Abstract
Gram-negative and Gram-positive bacteria release a variety of membrane vesicles through different formation routes. Knowledge of the structure, molecular cargo and function of bacterial extracellular vesicles (BEVs) is primarily obtained from bacteria cultured in laboratory conditions. BEVs in human body fluids have been less thoroughly investigated most probably due to the methodological challenges in separating BEVs from their matrix and host-derived eukaryotic extracellular vesicles (EEVs) such as exosomes and microvesicles. Here, we present a step-by-step procedure to separate and characterize BEVs from human body fluids. BEVs are separated through the orthogonal implementation of ultrafiltration, size-exclusion chromatography (SEC) and density-gradient centrifugation. Size separates BEVs from bacteria, flagella and cell debris in stool; and blood cells, high density lipoproteins (HDLs) and soluble proteins in blood. Density separates BEVs from fibers, protein aggregates and EEVs in stool; and low-density lipoproteins (LDLs), very-low-density lipoproteins (VLDLs), chylomicrons, protein aggregates and EEVs in blood. The procedure is label free, maintains the integrity of BEVs and ensures reproducibility through the use of automated liquid handlers. Post-separation BEVs are characterized using orthogonal biochemical endotoxin and Toll-like receptor-based reporter assays in combination with proteomics, electron microscopy and nanoparticle tracking analysis (NTA) to evaluate BEV quality, abundance, structure and molecular cargo. Separation and characterization of BEVs from body fluids can be done within 72 h, is compatible with EEV analysis and can be readily adopted by researchers experienced in basic molecular biology and extracellular vesicle analysis. We anticipate that this protocol will expand our knowledge on the biological heterogeneity, molecular cargo and function of BEVs in human body fluids and steer the development of laboratory research tools and clinical diagnostic kits.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
The computer code used for the robot-assisted steps of the protocol (Steps 20A and 27A) is available from the corresponding author upon reasonable request.
References
Hughes, D. T. & Sperandio, V. Inter-kingdom signalling: communication between bacteria and their hosts. Nat. Rev. Microbiol. 6, 111–120 (2008).
Gopalakrishnan, V., Helmink, B. A., Spencer, C. N., Reuben, A. & Wargo, J. A. The influence of the gut microbiome on cancer, immunity, and cancer immunotherapy. Cancer Cell 33, 570–580 (2018).
Zitvogel, L., Ma, Y., Raoult, D., Kroemer, G. & Gajewski, T. F. The microbiome in cancer immunotherapy: diagnostic tools and therapeutic strategies. Science 359, 1366–1370 (2018).
Brenchley, J. M. et al. Microbial translocation is a cause of systemic immune activation in chronic HIV infection. Nat. Med. 12, 1365–1371 (2006).
Tanoue, T. et al. A defined commensal consortium elicits CD8 T cells and anti-cancer immunity. Nature 565, 600–605 (2019).
Tulkens, J. et al. Increased levels of systemic LPS-positive bacterial extracellular vesicles in patients with intestinal barrier dysfunction. Gut (2018). https://doi.org/10.1136/gutjnl-2018-317726
Toyofuku, M., Nomura, N. & Eberl, L. Types and origins of bacterial membrane vesicles. Nat. Rev. Microbiol. 17, 13–24 (2019).
Schwechheimer, C. & Kuehn, M. J. Outer-membrane vesicles from Gram-negative bacteria: biogenesis and functions. Nat. Rev. Microbiol. 13, 605–619 (2015).
Kaparakis-Liaskos, M. & Ferrero, R. L. Immune modulation by bacterial outer membrane vesicles. Nat. Rev. Immunol. 15, 375–387 (2015).
Brown, L., Wolf, J. M., Prados-Rosales, R. & Casadevall, A. Through the wall: extracellular vesicles in Gram-positive bacteria, mycobacteria and fungi. Nat. Rev. Microbiol. 13, 620–30 (2015).
Dauros Singorenko, P. et al. Isolation of membrane vesicles from prokaryotes: a technical and biological comparison reveals heterogeneity. J. Extracell. Vesicles 6, 1324731 (2017).
De Wever, O. & Hendrix, A. A supporting ecosystem to mature extracellular vesicles into clinical application. EMBO J. 38, e101412 (2019).
