Abstract
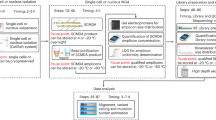
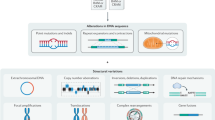
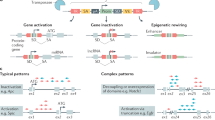
Mouse models of human cancer have transformed our ability to link genetics, molecular mechanisms and phenotypes. Both reverse and forward genetics in mice are currently gaining momentum through advances in next-generation sequencing (NGS). Methodologies to analyze sequencing data were, however, developed for humans and hence do not account for species-specific differences in genome structures and experimental setups. Here, we describe standardized computational pipelines specifically tailored to the analysis of mouse genomic data. We present novel tools and workflows for the detection of different alteration types, including single-nucleotide variants (SNVs), small insertions and deletions (indels), copy-number variations (CNVs), loss of heterozygosity (LOH) and complex rearrangements, such as in chromothripsis. Workflows have been extensively validated and cross-compared using multiple methodologies. We also give step-by-step guidance on the execution of individual analysis types, provide advice on data interpretation and make the complete code available online. The protocol takes 2–7 d, depending on the desired analyses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
















Similar content being viewed by others
Data availability
NGS data from mouse pancreatic cancer cell cultures are available from the European Nucleotide Archive using study accession no. PRJEB23787. The validation datasets generated during the current study are available from the corresponding author upon request.
Code availability
The source code for all pipelines is available for public use at https://github.com/roland-rad-lab/MoCaSeq under the MIT license. In addition, the main workflow described in this protocol is packaged as a Docker container, available at https://cloud.docker.com/repository/docker/rolandradlab/mocaseq.
References
Morse, H. C. III. Origins of Inbred Mice (Elsevier Science, 2012).
van der Weyden, L., Adams, D. J. & Bradley, A. Tools for targeted manipulation of the mouse genome. Physiol. Genomics 11, 133–164 (2002).
Jonkers, J. & Berns, A. Conditional mouse models of sporadic cancer. Nat. Rev. Cancer 2, 251–265 (2002).
Weber, J. & Rad, R. Engineering CRISPR mouse models of cancer. Curr. Opin. Genet. Dev. 54, 88–96 (2019).
Breschi, A., Gingeras, T. R. & Guigo, R. Comparative transcriptomics in human and mouse. Nat. Rev. Genet. 18, 425–440 (2017).
Mouse Genome Sequencing, Consortium et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
She, X., Cheng, Z., Zollner, S., Church, D. M. & Eichler, E. E. Mouse segmental duplication and copy number variation. Nat. Genet. 40, 909–914 (2008).
Egan, C. M., Sridhar, S., Wigler, M. & Hall, I. M. Recurrent DNA copy number variation in the laboratory mouse. Nat. Genet. 39, 1384–1389 (2007).
Keane, T. M. et al. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature 477, 289–294 (2011).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Lee, G. H. et al. Strain specific sensitivity to diethylnitrosamine-induced carcinogenesis is maintained in hepatocytes of C3H/HeN in equilibrium with C57BL/6N chimeric mice. Cancer Res. 51, 3257–3260 (1991).
Reilly, K. M., Loisel, D. A., Bronson, R. T., McLaughlin, M. E. & Jacks, T. Nf1;Trp53 mutant mice develop glioblastoma with evidence of strain-specific effects. Nat. Genet. 26, 109–113 (2000).
Moser, A. R., Hegge, L. F. & Cardiff, R. D. Genetic background affects susceptibility to mammary hyperplasias and carcinomas in Apc(min)/+ mice. Cancer Res. 61, 3480–3485 (2001).
Xu, X. et al. Induction of intrahepatic cholangiocellular carcinoma by liver-specific disruption of Smad4 and Pten in mice. J. Clin. Invest. 116, 1843–1852 (2006).
