Abstract
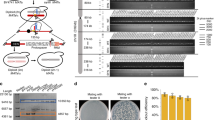
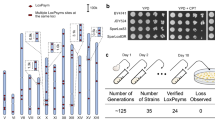
CRISPR–Cas9-facilitated functional chromosome fusion allows the generation of a series of yeast strains with progressively reduced chromosome numbers that are valuable resources for the study of fundamental concepts in chromosome biology, including replication, recombination and segregation. We created a new yeast strain with a single chromosome by using the protocol for chromosome fusion described herein. To ensure the accuracy of chromosome fusions in yeast, the long redundant repetitive sequences near linear chromosomal ends are deleted, and the fusion orders are correspondingly determined. Possible influence on gene expression is minimized to retain gene functionality. This protocol provides experimentally derived guidelines for the generation of functional chromosome fusions in yeast, especially for the deletion of repetitive sequences, the determination of the fusion order and cleavage sites, and primary evaluation of the functionality of chromosome fusions. Beginning with design, one round of typical chromosome fusion and functional verifications can be accomplished within 18 d.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
Data availability
The plasmids used in this protocol, including pCas9 (accession number 1.2624), pHIS426 (accession number 1.2623) and pXX11 (accession number 1.2613), can be obtained from the Registry and Database of Bioparts for Synthetic Biology (http://npbiosys.scbit.org/strainOrder) upon reasonable request. All relevant data are reported in the article.
References
Gordon, J. L., Byrne, K. P. & Wolfe, K. H. Mechanisms of chromosome number evolution in yeast. PLoS Genet. 7, e1002190 (2011).
McClintock, B. The stability of broken ends of chromosomes in Zea mays. Genetics 26, 234–282 (1941).
Hill, A. & Bloom, K. Genetic manipulation of centromere function. Mol. Cell. Biol. 7, 2397–2405 (1987).
Hill, A. & Bloom, K. Acquisition and processing of a conditional dicentric chromosome in Saccharomyces cerevisiae. Mol. Cell. Biol. 9, 1368–1370 (1989).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Shao, Y. et al. Creating a functional single-chromosome yeast. Nature 560, 331–335 (2018).
Goffeau, A. et al. Life with 6000 genes. Science 274, 546–567 (1996).
Qin, Z. & Cohen, S. N. Long palindromes formed in Streptomyces by nonrecombinational intra-strand annealing. Genes Dev. 14, 1789–1796 (2000).
Akgun, E. et al. Palindrome resolution and recombination in the mammalian germ line. Mol. Cell. Biol. 17, 5559–5570 (1997).
Leach, D. R. Long DNA palindromes, cruciform structures, genetic instability and secondary structure repair. BioEssays 16, 893–900 (1994).
Henikoff, S. & Henikoff, J. G. “Point” centromeres of Saccharomyces harbor single centromere-specific nucleosomes. Genetics 190, 1575–1577 (2012).
Jager, D. & Philippsen, P. Stabilization of dicentric chromosomes in Saccharomyces cerevisiae by telomere addition to broken ends or by centromere deletion. EMBO J. 8, 247–254 (1989).
Pobiega, S. & Marcand, S. Dicentric breakage at telomere fusions. Genes Dev. 24, 720–733 (2010).
Lopez, V. et al. Cytokinesis breaks dicentric chromosomes preferentially at pericentromeric regions and telomere fusions. Genes Dev. 29, 322–336 (2015).
Luo, J., Sun, X., Cormack, B. P. & Boeke, J. D. Karyotype engineering by chromosome fusion leads to reproductive isolation in yeast. Nature 560, 392–396 (2018).
Shao, Y., Lu, N., Qin, Z. & Xue, X. CRISPR–Cas9 facilitated multiple-chromosome fusion in Saccharomyces cerevisiae. ACS Synth. Biol 7, 2706–2708 (2018).
Xue, X. et al. MEGA (multiple essential genes assembling) deletion and replacement method for genome reduction in Escherichia coli. ACS Synth. Biol. 4, 700–706 (2015).
Gietz, R. D. & Schiestl, R. H. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 31–34 (2007).
Chung, C. T., Niemela, S. L. & Miller, R. H. One-step preparation of competent Escherichia coli: transformation and storage of bacterial cells in the same solution. Proc. Natl. Acad. Sci. USA 86, 2172–2175 (1989).
Acknowledgements
This research was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB19000000, Z.Q.; 153D31KYSB20160074, Z.Q.), the National Natural Science Foundation of China (31830105, Z.Q.; 31770099 and 31200059, X.X.), Shanghai Research Project (18JC1420200, Z.Q.) and China Postdoctoral Science Foundation (2018M640427 and 2019T120362, Y.S.)
Author information
Authors and Affiliations
Contributions
Z.Q. and X.X. designed and analyzed all the experiments. Y.S. constructed the chromosome fusion yeast strains and performed PCR verification. N.L. conducted the PFGE confirmation experiment and growth assays. X.X. and Y.S. wrote the primary manuscript with a substantial contribution from Z.Q.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Protocols thanks Meru Sadhu and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Shao, Y. et al. Nature 560, 331–335 (2018): https://doi.org/10.1038/s41586-018-0382-x
Shao, Y., Lu, N., Qin, Z. & Xue, X. ACS Synth. Biol. 7, 2706–2708 (2018): https://doi.org/10.1021/acssynbio.8b00397
Shao, Y. et al. Cell Res. 29, 87–89 (2019): https://doi.org/10.1038/s41422-018-0110-y
Supplementary information
Rights and permissions
About this article
Cite this article
Shao, Y., Lu, N., Xue, X. et al. Creating functional chromosome fusions in yeast with CRISPR–Cas9. Nat Protoc 14, 2521–2545 (2019). https://doi.org/10.1038/s41596-019-0192-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0192-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.