Sender, R., Fuchs, S. & Milo, R. Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 14, e1002533 (2016).
Namork, E. & Brandtzaeg, P. Fatal meningococcal septicaemia with “blebbing” meningococcus. Lancet 360, 1741 (2002).
Stephens, D. S., Edwards, K. M., Morris, F. & McGee, Z. A. Pili and outer membrane appendages on Neisseria meningitidis in the cerebrospinal fluid of an infant. J. Infect. Dis. 146, 568–568 (1982).
Park, K. S. et al. Sepsis-like systemic inflammation induced by nano-sized extracellular vesicles from feces. Front. Microbiol. 9, 1735 (2018).
Choi, H.-I. et al. Helicobacter pylori-derived extracellular vesicles increased in the gastric juices of gastric adenocarcinoma patients and induced inflammation mainly via specific targeting of gastric epithelial cells. Exp. Mol. Med. 49, e330 (2017).
Van Deun, J. et al. The impact of disparate isolation methods for extracellular vesicles on downstream RNA profiling. J. Extracell. Vesicles 3, 24858 (2014).
Simonsen, J. B. What are we looking at? Extracellular vesicles, lipoproteins, or both? Circ. Res. 121, 920–922 (2017).
Coumans, F. A. W. et al. Methodological guidelines to study extracellular vesicles. Circ. Res. 120, 1632–1648 (2017).
Van Deun, J. et al. EV-TRACK: transparent reporting and centralizing knowledge in extracellular vesicle research. Nat. Methods 14, 228–232 (2017).
Ford, T., Graham, J. & Rickwood, D. Iodixanol: a nonionic iso-osmotic centrifugation medium for the formation of self-generated gradients. Anal. Biochem. 220, 360–366 (1994).
Heintz-Buschart, A. et al. Small RNA profiling of low biomass samples: identification and removal of contaminants. BMC Biol. 16, 52 (2018).
Geeurickx, E. et al. The generation and use of recombinant extracellular vesicles as biological reference material. Nat. Commun. 10, 3288 (2019).
Théry, C. et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 7, 1535750 (2018).
Bauman, S. J. & Kuehn, M. J. Purification of outer membrane vesicles from Pseudomonas aeruginosa and their activation of an IL-8 response. Microbes Infect. 8, 2400–8 (2006).
Tashiro, Y. et al. Variation of physiochemical properties and cell association activity of membrane vesicles with growth phase in Pseudomonas aeruginosa. Appl. Environ. Microbiol. 76, 3732–3739 (2010).
Renelli, M., Matias, V., Lo, R. Y. & Beveridge, T. J. DNA-containing membrane vesicles of Pseudomonas aeruginosa PAO1 and their genetic transformation potential. Microbiology 150, 2161–2169 (2004).
Costea, P. I. et al. Towards standards for human fecal sample processing in metagenomic studies. Nat. Biotechnol. 35, 1069–1076 (2017).
Lacroix, R. et al. Impact of pre-analytical parameters on the measurement of circulating microparticles: towards standardization of protocol. J. Thromb. Haemost. 10, 437–46 (2012).
Witwer, K. W. et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J. Extracell. Vesicles 2, 20360 (2013).
Vandeputte, D. et al. Stool consistency is strongly associated with gut microbiota richness and composition, enterotypes and bacterial growth rates. Gut 65, 57–62 (2016).
Zhang, H. & Lyden, D. Asymmetric-flow field-flow fractionation technology for exomere and small extracellular vesicle separation and characterization. Nat. Protoc. 14, 1027–1053 (2019).
Zhang, H. et al. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nat. Cell Biol. 20, 332–343 (2018).
Vergauwen, G. et al. Confounding factors of ultrafiltration and protein analysis in extracellular vesicle research. Sci. Rep. 7, 2704 (2017).
Daneshian, M., von Aulock, S. & Hartung, T. Assessment of pyrogenic contaminations with validated human whole-blood assay. Nat. Protoc. 4, 1709–1721 (2009).
Vaara, M. Agents that increase the permeability of the outer membrane. Microbiol. Rev. 56, 395–411 (1992).
McCaig, W. D., Koller, A. & Thanassi, D. G. Production of outer membrane vesicles and outer membrane tubes by Francisella novicida. J. Bacteriol. 195, 1120–32 (2013).
