been provided in the Supplementary Software of this paper. The code in this paper has been peer-reviewed.
References
Lu, R., Neff, N. F., Quake, S. R. & Weissman, I. L. Tracking single hematopoietic stem cells in vivo using high-throughput sequencing in conjunction with viral genetic barcoding. Nat. Biotechnol. 29, 928–933 (2011).
Dick, J. E., Magli, M. C., Huszar, D., Phillips, R. A. & Bernstein, A. Introduction of a selectable gene into primitive stem cells capable of long-term reconstitution of the hemopoietic system of W/Wv mice. Cell 42, 71–79 (1985).
Keller, G., Paige, C., Gilboa, E. & Wagner, E. F. Expression of a foreign gene in myeloid and lymphoid cells derived from multipotent haematopoietic precursors. Nature 318, 149–154 (1985).
Lemischka, I. R., Raulet, D. H. & Mulligan, R. C. Developmental potential and dynamic behavior of hematopoietic stem cells. Cell 45, 917–927 (1986).
Schepers, K. et al. Dissecting T cell lineage relationships by cellular barcoding. J. Exp. Med 205, 2309–2318 (2008).
Lyne, A.-M. et al. A track of the clones: new developments in cellular barcoding. Exp. Hematol. 68, 15–20 (2018).
Naik, S. H. et al. Diverse and heritable lineage imprinting of early haematopoietic progenitors. Nature 496, 229–232 (2013).
Cheung, A. M. S. et al. Analysis of the clonal growth and differentiation dynamics of primitive barcoded human cord blood cells in NSG mice. Blood 122, 3129–3137 (2013).
Nguyen, L. V. et al. Clonal analysis via barcoding reveals diverse growth and differentiation of transplanted mouse and human mammary stem cells. Cell Stem Cell 14, 253–263 (2014).
Gerrits, A. et al. Cellular barcoding tool for clonal analysis in the hematopoietic system. Blood 115, 2610–2618 (2010).
Nguyen, L. et al. Functional compensation between hematopoietic stem cell clones in vivo. EMBO Rep. 19, e45702(2018).
Brewer, C., Chu, E., Chin, M. & Lu, R. Transplantation dose alters the differentiation program of hematopoietic stem cells. Cell Rep 15, 1848–1857 (2016).
Wu, C. et al. Clonal tracking of rhesus macaque hematopoiesis highlights a distinct lineage origin for natural killer cells. Cell Stem Cell 14, 486–499 (2014).
Lu, R., Czechowicz, A., Seita, J., Jiang, D. & Weissman, I. L. Clonal-level lineage commitment pathways of hematopoietic stem cells in vivo. Proc. Natl Acad. Sci. USA 116, 1447–1456 (2019).
Bystrykh, L. V., de Haan, G. & Verovskaya, E. Barcoded vector libraries and retroviral or lentiviral barcoding of hematopoietic stem cells in Hematopoietic Stem Cell Protocols (eds Bunting, K. D. & Qu, C.-K.) 345–360 (Springer New York, 2014).
Bystrykh, L. V. & Belderbos, M. E. Clonal analysis of cells with cellular barcoding: when numbers and sizes matter in Stem Cell Heterogeneity: Methods and Protocols (ed Turksen, K.) 57–89 (Springer New York, 2016).
Naik, S. H., Schumacher, T. N. & Perié, L. Cellular barcoding: a technical appraisal. Exp. Hematol. 42, 598–608 (2014).
Thielecke, L. et al. Limitations and challenges of genetic barcode quantification. Sci. Rep 7, 43249 (2017).
Woodworth, M. B., Girskis, K. M. & Walsh, C. A. Building a lineage from single cells: genetic techniques for cell lineage tracking. Nat. Rev. Genet. 18, 230 (2017).
Kebschull, J. M. & Zador, A. M. Cellular barcoding: lineage tracing, screening and beyond. Nat. Methods 15, 871–879 (2018).