Rad, R. et al. A genetic progression model of Braf(V600E)-induced intestinal tumorigenesis reveals targets for therapeutic intervention. Cancer Cell 24, 15–29 (2013).
Mueller, S. et al. Evolutionary routes and KRAS dosage define pancreatic cancer phenotypes. Nature 554, 62–68 (2018).
Cancer Genome Atlas Research Network. Integrated genomic characterization of pancreatic ductal adenocarcinoma. Cancer Cell 32, 185–203 e113 (2017).
de Ruiter, J. R., Wessels, L. F. A. & Jonkers, J. Mouse models in the era of large human tumour sequencing studies. Open Biol. 8, 180080 (2018).
McFadden, D. G. et al. Genetic and clonal dissection of murine small cell lung carcinoma progression by genome sequencing. Cell 156, 1298–1311 (2014).
McFadden, D. G. et al. Mutational landscape of EGFR-, MYC-, and Kras-driven genetically engineered mouse models of lung adenocarcinoma. Proc. Natl Acad. Sci. USA 113, E6409–E6417 (2016).
Koren, S. et al. PIK3CA(H1047R) induces multipotency and multi-lineage mammary tumours. Nature 525, 114–118 (2015).
Ferreira, R. M. M. et al. Duct- and acinar-derived pancreatic ductal adenocarcinomas show distinct tumor progression and marker expression. Cell Rep. 21, 966–978 (2017).
Chung, W. J. et al. Kras mutant genetically engineered mouse models of human cancers are genomically heterogeneous. Proc. Natl Acad. Sci. USA 114, E10947–E10955 (2017).
Winters, I. P., Murray, C. W. & Winslow, M. M. Towards quantitative and multiplexed in vivo functional cancer genomics. Nat. Rev. Genet. 19, 741–755 (2018).
Maronpot, R. R., Fox, T., Malarkey, D. E. & Goldsworthy, T. L. Mutations in the ras proto-oncogene: clues to etiology and molecular pathogenesis of mouse liver tumors. Toxicology 101, 125–156 (1995).
Quintanilla, M., Brown, K., Ramsden, M. & Balmain, A. Carcinogen-specific mutation and amplification of Ha-ras during mouse skin carcinogenesis. Nature 322, 78–80 (1986).
You, M., Candrian, U., Maronpot, R. R., Stoner, G. D. & Anderson, M. W. Activation of the Ki-ras protooncogene in spontaneously occurring and chemically induced lung tumors of the strain A mouse. Proc. Natl Acad. Sci. USA 86, 3070–3074 (1989).
McCreery, M. Q. et al. Evolution of metastasis revealed by mutational landscapes of chemically induced skin cancers. Nat. Med. 21, 1514–1520 (2015).
Nassar, D., Latil, M., Boeckx, B., Lambrechts, D. & Blanpain, C. Genomic landscape of carcinogen-induced and genetically induced mouse skin squamous cell carcinoma. Nat. Med. 21, 946–954 (2015).
Westcott, P. M. et al. The mutational landscapes of genetic and chemical models of Kras-driven lung cancer. Nature 517, 489–492 (2015).
Connor, F. et al. Mutational landscape of a chemically-induced mouse model of liver cancer. J. Hepatol. 69, 840–850 (2018).
Arora, K. et al. Deep sequencing of 3 cancer cell lines on 2 sequencing platforms. Preprint at bioRxiv https://doi.org/10.1101/623702 (2019).
Weirather, J. L. et al. Comprehensive comparison of pacific Biosciences and Oxford Nanopore Technologies and their applications to transcriptome analysis. F1000Res 6, 100 (2017).
Uchimura, A. et al. Germline mutation rates and the long-term phenotypic effects of mutation accumulation in wild-type laboratory mice and mutator mice. Genome Res. 25, 1125–1134 (2015).
Milholland, B. et al. Differences between germline and somatic mutation rates in humans and mice. Nat. Commun. 8, 15183 (2017).
Adewoye, A. B., Lindsay, S. J., Dubrova, Y. E. & Hurles, M. E. The genome-wide effects of ionizing radiation on mutation induction in the mammalian germline. Nat. Commun. 6, 6684 (2015).
