Abstract
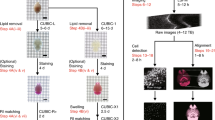
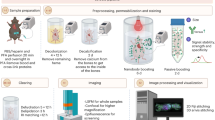
Understanding the structure–function relationships between diverse cell types in a complex organ environment requires detailed in situ reconstruction of cell-associated molecular properties in the context of 3D, macro-scale tissue architecture. We recently developed clearing-enhanced 3D (Ce3D), a simple and effective method for tissue clearing that achieves excellent transparency; preserves cell morphology, tissue architecture, and reporter molecule fluorescence; and is robustly compatible with direct immunolabeling. These characteristics permit high-quality multiplex fluorescence microscopy of large tissue volumes, as well as image analysis using advanced platforms such as volumetric histocytometry, collectively allowing quantitative characterization of cells with respect to their spatial positioning within tissues on the basis of phenotypic and functional markers. Ce3D clearing is fast, achieving robust transparency of most tissues within 24 h, albeit still necessitating additional time for staining, imaging, and analysis (1–2 weeks). Here, we provide detailed procedures for tissue clearing using Ce3D, including optimized workflows for tissue processing and staining, as well as treatment of difficult-to-clear organs such as the brain. We also describe a new procedure for RNA detection in Ce3D-treated tissues, as well as provide additional details for the volumetric histocytometry data processing steps. Finally, we discuss limitations and work-around strategies for improving antibody-based tissue immunolabeling, fluorophore multiplexing, large-volume microscopy, and computational analysis of large image datasets. Together, these detailed procedures and solutions for high-resolution volumetric microscopy with Ce3D enable quantitative visualization of cells and tissues at a high level of detail, allowing exploration of cellular spatial relationships in a variety of tissue settings.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The image datasets generated and/or analyzed during the current study are available from the corresponding authors upon reasonable request.
References
Spitzer, M. H. & Nolan, G. P. Mass cytometry: single cells, many features. Cell 165, 780–791 (2016).
Alcantara-Hernandez, M. et al. High-dimensional phenotypic mapping of human dendritic cells reveals interindividual variation and tissue specialization. Immunity 47, 1037–1050 (2017).
Papalexi, E. & Satija, R. Single-cell RNA sequencing to explore immune cell heterogeneity. Nat. Rev. Immunol. 18, 35–45 (2018).
Altschuler, S. J. & Wu, L. F. Cellular heterogeneity: do differences make a difference? Cell 141, 559–563 (2010).
Junttila, M. R. & de Sauvage, F. J. Influence of tumour micro-environment heterogeneity on therapeutic response. Nature 501, 346–354 (2013).
Cutrale, F. et al. Hyperspectral phasor analysis enables multiplexed 5D in vivo imaging. Nat. Methods 14, 149–152 (2017).
Gerner, M. Y., Kastenmuller, W., Ifrim, I., Kabat, J. & Germain, R. N. Histo-cytometry: a method for highly multiplex quantitative tissue imaging analysis applied to dendritic cell subset microanatomy in lymph nodes. Immunity 37, 364–376 (2012).
Lin, J. R., Fallahi-Sichani, M. & Sorger, P. K. Highly multiplexed imaging of single cells using a high-throughput cyclic immunofluorescence method. Nat. Commun. 6, 8390 (2015).
Goltsev, Y. et al. Deep profiling of mouse splenic architecture with CODEX multiplexed imaging. Cell 174, 968–981 (2018).
Keren, L. et al. A structured tumor-immune microenvironment in triple negative breast cancer revealed by multiplexed ion beam imaging. Cell 174, 1373–1387 (2018).
Gerner, M. Y., Torabi-Parizi, P. & Germain, R. N. Strategically localized dendritic cells promote rapid T cell responses to lymph-borne particulate antigens. Immunity 42, 172–185 (2015).
Vu Manh, T. P., Bertho, N., Hosmalin, A., Schwartz-Cornil, I. & Dalod, M. Investigating evolutionary conservation of dendritic cell subset identity and functions. Front. Immunol. 6, 260 (2015).