Nguyen, L. V. et al. DNA barcoding reveals diverse growth kinetics of human breast tumour subclones in serially passaged xenografts. Nat. Commun. 5, 5871 (2014).
Nguyen, L. V. et al. Barcoding reveals complex clonal dynamics of de novo transformed human mammary cells. Nature 528, 267 (2015).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR-Cas9 aystem. Science 343, 80 (2014).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84 (2014).
Dykstra, B. et al. Long-term propagation of distinct hematopoietic differentiation programs in vivo. Cell Stem Cell 1, 218–229 (2007).
Sieburg, H. B. et al. The hematopoietic stem compartment consists of a limited number of discrete stem cell subsets. Blood 107, 2311–2316 (2006).
Osawa, M., Hanada, K., Hamada, H. & Nakauchi, H. Long-term lymphohematopoietic reconstitution by a single CD34-low/negative hematopoietic stem cell. Science 273, 242 (1996).
Weber, K., Thomaschewski, M., Benten, D. & Fehse, B. RGB marking with lentiviral vectors for multicolor clonal cell tracking. Nat. Protoc. 7, 839 (2012).
Cornils, K. et al. Multiplexing clonality: combining RGB marking and genetic barcoding. Nucleic Acids Res. 42, e56–e56 (2014).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56 (2007).
Rios, A. C., Fu, N. Y., Lindeman, G. J. & Visvader, J. E. In situ identification of bipotent stem cells in the mammary gland. Nature 506, 322 (2014).
Schmidt, M. et al. High-resolution insertion-site analysis by linear amplification–mediated PCR (LAM-PCR). Nat. Methods 4, 1051 (2007).
Harkey, M. A. et al. Multiarm high-throughput integration site detection: limitations of LAM-PCR technology and optimization for clonal analysis. Stem Cells Dev 16, 381–392 (2007).
Wu, C. et al. High efficiency restriction enzyme-free linear amplification-mediated polymerase chain reaction approach for tracking lentiviral integration sites does not abrogate retrieval bias. Hum. Gene Ther. 24, 38–47 (2013).
Wu, C. et al. Tracking retroviral-integrated clones with modified non-restriction enzyme LAM-PCR technology. Mol. Ther. 19, S45 (2011).
Zhou, S. et al. Quantitative shearing linear amplification polymerase chain reaction: an improved method for quantifying lentiviral vector insertion sites in transplanted hematopoietic cell systems. Hum. Gene Ther. Methods 26, 4–12 (2014).
Sun, J. et al. Clonal dynamics of native haematopoiesis. Nature 514, 322–327 (2014).
Pei, W. et al. Polylox barcoding reveals haematopoietic stem cell fates realized in vivo. Nature 548, 456–460 (2017).
Frieda, K. L. et al. Synthetic recording and in situ readout of lineage information in single cells. Nature 541, 107 (2016).
Kalhor, R. et al. Developmental barcoding of whole mouse via homing CRISPR. Science 361, eaat9804 (2018).
Rüfer, A. W. & Sauer, B. Non-contact positions impose site selectivity on Cre recombinase. Nucleic Acids Res 30, 2764–2771 (2002).
Shen, M. W. et al. Predictable and precise template-free CRISPR editing of pathogenic variants. Nature 563, 646–651 (2018).
Lee-Six, H. et al. Population dynamics of normal human blood inferred from somatic mutations. Nature 561, 473–478 (2018).
Osorio, F. G. et al. Somatic mutations reveal lineage relationships and age-related mutagenesis in human hematopoiesis. Cell Rep. 25, 2308–2316.e4 (2018).
Chapal-Ilani, N. et al. Comparing algorithms that reconstruct cell lineage trees utilizing information on microsatellite mutations. PLoS Comput. Biol. 9, e1003297 (2013).
Wasserstrom, A. et al. Reconstruction of cell lineage trees in mice. PLoS ONE 3, e1939 (2008).