Paolini, L. et al. Residual matrix from different separation techniques impacts exosome biological activity. Sci. Rep. 6, 23550 (2016).
Yang, P.-C. & Mahmood, T. Western blot: technique, theory, and trouble shooting. N. Am. J. Med. Sci. 4, 429 (2012).
Wiśniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Kanehisa, M., Sato, Y. & Morishima, K. BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J. Mol. Biol. 428, 726–731 (2016).
Wolf, P. The nature and significance of platelet products in human plasma. Br. J. Haematol. 13, 269–288 (1967).
van der Vlist, E. J., Nolte-’t Hoen, E. N. M., Stoorvogel, W., Arkesteijn, G. J. A. & Wauben, M. H. M. Fluorescent labeling of nano-sized vesicles released by cells and subsequent quantitative and qualitative analysis by high-resolution flow cytometry. Nat. Protoc. 7, 1311–1326 (2012).
Acknowledgements
This work was supported by Krediet aan Navorsers and a fellowship (A.H.) from Research Foundation Flanders (FWO) and Concerted Research Action from Ghent University.
Author information
Authors and Affiliations
Contributions
J.T., O.D.W. and A.H. acquired and analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Cherie Blenkiron and Masanori Toyofuku for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Tulkens, J. et al. Gut (2018): https://doi.org/10.1136/gutjnl-2018-317726
EV-TRACK Consortium, Van Deun, J. et al. Nat. Methods 14, 228–232 (2017): https://doi.org/10.1038/nmeth.4185
Van Deun, J. et al. J. Extracell. Vesicles 3, 24858 (2014): https://doi.org/10.3402/jev.v3.24858
Integrated supplementary information
Supplementary Fig. 1 BEV as potent stimulators of PBMC.
After stimulation of PBMC with BEVs, immediate release of chemo- and cytokines can be observed after performing Luminex assay. As proof of concept, we show here IL-8 and TNF-α concentration as a function of time after BEV stimulation. In this study, collection of blood and stool samples was according to Ethical Committee of Ghent University Hospital and in accordance to relevant guidelines. The patients provided informed written consent. PBMC, peripheral blood mononuclear cells.
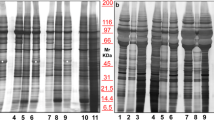
Supplementary Fig. 2 Evaluation of the performance of size exclusion chromatography.
At least 1×1010 EcN BEVs are spiked in 2 ml plasma or PBS and SEC is performed. LPS activity of the collected SEC fractions is measured by LAL assay. A first peak should be visible in SEC fractions 4-6 when the technique is performed correctly. Western blot (using anti-LPS (1:1,000) or anti-OmpA (1:5,000) antibodies) can further validate these results. Figure adapted from Tulkens et al.6. In this study, collection of blood samples was according to Ethical Committee of Ghent University Hospital and in accordance to relevant guidelines. The patients provided informed written consent. EcN, Escherichia coli Nissle 1917.
Supplementary information
Supplementary Information
Supplementary Figures 1 and 2
Supplementary Video 1
Robot-assisted preparation of density gradients.
Supplementary Video 2
Loading of a sample on a size-exclusion chromatography column.
Supplementary Video 3
Non-stimulated PBMCs recorded using a live-cell analysis system.
Supplementary Video 4
BEV-induced PBMC stimulation recorded using a live-cell analysis system.
Rights and permissions
About this article
Cite this article
Tulkens, J., De Wever, O. & Hendrix, A. Analyzing bacterial extracellular vesicles in human body fluids by orthogonal biophysical separation and biochemical characterization. Nat Protoc 15, 40–67 (2020). https://doi.org/10.1038/s41596-019-0236-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0236-5
This article is cited by
-
Exosome-based delivery strategies for tumor therapy: an update on modification, loading, and clinical application
Journal of Nanobiotechnology (2024)
-
Integrating automated liquid handling in the separation workflow of extracellular vesicles enhances specificity and reproducibility
Journal of Nanobiotechnology (2023)
-
Maternal microbiota communicates with the fetus through microbiota-derived extracellular vesicles
Microbiome (2023)
-
Composition and functions of bacterial membrane vesicles
Nature Reviews Microbiology (2023)
-
Extracellular vesicle analysis
Nature Reviews Methods Primers (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.