Einaga, N. et al. Assessment of the quality of DNA from various formalin-fixed paraffin-embedded (FFPE) tissues and the use of this DNA for next-generation sequencing (NGS) with no artifactual mutation. PLoS One 12, e0176280 (2017).
Shi, W. et al. Reliability of whole-exome sequencing for assessing intratumor genetic heterogeneity. Cell Rep. 25, 1446–1457 (2018).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Francis, J. C. et al. Whole-exome DNA sequence analysis of Brca2- and Trp53-deficient mouse mammary gland tumours. J. Pathol. 236, 186–200 (2015).
Ratnaparkhe, M. et al. Defective DNA damage repair leads to frequent catastrophic genomic events in murine and human tumors. Nat. Commun. 9, 4760 (2018).
Kim, S. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods 15, 591–594 (2018).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. Preprint at bioRxiv https://doi.org/10.1101/201178 (2018).
Ye, K., Schulz, M. H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Costello, M. et al. Discovery and characterization of artifactual mutations in deep coverage targeted capture sequencing data due to oxidative DNA damage during sample preparation. Nucleic Acids Res 41, e67 (2013).
Choi, Y. & Chan, A. P. PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 31, 2745–2747 (2015).
Dees, N. D. et al. MuSiC: identifying mutational significance in cancer genomes. Genome Res. 22, 1589–1598 (2012).
Gehring, J. S., Fischer, B., Lawrence, M. & Huber, W. SomaticSignatures: inferring mutational signatures from single-nucleotide variants. Bioinformatics 31, 3673–3675 (2015).
Kuilman, T. et al. CopywriteR: DNA copy number detection from off-target sequence data. Genome Biol. 16, 49 (2015).
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS Comput. Biol. 12, e1004873 (2016).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Korbel, J. O. & Campbell, P. J. Criteria for inference of chromothripsis in cancer genomes. Cell 152, 1226–1236 (2013).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, i333–i339 (2012).
Ha, G. et al. Integrative analysis of genome-wide loss of heterozygosity and monoallelic expression at nucleotide resolution reveals disrupted pathways in triple-negative breast cancer. Genome Res. 22, 1995–2007 (2012).
Choi, Y., Chan, A. P., Kirkness, E., Telenti, A. & Schork, N. J. Comparison of phasing strategies for whole human genomes. PLoS Genet. 14, e1007308 (2018).
Medvedev, P., Fiume, M., Dzamba, M., Smith, T. & Brudno, M. Detecting copy number variation with mated short reads. Genome Res. 20, 1613–1622 (2010).
Guillen, J. FELASA guidelines and recommendations. J. Am. Assoc. Lab Anim. Sci. 51, 311–321 (2012).
Slaoui, M. & Fiette, L. Histopathology procedures: from tissue sampling to histopathological evaluation. Methods Mol. Biol. 691, 69–82 (2011).
Friedrich, M. J. et al. Genome-wide transposon screening and quantitative insertion site sequencing for cancer gene discovery in mice. Nat Protoc. 12, 289–309 (2017).
Witkiewicz, A. K. et al. Whole-exome sequencing of pancreatic cancer defines genetic diversity and therapeutic targets. Nat. Commun. 6, 6744 (2015).
Acknowledgements
D.S. is supported by the European Research Council (Consolidator Grant 648521) and the Deutsche Forschungsgemeinschaft (SA1374/4-2; SFB 1321). I.V. is supported by the European Research Council (Starting Grant INTRAHETEROSEQ) and the Spanish Goverment (SAF2016-76758-R). R.R. is supported by the European Research Council (Consolidator Grants PACA-MET and MSCA-ITN-ETN PRECODE), the Deutsche Forschungsgemeinschaft (DFG RA1629/2-1; SFB1243; SFB1321; SFB1335), the German Cancer Consortium Joint Funding Program, and the Deutsche Krebshilfe (70112480).