Gerner, M. Y., Casey, K. A., Kastenmuller, W. & Germain, R. N. Dendritic cell and antigen dispersal landscapes regulate T cell immunity. J. Exp. Med. 214, 3105–3122 (2017).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Liu, Z. et al. Immune homeostasis enforced by co-localized effector and regulatory T cells. Nature 528, 225–230 (2015).
Petrovas, C. et al. Follicular CD8 T cells accumulate in HIV infection and can kill infected cells in vitro via bispecific antibodies. Sci. Transl. Med. 9, aag2285 (2017).
Petrovas, C. et al. CD4 T follicular helper cell dynamics during SIV infection. J. Clin. Invest. 122, 3281–3294 (2012).
Preite, S. et al. Hyperactivated PI3Kdelta promotes self and commensal reactivity at the expense of optimal humoral immunity. Nat. Immunol. 19, 986–1000 (2018).
Sayin, I. et al. Spatial distribution and function of T follicular regulatory cells in human lymph nodes. J. Exp. Med. 215, 1531–1542 (2018).
Mao, K. et al. Innate and adaptive lymphocytes sequentially shape the gut microbiota and lipid metabolism. Nature 554, 255–259 (2018).
Fonseca, D. M. et al. Microbiota-dependent sequelae of acute infection compromise tissue-specific immunity. Cell 163, 354–366 (2015).
Radtke, A. J. et al. Lymph-node resident CD8α+ dendritic cells capture antigens from migratory malaria sporozoites and induce CD8+ T cell responses. PLoS Pathog. 11, e1004637 (2015).
Steinert, E. M. et al. Quantifying memory CD8 T cells reveals regionalization of immunosurveillance. Cell 161, 737–749 (2015).
Masuda, A. et al. An improved method for isolation of epithelial and stromal cells from the human endometrium. J. Reprod. Dev. 62, 213–218 (2016).
Jabbari, A., Legge, K. L. & Harty, J. T. T cell conditioning explains early disappearance of the memory CD8 T cell response to infection. J. Immunol. 177, 3012–3018 (2006).
Richardson, D. S. & Lichtman, J. W. Clarifying tissue clearing. Cell 162, 246–257 (2015).
Tainaka, K., Kuno, A., Kubota, S. I., Murakami, T. & Ueda, H. R. Chemical principles in tissue clearing and staining protocols for whole-body cell profiling. Annu. Rev. Cell Dev. Biol. 32, 713–741 (2016).
Erturk, A. et al. Three-dimensional imaging of solvent-cleared organs using 3DISCO. Nat. Protoc. 7, 1983–1995 (2012).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Hama, H. et al. ScaleS: an optical clearing palette for biological imaging. Nat. Neurosci. 18, 1518–1529 (2015).
Hama, H. et al. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat. Neurosci. 14, 1481–1488 (2011).
Tainaka, K. et al. Whole-body imaging with single-cell resolution by tissue decolorization. Cell 159, 911–924 (2014).
Chung, K. et al. Structural and molecular interrogation of intact biological systems. Nature 497, 332–337 (2013).
Yang, B. et al. Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell 158, 945–958 (2014).
Murray, E. et al. Simple, scalable proteomic imaging for high-dimensional profiling of intact systems. Cell 163, 1500–1514 (2015).
Dodt, H. U. et al. Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain. Nat. Methods 4, 331–336 (2007).
Kuwajima, T. et al. ClearT: a detergent- and solvent-free clearing method for neuronal and non-neuronal tissue. Development 140, 1364–1368 (2013).
Ke, M. T., Fujimoto, S. & Imai, T. SeeDB: a simple and morphology-preserving optical clearing agent for neuronal circuit reconstruction. Nat. Neurosci. 16, 1154–1161 (2013).
Ke, M. T. et al. Super-resolution mapping of neuronal circuitry with an index-optimized clearing agent. Cell Rep. 14, 2718–2732 (2016).
