Abstract
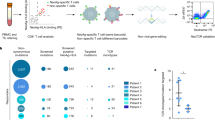
The identification of immunogenic neoantigens and their cognate T cells represents the most crucial and rate-limiting steps in the development of personalized cancer immunotherapies that are based on vaccination or on infusion of T cell receptor (TCR)-engineered T cells. Recent advances in deep-sequencing technologies and in silico prediction algorithms have allowed rapid identification of candidate neoepitopes. However, large-scale validation of putative neoepitopes and the isolation of reactive T cells are challenging because of the limited availablity of patient material and the low frequencies of neoepitope-specific T cells. Here we describe a standardized protocol for the induction of neoepitope-reactive T cells from healthy donor T cell repertoires, unaffected by the potentially immunosuppressive environment of the tumor-bearing host. Monocyte-derived dendritic cells (DCs) transfected with mRNA encoding candidate neoepitopes are used to prime autologous naive CD8+ T cells. Antigen-specific T cells that recognize endogenously processed and presented epitopes are detected using peptide–MHC (pMHC) multimers. Single multimer-positive T cells are sorted for the identification of TCR sequences, after an optional step that includes clonal expansion and functional characterization. The time required to identify neoepitope-specific T cells is 15 d, with an additional 2–4 weeks required for clonal expansion and downstream functional characterization. Identified neoepitopes and corresponding TCRs provide candidates for use in vaccination and TCR-based cancer immunotherapies, and datasets generated by this technology should be useful for improving algorithms to predict immunogenic neoantigens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Science Shaped



Similar content being viewed by others
References
Rosenberg, S. A. & Restifo, N. P. Adoptive cell transfer as personalized immunotherapy for human cancer. Science 348, 62–68 (2015).
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Tran, E. et al. Cancer immunotherapy based on mutation-specific CD4+ T cells in a patient with epithelial cancer. Science 344, 641–645 (2014).
Tran, E. et al. T-cell transfer therapy targeting mutant KRAS in cancer. N. Engl. J. Med. 375, 2255–2262 (2016).
Carreno, B. M. et al. A dendritic cell vaccine increases the breadth and diversity of melanoma neoantigen-specific T cells. Science 348, 803–808 (2015).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Ott, P. A. et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 547, 217–221 (2017).
Rizvi, N. A. et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Robbins, P. F. et al. Mining exomic sequencing data to identify mutated antigens recognized by adoptively transferred tumor-reactive T cells. Nat. Med. 19, 747–752 (2013).
van Rooij, N. et al. Tumor exome analysis reveals neoantigen-specific T-cell reactivity in an ipilimumab-responsive melanoma. J. Clin. Oncol. 31, e439–e442 (2013).
Linnemann, C. et al. High-throughput epitope discovery reveals frequent recognition of neo-antigens by CD4+ T cells in human melanoma. Nat. Med. 21, 81–85 (2015).
Tran, E. et al. Immunogenicity of somatic mutations in human gastrointestinal cancers. Science 350, 1387–1390 (2015).
Gubin, M. M. et al. Checkpoint blockade cancer immunotherapy targets tumour-specific mutant antigens. Nature 515, 577–581 (2014).
Matsushita, H. et al. Cancer exome analysis reveals a T-cell-dependent mechanism of cancer immunoediting. Nature 482, 400–404 (2012).
Yadav, M. et al. Predicting immunogenic tumour mutations by combining mass spectrometry and exome sequencing. Nature 515, 572–576 (2014).
Karpanen, T. & Olweus, J. The potential of donor T-cell repertoires in neoantigen-targeted cancer immunotherapy. Front. Immunol. 8, 1718 (2017).
Stronen, E. et al. Targeting of cancer neoantigens with donor-derived T cell receptor repertoires. Science 352, 1337–1341 (2016).
Qi, Q. et al. Diversity and clonal selection in the human T-cell repertoire. Proc. Natl Acad. Sci. USA 111, 13139–13144 (2014).
Arstila, T. P. et al. A direct estimate of the human αβ T cell receptor diversity. Science 286, 958–961 (1999).
Van Tendeloo, V. F. et al. Highly efficient gene delivery by mRNA electroporation in human hematopoietic cells: superiority to lipofection and passive pulsing of mRNA and to electroporation of plasmid cDNA for tumor antigen loading of dendritic cells. Blood 98, 49–56 (2001).