Author information
Authors and Affiliations
Contributions
S.L., T.E., M.Z., S.M., L.G.-S., I.V. and R.R. conceptualized, designed or developed analysis workflows, tools or procedures. S.L. integrated and validated bioinformatic workflows. S.M., R.M., M.J.F., R.B. and F.Y. performed wet-lab experiments. G.S., G.S.V. and D.S. provided biological resources and critical input during protocol development. S.L. and R.R. wrote the manuscript with input from T.E., S.M., R.M., M.J.F and I.V.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Malachi Griffith and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Mueller, S. et al. Nature 554, 62–68 (2018): https://doi.org/10.1038/nature25459
Rad, R. et al. Cancer Cell 24, 15–29 (2013): https://doi.org/10.1016/j.ccr.2013.05.014
Key data used in this protocol
Mueller, S. et al. Nature 554, 62–68 (2018): https://doi.org/10.1038/nature25459
Integrated supplementary information
Supplementary Figure 1 Performance of CNVKit calling copy number changes in mouse primary pancreatic cancer cell cultures.
Sensitivity and precision of CNVKit in primary pancreatic cancer cell cultures (n = 38). CNV segments were compared on the gene-level to corresponding reference aCGH data. Segments with a log2 ratio between -0.25 and +0.25 were regarded as copy number neutral. Samples are sorted by the fraction of the genome affected by CNV.
Supplementary Figure 2 Detection of an intragenic EGFR-deletion in human glioblastoma.
a and b, Copy number profiles generated by CopywriteR (a) and CNVKit (b) of a glioblastoma based on WES. Chr7 containing the EGFR locus is shown. DNA and RNA were extracted from FFPE slides and library preparation was performed using Agilent SureSelect Human V6 enrichment kit and Illumina TruSeq Stranded Total RNA kit respectively. Top, Copy number profile of Chr7. Bottom, zoom-in of Chr7 containing the EGFR locus. While CopywriteR detects the amplification of EGFR (~25 copies), CNVKit shows that only Exons 1 to 24 of the EGFR locus are amplified and that exons 25 to 28 remain in the copy number neutral state (arrow). Through RNA-Seq, the copy number neutral state of Exons 25 to 28 could be confirmed. Because CopywriteR only bins off-target” reads (small dots in the middle panel), this small copy number change is not detected. CNVKit correctly detects that Exons 25 to 28 are not included in the amplification by using both on-target and off-target reads.
Supplementary Figure 3 Comparison of CopywriteR, aCGH and M-FISH for sample R1035.
a-c, Copy number profile, generated from WES using CopywriteR (a), aCGH (b) and ten M-FISH karyotypes (c) for sample R1035, a murine primary pancreatic cancer cell culture.
Supplementary Figure 4 Comparison of CopywriteR, aCGH and M-FISH for sample 5123.
a–c, Copy number profile, generated from WES using CopywriteR (a), aCGH (b) and ten M-FISH karyotypes (c) for sample 5123, a murine primary pancreatic cancer cell culture.
Supplementary Figure 5 Comparison of CopywriteR, aCGH and M-FISH for sample S302.
a-c, Copy number profile, generated from WES using CopywriteR (a), aCGH (b) and ten M-FISH karyotypes (c) for sample S302, a murine primary pancreatic cancer cell culture.
Supplementary Figure 6 WGS-based inference of chromothripsis in mouse pancreatic cancer 8661.