Pan, C. et al. Shrinkage-mediated imaging of entire organs and organisms using uDISCO. Nat. Methods 13, 859–867 (2016).
Murakami, T. C. et al. A three-dimensional single-cell-resolution whole-brain atlas using CUBIC-X expansion microscopy and tissue clearing. Nat. Neurosci. 21, 625–637 (2018).
Tainaka, K. et al. Chemical landscape for tissue clearing based on hydrophilic reagents. Cell Rep. 24, 2196–2210 (2018).
Kubota, S. I. et al. Whole-body profiling of cancer metastasis with single-cell resolution. Cell Rep. 20, 236–250 (2017).
Li, W., Germain, R. N. & Gerner, M. Y. Multiplex, quantitative cellular analysis in large tissue volumes with clearing-enhanced 3D microscopy (Ce3D). Proc. Natl. Acad. Sci. USA 114, E7321–E7330 (2017).
Wang, F. et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn. 14, 22–29 (2012).
Katoh, K. Microwave-assisted tissue preparation for rapid fixation, decalcification, antigen retrieval, cryosectioning, and immunostaining. Int. J. Cell Biol. 2016, 7076910 (2016).
Beghein, E. & Gettemans, J. Nanobody technology: a versatile toolkit for microscopic imaging, protein-protein interaction analysis, and protein function exploration. Front. Immunol. 8, 771 (2017).
Liu, T. L. et al. Observing the cell in its native state: imaging subcellular dynamics in multicellular organisms. Science 360, eaaq1392 (2018).
Acknowledgements
We thank K. Mao for helping with gut and mammary gland tissue preparation. In addition, we thank all members of the Laboratory of Immune System Biology, National Institute of Allergy and Infectious Diseases (NIAID) at the NIH for many helpful comments during the course of these experiments. This research was supported by the Intramural Research Program, NIAID, NIH, and NIH grant no. R01AI134713 MYG.
Author information
Authors and Affiliations
Contributions
W.L., M.Y.G., and R.N.G. designed the experiments. W.L. and M.Y.G. performed the experiments and analysis. M.Y.G., W.L., and R.N.G. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
A patent for the methodology described in this paper was filed with the US Patent Office (PCT Patent Application PCT/US2017/049133, HHS reference no. E–168–2016, ‘Enhanced Tissue Clearing Solution, Clearing-Enhanced 3D (Ce3D), Compatible With Advanced Fluorescence Microscopy Imaging’).
Additional information
Journal peer review information: Nature Protocols thanks Michael Donovan and Constantinos Petrovas for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Li, W., Germain, R.N. and Gerner, M.Y. PNAS 114, E7321–E7330 (2017): https://www.pnas.org/content/114/35/E7321
Gerner, M.Y., Kastenmuller, W., Ifrim, I., Kabat, J. & Germain, R.N. Immunity 37, 364–376 (2012): https://www.cell.com/immunity/fulltext/S1074-7613(12)00321-4
Liu, Z. et al. Nature 528, 225–230 (2015): https://www.nature.com/articles/nature16169
Supplementary information
Rights and permissions
About this article
Cite this article
Li, W., Germain, R.N. & Gerner, M.Y. High-dimensional cell-level analysis of tissues with Ce3D multiplex volume imaging. Nat Protoc 14, 1708–1733 (2019). https://doi.org/10.1038/s41596-019-0156-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0156-4
This article is cited by
-
Endothelial sensing of AHR ligands regulates intestinal homeostasis
Nature (2023)
-
IBEX: a user-friendly and open-source solution for high-plex immunostaining
Nature Reviews Immunology (2023)
-
Formation of the Heart: Defining Cardiomyocyte Progenitors at Single-Cell Resolution
Current Cardiology Reports (2023)
-
Multitier mechanics control stromal adaptations in the swelling lymph node
Nature Immunology (2022)
-
Generation and characterization of hair-bearing skin organoids from human pluripotent stem cells
Nature Protocols (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.