Ponsaerts, P. et al. Messenger RNA electroporation of human monocytes, followed by rapid in vitro differentiation, leads to highly stimulatory antigen-loaded mature dendritic cells. J. Immunol. 169, 1669–1675 (2002).
Hui, S. W. Effects of pulse length and strength on electroporation efficiency. Methods Mol. Biol. 55, 29–40 (1995).
Livingston, B. D. et al. Optimization of epitope processing enhances immunogenicity of multiepitope DNA vaccines. Vaccine 19, 4652–4660 (2001).
Schubert, B. & Kohlbacher, O. Designing string-of-beads vaccines with optimal spacers. Genome Med. 8, 9 (2016).
Toussaint, N. C., Maman, Y., Kohlbacher, O. & Louzoun, Y. Universal peptide vaccines—optimal peptide vaccine design based on viral sequence conservation. Vaccine 29, 8475–8753 (2011).
Pittet, M. J. et al. High frequencies of naive Melan-A/MART-1-specific CD8+ T cells in a large proportion of human histocompatibility leukocyte antigen (HLA)-A2 individuals. J. Exp. Med. 190, 705–715 (1999).
Wolfl, M. & Greenberg, P. D. Antigen-specific activation and cytokine-facilitated expansion of naive, human CD8+ T cells. Nat. Protoc. 9, 950–966 (2014).
Hadrup, S. R. et al. Parallel detection of antigen-specific T-cell responses by multidimensional encoding of MHC multimers. Nat. Methods 6, 520–526 (2009).
Han, A., Glanville, J., Hansmann, L. & Davis, M. M. Linking T-cell receptor sequence to functional phenotype at the single-cell level. Nat. Biotechnol. 32, 684–692 (2014).
Tang, F. et al. RNA-seq analysis to capture the transcriptome landscape of a single cell. Nat. Protoc. 5, 516–535 (2010).
Davis, M. M. & Bjorkman, P. J. T-cell antigen receptor genes and T-cell recognition. Nature 334, 395–402 (1988).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Bentzen, A. K. et al. Large-scale detection of antigen-specific T cells using peptide-MHC-I multimers labeled with DNA barcodes. Nat. Biotechnol. 34, 1037–1045 (2016).
Andreatta, M. & Nielsen, M. Gapped sequence alignment using artificial neural networks: application to the MHC class I system. Bioinformatics 32, 511–517 (2016).
Bassani-Sternberg, M. et al. Direct identification of clinically relevant neoepitopes presented on native human melanoma tissue by mass spectrometry. Nat. Commun. 7, 13404 (2016).
Chheda, Z. S. et al. Novel and shared neoantigen derived from histone 3 variant H3.3K27M mutation for glioma T cell therapy. J. Exp. Med. 215, 141–157 (2018).
Kalaora, S. et al. Combined analysis of antigen presentation and T-cell recognition reveals restricted immune responses in melanoma. Cancer Discov. 8, 1366–1375 (2018).
Andersen, R. S. et al. Parallel detection of antigen-specific T cell responses by combinatorial encoding of MHC multimers. Nat. Protoc. 7, 891–902 (2012).
Toebes, M. et al. Design and use of conditional MHC class I ligands. Nat. Med. 12, 246–251 (2006).
Toebes, M., Rodenko, B., Ovaa, H. & Schumacher, T. N. Generation of peptide MHC class I monomers and multimers through ligand exchange. Curr. Protoc. Immunol. 87, 18.16.1–18.16.20 (2009).
Hadrup, S. R. et al. Cryopreservation of MHC multimers: recommendations for quality assurance in detection of antigen specific T cells. Cytometry A 87, 37–48 (2015).
Acknowledgements
We thank the Oslo University Hospital (OUH) flow cytometry core facility for excellent technical assistance. This work was supported by Stiftelsen Kristian Gerhard Jebsen (J.O. and T.N.S.), South-Eastern Regional Health Authority Norway, the Research Council of Norway, the Norwegian Cancer Society, the University of Oslo, Oslo University Hospital (all J.O.) and the Queen Wilhelmina Cancer Research Award (T.N.S.).