McKenzie, J. L., Gan, O. I., Doedens, M., Wang, J. C. Y. & Dick, J. E. Individual stem cells with highly variable proliferation and self-renewal properties comprise the human hematopoietic stem cell compartment. Nat. Immunol. 7, 1225 (2006).
Gonzalez-Murillo, A., Lozano, M. L., Montini, E., Bueren, J. A. & Guenechea, G. Unaltered repopulation properties of mouse hematopoietic stem cells transduced with lentiviral vectors. Blood 112, 3138–3147 (2008).
Wu, C. et al. Clonal expansion and compartmentalized maintenance of rhesus macaque NK cell subsets. Sci. Immunol. 3, aat9781 (2018).
Bystrykh, L. V., Verovskaya, E., Zwart, E., Broekhuis, M. & de Haan, G. Counting stem cells: methodological constraints. Nat. Methods 9, 567 (2012).
Merino, D. et al. Barcoding reveals complex clonal behavior in patient-derived xenografts of metastatic triple negative breast cancer. Nat. Commun. 10, 766 (2019).
Guernet, A. et al. CRISPR-barcoding for intratumor genetic heterogeneity modeling and functional analysis of oncogenic driver mutations. Mol. Cell 63, 526–538 (2016).
Verovskaya, E. et al. Heterogeneity of young and aged murine hematopoietic stem cells revealed by quantitative clonal analysis using cellular barcoding. Blood 122, 523–532 (2013).
Thielecke, L., Cornils, K. & Glauche, I. genBaRcode: a comprehensive R package for genetic barcode analysis. Bioinformatics https://doi.org/10.1093/bioinformatics/btz872 (2019).
Acknowledgements
We thank all members of the Lu lab for helping to optimize the protocol and C. Lytal for help editing the text. We thank the USC Stem Cell Flow Cytometry Facility and CHLA Sequencing Core for technical support. This research was funded primarily by a National Institutes of Health (NIH) R00 early investigator grant (NIH-R00-HL113104) and R01 grants (R01HL135292 and R01HL138225). R.L. is a Scholar of the Leukemia & Lymphoma Society and a Richard N. Merkin Assistant Professor. The project described was supported in part by award no. P30CA014089 from the National Cancer Institute. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Cancer Institute or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
R.L. conceived and developed the protocol. C.B., D.J., J.C., and A.N. optimized the barcode extraction protocol. D.J. and J.E. improved the Python code for analyzing high-throughput sequencing data. C.B., D.J., and R.L. prepared the manuscript. J.C. and A.N. provided assistance in manuscript text preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Leïla Périé and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Lu, R., Neff, N. R., Quake, S. R. & Weissman, I. L. Nat. Biotechnol. 29, 928–933 (2011): https://doi.org/10.1038/nbt.1977
Nguyen, L. et al. EMBO Rep. 19, e45702 (2018): https://doi.org/10.15252/embr.201745702
Brewer, C., Chu, E., Chin, M. & Lu, R. Cell Rep. 15, 1848–1857 (2016): https://doi.org/10.1016/j.celrep.2016.04.061
Supplementary information
Supplementary Software
The software for data analysis (Steps 121–127)
Rights and permissions
About this article
Cite this article
Bramlett, C., Jiang, D., Nogalska, A. et al. Clonal tracking using embedded viral barcoding and high-throughput sequencing. Nat Protoc 15, 1436–1458 (2020). https://doi.org/10.1038/s41596-019-0290-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0290-z
This article is cited by
-
Lineage tracking to reveal the fate of hematopoietic stem cells influenced by Flk2− multipotent progenitors after transplantation
Experimental & Molecular Medicine (2023)
-
Panhematopoietic RNA barcoding enables kinetic measurements of nucleate and anucleate lineages and the activation of myeloid clones following acute platelet depletion
Genome Biology (2023)
-
Connecting past and present: single-cell lineage tracing
Protein & Cell (2022)
-
Interrogation of clonal tracking data using barcodetrackR
Nature Computational Science (2021)
-
BAR-Seq clonal tracking of gene-edited cells
Nature Protocols (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.