a-f, The analysis workflow described in this protocol was used to perform testing of chromothripsis hallmarks from WGS data for sample 8661, a mouse pancreatic cancer primary cell culture. a, Clustering of breakpoints: The distribution of observed distances between breakpoints (n = 41) differs significantly from an exponential distribution (“expected”). P < 10−3; χ2 goodness-of-fit. b, Interspersed loss and retention of heterozygosity: Comparison of CNV and LOH plots for Chr4. Copy number changes cluster in the second half of the chromosome. Only three distinct copy number states (2, 1 and 0 copies) can be identified. The number of heterozygous germline variants is insufficient for LOH analysis. c, Regularity of oscillating copy number states: A Monte Carlo approach was used to simulate the sequential acquisition of observed rearrangements on Chr4 (n = 1000 simulations per number of structural variations). Black dots represent the mean copy number states. The associated 95% confidence interval are shown as black lines. Chr4 showed less copy number states than expected by sequential acquisition of observed rearrangements. d, Randomness of DNA fragment joins: All four types of structural variations are uniformly distributed in the chromothriptic chromosome. P = 0.82; χ2 goodness-of-fit. e, Randomness of DNA fragment order: Start and end positions of observed rearrangements (n = 42) were randomly reordered using a Monte Carlo approach (n = 1000 simulations) to generate a random background distribution. The segment order of sample 8661 is located within the null model of random distribution. Two-sided P = 0.56. f, Ability to walk the derivative chromosome: Rearrangement graph of Chr4 (n = 42 rearrangements). Each fragment is represented by two blocks, indicating the read-orientations (5’ or 3’, indicated in red or grey) for the start and end of each segment, when mapped to the reference genome. P < 10-5; Wald-Wolfowitz test. SV, structural variation.
Supplementary Figure 7 WGS-based inference of chromothripsis in mouse pancreatic cancer 5671.
a-f, The analysis workflow described in this protocol was used to perform testing of chromothripsis hallmarks from WGS data for sample 5671, a mouse pancreatic cancer primary cell culture. a, Clustering of breakpoints: The distribution of observed distances between breakpoints (n = 55) differs significantly from an exponential distribution (“expected”). P = 0.003; χ2 goodness-of-fit. b, Interspersed loss and retention of heterozygosity: Comparison of CNV and LOH plots for Chr15. Copy number changes cluster in the second half of the chromosome. Only three distinct copy number states (2 and 1 copies, ~20 copies for double minute chromosome) can be identified. Regions of loss and retention of heterozygosity alternate, with a very high overlap between regions of LOH and copy number loss. c, Regularity of oscillating copy number states: A Monte Carlo approach was used to simulate the sequential acquisition of observed rearrangements on Chr15 (n = 1000 simulations per number of structural variations). Black dots represent the mean copy number states. The associated 95% confidence interval are shown as black lines. Chr15 showed less copy number states than expected by sequential acquisition of observed rearrangements. d, Randomness of DNA fragment joins: All four types of structural variations are uniformly distributed in the chromothriptic chromosome. P = 0.23; χ2 goodness-of-fit. e, Randomness of DNA fragment order: Start and end positions of observed rearrangements (n = 56) were randomly reordered using a Monte Carlo approach (n = 1000 simulations) to generate a random background distribution. The segment order of sample 5671 is located within the null model of random distribution. Two-sided P = 0.2. f, Ability to walk the derivative chromosome: Rearrangement graph of Chr15 (n = 56 rearrangements). Each fragment is represented by two blocks, indicating the read-orientations (5’ or 3’, indicated in red or grey) for the start and end of each segment, when mapped to the reference genome. P = 0.004; Wald-Wolfowitz test. SV, structural variation.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7, Supplementary Methods, Supplementary Tables 1–3
Rights and permissions
About this article
Cite this article
Lange, S., Engleitner, T., Mueller, S. et al. Analysis pipelines for cancer genome sequencing in mice. Nat Protoc 15, 266–315 (2020). https://doi.org/10.1038/s41596-019-0234-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0234-7
This article is cited by
-
Towards accurate indel calling for oncopanel sequencing through an international pipeline competition at precisionFDA
Scientific Reports (2024)
-
An analysis pipeline for understanding 6-thioguanine effects on a mouse tumour genome
Cancer Immunology, Immunotherapy (2024)
-
Critically short telomeres derepress retrotransposons to promote genome instability in embryonic stem cells
Cell Discovery (2023)
-
Epigenetic dysregulation from chromosomal transit in micronuclei
Nature (2023)
-
Interferon signaling promotes tolerance to chromosomal instability during metastatic evolution in renal cancer
Nature Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.