Author information
Authors and Affiliations
Contributions
M.A., Z.F., E.G., M.-L.B. and J.O. conceived and designed the experiments. M.A., Z.F. and E.G. performed the experiments and analyzed the data. B.S., O.K., W.Y. and M.T. designed and synthesized mRNA minigenes and pMHC multimers. M.A., Z.F., E.G., E.S., T.N.S. and J.O. wrote the manuscript. All authors reviewed and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.O. has a collaboration with Kite Pharma and is a member of the Scientific Advisory Board of Intellia Therapeutics. J.O is the inventor on the patent WO2015071763 (CTL peptide epitopes and antigen-specific T cells, methods for their discovery, and uses thereof). T.N.S. is a consultant for Adaptive Biotechnologies, AIMM Therapeutics, Allogene Therapeutics, Amgen, Merus, Neon Therapeutics and Scenic Biotech; is a recipient of grant/research support from MSD, Bristol-Myers Squibb and Merck KGaA; is a stockholder in AIMM Therapeutics, Allogene Therapeutics, Merus, Neogene Therapeutics and Neon Therapeutics; and is a venture partner at Third Rock Ventures.
Additional information
Journal peer review information: Nature Protocols thanks Timothy Chan and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Strønen, E. et al. Science 352, 1337–1341 (2016): http://science.sciencemag.org/content/352/6291/1337
Kumari, S. et al. Proc. Natl. Acad. Sci. USA 111, 403–408 (2014): https://www.pnas.org/content/111/1/403
Integrated supplementary information
Supplementary Figure 1
Flow cytometric characterization of the purity of naive and memory CD8+ T cells isolated from the same donor.
Supplementary Figure 2 Different ratios of DCs to naive CD8+ T cells.
M2-minigene-electroporated DCs were cocultured with T cells at DC:T cell ratios of 1:8, 1:4, 1:2 and 1:1 (n = 4 donors, 3 cultures/donor), and cultures were analyzed for the presence of neo-1, -2, -3, -4 and -5 pMHC-multimer-reactive populations on day 12. Each dot represents the percentage of pMHC-multimer-positive cells among CD8+ cells identified by flow cytometry.
Supplementary Figure 3 Comparison of the ability of M2 and M3 minigenes to induce T cell responses.
Blood from 20 different donors was used to initiate cultures stimulated with either M2 or M3 minigene-electroporated DCs to compare the induction of T cell responses against neo-1, -2, -3, -4 and -5 epitopes (n = 10 donors/minigene, 3 cultures/donor). Circles designate data points from M2-induced cultures, and triangles designate data points from M3-induced cultures. Data plotted as mean ± s.e.m. *P < 0.05.
Supplementary Figure 4 Multiplexed functional analysis of T cell clones.
Clones were labeled as described in the legend of Fig. 4. To the right, the CD107a/b degranulation response of 16 individual clones against target cells pulsed with 10 nM neo-3 peptide is shown. Numbers in the lower left corner of plots correspond to individual clones in the dot plot to the left.
Supplementary Figure 5 Gating strategy for the identification of pMHC-multimer-positive CD8+ T cells.
Lymphocytes were identified, and doublets and dead cells were excluded with the help of forward (FSC) and side scatter (SSC) gates and live/dead fixable near-IR dead cell staining. From the live cell gate, CD8+ T cells were gated and pMHC-multimer-reactive cells were identified as double positive for PE- and APC-conjugated pMHC multimers.
Supplementary information
Supplementary Information
Supplementary Figures 1–5, Supplementary Note, Supplementary Methods and Supplementary Table 1
Rights and permissions
About this article
Cite this article
Ali, M., Foldvari, Z., Giannakopoulou, E. et al. Induction of neoantigen-reactive T cells from healthy donors. Nat Protoc 14, 1926–1943 (2019). https://doi.org/10.1038/s41596-019-0170-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0170-6
This article is cited by
-
Novel insights into TCR-T cell therapy in solid neoplasms: optimizing adoptive immunotherapy
Experimental Hematology & Oncology (2024)
-
Adoptive neoantigen-reactive T cell therapy: improvement strategies and current clinical researches
Biomarker Research (2023)
-
Peptide vaccine from cancer-testis antigen ODF2 can potentiate the cytotoxic T lymphocyte infiltration through IL-15 in non-MSI-H colorectal cancer
Cancer Immunology, Immunotherapy (2023)
-
A T cell receptor targeting a recurrent driver mutation in FLT3 mediates elimination of primary human acute myeloid leukemia in vivo
Nature Cancer (2023)
-
Adoptive cell therapies in thoracic malignancies
Cancer Immunology, Immunotherapy (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.