Abstract
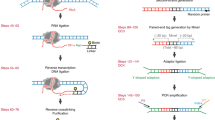
Methods developed to capture protein-anchored chromatin interactions (chromatin interaction analysis by paired-end tag sequencing and HiChIP) have yielded tremendous insights into the 3D folding principles of the genome, but are normalized by sequencing depth and therefore unable to accurately measure global changes in chromatin interactions and contact domain organization. We herein describe the protocol for absolute quantification of chromatin architecture (AQuA)–HiChIP, an advance that allows the absolute differences in protein-anchored chromatin interactions between samples to be determined. With our method, defined ratios of mouse and human fixed nuclei are mixed and subjected to endonuclease digestion. Chromatin contacts are captured by biotin-dATP incorporation and proximity ligation, followed by gentle shearing, ChIP, biotin capture and paired-end sequencing. 3D contacts are counted from paired-end tags (PETs) from the human genome and are normalized to the total PETs from the mouse genome. As orthogonal normalization allows observation of global changes, the approach will enable more quantitative insights into the topological determinants of transcriptional control and tissue-specific epigenetic memory. With our approach, we have discovered that rapid histone deacetylase inhibition disrupts super enhancer function by creating many new aberrant contacts. The code for data analysis is available at https://github.com/GryderArt/AQuA-HiChIP. This protocol reports both experimental and bioinformatic details to perform AQuA-HiChIP, going from cell culture to ranking chromatin interactions within 6 d.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Data generated by this protocol, and visualized herein, is available through Gene Expression Omnibus, accession number GSE120770.
Code availability
All code used herein is either provided by other research laboratories (see links throughout the protocol for any given step) or is custom scripted in R (available here: https://github.com/GryderArt/AQuA-HiChIP). The code in this protocol has been peer reviewed.
References
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860 (2001).
Roh, T.-y, Ngau, W. C., Cui, K., Landsman, D. & Zhao, K. High-resolution genome-wide mapping of histone modifications. Nat. Biotechnol. 22, 1013 (2004).
Roh, T.-Y., Cuddapah, S. & Zhao, K. Active chromatin domains are defined by acetylation islands revealed by genome-wide mapping. Genes Dev. 19, 542–552 (2005).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
ENCODE Project Consortium. The ENCODE (ENCyclopedia of DNA elements) project. Science 306, 636–640 (2004).
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
Pennisi, E. ENCODE Project writes eulogy for junk DNA. Science 337, 1159–1161 (2012).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Fullwood, M. J. et al. An oestrogen-receptor-α-bound human chromatin interactome. Nature 462, 58 (2009).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Mumbach, M. R. et al. Enhancer connectome in primary human cells identifies target genes of disease-associated DNA elements. Nat. Genet. 49, 1602 (2017).
Goldberg, A. D., Allis, C. D. & Bernstein, E. Epigenetics: a landscape takes shape. Cell 128, 635–638 (2007).
Orlando, D. A. et al. Quantitative ChIP-seq normalization reveals global modulation of the epigenome. Cell Rep. 9, 1163–1170 (2014).
Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Lai, B. et al. Trac-looping measures genome structure and chromatin accessibility. Nat. Methods 15, 741–747 (2018).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917 (2009).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640 (2007).
Nabet, B. et al. The dTAG system for immediate and target-specific protein degradation. Nat. Chem. Biol. 14, 431 (2018).
Pogo, B., Allfrey, V. & Mirsky, A. RNA synthesis and histone acetylation during the course of gene activation in lymphocytes. Proc. Natl Acad. Sci. USA 55, 805–812 (1966).
Gryder, B. E. et al. Histone hyperacetylation disrupts core gene regulatory architecture in rhabdomyosarcoma. Nat. Genet 51, 1714–1722 (2019).
Yohe, M. E. et al. MEK inhibition induces MYOG and remodels super-enhancers in RAS-driven rhabdomyosarcoma. Sci. Transl. Med. 10, eaan4470 (2018).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Gryder, B. E. et al. PAX3-FOXO1 establishes myogenic super enhancers and confers BET bromodomain vulnerability. Cancer Disco. 7, 884–899 (2017).
Stanton, B. Z. et al. Smarca4 ATPase mutations disrupt direct eviction of PRC1 from chromatin. Nat. Genet. 49, 282–288 (2017).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based Analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Stansfield, J. C., Cresswell, K. G., Vladimirov, V. I. & Dozmorov, M. G. HiCcompare: an R-package for joint normalization and comparison of HI-C datasets. BMC Bioinforma. 19, 279 (2018).
Acknowledgements
We gratefully acknowledge Y. Song and the NCI Sequencing Core for assistance. We are grateful to Tom Misteli and Justin Demmerle for helpful conversations relating to this work. We thank Emma Chory for technical advice. We wish to honor the lasting memory of Joseph P. Calarco. This work was facilitated by Biowulf High Performance Computing Systems and enabled by funding from the Division of Intramural Research from NIH NCI CCR.
Author information
Authors and Affiliations
Contributions
B.Z.S, B.E.G. and J.K. conceived of the project. B.Z.S. and B.E.G. performed AQuA-HiChIP experiments. B.E.G. built the AQuA analysis pipeline, B.Z.S. and B.E.G. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Gryder, B. E. et al. Nat. Genet. 51, 1714–1722 (2019): https://doi.org/10.1038/s41588-019-0534-4
Stanton, B. et al. Preprint at https://protocolexchange.researchsquare.com/article/nprot-7121/v1 (2018): https://doi.org/10.1038/protex.2018.130
Supplementary information
Rights and permissions
About this article
Cite this article
Gryder, B.E., Khan, J. & Stanton, B.Z. Measurement of differential chromatin interactions with absolute quantification of architecture (AQuA-HiChIP). Nat Protoc 15, 1209–1236 (2020). https://doi.org/10.1038/s41596-019-0285-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0285-9
This article is cited by
-
PAX3-FOXO1 uses its activation domain to recruit CBP/P300 and shape RNA Pol2 cluster distribution
Nature Communications (2023)
-
3D chromatin architecture and transcription regulation in cancer
Journal of Hematology & Oncology (2022)
-
Chromatin structure in cancer
BMC Molecular and Cell Biology (2022)
-
FOXF1 is required for the oncogenic properties of PAX3-FOXO1 in rhabdomyosarcoma
Oncogene (2021)
-
Interaction between SNAI2 and MYOD enhances oncogenesis and suppresses differentiation in Fusion Negative Rhabdomyosarcoma
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.